

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.5
(404 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
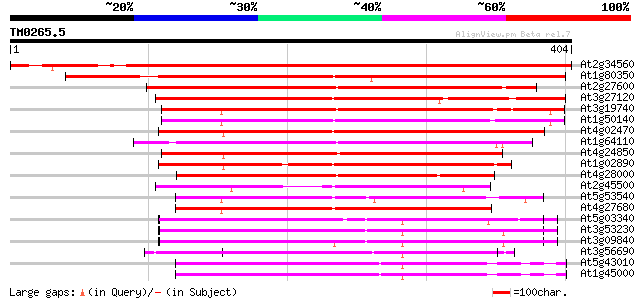
Score E
Sequences producing significant alignments: (bits) Value
At2g34560 katanin like protein 555 e-158
At1g80350 CAD ATPase (AAA1) 279 2e-75
At2g27600 putative AAA-type ATPase 261 4e-70
At3g27120 spastin protein, putative 235 4e-62
At3g19740 ATPase, putative 218 4e-57
At1g50140 hypothetical protein, 5' partial 214 7e-56
At4g02470 213 1e-55
At1g64110 unknown protein 209 3e-54
At4g24850 unknown protein (At4g24850) 207 1e-53
At1g02890 hypothetical protein 200 1e-51
At4g28000 putative protein 197 1e-50
At2g45500 hypothetical protein 189 2e-48
At5g53540 26S proteasome regulatory particle chain RPT6-like pro... 188 4e-48
At4g27680 unknown protein 186 3e-47
At5g03340 transitional endoplasmic reticulum ATPase 183 1e-46
At3g53230 CDC48 - like protein 181 5e-46
At3g09840 putative transitional endoplasmic reticulum ATPase 180 1e-45
At3g56690 calmodulin-binding protein 163 2e-40
At5g43010 26S proteasome AAA-ATPase subunit RPT4a (gb|AAF22524.1) 154 8e-38
At1g45000 26S proteasome AAA-ATPase subunit RPT4a like protein 154 8e-38
>At2g34560 katanin like protein
Length = 384
Score = 555 bits (1431), Expect = e-158
Identities = 289/409 (70%), Positives = 329/409 (79%), Gaps = 30/409 (7%)
Query: 1 MADEEPMPTRWSFQDFKLYYDSKFGRKKV----VQNGENADKAVGNGSSMSVVSNGNVHS 56
MA +EP TRWSF FGRKK+ V N + + NG++ V +N + +
Sbjct: 1 MATDEPSQTRWSFL---------FGRKKLPEEDVSNKDQPEDGSSNGNNGDVNNNSSPVT 51
Query: 57 KRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDIIR 116
+ + A+ NG N + E+P+KS+ PPFESAE RTLAESLSRDIIR
Sbjct: 52 NQDGNTAL--------ANG--------NVIREKPKKSMFPPFESAETRTLAESLSRDIIR 95
Query: 117 GSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKA 176
G+P++KWESIKGLENAK+LLKEAVVMPIKYP YF GLL+PWKGILLFGPPGTGKTMLAKA
Sbjct: 96 GNPNIKWESIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKA 155
Query: 177 VATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG-EA 235
VATEC TTFFNISASSVVSKWRGDSEKL++VLF LARHHAPSTIFLDEIDAIISQRG E
Sbjct: 156 VATECNTTFFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEG 215
Query: 236 RSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPE 295
RSEHEASRRLKTELLIQMDGL +T+ELVFVLAATNLPWELDAAMLRRLEKRILVPLP+PE
Sbjct: 216 RSEHEASRRLKTELLIQMDGLQKTNELVFVLAATNLPWELDAAMLRRLEKRILVPLPDPE 275
Query: 296 ARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRED 355
AR MFE L+P QP +E +P+D+LV ++EGYSGSDIR+LCKE AMQPLRR ++ LE RED
Sbjct: 276 ARRGMFEMLIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLAILEDRED 335
Query: 356 LVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQILQ 404
+VPE+ELPK+GPI PEDI AL NTRPSAHLHAH YDKFN DYGSQIL+
Sbjct: 336 VVPEDELPKIGPILPEDIDRALSNTRPSAHLHAHLYDKFNDDYGSQILK 384
>At1g80350 CAD ATPase (AAA1)
Length = 523
Score = 279 bits (713), Expect = 2e-75
Identities = 149/368 (40%), Positives = 229/368 (61%), Gaps = 21/368 (5%)
Query: 41 GNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFES 100
G G + S + G S A + +++ NG + S + E P + L
Sbjct: 168 GRGGATSKSTAGARSSTAGKKGAASKSNKAESMNGDAEDGKSKRGLYEGPDEDL------ 221
Query: 101 AEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGI 160
A L RD++ +P V+W+ + GL AKRLL+EAVV+P+ P+YF G+ PWKG+
Sbjct: 222 ------AAMLERDVLDSTPGVRWDDVAGLSEAKRLLEEAVVLPLWMPEYFQGIRRPWKGV 275
Query: 161 LLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTI 220
L+FGPPGTGKT+LAKAVATEC TTFFN+S++++ SKWRG+SE++V+ LF LAR +APSTI
Sbjct: 276 LMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTI 335
Query: 221 FLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRT-------DELVFVLAATNLPW 273
F+DEID++ + RG EHE+SRR+K+ELL+Q+DG++ T ++V VLAATN PW
Sbjct: 336 FIDEIDSLCNSRG-GSGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPW 394
Query: 274 ELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRL 333
++D A+ RRLEKRI +PLP+ E+R A+ L + + + +TEGYSG D+
Sbjct: 395 DIDEALRRRLEKRIYIPLPDFESRKALININLRTVEVASDVNIEDVARRTEGYSGDDLTN 454
Query: 334 LCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHAHKYD 392
+C++ +M +RR ++ + E ++ P+ D + A++ +PS + K++
Sbjct: 455 VCRDASMNGMRRKIAGKTRDEIKNMSKDDISNDPVAMCDFEEAIRKVQPSVSSSDIEKHE 514
Query: 393 KFNADYGS 400
K+ +++GS
Sbjct: 515 KWLSEFGS 522
>At2g27600 putative AAA-type ATPase
Length = 435
Score = 261 bits (668), Expect = 4e-70
Identities = 135/282 (47%), Positives = 190/282 (66%), Gaps = 4/282 (1%)
Query: 99 ESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWK 158
E E L L+ I+R P++KW + GLE+AK+ L+EAV++P+K+P++FTG PW+
Sbjct: 107 EDPEQSKLRAGLNSAIVREKPNIKWSDVAGLESAKQALQEAVILPVKFPQFFTGKRRPWR 166
Query: 159 GILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPS 218
LL+GPPGTGK+ LAKAVATE +TFF++S+S +VSKW G+SEKLV LF++AR APS
Sbjct: 167 AFLLYGPPGTGKSYLAKAVATEADSTFFSVSSSDLVSKWMGESEKLVSNLFEMARESAPS 226
Query: 219 TIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAA 278
IF+DEID++ RGE +E EASRR+KTELL+QM G+ DE V VLAATN P+ LD A
Sbjct: 227 IIFVDEIDSLCGTRGEG-NESEASRRIKTELLVQMQGVGHNDEKVLVLAATNTPYALDQA 285
Query: 279 MLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIP-YDLLVNQTEGYSGSDIRLLCKE 337
+ RR +KRI +PLPE +AR MF+ L P + P ++ L +TEG+SGSD+ + K+
Sbjct: 286 IRRRFDKRIYIPLPEAKARQHMFKVHLGDTPHNLTEPDFEYLGQKTEGFSGSDVSVCVKD 345
Query: 338 VAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKN 379
V +P+R+ + + P+ GP P IQ +++
Sbjct: 346 VLFEPVRKTQDAMFFFKS--PDGTWMPCGPRHPGAIQTTMQD 385
>At3g27120 spastin protein, putative
Length = 327
Score = 235 bits (599), Expect = 4e-62
Identities = 131/301 (43%), Positives = 189/301 (62%), Gaps = 14/301 (4%)
Query: 106 LAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGP 165
L E +S +I+ P+V+W+ I GLE+AK+ + E V+ P+ P F G SP KG+LLFGP
Sbjct: 32 LIEHVSNEIMDRDPNVRWDDIAGLEHAKKCVTEMVIWPLLRPDIFKGCRSPGKGLLLFGP 91
Query: 166 PGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEI 225
PGTGKTM+ KA+A E K TFF ISASS+ SKW G+ EKLV+ LF +A P+ IF+DEI
Sbjct: 92 PGTGKTMIGKAIAGEAKATFFYISASSLTSKWIGEGEKLVRALFGVASCRQPAVIFVDEI 151
Query: 226 DAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEK 285
D+++SQR ++ EHE+SRRLKT+ LI+M+G E + ++ ATN P ELD A RRL K
Sbjct: 152 DSLLSQR-KSDGEHESSRRLKTQFLIEMEGFDSGSEQILLIGATNRPQELDEAARRRLTK 210
Query: 286 RILVPLPEPEARVAMFEELLPPQ-----PDEESIPYDLLVNQTEGYSGSDIRLLCKEVAM 340
R+ +PLP EAR + + LL D++ +++ N TEGYSGSD++ L K+ M
Sbjct: 211 RLYIPLPSSEARAWIIQNLLKKDGLFTLSDDD---MNIICNLTEGYSGSDMKNLVKDATM 267
Query: 341 QPLRRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHAHKYDKFNADYG 399
PLR + + +L ++ + + +D + AL+ RPS + Y+ +N +G
Sbjct: 268 GPLREALKRGIDITNLTKDD----MRLVTLQDFKDALQEVRPSVSQNELGIYENWNNQFG 323
Query: 400 S 400
S
Sbjct: 324 S 324
>At3g19740 ATPase, putative
Length = 439
Score = 218 bits (556), Expect = 4e-57
Identities = 124/296 (41%), Positives = 181/296 (60%), Gaps = 10/296 (3%)
Query: 110 LSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILLFGPPG 167
+S + G VK++ I LE+ K+ L E V++P++ P+ FT LL P KGILLFGPPG
Sbjct: 136 VSAVVAPGEIGVKFDDIGALEHVKKTLNELVILPMRRPELFTRGNLLRPCKGILLFGPPG 195
Query: 168 TGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDA 227
TGKT+LAKA+ATE F +I+ S++ SKW GD+EKL K LF A AP IF+DE+D+
Sbjct: 196 TGKTLLAKALATEAGANFISITGSTLTSKWFGDAEKLTKALFSFASKLAPVIIFVDEVDS 255
Query: 228 IISQRGEARSEHEASRRLKTELLIQMDGLTRTD-ELVFVLAATNLPWELDAAMLRRLEKR 286
++ RG A EHEA+RR++ E + DGL D + + +L ATN P++LD A++RRL +R
Sbjct: 256 LLGARGGA-FEHEATRRMRNEFMAAWDGLRSKDSQRILILGATNRPFDLDDAVIRRLPRR 314
Query: 287 ILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRL 346
I V LP+ E R+ + + L P+ E +D L +TEGYSGSD++ LC A +P++ L
Sbjct: 315 IYVDLPDAENRLKILKIFLTPENLETGFEFDKLAKETEGYSGSDLKNLCIAAAYRPVQEL 374
Query: 347 MSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSAHLHA---HKYDKFNADYG 399
+ E+ +D V P + P+ +D + PS A ++ K+N YG
Sbjct: 375 LQ--EENKDSVTNAS-PDLRPLSLDDFIQSKAKVSPSVAYDATTMNELRKWNEQYG 427
>At1g50140 hypothetical protein, 5' partial
Length = 601
Score = 214 bits (545), Expect = 7e-56
Identities = 122/296 (41%), Positives = 178/296 (59%), Gaps = 10/296 (3%)
Query: 110 LSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILLFGPPG 167
+S + G VK+E I LE+ K+ L E V++P++ P+ F LL P KGILLFGPPG
Sbjct: 298 VSAVVAPGEIGVKFEDIGALEDVKKALNELVILPMRRPELFARGNLLRPCKGILLFGPPG 357
Query: 168 TGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDA 227
TGKT+LAKA+ATE F +I+ S++ SKW GD+EKL K LF A AP IF+DEID+
Sbjct: 358 TGKTLLAKALATEAGANFISITGSTLTSKWFGDAEKLTKALFSFATKLAPVIIFVDEIDS 417
Query: 228 IISQRGEARSEHEASRRLKTELLIQMDGLTRTD-ELVFVLAATNLPWELDAAMLRRLEKR 286
++ RG SEHEA+RR++ E + DGL D + + +L ATN P++LD A++RRL +R
Sbjct: 418 LLGARG-GSSEHEATRRMRNEFMAAWDGLRSKDSQRILILGATNRPFDLDDAVIRRLPRR 476
Query: 287 ILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRL 346
I V LP+ E R+ + + L P+ E ++ L +TEGYSGSD++ LC A +P++ L
Sbjct: 477 IYVDLPDAENRLKILKIFLTPENLESDFQFEKLAKETEGYSGSDLKNLCIAAAYRPVQEL 536
Query: 347 MSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSAHLHA---HKYDKFNADYG 399
L++ + E P + + +D + PS A ++ K+N YG
Sbjct: 537 ---LQEEQKGARAEASPGLRSLSLDDFIQSKAKVSPSVAYDATTMNELRKWNEQYG 589
>At4g02470
Length = 371
Score = 213 bits (543), Expect = 1e-55
Identities = 120/282 (42%), Positives = 171/282 (60%), Gaps = 5/282 (1%)
Query: 108 ESLSRDIIRGSP-DVKWESIKGLENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFG 164
+ L D+I S V ++ I LEN K LKE V++P++ P+ F L P KGILLFG
Sbjct: 52 KKLLSDVIPPSDIGVSFDDIGALENVKETLKELVMLPLQRPELFDKGQLTKPTKGILLFG 111
Query: 165 PPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDE 224
PPGTGKTMLAKAVATE F NIS SS+ SKW G+ EK VK +F LA APS IF+DE
Sbjct: 112 PPGTGKTMLAKAVATEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKIAPSVIFVDE 171
Query: 225 IDAIISQRGEARSEHEASRRLKTELLIQMDGL-TRTDELVFVLAATNLPWELDAAMLRRL 283
+D+++ +R E EHEA R++K E ++ DGL T+ E V VLAATN P++LD A++RRL
Sbjct: 172 VDSMLGRR-ENPGEHEAMRKMKNEFMVNWDGLRTKDRERVLVLAATNRPFDLDEAVIRRL 230
Query: 284 EKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPL 343
+R++V LP+ R + +L + + + + N T+GYSGSD++ LC A P+
Sbjct: 231 PRRLMVNLPDATNRSKILSVILAKEEIAPDVDLEAIANMTDGYSGSDLKNLCVTAAHFPI 290
Query: 344 RRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSAH 385
R ++ + ++ + E P D+++ N +AH
Sbjct: 291 REILEKEKKEKTAAQAENRPTPPLYSCTDVRSLTMNDFKAAH 332
>At1g64110 unknown protein
Length = 824
Score = 209 bits (531), Expect = 3e-54
Identities = 124/304 (40%), Positives = 180/304 (58%), Gaps = 22/304 (7%)
Query: 90 PQKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKY 149
P+ + P E R E + + I +V ++ I L+ K L+E V++P++ P
Sbjct: 486 PKAPEVAPDNEFEKRIRPEVIPAEEI----NVTFKDIGALDEIKESLQELVMLPLRRPDL 541
Query: 150 FTG-LLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVL 208
FTG LL P +GILLFGPPGTGKTMLAKA+A E +F N+S S++ SKW G+ EK V+ L
Sbjct: 542 FTGGLLKPCRGILLFGPPGTGKTMLAKAIAKEAGASFINVSMSTITSKWFGEDEKNVRAL 601
Query: 209 FQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGL-TRTDELVFVLA 267
F LA +P+ IF+DE+D+++ QR EHEA R++K E + DGL T+ E + VLA
Sbjct: 602 FTLASKVSPTIIFVDEVDSMLGQRTRV-GEHEAMRKIKNEFMSHWDGLMTKPGERILVLA 660
Query: 268 ATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYS 327
ATN P++LD A++RR E+RI+V LP E R + LL + +E++ Y L TEGY+
Sbjct: 661 ATNRPFDLDEAIIRRFERRIMVGLPAVENREKILRTLLAKEKVDENLDYKELAMMTEGYT 720
Query: 328 GSDIRLLCKEVAMQPLRRLMSQ--------LEQR-------EDLVPEEELPKVGPIRPED 372
GSD++ LC A +P+R L+ Q +QR ED EE + + P+ +D
Sbjct: 721 GSDLKNLCTTAAYRPVRELIQQERIKDTEKKKQREPTKAGEEDEGKEERVITLRPLNRQD 780
Query: 373 IQAA 376
+ A
Sbjct: 781 FKEA 784
>At4g24850 unknown protein (At4g24850)
Length = 442
Score = 207 bits (526), Expect = 1e-53
Identities = 111/249 (44%), Positives = 160/249 (63%), Gaps = 4/249 (1%)
Query: 110 LSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFGPPG 167
LS I+ DV ++ I LE K +LKE V++P++ P+ F L P KGILLFGPPG
Sbjct: 126 LSDVILPSDIDVTFDDIGALEKVKDILKELVMLPLQRPELFCKGELTKPCKGILLFGPPG 185
Query: 168 TGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDA 227
TGKTMLAKAVA E F NIS SS+ SKW G+ EK VK +F LA +PS IF+DE+D+
Sbjct: 186 TGKTMLAKAVAKEADANFINISMSSITSKWFGEGEKYVKAVFSLASKMSPSVIFVDEVDS 245
Query: 228 IISQRGEARSEHEASRRLKTELLIQMDGLTRTD-ELVFVLAATNLPWELDAAMLRRLEKR 286
++ +R R EHEASR++K E ++ DGLT + E V VLAATN P++LD A++RRL +R
Sbjct: 246 MLGRREHPR-EHEASRKIKNEFMMHWDGLTTQERERVLVLAATNRPFDLDEAVIRRLPRR 304
Query: 287 ILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRL 346
++V LP+ R + + +L + + + + T GYSGSD++ LC A +P++ +
Sbjct: 305 LMVGLPDTSNRAFILKVILAKEDLSPDLDIGEIASMTNGYSGSDLKNLCVTAAHRPIKEI 364
Query: 347 MSQLEQRED 355
+ + ++ D
Sbjct: 365 LEKEKRERD 373
>At1g02890 hypothetical protein
Length = 1240
Score = 200 bits (508), Expect = 1e-51
Identities = 118/258 (45%), Positives = 162/258 (62%), Gaps = 10/258 (3%)
Query: 108 ESLSRDIIRGSP-DVKWESIKGLENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFG 164
+ L D+I S V + I LEN K LKE V++P++ P+ F L P KGILLFG
Sbjct: 925 KKLLSDVIPPSDIGVSFSDIGALENVKDTLKELVMLPLQRPELFGKGQLTKPTKGILLFG 984
Query: 165 PPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDE 224
PPGTGKTMLAKAVATE F NIS SS+ SK EK VK +F LA APS IF+DE
Sbjct: 985 PPGTGKTMLAKAVATEAGANFINISMSSITSK----GEKYVKAVFSLASKIAPSVIFVDE 1040
Query: 225 IDAIISQRGEARSEHEASRRLKTELLIQMDGL-TRTDELVFVLAATNLPWELDAAMLRRL 283
+D+++ +R E EHEA R++K E +I DGL T+ E V VLAATN P++LD A++RRL
Sbjct: 1041 VDSMLGRR-ENPGEHEAMRKMKNEFMINWDGLRTKDKERVLVLAATNRPFDLDEAVIRRL 1099
Query: 284 EKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPL 343
+R++V LP+ R + +L + E + + + N T+GYSGSD++ LC A P+
Sbjct: 1100 PRRLMVNLPDSANRSKILSVILAKEEMAEDVDLEAIANMTDGYSGSDLKNLCVTAAHLPI 1159
Query: 344 RRLMSQLEQREDLVPEEE 361
R ++ + E++E V + E
Sbjct: 1160 REILEK-EKKERSVAQAE 1176
>At4g28000 putative protein
Length = 726
Score = 197 bits (500), Expect = 1e-50
Identities = 103/231 (44%), Positives = 150/231 (64%), Gaps = 4/231 (1%)
Query: 121 VKWESIKGLENAKRLLKEAVVMPIKYPKYFTG-LLSPWKGILLFGPPGTGKTMLAKAVAT 179
V + I L+ K L+E V++P++ P F G LL P +GILLFGPPGTGKTM+AKA+A
Sbjct: 411 VTFADIGSLDETKESLQELVMLPLRRPDLFKGGLLKPCRGILLFGPPGTGKTMMAKAIAN 470
Query: 180 ECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEH 239
E +F N+S S++ SKW G+ EK V+ LF LA +P+ IF+DE+D+++ QR EH
Sbjct: 471 EAGASFINVSMSTITSKWFGEDEKNVRALFTLAAKVSPTIIFVDEVDSMLGQRTRV-GEH 529
Query: 240 EASRRLKTELLIQMDGL-TRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARV 298
EA R++K E + DGL + + + VLAATN P++LD A++RR E+RI+V LP E+R
Sbjct: 530 EAMRKIKNEFMTHWDGLMSNAGDRILVLAATNRPFDLDEAIIRRFERRIMVGLPSVESRE 589
Query: 299 AMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQ 349
+ LL + E++ + L T+GYSGSD++ C A +P+R L+ Q
Sbjct: 590 KILRTLLSKE-KTENLDFQELAQMTDGYSGSDLKNFCTTAAYRPVRELIKQ 639
>At2g45500 hypothetical protein
Length = 481
Score = 189 bits (480), Expect = 2e-48
Identities = 119/268 (44%), Positives = 152/268 (56%), Gaps = 56/268 (20%)
Query: 106 LAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWK-----GI 160
L E ++ I+ SP VKW+ + GL AK+ L E V++P K FTGL P + G+
Sbjct: 198 LVEMINTTIVDRSPSVKWDDVAGLNGAKQALLEMVILPAKRRDLFTGLRRPARVTSLLGL 257
Query: 161 LLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTI 220
LLFGPPG GKTMLAKAVA+E + TFFN+SASS+ SKW
Sbjct: 258 LLFGPPGNGKTMLAKAVASESQATFFNVSASSLTSKW----------------------- 294
Query: 221 FLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLT-RTDELVFVLAATNLPWELDAAM 279
ID+I+S R + SE+EASRRLK+E LIQ DG+T D+LV ++ ATN P ELD A+
Sbjct: 295 ----IDSIMSTR--STSENEASRRLKSEFLIQFDGVTSNPDDLVIIIGATNKPQELDDAV 348
Query: 280 LRRLEKRILVPLPEPEARVAMFEELLPPQPDEESI-PYDLLVNQTEG------------- 325
LRRL KRI VPLP+ R +F+ L QP S D +V +TEG
Sbjct: 349 LRRLVKRIYVPLPDSNVRKLLFKTKLKCQPHSLSDGDIDKIVKETEGKLYRLCIKKHRFI 408
Query: 326 -------YSGSDIRLLCKEVAMQPLRRL 346
YSGSD++ LC+E AM P+R L
Sbjct: 409 SQVTDKRYSGSDLQALCEEAAMMPIREL 436
>At5g53540 26S proteasome regulatory particle chain RPT6-like
protein
Length = 403
Score = 188 bits (478), Expect = 4e-48
Identities = 115/275 (41%), Positives = 164/275 (58%), Gaps = 22/275 (8%)
Query: 120 DVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILLFGPPGTGKTMLAKAV 177
DV++ SI GLE+ K+ L E V++P+K P+ F LL P KG+LL+GPPGTGKTMLAKA+
Sbjct: 83 DVEFGSIGGLESIKQALYELVILPLKRPELFAYGKLLGPQKGVLLYGPPGTGKTMLAKAI 142
Query: 178 ATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARS 237
A E + F N+ S+++SKW GD++KLV +F LA P+ IF+DE+D+ + QR +
Sbjct: 143 ARESEAVFINVKVSNLMSKWFGDAQKLVSAVFSLAYKLQPAIIFIDEVDSFLGQR--RST 200
Query: 238 EHEASRRLKTELLIQMDGLTRTDE--LVFVLAATNLPWELDAAMLRRLEKRILVPLPEPE 295
++EA +KTE + DG T TD+ V VLAATN P ELD A+LRR + + +P+ +
Sbjct: 201 DNEAMSNMKTEFMALWDGFT-TDQNARVMVLAATNRPSELDEAILRRFPQSFEIGMPDCQ 259
Query: 296 ARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRED 355
R + + +L + E I YD + E Y+GSDI LCK+ A P+ RE
Sbjct: 260 ERAQILKVVLKGESVESDINYDRIARLCEDYTGSDIFELCKKAAYFPI---------REI 310
Query: 356 LVPEEELPKVGPIRP------EDIQAALKNTRPSA 384
L E+E +V RP E + A K T+ +A
Sbjct: 311 LEAEKEGKRVSVPRPLTQLDLEKVLATSKKTQVAA 345
>At4g27680 unknown protein
Length = 398
Score = 186 bits (471), Expect = 3e-47
Identities = 102/231 (44%), Positives = 144/231 (62%), Gaps = 5/231 (2%)
Query: 120 DVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILLFGPPGTGKTMLAKAV 177
DV++ SI GLE K+ L E V++P+K P+ F LL P KG+LL+GPPGTGKTMLAKA+
Sbjct: 80 DVEFGSIGGLETIKQALYELVILPLKRPELFAYGKLLGPQKGVLLYGPPGTGKTMLAKAI 139
Query: 178 ATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARS 237
A E F N+ S+++SKW GD++KLV +F LA P+ IF+DE+++ + QR +
Sbjct: 140 AKESGAVFINVRVSNLMSKWFGDAQKLVSAVFSLAYKLQPAIIFIDEVESFLGQR--RST 197
Query: 238 EHEASRRLKTELLIQMDGL-TRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEA 296
+HEA +KTE + DG T V VLAATN P ELD A+LRRL + + +P+
Sbjct: 198 DHEAMANMKTEFMALWDGFSTDPHARVMVLAATNRPSELDEAILRRLPQAFEIGIPDRRE 257
Query: 297 RVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLM 347
R + + L + E I +D + EGY+GSDI LCK+ A P+R ++
Sbjct: 258 RAEILKVTLKGERVEPDIDFDHIARLCEGYTGSDIFELCKKAAYFPIREIL 308
>At5g03340 transitional endoplasmic reticulum ATPase
Length = 810
Score = 183 bits (465), Expect = 1e-46
Identities = 107/303 (35%), Positives = 170/303 (55%), Gaps = 22/303 (7%)
Query: 109 SLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 167
S R+ + P+V WE I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 465 SALRETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 524
Query: 168 TGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDA 227
GKT+LAKA+A EC+ F ++ +++ W G+SE V+ +F AR AP +F DE+D+
Sbjct: 525 CGKTLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDS 584
Query: 228 IISQRGEARSE-HEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLE 284
I +QRG + + A+ R+ +LL +MDG+ + VF++ ATN P +D+A+LR RL+
Sbjct: 585 IATQRGNSAGDAGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDIIDSALLRPGRLD 643
Query: 285 KRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPL- 343
+ I +PLP+ ++R+ +F+ L P + + L T+G+SG+DI +C+ +
Sbjct: 644 QLIYIPLPDEDSRLNIFKACLRKSPVAKDVDVTALAKYTQGFSGADITEICQRACKYAIR 703
Query: 344 -----------RRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHAHKY 391
RR + ED+V +E V IR + ++K R S + KY
Sbjct: 704 ENIEKDIENERRRSQNPEAMEEDMVDDE----VSEIRAAHFEESMKYARRSVSDADIRKY 759
Query: 392 DKF 394
F
Sbjct: 760 QAF 762
Score = 154 bits (390), Expect = 6e-38
Identities = 94/280 (33%), Positives = 156/280 (55%), Gaps = 7/280 (2%)
Query: 108 ESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 166
E + R+ +V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 191 EPVKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 250
Query: 167 GTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEID 226
G+GKT++A+AVA E FF I+ ++SK G+SE ++ F+ A +APS IF+DEID
Sbjct: 251 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 310
Query: 227 AIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLE 284
+I +R + E E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R R +
Sbjct: 311 SIAPKREKTNGEVE--RRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRRFGRFD 367
Query: 285 KRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLR 344
+ I + +P+ R+ + E + + + T GY G+D+ LC E A+Q +R
Sbjct: 368 REIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERISKDTHGYVGADLAALCTEAALQCIR 427
Query: 345 RLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSA 384
M ++ +D + E L + + E AL N+ PSA
Sbjct: 428 EKMDVIDLEDDSIDAEILNSMA-VSNEHFHTALGNSNPSA 466
>At3g53230 CDC48 - like protein
Length = 815
Score = 181 bits (460), Expect = 5e-46
Identities = 105/297 (35%), Positives = 169/297 (56%), Gaps = 12/297 (4%)
Query: 109 SLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 167
S R+ + P+V WE I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 466 SALRETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 525
Query: 168 TGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDA 227
GKT+LAKA+A EC+ F +I +++ W G+SE V+ +F AR AP +F DE+D+
Sbjct: 526 CGKTLLAKAIANECQANFISIKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDS 585
Query: 228 IISQRGEARSE-HEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLE 284
I +QRG + + A+ R+ +LL +MDG+ + VF++ ATN P +D A+LR RL+
Sbjct: 586 IATQRGNSVGDAGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDIIDPALLRPGRLD 644
Query: 285 KRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLR 344
+ I +PLP+ E+R +F+ L P + + L T+G+SG+DI +C+ +R
Sbjct: 645 QLIYIPLPDEESRYQIFKSCLRKSPVAKDVDLRALAKYTQGFSGADITEICQRSCKYAIR 704
Query: 345 RLMSQLEQRE------DLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHAHKYDKF 394
+ + ++E EE+ ++ I+ + ++K R S + KY F
Sbjct: 705 ENIEKDIEKERKRAESPEAMEEDEEEIAEIKAGHFEESMKYARRSVSDADIRKYQAF 761
Score = 152 bits (385), Expect = 2e-37
Identities = 93/280 (33%), Positives = 157/280 (55%), Gaps = 7/280 (2%)
Query: 108 ESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 166
E + R+ +V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 192 EPIKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 251
Query: 167 GTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEID 226
G+GKT++A+AVA E FF I+ ++SK G+SE ++ F+ A +APS IF+DEID
Sbjct: 252 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 311
Query: 227 AIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLE 284
+I +R + E E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R R +
Sbjct: 312 SIAPKREKTHGEVE--RRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRRFGRFD 368
Query: 285 KRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLR 344
+ I + +P+ R+ + E + + + T GY G+D+ LC E A+Q +R
Sbjct: 369 REIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERVSKDTHGYVGADLAALCTEAALQCIR 428
Query: 345 RLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSA 384
M ++ ++ + E L + + + Q AL N+ PSA
Sbjct: 429 EKMDVIDLDDEEIDAEILNSMA-VSNDHFQTALGNSNPSA 467
>At3g09840 putative transitional endoplasmic reticulum ATPase
Length = 809
Score = 180 bits (456), Expect = 1e-45
Identities = 102/299 (34%), Positives = 170/299 (56%), Gaps = 14/299 (4%)
Query: 109 SLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 167
S R+ + P+V W I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 465 SALRETVVEVPNVSWNDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 524
Query: 168 TGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDA 227
GKT+LAKA+A EC+ F ++ +++ W G+SE V+ +F AR AP +F DE+D+
Sbjct: 525 CGKTLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDS 584
Query: 228 IISQR--GEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RL 283
I +QR G A+ R+ +LL +MDG+ + VF++ ATN P +D+A+LR RL
Sbjct: 585 IATQRGGGSGGDGGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDIIDSALLRPGRL 643
Query: 284 EKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPL 343
++ I +PLP+ ++R+ +F+ L P + + L T+G+SG+DI +C+ +
Sbjct: 644 DQLIYIPLPDEDSRLNIFKAALRKSPIAKDVDIGALAKYTQGFSGADITEICQRACKYAI 703
Query: 344 RRLMSQLEQRE-------DLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHAHKYDKF 394
R + + ++E + + E+ + +V I+ + ++K R S + KY F
Sbjct: 704 RENIEKDIEKEKRRSENPEAMEEDGVDEVSEIKAAHFEESMKYARRSVSDADIRKYQAF 762
Score = 156 bits (394), Expect = 2e-38
Identities = 95/280 (33%), Positives = 156/280 (54%), Gaps = 7/280 (2%)
Query: 108 ESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 166
E + R+ DV ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 191 EPVKREDEERLDDVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 250
Query: 167 GTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEID 226
G+GKT++A+AVA E FF I+ ++SK G+SE ++ F+ A +APS IF+DEID
Sbjct: 251 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 310
Query: 227 AIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLE 284
+I +R + E E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R R +
Sbjct: 311 SIAPKREKTNGEVE--RRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRRFGRFD 367
Query: 285 KRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLR 344
+ I + +P+ R+ + E + + + T GY G+D+ LC E A+Q +R
Sbjct: 368 REIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERISKDTHGYVGADLAALCTEAALQCIR 427
Query: 345 RLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSA 384
M ++ +D + E L + + E AL N+ PSA
Sbjct: 428 EKMDVIDLEDDSIDAEILNSMA-VTNEHFHTALGNSNPSA 466
>At3g56690 calmodulin-binding protein
Length = 1022
Score = 163 bits (412), Expect = 2e-40
Identities = 100/257 (38%), Positives = 147/257 (56%), Gaps = 5/257 (1%)
Query: 98 FESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSP 156
FE+A+ + + S R++I P V WE + G K L EAV P K+ F + P
Sbjct: 699 FENAKTK-IRPSAMREVILEVPKVNWEDVGGQNEVKNQLMEAVEWPQKHQDAFKRIGTRP 757
Query: 157 WKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHA 216
GIL+FGPPG KT++A+AVA+E K F + + SKW G+SEK V+ LF AR +A
Sbjct: 758 PSGILMFGPPGCSKTLMARAVASEAKLNFLAVKGPELFSKWVGESEKAVRSLFAKARANA 817
Query: 217 PSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELD 276
PS IF DEID++ S RG+ S R+ ++LL+++DGL + V V+AATN P ++D
Sbjct: 818 PSIIFFDEIDSLASIRGKENDGVSVSDRVMSQLLVELDGLHQRVG-VTVIAATNRPDKID 876
Query: 277 AAMLR--RLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLL 334
+A+LR R ++ + V P R A+ + L P I L + T+GY+G+DI L+
Sbjct: 877 SALLRPGRFDRLLYVGPPNETDREAILKIHLRKIPCSSDICLKELASITKGYTGADISLI 936
Query: 335 CKEVAMQPLRRLMSQLE 351
C+E A+ L + E
Sbjct: 937 CREAAIAALEESLEMEE 953
Score = 125 bits (314), Expect = 4e-29
Identities = 78/213 (36%), Positives = 120/213 (55%), Gaps = 7/213 (3%)
Query: 154 LSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLAR 213
L P KG+L+ GPPGTGKT LA+ A FF+++ ++S++ G+SEK + +F+ A
Sbjct: 415 LRPTKGVLIHGPPGTGKTSLARTFARHSGVNFFSVNGPEIISQYLGESEKALDEVFRSAS 474
Query: 214 HHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPW 273
+ P+ +F+D++DAI R E E S+R+ LL MDG++RTD +V V+AATN P
Sbjct: 475 NATPAVVFIDDLDAIAPARKE--GGEELSQRMVATLLNLMDGISRTDGVV-VIAATNRPD 531
Query: 274 ELDAAMLR--RLEKRILVPLPEPEARVAMFEELLPPQPDE-ESIPYDLLVNQTEGYSGSD 330
++ A+ R RL++ I + +P R + +L +I + L T G+ G+D
Sbjct: 532 SIEPALRRPGRLDREIEIGVPSSTQRSDILHIILRGMRHSLSNIQVEQLAMATHGFVGAD 591
Query: 331 IRLLCKEVAMQPLRRLMSQLEQREDLVPEEELP 363
+ LC E A LRR + Q +L P EE P
Sbjct: 592 LSALCCEAAFVCLRRHLDQSSSSSNL-PLEEAP 623
>At5g43010 26S proteasome AAA-ATPase subunit RPT4a (gb|AAF22524.1)
Length = 399
Score = 154 bits (389), Expect = 8e-38
Identities = 92/286 (32%), Positives = 161/286 (56%), Gaps = 25/286 (8%)
Query: 120 DVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAVA 178
++ + ++ GL + R L+E++ +P+ P+ F + + P KG+LL+GPPGTGKT+LA+A+A
Sbjct: 135 NISYSAVGGLGDQIRELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLARAIA 194
Query: 179 TECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQR-GEARS 237
+ F + +S+++ K+ G+S +L++ +F AR H P IF+DEIDAI +R E S
Sbjct: 195 SNIDANFLKVVSSAIIDKYIGESARLIREMFNYAREHQPCIIFMDEIDAIGGRRFSEGTS 254
Query: 238 EHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVPLPEPE 295
+R ELL Q+DG ++ ++ ATN P LD A+LR RL+++I +PLP +
Sbjct: 255 ADREIQRTLMELLNQLDGFDNLGKVKMIM-ATNRPDVLDPALLRPGRLDRKIEIPLPNEQ 313
Query: 296 ARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRED 355
+R+ + + I Y+ +V EG++G+D+R +C E M +R +R+
Sbjct: 314 SRMDILKIHAAGIAKHGEIDYEAIVKLAEGFNGADLRNICTEAGMFAIR------AERDY 367
Query: 356 LVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQ 401
++ E+ + V + + K S+H +NAD+G +
Sbjct: 368 VIHEDFMKAVRKL------SEAKKLESSSH--------YNADFGKE 399
>At1g45000 26S proteasome AAA-ATPase subunit RPT4a like protein
Length = 399
Score = 154 bits (389), Expect = 8e-38
Identities = 92/286 (32%), Positives = 162/286 (56%), Gaps = 25/286 (8%)
Query: 120 DVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAVA 178
++ + ++ GL + R L+E++ +P+ P+ F + + P KG+LL+GPPGTGKT+LA+A+A
Sbjct: 135 NISYSAVGGLGDQIRELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLARAIA 194
Query: 179 TECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQR-GEARS 237
+ F + +S+++ K+ G+S +L++ +F AR H P IF+DEIDAI +R E S
Sbjct: 195 SNIDANFLKVVSSAIIDKYIGESARLIREMFNYAREHQPCIIFMDEIDAIGGRRFSEGTS 254
Query: 238 EHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVPLPEPE 295
+R ELL Q+DG + ++ ++ ATN P LD A+LR RL+++I +PLP +
Sbjct: 255 ADREIQRTLMELLNQLDGFDQLGKVKMIM-ATNRPDVLDPALLRPGRLDRKIEIPLPNEQ 313
Query: 296 ARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRED 355
+R+ + + I Y+ +V EG++G+D+R +C E M +R +R+
Sbjct: 314 SRMEILKIHASGIAKHGEIDYEAIVKLGEGFNGADLRNICTEAGMFAIR------AERDY 367
Query: 356 LVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQ 401
++ E+ + V + + K S+H +NAD+G +
Sbjct: 368 VIHEDFMKAVRKL------SEAKKLESSSH--------YNADFGKE 399
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.133 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,264,197
Number of Sequences: 26719
Number of extensions: 399097
Number of successful extensions: 1688
Number of sequences better than 10.0: 161
Number of HSP's better than 10.0 without gapping: 125
Number of HSP's successfully gapped in prelim test: 36
Number of HSP's that attempted gapping in prelim test: 1371
Number of HSP's gapped (non-prelim): 189
length of query: 404
length of database: 11,318,596
effective HSP length: 102
effective length of query: 302
effective length of database: 8,593,258
effective search space: 2595163916
effective search space used: 2595163916
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0265.5


