

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.11
(1543 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
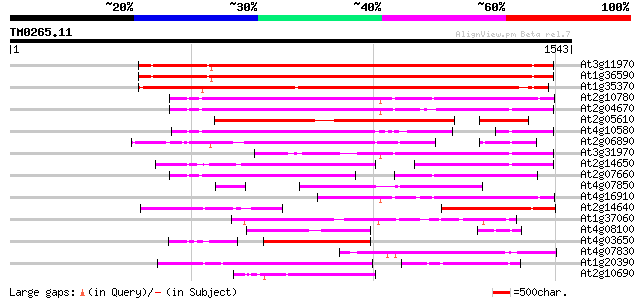
Score E
Sequences producing significant alignments: (bits) Value
At3g11970 hypothetical protein 1008 0.0
At1g36590 hypothetical protein 1004 0.0
At1g35370 hypothetical protein 931 0.0
At2g10780 pseudogene 694 0.0
At2g04670 putative retroelement pol polyprotein 632 0.0
At2g05610 putative retroelement pol polyprotein 577 e-164
At4g10580 putative reverse-transcriptase -like protein 498 e-141
At2g06890 putative retroelement integrase 449 e-126
At3g31970 hypothetical protein 444 e-124
At2g14650 putative retroelement pol polyprotein 425 e-118
At2g07660 putative retroelement pol polyprotein 392 e-109
At4g07850 putative polyprotein 385 e-106
At4g16910 retrotransposon like protein 352 7e-97
At2g14640 putative retroelement pol polyprotein 310 4e-84
At1g37060 Athila retroelment ORF 1, putative 304 3e-82
At4g08100 putative polyprotein 292 9e-79
At4g03650 putative reverse transcriptase 277 4e-74
At4g07830 putative reverse transcriptase 266 7e-71
At1g20390 hypothetical protein 224 2e-58
At2g10690 putative retroelement pol polyprotein 202 1e-51
>At3g11970 hypothetical protein
Length = 1499
Score = 1008 bits (2607), Expect = 0.0
Identities = 506/1152 (43%), Positives = 730/1152 (62%), Gaps = 17/1152 (1%)
Query: 354 MDEDGEIVMLEEEAEEIEEEETVECKLMGVLGCMGEHQTQTMKLAGQVANVELLILIDSG 413
MD D E EE ++E + + V G G +TM++ G + ILIDSG
Sbjct: 352 MDVDEEFEDAREELVNDDDEHMPQISVNAVSGIAGY---KTMRVKGTYDKKIIFILIDSG 408
Query: 414 ASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLE 473
++HNF+ P + LG + + + DG K+ G KL + D L++
Sbjct: 409 STHNFLDPNTAAKLGCKVDTAGLTRVSVADGRKLRVEGKVTDFSWKLQTTTFQSDILLIP 468
Query: 474 LGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTH 533
L G+D+VLGV WL TLG++ ++K L M+F + Q V L G+ +
Sbjct: 469 LQGIDMVLGVQWLETLGRISWEFKKLEMRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQE 528
Query: 534 GREQMEWWGPQLQVVEATKKG--------HEIPQE--LRAVLADFHEVFTKKIQLPPVRS 583
+ Q+ Q +V E+T+ E+ +E + VL ++ ++F + LPP R
Sbjct: 529 DQVQLAMLCVQ-EVSESTEGELCTINALTSELGEESVVEEVLNEYPDIFIEPTALPPFRE 587
Query: 584 RV-HQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVKKK 642
+ H+I L P+N RPYRY HQK EI++ V +LL G ++ S S ++SPV+LVKKK
Sbjct: 588 KHNHKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSPYASPVVLVKKK 647
Query: 643 DKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIP 702
D +WR+CVDYR LN T+ D +PIP++++L+DEL GAVIFSKIDL++GYHQ+R+ +DI
Sbjct: 648 DGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQ 707
Query: 703 KTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEH 762
KT F+TH+GH+EYLVMPFGL NAPATFQ +MN IF+P+LRKFVLVFFDDIL+YS SL EH
Sbjct: 708 KTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEH 767
Query: 763 RDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKN 822
R HLK V V+ +N A SKC F +++YLGH IS G+ DP K++ + EWP+P
Sbjct: 768 RQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTT 827
Query: 823 VKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMTSSPVL 882
+K +RGFLGL GYYR+F++++G +A PL LTK D F W A+QAF LK + +PVL
Sbjct: 828 LKQLRGFLGLAGYYRRFVRSFGVIAGPLHALTKTDAFEWTAVAQQAFEDLKAALCQAPVL 887
Query: 883 ALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQ 942
+LP FDK F VE DA G+GIGAVLMQ+ P+AY S+ L L S+YEKEL+A++ ++
Sbjct: 888 SLPLFDKQFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVR 947
Query: 943 HWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADA 1002
WRHYLL F + TD +SLK+ L+Q++++P QQ WL KLL + +E++Y+ G+EN ADA
Sbjct: 948 KWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQYRQGKENVVADA 1007
Query: 1003 LSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLL 1062
LSR ++ D K + A+D +++++ +Q+DP SK FS Q +L
Sbjct: 1008 LSRVEGSEVLHMAMTVVECDLLKDIQAGYANDSQLQDIITALQRDPDSKKYFSWSQNILR 1067
Query: 1063 YQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACD 1122
+ ++V+ +L HGS GGHSG T++R+ YW GM + ++ ++R+C
Sbjct: 1068 RKSKIVVPANDNIKNTILLWLHGSGVGGHSGRDVTHQRVKGLFYWKGMIKDIQAYIRSCG 1127
Query: 1123 TCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFI 1182
TCQ+ K + GLLQP P+P+ +W ++S+DFI GLP S G I+VVVDRLSK +HFI
Sbjct: 1128 TCQQCKSDPAASPGLLQPLPIPDTIWSEVSMDFIEGLPVSGGKTVIMVVVDRLSKAAHFI 1187
Query: 1183 LLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYH 1242
L HPY+A ++A ++ + +LHG P ++ SDRD +F S FW E F LQG LK++S+YH
Sbjct: 1188 ALSHPYSALTVAHAYLDNVFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGVALKLTSAYH 1247
Query: 1243 PETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRK 1302
P++DGQTEV+NRCLE+YLRC D P+ WS W+ AE+WYN+ +H S TPFE+VYG+
Sbjct: 1248 PQSDGQTEVVNRCLETYLRCMCHDRPQLWSKWLALAEYWYNTNYHSSSRMTPFEIVYGQV 1307
Query: 1303 APPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEW 1362
P + +L E+KV VA L ER++ L L+ HL RAQ +M +A++ R + F+IG++
Sbjct: 1308 PPVHLPYLPGESKVAVVARSLQEREDMLLFLKFHLMRAQHRMKQFADQHRTEREFEIGDY 1367
Query: 1363 VFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLK 1422
V++KL+P+RQ SVV R NQKL+ +++GP++I D+ G VAYKL LP S++HPVFHVS LK
Sbjct: 1368 VYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAYKLALPSYSQVHPVFHVSQLK 1427
Query: 1423 RAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWE 1482
+GN LP+ + ++ + PE ++ +++ + V + L+KW N +++ TWE
Sbjct: 1428 VLVGNVSTTVHLPSVM--QDVFEKVPEKVVERKMVNRQGKAVTKVLVKWSNEPLEEATWE 1485
Query: 1483 DNEVLAGQFPEF 1494
L FPEF
Sbjct: 1486 FLFDLQKTFPEF 1497
>At1g36590 hypothetical protein
Length = 1499
Score = 1004 bits (2595), Expect = 0.0
Identities = 505/1152 (43%), Positives = 730/1152 (62%), Gaps = 17/1152 (1%)
Query: 354 MDEDGEIVMLEEEAEEIEEEETVECKLMGVLGCMGEHQTQTMKLAGQVANVELLILIDSG 413
MD D E EE ++E + + V G G +TM++ G + ILIDSG
Sbjct: 352 MDVDEEFEDAREELVNDDDEHMPQISVNAVSGIAGY---KTMRVKGTYDKKIIFILIDSG 408
Query: 414 ASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLE 473
++HNF+ P + LG + + + DG K+ G KL + D L++
Sbjct: 409 STHNFLDPNTAAKLGCKVDTAGLTRVSVADGRKLRVEGKVTDFSWKLQTTTFQSDILLIP 468
Query: 474 LGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTH 533
L G+D+VLGV WL TLG++ ++K L M+F + Q V L G+ +
Sbjct: 469 LQGIDMVLGVQWLETLGRISWEFKKLEMRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQE 528
Query: 534 GREQMEWWGPQLQVVEATKKG--------HEIPQE--LRAVLADFHEVFTKKIQLPPVRS 583
+ Q+ Q +V E+T+ E+ +E + VL ++ ++F + LPP R
Sbjct: 529 DQVQLAMLCVQ-EVSESTEGELCTINALTSELGEESVVEEVLNEYPDIFIEPTALPPFRE 587
Query: 584 RV-HQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVKKK 642
+ H+I L P+N RPYRY HQK EI++ V +LL G ++ S S ++SPV+LVKKK
Sbjct: 588 KHNHKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSPYASPVVLVKKK 647
Query: 643 DKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIP 702
D +WR+CVDYR LN T+ D +PIP++++L+DEL GAVIFSKIDL++GYHQ+R+ +DI
Sbjct: 648 DGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQ 707
Query: 703 KTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEH 762
KT F+TH+GH+EYLVMPFGL NAPATFQ +MN IF+P+LRKFVLVFFDDIL+YS SL EH
Sbjct: 708 KTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEH 767
Query: 763 RDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKN 822
R HLK V V+ +N A SKC F +++YLGH IS G+ DP K++ + EWP+P
Sbjct: 768 RQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTT 827
Query: 823 VKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMTSSPVL 882
+K +RGFLGL GYYR+F++++G +A PL LTK D F W A+QAF LK + +PVL
Sbjct: 828 LKQLRGFLGLAGYYRRFVRSFGVIAGPLHALTKTDAFEWTAVAQQAFEDLKAALCQAPVL 887
Query: 883 ALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQ 942
+LP FDK F VE DA G+GIGAVLMQ+ P+AY S+ L L S+YEKEL+A++ ++
Sbjct: 888 SLPLFDKQFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVR 947
Query: 943 HWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADA 1002
WRHYLL F + TD +SLK+ L+Q++++P QQ WL KLL + +E++Y+ G+EN ADA
Sbjct: 948 KWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQYRQGKENVVADA 1007
Query: 1003 LSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLL 1062
LSR ++ D K + A+D +++++ +Q+DP SK FS Q +L
Sbjct: 1008 LSRVEGSEVLHMAMTVVECDLLKDIQAGYANDSQLQDIITALQRDPDSKKYFSWSQNILR 1067
Query: 1063 YQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACD 1122
+ ++V+ +L HGS GGHSG T++R+ Y GM + ++ ++R+C
Sbjct: 1068 RKSKIVVPANDNIKNTILLWLHGSGVGGHSGRDVTHQRVKGLFYSKGMIKDIQAYIRSCG 1127
Query: 1123 TCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFI 1182
TCQ+ K + GLLQP P+P+ +W ++S+DFI GLP S G I+VVVDRLSK +HFI
Sbjct: 1128 TCQQCKSDPAASPGLLQPLPIPDTIWSEVSMDFIEGLPVSGGKTVIMVVVDRLSKAAHFI 1187
Query: 1183 LLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYH 1242
L HPY+A ++A+ ++ + +LHG P ++ SDRD +F S FW E F LQG LK++S+YH
Sbjct: 1188 ALSHPYSALTVAQAYLDNVFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGVALKLTSAYH 1247
Query: 1243 PETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRK 1302
P++DGQTEV+NRCLE+YLRC D P+ WS W+ AE+WYN+ +H S TPFE+VYG+
Sbjct: 1248 PQSDGQTEVVNRCLETYLRCMCHDRPQLWSKWLALAEYWYNTNYHSSSRMTPFEIVYGQV 1307
Query: 1303 APPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEW 1362
P + +L E+KV VA L ER++ L L+ HL RAQ +M +A++ R + F+IG++
Sbjct: 1308 PPVHLPYLPGESKVAVVARSLQEREDMLLFLKFHLMRAQHRMKQFADQHRTEREFEIGDY 1367
Query: 1363 VFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLK 1422
V++KL+P+RQ SVV R NQKL+ +++GP++I D+ G VAYKL LP S++HPVFHVS LK
Sbjct: 1368 VYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAYKLALPSYSQVHPVFHVSQLK 1427
Query: 1423 RAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWE 1482
+GN LP+ + ++ + PE ++ +++ + V + L+KW N +++ TWE
Sbjct: 1428 VLVGNVSTTVHLPSVM--QDVFEKVPEKVVERKMVNRQGKAVTKVLVKWSNEPLEEATWE 1485
Query: 1483 DNEVLAGQFPEF 1494
L FPEF
Sbjct: 1486 FLFDLQKTFPEF 1497
>At1g35370 hypothetical protein
Length = 1447
Score = 931 bits (2405), Expect = 0.0
Identities = 486/1142 (42%), Positives = 701/1142 (60%), Gaps = 50/1142 (4%)
Query: 354 MDEDGEIVMLEEEAEEIEEEETVECKLM---GVLGCMGEHQTQTMKLAGQVANVELLILI 410
MD D E E+A E+ ++ E K M V G +TM + G V +L ILI
Sbjct: 329 MDVDEEF----EDAVEVLSDDDHEQKPMPQISVNAVSGISGYKTMGVKGTVDKRDLFILI 384
Query: 411 DSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDAL 470
DSG++HNFI + + LG + + + DG K+ G +G KL S D L
Sbjct: 385 DSGSTHNFIDSTVAAKLGCHVESAGLTKVAVADGRKLNVDGQIKGFTWKLQSTTFQSDIL 444
Query: 471 VLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSF-- 528
++ L G+D+VLGV WL TLG++ ++K L MQF Y Q V L G+ H
Sbjct: 445 LIPLQGVDMVLGVQWLETLGRISWEFKKLEMQFFYKNQRVWLHGIITGSVRDIKAHKLQK 504
Query: 529 -------LSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLADFHEVFTKKIQLPPV 581
L+ RE + ++ + A ++ ++ +F +VF + LPP
Sbjct: 505 TQADQIQLAMVCVREVVSDEEQEIGSISALTSDVVEESVVQNIVEEFPDVFAEPTDLPPF 564
Query: 582 RSRV-HQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVK 640
R + H+I L P+N RPYRY HQK+EI++ V +++++G I+ S S F+SPV+LVK
Sbjct: 565 REKHDHKIKLLEGANPVNQRPYRYVVHQKDEIDKIVQDMIKSGTIQVSSSPFASPVVLVK 624
Query: 641 KKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEED 700
KKD +WR+CVDY LN T+ D++ IP++++L+DEL G+V+FSKIDL++GYHQ+R+ +D
Sbjct: 625 KKDGTWRLCVDYTELNGMTVKDRFLIPLIEDLMDELGGSVVFSKIDLRAGYHQVRMDPDD 684
Query: 701 IPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLL 760
I KT F+THNGH+EYLVM FGL NAPATFQ++MN +FR +LRKFVLVFFDDILIYS S+
Sbjct: 685 IQKTAFKTHNGHFEYLVMLFGLTNAPATFQSLMNSVFRDFLRKFVLVFFDDILIYSSSIE 744
Query: 761 EHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEP 820
EH++HL+LV V+ + A SK ++LGH IS + DP K+Q + EWP P
Sbjct: 745 EHKEHLRLVFEVMRLHKLFAKGSK--------EHLGHFISAREIETDPAKIQAVKEWPTP 796
Query: 821 KNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMTSSP 880
VK VRGFLG GYYR+F++N+G +A PL LTK D F W EA+ AF LK V+ ++P
Sbjct: 797 TTVKQVRGFLGFAGYYRRFVRNFGVIAGPLHALTKTDGFCWSLEAQSAFDTLKAVLCNAP 856
Query: 881 VLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLC 940
VLALP FDK F VE DA G+GI AVLMQ+ P+AY S+ L L S+YEKEL+A +
Sbjct: 857 VLALPVFDKQFMVETDACGQGIRAVLMQKGHPLAYISRQLKGKQLHLSIYEKELLAFIFA 916
Query: 941 IQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAA 1000
++ WRHYLL F + TD +SLK+ L+Q++++P QQ WL KLL + +E++Y+ G+EN A
Sbjct: 917 VRKWRHYLLPSHFIIKTDQRSLKYLLEQRLNTPVQQQWLPKLLEFDYEIQYRQGKENLVA 976
Query: 1001 DALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGV 1060
DALSR +S D K++ SD +K+++ +QQ P +K +S Q +
Sbjct: 977 DALSRVEGSEVLHMALSIVECDFLKEIQVAYESDGVLKDIISALQQHPDAKKHYSWSQDI 1036
Query: 1061 LLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRA 1120
L + ++V+ LLQ H S GG SG +++R+ YW GM + ++ F+R+
Sbjct: 1037 LRRKSKIVVPNDVEITNKLLQWLHCSGMGGRSGRDASHQRVKSLFYWKGMVKDIQAFIRS 1096
Query: 1121 CDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSH 1180
C TCQ+ K + GLLQP P+P+ +W D+S+DFI GLP S G I+VVVDRLSK +H
Sbjct: 1097 CGTCQQCKSDNAAYPGLLQPLPIPDKIWCDVSMDFIEGLPNSGGKSVIMVVVDRLSKAAH 1156
Query: 1181 FILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSS 1240
F+ L HPY+A ++A+ F+ + + HG P ++ SDRD +F S FW E FKLQG +L+MSS+
Sbjct: 1157 FVALAHPYSALTVAQAFLDNVYKHHGCPTSIVSDRDVLFTSDFWKEFFKLQGVELRMSSA 1216
Query: 1241 YHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYG 1300
YHP++DGQTEV+NRCLE+YLRC P W+ W+P AE+WYN+ +H S TPFE+VYG
Sbjct: 1217 YHPQSDGQTEVVNRCLENYLRCMCHARPHLWNKWLPLAEYWYNTNYHSSSQMTPFELVYG 1276
Query: 1301 RKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIG 1360
+ P + +L ++KV VA L ER+ L L+ HL RAQ +M +A++ R + +FDIG
Sbjct: 1277 QAPPIHLPYLPGKSKVAVVARSLQERENMLLFLKFHLMRAQHRMKQFADQHRTERTFDIG 1336
Query: 1361 EWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSL 1420
++V++KL+P+RQ SVV R+NQKL+ +++GP++I +K G V
Sbjct: 1337 DFVYVKLQPYRQQSVVLRVNQKLSPKYFGPYKIIEKCGEV-------------------- 1376
Query: 1421 LKRAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVT 1480
+GN QLP+ L + + PE IL +++++ L+KW +++ T
Sbjct: 1377 ---MVGNVTTSTQLPSVL--PDIFEKAPEYILERKLVKRQGRAATMVLVKWIGEPVEEAT 1431
Query: 1481 WE 1482
W+
Sbjct: 1432 WK 1433
>At2g10780 pseudogene
Length = 1611
Score = 694 bits (1791), Expect = 0.0
Identities = 404/1095 (36%), Positives = 611/1095 (54%), Gaps = 53/1095 (4%)
Query: 439 IKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKL 498
++ G + G+ RGI + + + D +++ L D++LG+ WL K +D
Sbjct: 507 VRAAGGQAMYPTGLVRGISVVVNGVNMPADLIIVPLKKHDVILGMDWLGKY-KAHIDCHR 565
Query: 499 LTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIP 558
+QF E +++ QG+R S + E+M G + + T K
Sbjct: 566 GRIQFERDEGMLKFQGIRTTSG------SLVISAIQAERMLGKGCEAYLATITTKEVGAS 619
Query: 559 QELR--AVLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQV 616
EL+ ++ +F +VF +PP RS I L P PI+ PYR + ++++Q+
Sbjct: 620 AELKDIPIVNEFSDVFAAVSGVPPDRSDPFTIELEPGTTPISKAPYRMAPAEMAKLKKQL 679
Query: 617 TELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDEL 676
ELL+ G IRPS S + +PV+ VKKKD S+R+C+DYR LNK T+ +KYP+P +DEL+D+L
Sbjct: 680 EELLDKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQL 739
Query: 677 NGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDI 736
GA FSKIDL SGYHQI + D+ KT FRT H+E++VMPFGL NAPA F +MN +
Sbjct: 740 GGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYDHFEFVVMPFGLTNAPAAFMKMMNGV 799
Query: 737 FRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLG 796
FR +L +FV++F +DIL+YSKS H++HL+ VL L + A SKC F + +LG
Sbjct: 800 FRDFLDEFVIIFINDILVYSKSWEAHQEHLRAVLERLREHELFAKLSKCSFWQRSVGFLG 859
Query: 797 HIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKK 856
H+IS GVSVDPEK++ I EWP P+N +R FLGL GYYR+F+ ++ MA+PLT LT K
Sbjct: 860 HVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQPLTRLTGK 919
Query: 857 DN-FTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAY 915
D F W +E +++F +LK ++T++PVL LP+ + + V DA+ G+G VLMQ+ IAY
Sbjct: 920 DTAFNWSDECEKSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGSVIAY 979
Query: 916 FSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQ 975
S+ L ++ E+ A+V ++ WR YL G +YTDHKSLK+ Q + Q
Sbjct: 980 ASRQLRKHEKNYPTHDLEMAAVVFFLKIWRSYLYGAKVQIYTDHKSLKYIFTQPELNLRQ 1039
Query: 976 QCWLAKLLGYQFEVKYKPGQENKAADALSRRGDESDF----FSLVSF------------- 1018
+ W+ + Y ++ Y PG+ N+ ADALSRR E + LV+
Sbjct: 1040 RRWMELVADYNLDIAYHPGKANQVADALSRRRSEVEAERSQVDLVNMMGTLHVNALSKEV 1099
Query: 1019 -PLW---DDRKKLLEEIA----SDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLS 1070
PL D+ LL I D IK Q+ + + Q+ G ++ GR+ +
Sbjct: 1100 EPLGLGAADQADLLSRIRLAQERDEEIKGWAQNNKTEYQTSNN-----GTIVVNGRVCVP 1154
Query: 1071 PTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYS 1130
+L+E H S H G + YR L +WVGM++ V +V C TCQ K
Sbjct: 1155 NDRALKEEILREAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWVAKCPTCQLVKAE 1214
Query: 1131 AMSPGGLLQPFPVPNAVWEDLSLDFIIGLP---KSKGYEAILVVVDRLSKYSHFILLKHP 1187
P GLLQ P+P W+ +++DF+ GLP KSK + A+ VVVDRL+K +HF+ +
Sbjct: 1215 HQVPSGLLQNLPIPEWKWDHITMDFVTGLPTGIKSK-HNAVWVVVDRLTKSAHFMAISDK 1273
Query: 1188 YTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDG 1247
A+ IAE ++ EIVRLHGIP ++ SDRD F S FW K GT++ +S++YHP+TD
Sbjct: 1274 DGAEIIAEKYIDEIVRLHGIPVSIVSDRDTRFTSKFWKAFQKALGTRVNLSTAYHPQTDE 1333
Query: 1248 QTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKA-PPI 1306
Q+E + LE LR D W ++ EF YN++F SIG +P+E +YGR P+
Sbjct: 1334 QSERTIQTLEDMLRACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPYEALYGRACRTPL 1393
Query: 1307 IRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLK 1366
E ++ + + E E +K L+ L+ AQ++ SYANK+R++L F +G+ V+LK
Sbjct: 1394 CWTPVGERRLFGPTI-VDETTERMKFLKIKLKEAQDRQKSYANKRRKELEFQVGDLVYLK 1452
Query: 1367 LRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVE-SRIHPVFHVSLLKRAI 1425
++ +KL+ R+ GP+++ +++G VAYKL LP + + H VFHVS L++ +
Sbjct: 1453 AMTYKGAGRFTS-RKKLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCL 1511
Query: 1426 GNYQVQGQ-LPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIK--WKNRSIDDVTWE 1482
+ + + +P L + +P I+ + +G + L+K W R ++ TWE
Sbjct: 1512 SDQEESVEDIPPGLKENMTVEAWPVRIMDR--MTKGTRGKARDLLKVLWNCRGREEYTWE 1569
Query: 1483 DNEVLAGQFPEFCLE 1497
+ FPE+ E
Sbjct: 1570 TENKMKANFPEWFKE 1584
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 632 bits (1630), Expect = 0.0
Identities = 382/1083 (35%), Positives = 577/1083 (53%), Gaps = 70/1083 (6%)
Query: 439 IKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKL 498
+K+ G + G +G+ ++ + D ++ + D++LG+ WL +V +D
Sbjct: 365 VKVAGGQFLAVLGRAKGVDIQIAGESMPADLIISPVELYDVILGMDWLDHY-RVHLDCHR 423
Query: 499 LTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKK---GH 555
+ F E + QG+R S + ++M G + +V + G
Sbjct: 424 GRVSFERPEGRLVYQGVRPTSG------SLVISAVQAKKMIEKGCEAYLVTISMPESLGQ 477
Query: 556 EIPQELRAVLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQ 615
++R V+ +F +VF LPP RS I L P P++ PYR + E+++Q
Sbjct: 478 VAVSDIR-VIQEFEDVFQSLQGLPPSRSDPFTIELEPGTAPLSKAPYRMAPAEMTELKKQ 536
Query: 616 VTELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDE 675
+ +LL G IRPS S + +PV+ VKKKD S+R+C+DYR LN T+ +KYP+P +DELLD+
Sbjct: 537 LEDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQ 596
Query: 676 LNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMND 735
L GA FSKIDL SGYHQI + E D+ KT FRT GH+E++VMPF L NAPA F +MN
Sbjct: 597 LRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRYGHFEFVVMPFALTNAPAAFMRLMNS 656
Query: 736 IFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYL 795
+F+ +L +FV++F DDIL+YSKS EH HL+ V+ L A SKC F +I +L
Sbjct: 657 VFQEFLDEFVIIFIDDILVYSKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREIGFL 716
Query: 796 GHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTK 855
GHI+S GVSVDPEK++ I +WP P N +R FL LTGYYR+F+K + MA+P+T+LT
Sbjct: 717 GHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFLRLTGYYRRFVKGFASMAQPMTKLTG 776
Query: 856 KD-NFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIA 914
KD F W E ++ F LK ++TS+PVLALP+ + + V DA+ G+G VLMQ+ + IA
Sbjct: 777 KDVPFVWSPECEEGFVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQRGKVIA 836
Query: 915 YFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPD 974
Y S+ L ++ E+ ++ ++ WR YL G V+TDHKSLK+ Q +
Sbjct: 837 YASRQLRKHEGNYPTHDLEMAVVIFALKIWRSYLYGGKVQVFTDHKSLKYIFNQPELNLR 896
Query: 975 QQCWLAKLLGYQFEVKYKPGQENKAADALSRR----GDESDFFSLVS------------F 1018
Q W+ + Y E+ Y PG+ N ADALSR+ +LVS
Sbjct: 897 QMRWMELVADYDLEIAYHPGKANVVADALSRKRVGAAPGQSVEALVSEIGALRLCVVARE 956
Query: 1019 PLW---DDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPS 1075
PL DR LL L + S+ F+ G + GR+ +
Sbjct: 957 PLGLEAVDRADLLTRARLAQEKDEGLIAASKAEGSEYQFAA-NGTIFVYGRVCVPKDEEL 1015
Query: 1076 IPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPG 1135
+L E H S H G + YR L WVGM+R V ++V CD CQ K
Sbjct: 1016 RREILSEAHASMFSIHPGATKMYRDLKRYYQWVGMKRDVANWVAECDVCQLVK------- 1068
Query: 1136 GLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAE 1195
VP+A+W V++DRL+K +HF+ ++ A +A+
Sbjct: 1069 ---AEHQVPDAIW---------------------VIMDRLTKSAHFLAIRKTDGAAVLAK 1104
Query: 1196 VFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRC 1255
+V EIV+LHG+P ++ SDRD F FW GTK++MS++YHP+TDGQ+E +
Sbjct: 1105 KYVSEIVKLHGVPVSIVSDRDSKFTFAFWRAFQAKMGTKVQMSTAYHPQTDGQSERTIQT 1164
Query: 1256 LESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETK 1315
LE LR D W+ + EF YN+++ SIG PFE +YGR +R+ E +
Sbjct: 1165 LEDMLRMCVLDWGGHWADHLSLVEFAYNNSYQASIGMAPFEALYGRPCWTPLRWTQVEER 1224
Query: 1316 VVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHR--QH 1373
+ A + E E ++ L+ +++ AQ + SYA+K+RR+L F++G+ V+LK+ R
Sbjct: 1225 SIYGADYVQETTERIRVLKLNMKEAQARQRSYADKRRRELEFEVGDRVYLKMAMLRGPNR 1284
Query: 1374 SVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLP-VESRIHPVFHVSLLKRAI-GNYQVQ 1431
S+ + KL+ R+ GPF+I +++G VAY+L+LP V H VFHV +L++ + + +V
Sbjct: 1285 SI---LETKLSPRYMGPFRIVERVGPVAYRLELPDVMRAFHKVFHVLMLRKCLHKDDEVL 1341
Query: 1432 GQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQF 1491
++P DL + P +L RI ++ + W + + TWE + +F
Sbjct: 1342 VKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLWDCDGVTEETWEPEARMKARF 1401
Query: 1492 PEF 1494
++
Sbjct: 1402 KKW 1404
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 577 bits (1488), Expect = e-164
Identities = 287/660 (43%), Positives = 414/660 (62%), Gaps = 52/660 (7%)
Query: 564 VLADFHEVFTKKIQLPPVRS-RVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEA 622
V+ +F ++F + LPP R+ H+I L + P+N RPYRY HQK EI++ V ++L +
Sbjct: 10 VVTEFPDIFVEPTDLPPFRAPHDHKIELLEDSNPVNQRPYRYVVHQKNEIDKIVDDMLAS 69
Query: 623 GIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIF 682
G I+ S S ++SPV+LVKKKD +WR+CVDYR LN T+ D++PIP++++L+DEL G+ ++
Sbjct: 70 GTIQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDRFPIPLIEDLMDELGGSNVY 129
Query: 683 SKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLR 742
SKIDL++GYHQ+R+ DI KT F+THNGHYEYLVMPFGL NAPA+FQ++MN F+P+LR
Sbjct: 130 SKIDLRAGYHQVRMDPLDIHKTAFKTHNGHYEYLVMPFGLTNAPASFQSLMNSFFKPFLR 189
Query: 743 KFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGA 802
KFVLVFFDDILIYS S+ EH+ HL+ V V+ + A SKC F +++YLGH ISG
Sbjct: 190 KFVLVFFDDILIYSTSMEEHKKHLEAVFEVMRVHHLFAKMSKCAFAVPRVEYLGHFISGE 249
Query: 803 GVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWG 862
G++ DP K++ + +WP P N+K + GFLGLTGYYR+F
Sbjct: 250 GIATDPAKIKAVQDWPVPVNLKQLCGFLGLTGYYRRF----------------------- 286
Query: 863 EEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSN 922
F VE DA G GIGAVLMQ+ P+A+ S+ L
Sbjct: 287 ----------------------------FVVEMDACGHGIGAVLMQEGHPLAFISRQLKG 318
Query: 923 GNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKL 982
L S+YEKEL+A++ ++ WRHYL+ F + TD +SLK+ L+Q++++P QQ WL KL
Sbjct: 319 KQLHLSIYEKELLAVIFVVRKWRHYLISSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKL 378
Query: 983 LGYQFEVKYKPGQENKAADALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQ 1042
L + +E++YK G+EN ADALSR +S D K++ +D I+ ++
Sbjct: 379 LEFDYEIQYKQGKENLVADALSRVEGSEVLHMALSVVECDLLKEIQAGYVTDGDIQGIIT 438
Query: 1043 DVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLA 1102
+QQ SK ++ QGVL + ++V+ S +L+ H S GGHSG T++R+
Sbjct: 439 ILQQQADSKKHYTWSQGVLRRKNKIVVPNNSGIKDTILRWLHCSGMGGHSGKEVTHQRVK 498
Query: 1103 ESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKS 1162
YW M + ++ F+R+C TCQ+ K + GLLQP P+P+ +W D+S+DFI GLP S
Sbjct: 499 GLFYWKSMVKDIQAFIRSCGTCQQCKSDNAASPGLLQPLPIPDRIWSDVSMDFIDGLPLS 558
Query: 1163 KGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSH 1222
G I+VVVDRLSK +HFI L HPY+A ++A+ ++ + +LHG P+++ + V H
Sbjct: 559 NGKTVIMVVVDRLSKAAHFIALAHPYSAMTVAQAYLDNVFKLHGCPSSIVVSSRRVLVQH 618
Score = 131 bits (329), Expect = 3e-30
Identities = 60/134 (44%), Positives = 93/134 (68%)
Query: 1293 TPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKR 1352
TP+E VYG+ P + +L E+KV VA + ER+ + L+ HL RAQ +M A++
Sbjct: 626 TPYEAVYGQPPPLHLPYLPGESKVAVVARSMQERESMILFLKFHLMRAQHRMKQLADQHI 685
Query: 1353 RDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRI 1412
+ F++G++VF+KL+P+RQ SVV R QKL+ +++GP+++ D+ G VAYKL+LP S++
Sbjct: 686 TEREFEVGDYVFVKLQPYRQQSVVMRSTQKLSPKYFGPYKVIDRCGEVAYKLQLPANSQV 745
Query: 1413 HPVFHVSLLKRAIG 1426
HPVFHVS L+ +G
Sbjct: 746 HPVFHVSQLRVLVG 759
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 498 bits (1283), Expect = e-141
Identities = 290/779 (37%), Positives = 430/779 (54%), Gaps = 48/779 (6%)
Query: 444 GHKVVTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQF 503
G + G +G+ ++ + D ++ + D++LG+ WL +V +DW + F
Sbjct: 344 GQFLAVLGRAKGVDIQIAGESMPADLIISPVELYDVILGMDWLDYY-RVHLDWHRGRVFF 402
Query: 504 VYGEQVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKK---GHEIPQE 560
E + QG+R S + E+M G + +V + G +
Sbjct: 403 ERPEGRLVYQGVRPISG------SLVISAVQAEKMIEKGCEAYLVTISMPESVGQVAVSD 456
Query: 561 LRAVLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELL 620
+R V+ +F +VF LPP +S I L P P++ PYR + E+++Q+ +LL
Sbjct: 457 IR-VVQEFQDVFQSLQGLPPSQSDPFTIELEPGTAPLSKAPYRMAPAEMAELKKQLKDLL 515
Query: 621 EAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAV 680
G IRPS S + +PV+ VKKKD S+R+C+DYR LN+ T+ ++YP+P +DELLD+L GA
Sbjct: 516 GKGFIRPSTSPWGAPVLFVKKKDGSFRLCIDYRELNRVTVKNRYPLPRIDELLDQLRGAT 575
Query: 681 IFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPY 740
FSKIDL SGYHQI + E D+ KT FRT GH+E++VMPFGL NAPA F +MN +F+ +
Sbjct: 576 CFSKIDLTSGYHQIPIAEADVRKTAFRTRYGHFEFVVMPFGLTNAPAVFMRLMNSVFQEF 635
Query: 741 LRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIIS 800
L +FV++F DDIL+YSKS E HL+ V+ L A SKC F ++ +LGHI+S
Sbjct: 636 LDEFVIIFIDDILVYSKSPEEQEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVS 695
Query: 801 GAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKD-NF 859
GVSVDPEK++ I +WP P N +R FLG GYYR+F+K + MA+P+T+LT KD F
Sbjct: 696 AEGVSVDPEKIEAIRDWPRPTNATEIRSFLGWAGYYRRFVKGFASMAQPMTKLTGKDVPF 755
Query: 860 TWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKA 919
W +E ++ F LK ++TS+PVLALP+ + + V DA+ G+G VLMQ + IAY S+
Sbjct: 756 VWSQECEEGFVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQHGKVIAYASRQ 815
Query: 920 LSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWL 979
L ++ E+ A++ ++ WR YL G V+TDHKSLK+ Q + Q+ W+
Sbjct: 816 LMKHEGNYPTHDLEMAAVIFALKIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWM 875
Query: 980 AKLLGYQFEVKYKPGQENKAADALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKN 1039
+ Y E+ Y PG+ N DALSR+ V L + L+ EI +
Sbjct: 876 ELVADYDLEIAYHPGKANVVVDALSRK--------RVGAALGQSVEVLVSEIGA-----L 922
Query: 1040 LLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRT-- 1097
L V ++P +V + LL + RL G T
Sbjct: 923 RLCAVAREPLGLE--AVDRADLLTRVRLAQKK-------------------DEGLRATKM 961
Query: 1098 YRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFII 1157
YR L WVGM+ V ++V CD CQ K GG+LQ P+P W+ +++D ++
Sbjct: 962 YRDLKRYYQWVGMKMDVANWVAECDVCQLVKAEHQVLGGMLQSLPIPEWKWDFITMDLVV 1021
Query: 1158 GLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRD 1216
GL S+ +AI V+VDRL+K +HF+ ++ A +A+ FV EIV+LHG+P +K +D
Sbjct: 1022 GLRVSRTKDAIWVIVDRLTKSAHFLAIRKTDGAAVLAKKFVSEIVKLHGVPLNMKEAQD 1080
Score = 85.5 bits (210), Expect = 2e-16
Identities = 50/165 (30%), Positives = 91/165 (54%), Gaps = 11/165 (6%)
Query: 1336 HLQRAQEQMASYANKKRRDLSFDIGEWVFLKLR----PHRQHSVVKRINQKLAARFYGPF 1391
+++ AQ++ SYA+K+RR+L F++G+ V+LK+ P+R S KL+ R+ GPF
Sbjct: 1074 NMKEAQDRQRSYADKRRRELEFEVGDRVYLKMAMLRGPNRSIS-----ETKLSPRYMGPF 1128
Query: 1392 QIEDKIGTVAYKLKLP-VESRIHPVFHVSLLKRAI-GNYQVQGQLPTDLGIEEATDIYPE 1449
+I +++ VAY+L+LP V H VFHVS+L++ + + + ++P DL + P
Sbjct: 1129 KIVERVEPVAYRLELPDVMRAFHKVFHVSMLRKCLHKDDEALAKIPEDLQPNMTLEARPV 1188
Query: 1450 AILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEF 1494
+L RI ++ + W + + TWE + +F ++
Sbjct: 1189 RVLERRIKELRQKKIPLIKVLWDCDGVTEETWEPEARMKARFKKW 1233
>At2g06890 putative retroelement integrase
Length = 1215
Score = 449 bits (1155), Expect = e-126
Identities = 301/866 (34%), Positives = 444/866 (50%), Gaps = 79/866 (9%)
Query: 336 HKCPERAMRVLILGDGETMDEDGEIV----MLEEEAEEIEEEETVECKLMGVLGCMGEHQ 391
++CP + RV+IL D ++ + EI L+E E + E + + + + Q
Sbjct: 170 NECPNK--RVMILLDNGEIEPEEEIPDSPSSLKENEELPAQGELLVARRTLSVQTKTDEQ 227
Query: 392 TQTMKLAGQVANVE---LLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKV- 447
Q L +V ++ID G+ N S + LGL L D K+
Sbjct: 228 EQRKNLFHTRCHVHGKVCSLIIDGGSCTNVASETMVKKLGLKW---------LNDSGKMR 278
Query: 448 VTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGE 507
V + I EI D L +E G ++LG W S KV+ D F +
Sbjct: 279 VKNQVVVPIVIGKYEDEILCDVLPMEAG--HILLGRPWQSDR-KVMHDGFTNRHSFEFKG 335
Query: 508 QVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEA----------------- 550
L M + YQ +H +Q ++ +V A
Sbjct: 336 GKTILVSMTPHEVYQDQIHLKQKKEQVVKQPNFFAKSGEVKSAYSSKQPMLLFVFKEALT 395
Query: 551 --TKKGHEIPQELRAVLADFHEVFTKKIQ--LPPVRSRVHQITLNPEHGPINVRPYRYPH 606
T +P E+ ++L D+ +VF + LPP+R HQI P N YR
Sbjct: 396 SLTNFAPVLPSEMTSLLQDYKDVFPEDNPKGLPPIRGIEHQIDFVPGASLPNRPAYRTNP 455
Query: 607 HQKEEIERQVTELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPI 666
+ +E++RQ KD SWRMC D RA+N T+ +PI
Sbjct: 456 VETKELQRQ--------------------------KDGSWRMCFDCRAINNVTVKYCHPI 489
Query: 667 PIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAP 726
P +D++LDEL+G+ IFSKIDLKSGYHQIR++E D KT F+T +G YE+LVMPFGL +AP
Sbjct: 490 PRLDDMLDELHGSSIFSKIDLKSGYHQIRMNEGDEWKTAFKTKHGLYEWLVMPFGLTHAP 549
Query: 727 ATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCK 786
+TF +MN + R ++ FV+V+FDDIL+YS+SL EH +HL VL VL AN KC
Sbjct: 550 STFMRLMNHVLRAFIGIFVIVYFDDILVYSESLREHIEHLDSVLNVLRKEELYANLKKCT 609
Query: 787 FGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKM 846
F + +LG ++S GV VD EKV+ I +WP PK V VR F GL G+YR+F K++ +
Sbjct: 610 FCTDNLVFLGFVVSADGVKVDEEKVKAIRDWPSPKTVGEVRSFHGLAGFYRRFFKDFSTI 669
Query: 847 AKPLTELTKKD-NFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAV 905
PLTE+ KKD F W + ++AF LK +T++PVL L +F K+FE+ECDA+G GIGAV
Sbjct: 670 VAPLTEVMKKDVGFKWEKAQEEAFQSLKDKLTNAPVLILSEFLKTFEIECDASGIGIGAV 729
Query: 906 LMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHF 965
LMQ ++ IA+FS+ L L Y+KEL ALV +Q W+HYL K F ++TDH+SLKH
Sbjct: 730 LMQDQKLIAFFSEKLGGATLNYPTYDKELYALVRALQRWQHYLWPKVFVIHTDHESLKHL 789
Query: 966 LQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADALSRRGDESDFFSLVSFPLWDDRK 1025
Q+ + W+ + + + +KYK G++N ADALS+R S ++ L +
Sbjct: 790 KGQQKLNKRHARWVEFIETFAYVIKYKKGKDNVVADALSQR---YTLLSTLNVKLM-GFE 845
Query: 1026 KLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHG 1085
++ E +D + + + ++ + + L Y+ RL + P ++E HG
Sbjct: 846 QIKEVYETDHDFQEVYKACEKFASGR--YFRQDKFLFYENRLCV-PNCSLRDLFVREAHG 902
Query: 1086 SPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPN 1145
GH G +T + E W M+ V+ C+TC++ K S + P GL P P+P
Sbjct: 903 GGLMGHFGIAKTLEVMTEHFRWPHMKCDVKRICGRCNTCKQAK-SKIQPNGLYTPLPIPK 961
Query: 1146 AVWEDLSLDFIIGLPKSKGYEAILVV 1171
W D+S+DF++GLP++ G ++I VV
Sbjct: 962 HPWNDISMDFVMGLPRT-GKDSIFVV 986
Score = 49.7 bits (117), Expect = 1e-05
Identities = 48/166 (28%), Positives = 82/166 (48%), Gaps = 14/166 (8%)
Query: 1293 TPFEVVYGRKAPPIIRF-LTNETKVVAVALELSERDEALKQLRTHLQRAQEQM----ASY 1347
+PF++VYG PI F L V+++ ++ E ++Q+ + +R E+ A
Sbjct: 988 SPFQIVYGFN--PISPFDLIPLPLSERVSIDGKKKAELVQQIHENARRNIEEKTKLYAKQ 1045
Query: 1348 ANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLP 1407
ANK RR+ F++G+ V++ LR R + K KL R GPF+I +I AY+L L
Sbjct: 1046 ANKGRREQIFEVGDMVWIHLRKERFPAQRK---SKLMPRIDGPFKIIKRINDNAYQLDLQ 1102
Query: 1408 VES--RIHPVFH--VSLLKRAIGNYQVQGQLPTDLGIEEATDIYPE 1449
E R +P ++ +I + + + +L +L EE + E
Sbjct: 1103 DEPDLRSNPFQEGGDDMIMDSINDMEHEPELERELVAEEGAKLVAE 1148
>At3g31970 hypothetical protein
Length = 1329
Score = 444 bits (1143), Expect = e-124
Identities = 284/844 (33%), Positives = 429/844 (50%), Gaps = 95/844 (11%)
Query: 673 LDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAI 732
+ E G V S I + + I + E ++ KT FRT GH+E++VMPFGL N P F +
Sbjct: 552 MPESVGQVAVSDIRVVHEFEDIPIAEANVRKTAFRTRYGHFEFVVMPFGLTNIPTAFMRL 611
Query: 733 MNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQI 792
MN +F+ +L +FV++F DDIL+YSKS EH + +S + A++
Sbjct: 612 MNSVFQEFLDEFVIIFIDDILVYSKSPEEHEVQ--------------SEESDGEAAGAEV 657
Query: 793 DYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTE 852
GVSVD EK++ I +WP P N +R FLGL GYY++F+K + MA+P+T+
Sbjct: 658 ----------GVSVDLEKIEAIRDWPRPTNATEIRSFLGLAGYYKRFVKVFASMAQPMTK 707
Query: 853 LTKKD-NFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQ 911
LT KD F W E + F LK ++TS+PVLALP+ + + V DA+ G+ VLMQ+
Sbjct: 708 LTGKDVPFVWSPECDEGFMSLKEMLTSTPVLALPEHGEPYMVYTDASRVGLDCVLMQR-- 765
Query: 912 PIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKIS 971
GK V+TDHKSLK+ Q
Sbjct: 766 --------------------------------------GK---VFTDHKSLKYIFTQPEL 784
Query: 972 SPDQQCWLAKLLGYQFEVKYKPGQENKAADALSRRGDESDFFSLVSF------------P 1019
+ Q+ W+ + Y E+ Y G+ N ADALSR+ +LVS P
Sbjct: 785 NLRQRRWMELVADYDLEIAYHSGKANVVADALSRKRVGGSVEALVSEIGALRLCVMAQEP 844
Query: 1020 LW---DDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSI 1076
L DR LL + L + S+ F+ G +L GR+ +
Sbjct: 845 LGLEAVDRADLLTRVRLAQEKDEGLIAASKADGSEYQFAA-NGTILVHGRVCVPKDEELR 903
Query: 1077 PWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGG 1136
+L E H S H + YR L WVGM+R V ++V CD CQ K PGG
Sbjct: 904 REILSEAHASMFSIHPRATKMYRDLKRYYQWVGMKRDVANWVTECDVCQLVKAEHQVPGG 963
Query: 1137 LLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEV 1196
LLQ P+ W+ +++DF++GLP S+ +AI V+VDRL+K +HF+ ++ A +A+
Sbjct: 964 LLQSLPILEWKWDFITMDFVVGLPVSRTKDAIWVIVDRLTKSAHFLAIRKTDGAVLLAKK 1023
Query: 1197 FVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCL 1256
+V EIV LHG+P ++ SDRD F S FW GTK++MS++YHP+TDGQ+E + L
Sbjct: 1024 YVSEIVELHGVPVSIVSDRDSKFTSAFWRAFQGEMGTKVQMSTAYHPQTDGQSERTIQTL 1083
Query: 1257 ESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKV 1316
E LR D W+ + EF YN+++ SI PFE +YGR + + +
Sbjct: 1084 EDMLRMCVLDRGGHWADHLSLVEFAYNNSYQASIRMAPFEALYGRPCRTPLCWTQVGERS 1143
Query: 1317 VAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLR----PHRQ 1372
+ A + E E ++ L+ +++ AQ++ SYA+K+RR+L F++G+ V+LK+ P+R
Sbjct: 1144 IYGADYVLETTERIRVLKLNMKEAQDRQRSYADKRRRELEFEVGDRVYLKMAMLRGPNRS 1203
Query: 1373 HSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLP-VESRIHPVFHVSLLKRAI-GNYQV 1430
S KL R+ GPF+I +++G VAY+L+LP V H VFHVS+L++ + + +V
Sbjct: 1204 IS-----ETKLTPRYMGPFRIVERVGPVAYRLELPDVMRAFHKVFHVSMLRKCLHKDDEV 1258
Query: 1431 QGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQ 1490
++ DL + P IL RI ++ + W + + TWE +
Sbjct: 1259 LAKILEDLQPNMTLEARPVRILERRIKELRRKKIPLIKVLWNCDGVTEETWEPEARMKAS 1318
Query: 1491 FPEF 1494
F ++
Sbjct: 1319 FKKW 1322
Score = 43.1 bits (100), Expect = 0.001
Identities = 25/98 (25%), Positives = 49/98 (49%), Gaps = 1/98 (1%)
Query: 390 HQTQTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITP-ITTRSIKLGDGHKVV 448
+ T+ + + A E +L DSGASH FI+P+ S + P ++K+ G +
Sbjct: 396 YTTREIGSTSKRAITEAHVLFDSGASHCFITPESASRGNIRGDPGEQLGAVKVAGGQFLA 455
Query: 449 TRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWL 486
G +G+ ++ + +D ++ + D++LG+ WL
Sbjct: 456 VLGRAKGVDIQIAGESMPVDLIISPVELYDVILGMDWL 493
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 425 bits (1092), Expect = e-118
Identities = 241/608 (39%), Positives = 343/608 (55%), Gaps = 40/608 (6%)
Query: 401 VANVELLILIDSGASHNFISPKITSALGLVITP-ITTRSIKLGDGHKVVTRGICRGIKAK 459
V VE +L DSGASH FI+P+ S + P ++K+ G V G +G+ +
Sbjct: 308 VGGVEAHVLFDSGASHCFITPESASRGNIRGDPGEQLGAVKVAGGQFVAVLGRTKGVDIQ 367
Query: 460 LGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKD 519
+ + D ++ + D++LG+ WL +V +D + F E + QG+R
Sbjct: 368 IAGESMPADLIISPVELYDVILGMDWLDHY-RVHLDCHRGRVSFERPEGSLVYQGVR--- 423
Query: 520 SYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLADFHEVFTKKIQLP 579
T G V+ A + I + A L +F +VF LP
Sbjct: 424 -----------PTSGS----------LVISAVQAEKMIEKGCEAYL-EFEDVFQSLQGLP 461
Query: 580 PVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILV 639
P RS I L P P++ PYR + E+++Q+ +LL A PV+ V
Sbjct: 462 PSRSDPFTIELEPGTAPLSKAPYRMAPAEMAELKKQLEDLLGA------------PVLFV 509
Query: 640 KKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEE 699
KKKD S+R+C+DYR LN T+ +KYP+P +DELLD+L GA FSKIDL SGYH I + E
Sbjct: 510 KKKDGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEA 569
Query: 700 DIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSL 759
D+ KT FRT GH+E++VMPFGL NAPA F +MN +F+ L +FV++F DDIL+YSKSL
Sbjct: 570 DVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSKSL 629
Query: 760 LEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPE 819
EH HL+ V+ L A SKC F ++ +LGHI+S GVSVDPEK++ I +W
Sbjct: 630 EEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHT 689
Query: 820 PKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKD-NFTWGEEAKQAFHKLKTVMTS 878
P N +R FLGL GYYR+F+K + MA+P+T+LT KD F W E ++ F LK ++TS
Sbjct: 690 PTNATEIRSFLGLAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTS 749
Query: 879 SPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALV 938
+PVLALP+ + + V DA+G G+G VLMQ+ + IAY S+ L ++ E+ A++
Sbjct: 750 TPVLALPEHGEPYMVYTDASGVGLGCVLMQRGKVIAYASRQLRKHEGNYPTHDLEMAAVI 809
Query: 939 LCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENK 998
++ WR YL G V+TDHKSLK+ Q + Q+ W+ + Y E+ Y PG+ N
Sbjct: 810 FALKIWRSYLYGGNVQVFTDHKSLKYIFTQPELNLRQRQWMELVADYDLEIAYHPGKANV 869
Query: 999 AADALSRR 1006
ADALS +
Sbjct: 870 VADALSHK 877
Score = 243 bits (619), Expect = 8e-64
Identities = 135/387 (34%), Positives = 220/387 (55%), Gaps = 11/387 (2%)
Query: 1114 VRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVD 1173
V ++V CD CQ K PGG+LQ P+P W+ +++DF++GLP S+ +AI V+VD
Sbjct: 943 VANWVAECDVCQLVKAEHQVPGGMLQSLPIPEWKWDFITIDFVVGLPVSRTKDAIWVIVD 1002
Query: 1174 RLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGT 1233
RL+K +HF+ ++ A +A+ +V EIV+LHG+P ++ SDRD F S FW GT
Sbjct: 1003 RLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGT 1062
Query: 1234 KLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQT 1293
K++MS++YHP+T GQ+E + LE LR D W+ + EF YN+++ SIG
Sbjct: 1063 KVQMSTAYHPQTYGQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYPASIGMA 1122
Query: 1294 PFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRR 1353
PFE +Y R + + + A + E E ++ L+ +++ AQ++ SYA+K+RR
Sbjct: 1123 PFEALYERPCRTPLCLTQVGERSIYGADYVQETTERIRVLKLNMKEAQDRQRSYADKRRR 1182
Query: 1354 DLSFDIGEWVFLKLR----PHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLP-V 1408
+L F++G+ V+LK+ P+R S KL+ R+ GPF+I +++G VAY+L+LP V
Sbjct: 1183 ELEFEVGDRVYLKMAMLRGPNRSIS-----ETKLSPRYMGPFRIVERVGPVAYRLELPDV 1237
Query: 1409 ESRIHPVFHVSLLKRAI-GNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQS 1467
H VFHVS+L++ + + +V ++P DL + P +L RI ++
Sbjct: 1238 MRAFHKVFHVSMLRKCLHKDDEVLAKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLI 1297
Query: 1468 LIKWKNRSIDDVTWEDNEVLAGQFPEF 1494
+ W + TWE + +F ++
Sbjct: 1298 KVLWDCDGVTKETWEPEARMKARFKKW 1324
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 392 bits (1008), Expect = e-109
Identities = 209/514 (40%), Positives = 307/514 (59%), Gaps = 10/514 (1%)
Query: 439 IKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKL 498
++ G + + RGI + + + D +++ L D++LG+ WL K +D
Sbjct: 2 VRAAGGQAMYPTRLVRGISVVVNGVNMPADLIIVPLKKHDVILGMDWLGKY-KGHIDCHR 60
Query: 499 LTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIP 558
+QF E +++ QG+R S + E+M G + + T K
Sbjct: 61 ERIQFERDEGMLKFQGIRTTSG------SLVISAIQAERMLGKGCEAYLATITTKEVGAS 114
Query: 559 QELRAVLA--DFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQV 616
EL+ +L +F +VF +PP RS I L P PI+ PYR + E+++Q+
Sbjct: 115 AELKDILIVNEFSDVFAAVSGVPPDRSDPFTIELEPGTTPISKAPYRMAPAEMAELKKQL 174
Query: 617 TELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDEL 676
ELL G IRPS S + +PV+ VKKKD S+R+C+DYR LNK T+ +KYP+P +DEL+D+L
Sbjct: 175 EELLAKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQL 234
Query: 677 NGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDI 736
GA FSKIDL SGYHQI + D+ KT FRT GH+E++VMPFGL NAPA F +MN +
Sbjct: 235 GGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMKMMNGV 294
Query: 737 FRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLG 796
FR +L +FV++F DDIL++SKS H++HL+ VL L + A SK F + +LG
Sbjct: 295 FRDFLDEFVIIFIDDILVHSKSWEAHQEHLRAVLERLREHELFAKLSKFSFWQRSVGFLG 354
Query: 797 HIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKK 856
H+IS GVSVDPEK++ I EWP P+N +R FLGL GYYR+F+ ++ MA+PLT LT K
Sbjct: 355 HVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQPLTRLTGK 414
Query: 857 DN-FTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAY 915
D F W +E +++F +LK ++ ++PVL LP+ + + V DA+ G+G VLMQ+ IAY
Sbjct: 415 DTAFNWSDECEKSFLELKAMLINAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGSVIAY 474
Query: 916 FSKALSNGNLTKSVYEKELMALVLCIQHWRHYLL 949
S+ L ++ E+ A+V ++ WR YL+
Sbjct: 475 ASRQLRKHEKNYPTHDLEMAAVVFALKIWRSYLI 508
Score = 250 bits (638), Expect = 5e-66
Identities = 143/400 (35%), Positives = 227/400 (56%), Gaps = 9/400 (2%)
Query: 1059 GVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFV 1118
G ++ GR+ + +L+E H S H G + YR L +WVGM++ V +V
Sbjct: 535 GTIVVNGRVCVPNDRALKEEILREAHQSKFSIHPGSNKMYRDLKRYYHWVGMRKDVARWV 594
Query: 1119 RACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLP---KSKGYEAILVVVDRL 1175
C TCQ K P GLLQ P+ W+ +++DF+ LP KSK + A+ VVVDRL
Sbjct: 595 AKCPTCQLVKAEHQVPSGLLQNLPISEWKWDHITMDFVTRLPTGIKSK-HNAVWVVVDRL 653
Query: 1176 SKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKL 1235
+K +HF+ + A+ IAE ++ EI+RLHGIP ++ SDRD F S FW+ K GT++
Sbjct: 654 TKSAHFMAISDKDGAEIIAEKYIDEIMRLHGIPVSIVSDRDTRFTSKFWNAFQKALGTRV 713
Query: 1236 KMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPF 1295
+S++YHP+TDGQ+E + LE LR D W ++ EF YN++F SIG +P+
Sbjct: 714 NLSTAYHPQTDGQSERTIQTLEDMLRACVLDWGGNWEKYLRLIEFAYNNSFQASIGMSPY 773
Query: 1296 EVVYGRKA-PPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRD 1354
E +YGR P+ E ++ + + E E +K L+ L+ AQ++ SYANK+R++
Sbjct: 774 EALYGRACRTPLCWTPVGERRLFGPTI-VDETTERMKFLKIKLKEAQDRQKSYANKRRKE 832
Query: 1355 LSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRI-H 1413
L F +G+ V+LK ++ +KL+ R+ GP+++ +++G VAYKL LP + + H
Sbjct: 833 LEFQVGDLVYLKAMTYKGAGRFTS-RKKLSPRYVGPYKVIERVGAVAYKLDLPPKLNVFH 891
Query: 1414 PVFHVSLLKRAIGNYQVQGQ-LPTDLGIEEATDIYPEAIL 1452
VFHVS L++ + + + + +P L + +P I+
Sbjct: 892 NVFHVSQLRKYLSDQEESVEDIPPGLKENMTVEAWPVRIM 931
>At4g07850 putative polyprotein
Length = 1138
Score = 385 bits (988), Expect = e-106
Identities = 207/505 (40%), Positives = 287/505 (55%), Gaps = 54/505 (10%)
Query: 798 IISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKD 857
+ S GV VD EKV+ I EWP PK+V VR F GL G+YR+F++++ +A PLTE+ KK+
Sbjct: 531 VASTDGVKVDEEKVKAIREWPSPKSVGKVRSFHGLAGFYRRFVRDFSTLAAPLTEVIKKN 590
Query: 858 -NFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYF 916
F W + + AF LK +T +PVL+LPDF K+FE+ECDA G GIGAVLMQ ++PIAYF
Sbjct: 591 VGFKWEQAPEDAFQALKEKLTHAPVLSLPDFLKTFEIECDAPGVGIGAVLMQDKKPIAYF 650
Query: 917 SKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQ 976
S+ L L Y+KEL ALV +Q W+HYL K F ++TDH+SLKH Q+ +
Sbjct: 651 SEKLGGATLNYPTYDKELYALVRALQTWQHYLWPKEFVIHTDHESLKHLKGQQKLNKRHA 710
Query: 977 CWLAKLLGYQFEVKYKPGQENKAADALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPY 1036
W+ + + + +KYK G++N ADALSRR
Sbjct: 711 RWVEFIETFPYVIKYKKGKDNVVADALSRRH----------------------------- 741
Query: 1037 IKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLR 1096
+G L Y RL + P S ++E HG GH G +
Sbjct: 742 ---------------------EGFLFYDNRLCI-PNSSLRELFIREAHGGGLMGHFGVSK 779
Query: 1097 TYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFI 1156
T + + + +W M+R V C TC++ K + P GL P P+P W D+S+DF+
Sbjct: 780 TLKVMQDHFHWPHMKRDVERMCERCTTCKQAK-AKSQPHGLCTPLPIPLHPWNDISMDFV 838
Query: 1157 IGLPKSK-GYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDR 1215
+GLP+++ G ++I VVVDR SK +HFI A IA +F RE+VRLHG+P T+ SDR
Sbjct: 839 VGLPRTRTGKDSIFVVVDRFSKMAHFIPCHKTDDAMHIANLFFREVVRLHGMPKTIVSDR 898
Query: 1216 DPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWV 1275
D F+S+FW L+ GTKL S++ HP+TDGQTEV+NR L + LR + KTW +
Sbjct: 899 DTKFLSYFWKTLWSKLGTKLLFSTTCHPQTDGQTEVVNRTLSTLLRALIKKNLKTWEDCL 958
Query: 1276 PWAEFWYNSTFHCSIGQTPFEVVYG 1300
P EF YN + H + +PF++VYG
Sbjct: 959 PHVEFAYNHSVHSATKFSPFQIVYG 983
Score = 60.5 bits (145), Expect = 8e-09
Identities = 36/83 (43%), Positives = 48/83 (57%), Gaps = 2/83 (2%)
Query: 567 DFHEVFTKKIQ--LPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGI 624
D+ +VF ++ LPP+R HQI P N YR + +E+E+QV EL+E G
Sbjct: 447 DYQDVFPEENPEGLPPIRGIEHQIDFVPGASLPNRPAYRTNPVETKELEKQVNELMERGH 506
Query: 625 IRPSMSSFSSPVILVKKKDKSWR 647
I SMS + PV+LV KKD SWR
Sbjct: 507 ICESMSPCAVPVLLVPKKDGSWR 529
>At4g16910 retrotransposon like protein
Length = 687
Score = 352 bits (904), Expect = 7e-97
Identities = 231/687 (33%), Positives = 362/687 (52%), Gaps = 54/687 (7%)
Query: 846 MAKPLTELTKKDN-FTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGA 904
MA+P+T+LT KD F W EE +++F +LK ++T++PVL LP+ + + V DA+ G+G
Sbjct: 1 MAQPMTQLTGKDTAFNWSEECERSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGC 60
Query: 905 VLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKH 964
VL+Q+ IAY S+ L + E+ A+V ++ WR YL G ++TDHKSLK+
Sbjct: 61 VLIQKGSVIAYASRQLRKHEKNYPTNDLEMAAVVFALKIWRSYLYGAKVQIFTDHKSLKY 120
Query: 965 FLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADALSRRGD----ESDFFSLVSF-- 1018
Q + Q+ W+ + Y + Y PG+ N+ DALSR E + +LV+
Sbjct: 121 IFTQPELNLRQRRWIKLVADYDLNIAYHPGKANQVVDALSRLRTVVEAERNQVNLVNMMG 180
Query: 1019 ------------PLW---DDRKKLLEEIAS----DPYIKNLLQDVQQDPQSKPGFSVLQG 1059
PL ++ LL I S D IK Q+ + + QS G
Sbjct: 181 TLHLNALSKEVEPLGLRAANQADLLSRIRSAQERDEEIKGWAQNNKTEYQSSNN-----G 235
Query: 1060 VLLYQGRLVLSPTSPSIPW-LLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFV 1118
++ GR V P ++ +L+E H S H G + YR L +WVGM++ V +V
Sbjct: 236 TIVVNGR-VCGPNDKALKEEILKEAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWV 294
Query: 1119 RACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLP---KSKGYEAILVVVDRL 1175
K P G+LQ P+P W+ + +DF+ GLP KSK + A+ VVVDRL
Sbjct: 295 A--------KEEHQVPSGMLQNLPIPEWKWDHIMMDFVTGLPTGIKSK-HNAVWVVVDRL 345
Query: 1176 SKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKL 1235
+K +HF+ + A+ IAE ++ EIVRLHGIP ++ SDRD F S FW K+ GT++
Sbjct: 346 TKSAHFMAISDKDAAEIIAEKYIDEIVRLHGIPVSIVSDRDTRFTSKFWKPFQKVLGTRV 405
Query: 1236 KMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPF 1295
+S++YHP+TDGQ+E + LE LR D W ++ EF YN++F SIG +P+
Sbjct: 406 NLSTAYHPQTDGQSERTIQTLEDMLRACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPY 465
Query: 1296 EVVYGRKA-PPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRD 1354
E +YGR P+ E ++ A+ + E + +K L+ L+ AQ++ SYANK+R++
Sbjct: 466 EALYGRAGRTPLCWTPVGERRLFGPAV-VDETTKKMKFLKIKLKEAQDRQKSYANKRRKE 524
Query: 1355 LSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVE-SRIH 1413
L F +G+ V+LK ++ +KL R+ GP+++ +++G VAYKL LP + H
Sbjct: 525 LEFQVGDLVYLKAMTYKGAGRFTS-RKKLRPRYVGPYKVIERVGAVAYKLDLPPKLDAFH 583
Query: 1414 PVFHVSLLKRAIGNYQVQGQ-LPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIK-- 1470
VFHVS L++ + + + +P L + +P I+ +++G L+K
Sbjct: 584 NVFHVSQLRKCLSEQEESMEDVPPGLKENMTVEAWPVRIMDQ--MKKGTRGKSMDLLKIL 641
Query: 1471 WKNRSIDDVTWEDNEVLAGQFPEFCLE 1497
W ++ TWE + FPE+ E
Sbjct: 642 WNCGGREEYTWETETKMKANFPEWFKE 668
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 310 bits (794), Expect = 4e-84
Identities = 152/316 (48%), Positives = 208/316 (65%), Gaps = 4/316 (1%)
Query: 1187 PYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETD 1246
P ++S+A FV +V+LHGIP ++ SD DPIF+S FW E +KL TKL MS++YHP+TD
Sbjct: 628 PLLSRSVAAKFVSHVVKLHGIPRSIVSDCDPIFMSLFWQEFWKLSRTKLWMSTAYHPQTD 687
Query: 1247 GQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPI 1306
GQTEV+NRC+E +LRCF HPK WS ++PWAE+WYN+TFH S G TPF+ +YGR PI
Sbjct: 688 GQTEVVNRCIEQFLRCFVHYHPKQWSSFIPWAEYWYNTTFHASTGMTPFQALYGRPPSPI 747
Query: 1307 IRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLK 1366
+ + +++ RDE L +L+ HL A M A+ + RD+SF +G+WV L+
Sbjct: 748 PAYELGSVVCGELNEQMAARDELLAELKQHLVTANNCMKQQADSRLRDVSFQVGDWVLLR 807
Query: 1367 LRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLKRAIG 1426
++P+RQ ++ +R +QKL+ RFYGPFQ+ K G VAY+L LP +RIHPVFHVSLLK +G
Sbjct: 808 IQPYRQKTLFRRSSQKLSHRFYGPFQVASKHGEVAYRLTLPEGTRIHPVFHVSLLKPWVG 867
Query: 1427 NYQV-QGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNE 1485
+ + GQLP L + P A+L R Q V L++W+ I+D TWE+ +
Sbjct: 868 DGEPDMGQLP-PLRNNGELKLQPTAVLEVRWRSQDKKRVADLLVQWEGLHIEDATWEEYD 926
Query: 1486 VLAGQFPEFC--LEDK 1499
LA FPEF LEDK
Sbjct: 927 QLAASFPEFVLNLEDK 942
Score = 244 bits (623), Expect = 3e-64
Identities = 145/392 (36%), Positives = 212/392 (53%), Gaps = 37/392 (9%)
Query: 361 VMLEEEAEEIEEEETVECK---LMGVLGCMGEHQTQTMKLAGQVANVELLILIDSGASHN 417
VM E +E EEE+ E + + + G T T+++ + L+ LI+SG++HN
Sbjct: 281 VMKGENDDEAVEEESTEAEGKPEITLYALEGVDTTSTIRVRATIHRNRLIALINSGSTHN 340
Query: 418 FISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLELGGL 477
FI K + L T ++++ +G +V R I +G + + L L GL
Sbjct: 341 FIGEKAVRGMNLKATTTKPFTVRVVNGMPLVCRSRYEAIPVVMGGVVFPVTLYALPLMGL 400
Query: 478 DLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTHGREQ 537
DL +GV WLSTLG + +WK T+QF + V+L G++ + H ++ + +
Sbjct: 401 DLAMGVQWLSTLGPTLCNWKEQTLQFHWAGDEVRLMGIK-PTGLRGVEHKTITK---KAR 456
Query: 538 MEWWGPQLQVVEATKKGHEIPQELRAVLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPI 597
M + + + + E++ ++ +F +F QLP R VH+I L P+
Sbjct: 457 MGHTIFAITMAHNSSDPNTPDGEIKKLIEEFAGLFETPTQLPSERPIVHRIALKKGTDPV 516
Query: 598 NVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNK 657
NVRPYRY +QK+E+ R AGIIRPS S F SPV+L
Sbjct: 517 NVRPYRYAFYQKDEMSR-------AGIIRPSSSPFLSPVLL------------------- 550
Query: 658 ATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLV 717
+++PIP VD++LDELNGAV F+K+DL +GY Q+R+H DIPK F+THNGHYEYLV
Sbjct: 551 ----ERFPIPTVDDMLDELNGAVYFTKLDLTAGYQQVRMHSPDIPKIAFQTHNGHYEYLV 606
Query: 718 MPFGLMNAPATFQAIMNDIFRPYLRKFVLVFF 749
MPFGL NAP+TFQA+MN+IF P L + V F
Sbjct: 607 MPFGLCNAPSTFQALMNEIFWPLLSRSVAAKF 638
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 304 bits (778), Expect = 3e-82
Identities = 233/839 (27%), Positives = 373/839 (43%), Gaps = 118/839 (14%)
Query: 609 KEEIERQVTELLEAGIIRP-SMSSFSSPVILVKKKD------------------KSWRMC 649
KE +++++ +LL+AG+I P S S++ SPV V KK RMC
Sbjct: 912 KEVVKKEILKLLDAGVIYPISDSTWVSPVHCVPKKGGMTVVKNSKDELIPTRTITGHRMC 971
Query: 650 VDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTH 709
++YR LN A+ + +P+P +D +L+ L + +D SG+ QI +H D KTTF
Sbjct: 972 IEYRKLNVASRKEHFPLPFIDHMLERLANHPYYCFLDSYSGFFQIPIHPNDQGKTTFTCP 1031
Query: 710 NGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLV 769
G + Y MPFGL NAPATFQ M IF + + V VF DD +Y S +L V
Sbjct: 1032 YGTFAYKRMPFGLCNAPATFQRCMTSIFSDLIEEMVEVFMDDFSVYGSSFSSCLLNLCRV 1091
Query: 770 LTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGF 829
L + V N KC F + LG IS G+ VD K+ +++ PK VK +R F
Sbjct: 1092 LKRCEETNLVLNWEKCHFMVREGIVLGRKISEEGIEVDKAKIDVMMQLQPPKTVKDIRSF 1151
Query: 830 LGLTGYYRKFIKNYGKMAKPLTE-LTKKDNFTWGEEAKQAFHKLKTVMTSSPVLALPDFD 888
LG G+YR FIK++ K+A+PLT L K+ F + +E AF +K + ++P++ P++D
Sbjct: 1152 LGHAGFYRIFIKDFSKLARPLTRLLCKETEFAFDDECLTAFKLIKEALITAPIVQAPNWD 1211
Query: 889 KSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYL 948
FE+ + + + + EKEL+A+V + +R YL
Sbjct: 1212 FPFEI------------------------ITMDDAQVRYATTEKELLAVVFAFEKFRSYL 1247
Query: 949 LGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADALSRRGD 1008
+G T+YTDH +L+H +K + P W+ L + E+ K G EN AD LSR
Sbjct: 1248 VGSKVTIYTDHAALRHIYAKKDTKPRLLRWILLLQEFDMEIVDKKGIENGVADHLSRMRI 1307
Query: 1009 ESDF----------------FSLVSFPLWDDRKKLLEEIASDPYI-----KNLLQDVQQD 1047
E + + P + D L P + K +D+
Sbjct: 1308 EDEVLIDDSMPEEQLMAIQQLNEKKLPWYADHVNYLVSGEEPPNLSSYEKKKFFKDINHF 1367
Query: 1048 PQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAES-LY 1106
+P L +Y+ + I +L HG GGH +T ++ ++ +
Sbjct: 1368 YWDEPYLYTLCKDKIYR----TCVSEDEIEGILLHCHGFAYGGHFATFKTMSKILQAGFW 1423
Query: 1107 WVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYE 1166
W M + ++F+ CD+ + VW +DF+ P S G +
Sbjct: 1424 WPSMFKDAQEFISKCDSFENFD------------------VW---GIDFMGPFPSSYGNK 1462
Query: 1167 AILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSE 1226
ILV +D +SK+ I H A+ + ++F I G+P V SD F++ +
Sbjct: 1463 YILVAIDYVSKWVEAI-ASHTNDARVVLKLFKTIIFPRFGVPRIVISDGGKHFINKGFEN 1521
Query: 1227 LFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTF 1286
L K G K K T GQ E+ NR +++ L K WS + + Y + F
Sbjct: 1522 LLKKHGVKHK--------TSGQVEISNREIKAILEKTVGSTRKDWSAKLNDTLWAYRTAF 1573
Query: 1287 HCSIGQTPFEVVYGR----------KAPPIIRFLTNETKVVAV--ALELSERDEALKQLR 1334
IG TPF ++YG+ KA ++ L + K ++L++ ++ +
Sbjct: 1574 KTPIGTTPFNLLYGKSCHLPVELEYKAMWAVKLLNFDIKTAEEKRLIQLNDLNKIRLEAY 1633
Query: 1335 THLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQI 1393
+ +E+ S+ +KK F +G+ V L +S ++ KL +R+ GPF +
Sbjct: 1634 ESSKIYKERTKSFHDKKIVSRDFKVGDQVLL------FNSRLRLFPGKLKSRWSGPFSV 1686
>At4g08100 putative polyprotein
Length = 1054
Score = 292 bits (748), Expect = 9e-79
Identities = 155/342 (45%), Positives = 204/342 (59%), Gaps = 29/342 (8%)
Query: 651 DYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHN 710
D RA+N T+ ++PIP +D+ LD+L+G+ IFSKIDLKSGYHQ R+ E D KT +T
Sbjct: 527 DCRAINNITVKYRHPIPRLDDTLDKLHGSSIFSKIDLKSGYHQTRMKEGDEWKTAIKTKQ 586
Query: 711 GHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVL 770
YE+LVMPFGL NAP TF +MN + R ++ FV+V+FDDIL+YSK+L H HLKLVL
Sbjct: 587 RLYEWLVMPFGLTNAPNTFMRLMNHVLRKHIGVFVIVYFDDILVYSKNLEYHVMHLKLVL 646
Query: 771 TVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFL 830
+L AN KC F + +LG ++S G+ VD EKV+ I EWP PKNV
Sbjct: 647 DLLRKEKLYANLKKCTFCTDNLVFLGFVVSADGIKVDEEKVKAIREWPNPKNVS------ 700
Query: 831 GLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKS 890
K F W + + AF LK +T+S V LP+F KS
Sbjct: 701 -----------------------EKDIGFKWEDAQENAFQALKEKLTNSSVPILPNFMKS 737
Query: 891 FEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLG 950
FE+ECDA+G GIGAVLMQ +PIAYFS+ L L Y+KEL ALV +Q W+HYL
Sbjct: 738 FEIECDASGLGIGAVLMQDHKPIAYFSEKLGGATLNYPTYDKELYALVRALQTWQHYLWP 797
Query: 951 KAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYK 992
K F ++TDH+SLKH Q+ + W+ + + + +KYK
Sbjct: 798 KKFVIHTDHESLKHLKGQQKLNKRHARWVEFIETFPYVIKYK 839
Score = 52.8 bits (125), Expect = 2e-06
Identities = 37/120 (30%), Positives = 65/120 (53%), Gaps = 5/120 (4%)
Query: 1287 HCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMAS 1346
H + +PFE+VYG K + + + +L+ ++D+ ++Q+ +L+ +Q
Sbjct: 851 HSATKFSPFEIVYGFKPTSPLDLIPLPLSERS-SLDGKKKDDLVQQV-LNLEARTKQYKK 908
Query: 1347 YANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKL 1406
YANK R+++ F+ G+ V++ LR R V + KL R GPF++ +I AYKL L
Sbjct: 909 YANKGRKEVIFNEGDQVWVHLRKKRFPEVR---SSKLMPRIDGPFKVLKRINNNAYKLDL 965
>At4g03650 putative reverse transcriptase
Length = 839
Score = 277 bits (708), Expect = 4e-74
Identities = 134/295 (45%), Positives = 188/295 (63%), Gaps = 1/295 (0%)
Query: 698 EEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSK 757
E D+ KT FRT GH+E++VMPFGL NAPA F +MN +F+ +L +FV++F DDIL+YSK
Sbjct: 518 EADVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSK 577
Query: 758 SLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEW 817
S EH HL+ V+ L A SKC F ++ +LGHI+S GVSVDPEK++ I +W
Sbjct: 578 SPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDW 637
Query: 818 PEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKD-NFTWGEEAKQAFHKLKTVM 876
P P N +R FLGL GYYR+FIK + MA+P+T+LT KD F W E ++ F LK ++
Sbjct: 638 PRPTNATEIRSFLGLAGYYRRFIKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEML 697
Query: 877 TSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMA 936
TS+PVLALP+ + + V DA+G G+G LMQ+ + IAY S+ L ++ E+ A
Sbjct: 698 TSTPVLALPEHGEPYMVYTDASGVGLGCALMQRGKVIAYASRQLRKHEGNYPTHDLEMAA 757
Query: 937 LVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKY 991
++ ++ WR YL G V+TDHKSLK+ Q + Q+ W+ + Y E+ Y
Sbjct: 758 VIFALKIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLEIAY 812
Score = 48.9 bits (115), Expect = 2e-05
Identities = 42/189 (22%), Positives = 79/189 (41%), Gaps = 23/189 (12%)
Query: 438 SIKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWK 497
++K+ G + G +G+ ++ + D ++ + D++LG+ WL +V +D
Sbjct: 357 AVKVAGGQFLAVLGGAKGVDIQIAGESLPADLIISHVELYDVILGMDWLDHY-RVHLDCH 415
Query: 498 LLTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEI 557
+ F E G G+++ + L D + A G
Sbjct: 416 RGRVSFERPE------GSAGRENDREGLRGLPGDD---------------IYAGVCGQVA 454
Query: 558 PQELRAVLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVT 617
++R V+ +F VF LPP RS I L P+ P++ PYR + E+++Q+
Sbjct: 455 VSDIR-VVQEFQYVFQSLQGLPPSRSDPFTIELEPKTAPLSKAPYRMAPAKMAELKKQLE 513
Query: 618 ELLEAGIIR 626
+LL +R
Sbjct: 514 DLLAEADVR 522
Score = 35.8 bits (81), Expect = 0.20
Identities = 13/19 (68%), Positives = 19/19 (99%)
Query: 1232 GTKLKMSSSYHPETDGQTE 1250
GTK++MS++YHP+TDGQ+E
Sbjct: 818 GTKVQMSTTYHPQTDGQSE 836
>At4g07830 putative reverse transcriptase
Length = 611
Score = 266 bits (680), Expect = 7e-71
Identities = 188/616 (30%), Positives = 293/616 (47%), Gaps = 61/616 (9%)
Query: 907 MQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFL 966
MQ+ + IAY S+ L ++ E+ A V+TDHKSLK+
Sbjct: 1 MQRGKVIAYGSRQLKKHEGNYPTHDLEMAA------------------VFTDHKSLKYIF 42
Query: 967 QQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADALSRRGDESDFFSLVSFPLWDDRKK 1026
Q + + W+ + Y E+ Y PG+ N DALSR+ + V + +
Sbjct: 43 TQLELNLRLRRWMKLVADYDLEIAYHPGKANVVTDALSRKRVGAAPGQSVETLVIEIGAL 102
Query: 1027 LLEEIASDPY------IKNLLQDVQQDPQSKPGFSVLQ------------GVLLYQGRLV 1068
L +A +P +LL VQ + G G +L GR+
Sbjct: 103 RLCAVAREPLGLEAVDQTDLLSRVQLAQEKDEGLIAASKAEGFEYQFAANGTILVHGRVC 162
Query: 1069 LSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQK 1128
+ +L E H S H G + YR L WVGM+R V ++V CD CQ K
Sbjct: 163 VPKDKELRQEILSEAHASMFSIHPGATKMYRDLKRYYQWVGMKRDVGNWVEECDVCQLVK 222
Query: 1129 YSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPY 1188
G LLQ P+P W+ +++DF++GLP S+ +AI V+VDRL+K +HF+ ++
Sbjct: 223 IEHQVSGSLLQSLPIPEWKWDFITMDFVVGLPVSRTKDAIWVIVDRLTKSAHFLAIRKTD 282
Query: 1189 TAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQ 1248
A +A+ +V EIV+LHG+P ++ SDRD F S FW GTK++MS++YHP+TDGQ
Sbjct: 283 GAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGTKVQMSTAYHPQTDGQ 342
Query: 1249 TEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIR 1308
+E + LE LR D W+ + EF YN+++ SIG PFEV+YGR +
Sbjct: 343 SERTIQTLEDMLRMCVLDWRGHWADHLSLVEFAYNNSYQASIGMAPFEVLYGRPCRTLCW 402
Query: 1309 FLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLR 1368
E + A + E E ++ L+ +++ AQ + SYA+K+R++L F++G+ V
Sbjct: 403 TQVGERSIYG-ADYVQEITERIRVLKLNMKEAQNRQRSYADKRRKELEFEVGDSV----- 456
Query: 1369 PHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKL-PVESRIHPVFHVSLLKRAI-G 1426
Q V R E ++G VA++L+L V H VFHVS+L++ +
Sbjct: 457 --SQDGHVAR--------------SEQRVGPVAFRLELSDVMRAFHKVFHVSMLRKCLHK 500
Query: 1427 NYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEV 1486
+ +V ++P DL + P +L RI ++ + + + TWE
Sbjct: 501 DDEVLAKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLRNCDGVTEETWEPEAR 560
Query: 1487 LAGQFPEFCLEDKAVS 1502
L +F ++ E K VS
Sbjct: 561 LKARFKKW-FEKKDVS 575
>At1g20390 hypothetical protein
Length = 1791
Score = 224 bits (572), Expect = 2e-58
Identities = 162/605 (26%), Positives = 294/605 (47%), Gaps = 19/605 (3%)
Query: 408 ILIDSGASHNFISPKITSALGLVITPITTRSIKLG--DGHKVVTRGICRGIKAKLGSIEI 465
+LID+G+S + I + +A+ + I S L DG V+T G + + +G +
Sbjct: 585 VLIDTGSSVDLIFKDVLTAMNITDRQIKPVSKPLAGFDGDFVMTIGTIK-LPIFVGGLIA 643
Query: 466 TIDALVLELGGL-DLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMR-GKDSYQS 523
+ +V+ + +++LG W+ + + + ++F + L+ + K +S
Sbjct: 644 WVKFVVIGKPAVYNVILGTPWIHQMQAIPSTYHQC-VKFPTHNGIFTLRAPKEAKTPSRS 702
Query: 524 NLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLADFHEVFTKKIQ----LP 579
S L T ++ P V + I EL A+L + F I+ +
Sbjct: 703 YEESELCRTE-MVNIDESDPTRCVGVGAEISPSIRLELIALLKRNSKTFAWSIEDMKGID 761
Query: 580 PVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAG-IIRPSMSSFSSPVIL 638
P + H++ ++P P+ + + + + +V +LL+AG II + + ++
Sbjct: 762 PAIT-AHELNVDPTFKPVKQKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVV 820
Query: 639 VKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHE 698
VKKK+ WR+CVDY LNKA D YP+P +D L++ +G + S +D SGY+QI +H+
Sbjct: 821 VKKKNGKWRVCVDYTDLNKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGYNQILMHK 880
Query: 699 EDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKS 758
+D KT+F T G Y Y VM FGL NA AT+Q +N + + + V V+ DD+L+ S
Sbjct: 881 DDQEKTSFVTDRGTYCYKVMSFGLKNAGATYQRFVNKMLADQIGRTVEVYIDDMLVKSLK 940
Query: 759 LLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWP 818
+H +HL VL + N +KC FG ++LG++++ G+ +P++++ I+E P
Sbjct: 941 PEDHVEHLSKCFDVLNTYGMKLNPTKCTFGVTSGEFLGYVVTKRGIEANPKQIRAILELP 1000
Query: 819 EPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTK-KDNFTWGEEAKQAFHKLKTVMT 877
P+N + V+ G +FI P L K + F W +++++AF KLK ++
Sbjct: 1001 SPRNAREVQRLTGRIAALNRFISRSTDKCLPFYNLLKRRAQFDWDKDSEEAFEKLKDYLS 1060
Query: 878 SSPVLALPDFDKSFEVECDAAGRGIGAVLMQ----QRQPIAYFSKALSNGNLTKSVYEKE 933
+ P+L P+ ++ + + + +VL++ +++PI Y SK+L V EK
Sbjct: 1061 TPPILVKPEVGETLYLYIAVSDHAVSSVLVREDRGEQRPIFYTSKSLVEAETRYPVIEKA 1120
Query: 934 LMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKP 993
+A+V + R Y V TD + L+ L S W +L Y + + +P
Sbjct: 1121 ALAVVTSARKLRPYFQSHTIAVLTD-QPLRVALHSPSQSGRMTKWAVELSEYDIDFRPRP 1179
Query: 994 GQENK 998
+++
Sbjct: 1180 AMKSQ 1184
Score = 85.1 bits (209), Expect = 3e-16
Identities = 81/344 (23%), Positives = 148/344 (42%), Gaps = 30/344 (8%)
Query: 1079 LLQEYHGSPTGGHSGFLRTYRRLAE-SLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGL 1137
+++E H G HSG +L + YW M + F C+ CQR + P L
Sbjct: 1438 IMKETHEGAAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAKCEQCQRHAPTIHQPTEL 1497
Query: 1138 LQP--FPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKY---SHFILLKHPYTAKS 1192
L+ P P W ++D + +P S+ ILV+ D +K+ + ++ A
Sbjct: 1498 LRAGVAPYPFMRW---AMDIVGPMPASRQKRFILVMTDYFTKWVEAESYATIR----AND 1550
Query: 1193 IAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVI 1252
+ + I+ HG+P + +D F+S + +L S+ +P+ +GQ E
Sbjct: 1551 VQNFVWKFIICRHGLPYEIITDNGSQFISLSFENFCASWKIRLNKSTPRYPQGNGQAEAT 1610
Query: 1253 NRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRK--APPIIRFL 1310
N+ + S L+ + W+ + + Y +T + QTPF YG + AP + +
Sbjct: 1611 NKTILSGLKKRLDEKKGAWADELDGVLWSYRTTPRSATDQTPFAHAYGMEAMAPAEVGYS 1670
Query: 1311 TNETKVVAVALELSERD-----EALKQLRT----HLQRAQEQMASYANKKRRDLSFDIGE 1361
+ ++ EL++R + L+++R +Q Q A + N+K + FD+G+
Sbjct: 1671 SLRRSMMVKNPELNDRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVHNRHFDVGD 1730
Query: 1362 WVFLKLRPHRQHSVVKRINQ-KLAARFYGPFQIEDKIGTVAYKL 1404
V K+ + IN KL A + G +Q+ + Y+L
Sbjct: 1731 LVLRKVFEN-----TAEINAGKLGANWEGSYQVSKIVRPGDYEL 1769
>At2g10690 putative retroelement pol polyprotein
Length = 622
Score = 202 bits (514), Expect = 1e-51
Identities = 140/403 (34%), Positives = 201/403 (49%), Gaps = 28/403 (6%)
Query: 617 TELLEAG---IIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELL 673
TE+L+ G I +S +P + +K K R Y L D YP+ I D L
Sbjct: 80 TEMLDHGGENISSDDLSELKAPKLDLKPLPKGLR----YAFLGPN---DTYPVIINDGLS 132
Query: 674 DELNGAVIFSKIDLKSGYHQIRVHEE------DIPKTTFRTHNGHYEYLVMPFGLMNAPA 727
DE ++ +L+ Q+ V + KTTF NG + Y M FGL NAP
Sbjct: 133 DEQVNQLLN---ELRKYRGQLVVSFKLQFTLMIKKKTTFTCPNGTFAYKRMSFGLCNAPG 189
Query: 728 TFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKF 787
TFQ M IF ++ + + VF DD +S LL +L VL + V N KC F
Sbjct: 190 TFQRSMTSIFSDFIEEIMEVFMDDFSGFSSCLL----NLCRVLERCEETNLVLNWEKCHF 245
Query: 788 GCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMA 847
+ LGH ISG G+ VD K+ +++ PK VK +R FLG +YR+FIK++ K+A
Sbjct: 246 MVHEGIVLGHKISGKGIEVDKAKIDVMIQLQPPKTVKDIRSFLGHAEFYRRFIKDFSKIA 305
Query: 848 KPLTE-LTKKDNFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVL 906
+PLT L K+ F + E+ +AFH +K + S+P+ P++D FE+ CDA +GAVL
Sbjct: 306 RPLTRLLCKETEFNFDEDCLKAFHLIKETLVSAPIFQAPNWDHPFEIMCDAFDYAVGAVL 365
Query: 907 MQ----QRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSL 962
Q + I Y S+ + + EKEL+A+V + R YL+G VY DH +L
Sbjct: 366 DQKIDDKLHVIYYASRTMDEAQTRYATIEKELLAVVFAFEKIRSYLVGFKVKVYIDHAAL 425
Query: 963 KHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADALSR 1005
+H +K + W+ L + E+ K G EN AD LSR
Sbjct: 426 RHIYAKKETKSRLLRWILLLQEFDMEIIDKKGVENGVADHLSR 468
Score = 37.7 bits (86), Expect = 0.052
Identities = 18/57 (31%), Positives = 30/57 (52%)
Query: 1245 TDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGR 1301
T GQ EV NR ++ L + WS + + Y + + IG+TPF+++YG+
Sbjct: 524 TSGQVEVSNRQMKDILTKVVGVSKRDWSTKLDETLWAYRTAYKTPIGRTPFQMLYGK 580
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,307,647
Number of Sequences: 26719
Number of extensions: 1568492
Number of successful extensions: 4379
Number of sequences better than 10.0: 96
Number of HSP's better than 10.0 without gapping: 70
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 4047
Number of HSP's gapped (non-prelim): 199
length of query: 1543
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1431
effective length of database: 8,326,068
effective search space: 11914603308
effective search space used: 11914603308
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0265.11


