

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
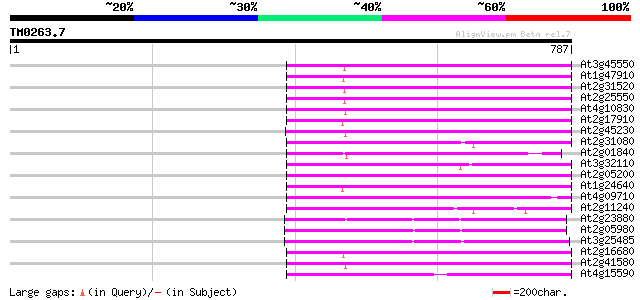
Query= TM0263.7
(787 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g45550 putative protein 272 5e-73
At1g47910 reverse transcriptase, putative 272 5e-73
At2g31520 putative non-LTR retroelement reverse transcriptase 271 8e-73
At2g25550 putative non-LTR retroelement reverse transcriptase 271 8e-73
At4g10830 putative protein 271 1e-72
At2g17910 putative non-LTR retroelement reverse transcriptase 262 5e-70
At2g45230 putative non-LTR retroelement reverse transcriptase 257 2e-68
At2g31080 putative non-LTR retroelement reverse transcriptase 257 2e-68
At2g01840 putative non-LTR retroelement reverse transcriptase 256 3e-68
At3g32110 non-LTR reverse transcriptase, putative 254 1e-67
At2g05200 putative non-LTR retroelement reverse transcriptase 254 1e-67
At1g24640 hypothetical protein 252 5e-67
At4g09710 RNA-directed DNA polymerase -like protein 250 2e-66
At2g11240 pseudogene 246 4e-65
At2g23880 putative non-LTR retroelement reverse transcriptase 243 4e-64
At2g05980 putative non-LTR retroelement reverse transcriptase 243 4e-64
At3g25485 unknown protein 242 5e-64
At2g16680 putative non-LTR retroelement reverse transcriptase 242 5e-64
At2g41580 putative non-LTR retroelement reverse transcriptase 238 1e-62
At4g15590 reverse transcriptase like protein 236 3e-62
>At3g45550 putative protein
Length = 851
Score = 272 bits (696), Expect = 5e-73
Identities = 149/404 (36%), Positives = 231/404 (56%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I +IPK++NPQ L +YRPI+L +YK+++K + RL+ L+ ++ D Q AFI GR
Sbjct: 106 NHTNICMIPKIQNPQTLSDYRPIALCNVLYKVISKCMVNRLKAHLNSIVSDSQAAFIPGR 165
Query: 449 FMLDSVVVANEVVEEAKRRK---KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++A+E++ K RK K K D KAYD V+W+FL M GF KWI
Sbjct: 166 IINDNVMIAHEIMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIG 225
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI ++S SVL+NGSP + RG+RQGDPL+P+LF++ + L+ L++
Sbjct: 226 WIMAAVKSVHYSVLINGSPHGYISPTRGIRQGDPLSPYLFILCGDILSHLIKVKASSGDI 285
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G R+G I+ LQFADD+LFF +A+ +N +K + +E SG K+N KS +
Sbjct: 286 RGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSLITFG 345
Query: 626 SMVSRETQI-FAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S V TQ +LN G YLG+P +K + +I +++ + + K L
Sbjct: 346 SRVYGSTQTRLKTLLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFL 405
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S G+ L+KSV ++P++ S FK P+ I+ + + F W RG+ WV W+ +
Sbjct: 406 SPAGKEILLKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNKRGIPWVAWKRL 465
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K+EGGLG +DL FN ALL K WR+++ L+ +V++++
Sbjct: 466 QYSKKEGGLGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMKAR 509
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 272 bits (696), Expect = 5e-73
Identities = 145/404 (35%), Positives = 229/404 (55%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I LIPK E P + RPISL YK+++K+L RL+ VL +I + Q+AF+ GR
Sbjct: 269 NTTNICLIPKTERPTRMTELRPISLCNVGYKVISKILCQRLKTVLPNLISETQSAFVDGR 328
Query: 449 FMLDSVVVANEVVEEAKRR---KKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D++++A E+ + K + K D KAYD V+WNF+ ++ ++GF KWI
Sbjct: 329 LISDNILIAQEMFHGLRTNSSCKDKFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKWIS 388
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI CI + VL+NG P RGLRQGDPL+P+LF++ E L +R+A +Q L
Sbjct: 389 WIMWCITTVQYKVLINGQPKGLIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQNLI 448
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA-G 624
TG +V +S L FADD+LFF +A+ + ++ IL+ +E++SG ++NF KS + G
Sbjct: 449 TGIKVATPSPAVSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQFG 508
Query: 625 ISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
+ +L +G +YLG+P K + V ++Q +++ K L
Sbjct: 509 HKVEDSIKADIKLILGIHNLGGMGSYLGLPESLGGSKTKVFSFVRDRLQSRINGWSAKFL 568
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKWEEI 743
S GG+ +IKSV ++LP + S F+ PK I + KF W + RG+ W+ W+++
Sbjct: 569 SKGGKEVMIKSVAATLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWDKL 628
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
C K +GGLG ++++ FN ALL K WRL+ + L+ +V + +
Sbjct: 629 CSSKSDGGLGFRNVDDFNSALLAKQLWRLITAPDSLFAKVFKGR 672
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 271 bits (694), Expect = 8e-73
Identities = 151/404 (37%), Positives = 229/404 (56%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I +IPK+ NP L +YRPI+L +YK+++K L RL+ L+ ++ D Q AFI GR
Sbjct: 641 NHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGR 700
Query: 449 FMLDSVVVANEVVEEAKRRK---KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++A+EV+ K RK K K D KAYD V+W+FL M GF +KWI
Sbjct: 701 IINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIG 760
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI ++S SVL+NGSP RG+RQGDPL+P+LF++ + L+ L+
Sbjct: 761 WIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDL 820
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G R+G I+ LQFADD+LFF +A+ +N +K + +E SG K+N KS +
Sbjct: 821 RGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFG 880
Query: 626 SMVSRETQI-FAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S V TQ +L G YLG+P +K +E +I +++++ S + L
Sbjct: 881 SRVYGSTQSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFL 940
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S G+ ++KSV ++P++ S FK PK I+ + + F W RG+ WV W+ +
Sbjct: 941 SPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRL 1000
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K+EGGLG +DL FN ALL K WRL++ L+ +V++++
Sbjct: 1001 QYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKAR 1044
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 271 bits (694), Expect = 8e-73
Identities = 151/404 (37%), Positives = 229/404 (56%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I +IPK+ NP L +YRPI+L +YK+++K L RL+ L+ ++ D Q AFI GR
Sbjct: 867 NHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGR 926
Query: 449 FMLDSVVVANEVVEEAKRRK---KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++A+EV+ K RK K K D KAYD V+W+FL M GF +KWI
Sbjct: 927 IINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIG 986
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI ++S SVL+NGSP RG+RQGDPL+P+LF++ + L+ L+
Sbjct: 987 WIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDL 1046
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G R+G I+ LQFADD+LFF +A+ +N +K + +E SG K+N KS +
Sbjct: 1047 RGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFG 1106
Query: 626 SMVSRETQI-FAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S V TQ +L G YLG+P +K +E +I +++++ S + L
Sbjct: 1107 SRVYGSTQSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFL 1166
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S G+ ++KSV ++P++ S FK PK I+ + + F W RG+ WV W+ +
Sbjct: 1167 SPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRL 1226
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K+EGGLG +DL FN ALL K WRL++ L+ +V++++
Sbjct: 1227 QYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKAR 1270
>At4g10830 putative protein
Length = 1294
Score = 271 bits (693), Expect = 1e-72
Identities = 149/404 (36%), Positives = 232/404 (56%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I +IPK+ NP+ L +YRPI+L +YKI++K L RL+ L ++ D Q AFI GR
Sbjct: 847 NHTNICMIPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGR 906
Query: 449 FMLDSVVVANEVVEEAKRRKK---ECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++A+E++ K RK+ K D KAYD V+WNFL M GFS WI
Sbjct: 907 LVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIK 966
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI G ++S + SVLVNG P RG+RQGDPL+P+LF++ A+ LN L++ V +
Sbjct: 967 WIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEGDI 1026
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G R+G ++ LQFADD+LFF +++ +N +K + +E SG K+N KS +
Sbjct: 1027 RGIRIGNGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFG 1086
Query: 626 SMVSRETQ-IFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S V TQ +L + G YLG+P +K + +I++++++ S K L
Sbjct: 1087 SRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYL 1146
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S G+ ++KSV S+P++ S FK P I+ + + F W + R + W+ W+ +
Sbjct: 1147 SPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRL 1206
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K+EGGLG +DL FN ALL K WR++ L+ ++++++
Sbjct: 1207 QYSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKAR 1250
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 262 bits (670), Expect = 5e-70
Identities = 144/404 (35%), Positives = 233/404 (57%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I LIPK+ +PQ + + RPISL +YKI++K+L+ RL++ L ++ Q+AF+ R
Sbjct: 463 NHTHICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQR 522
Query: 449 FMLDSVVVANEVVEEAK---RRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+++VA+E++ + R KE FK D KAYD V+W FL MM+ +GF++KWI
Sbjct: 523 LISDNILVAHEMIHSLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWIS 582
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI C+ S S SVL+NG P RG+RQGDPL+P LF++ E L ++ +A Q
Sbjct: 583 WIMNCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKI 642
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA-G 624
TG + +V ++ L FADDTL +A+ Q + L + +SG +N KS + G
Sbjct: 643 TGIQFQDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFG 702
Query: 625 ISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
++ + + + G YLG+P + K + + +K+Q +L+ K+L
Sbjct: 703 KNVDIQIKDWIKSRSGISLEGGTGKYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTL 762
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGG-EEGRGVAWVKWEEI 743
S GG+ L+KS+ +LP++ S FK PK + + + F W ++ R + W+ W+ +
Sbjct: 763 SQGGKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRL 822
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K++GG G KDL+ FN+ALL K WR+L+E+ L+ +V +S+
Sbjct: 823 TLPKDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSR 866
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 257 bits (657), Expect = 2e-68
Identities = 144/406 (35%), Positives = 229/406 (55%), Gaps = 5/406 (1%)
Query: 387 GGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIG 446
G N + I LIPK+ + + ++RPISL +YK++ KL++ RL+++L +I + Q AF+
Sbjct: 483 GMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANRLKKILPSLISETQAAFVK 542
Query: 447 GRFMLDSVVVANEVVEEAKRRKK---ECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKW 503
GR + D++++A+E++ K E K D KAYD V+W FL M +GF+ W
Sbjct: 543 GRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRVEWPFLEKAMRGLGFADHW 602
Query: 504 IHWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQK 563
I I C++S VL+NG+P E RGLRQGDPL+P+LF+I E L +++ A Q+
Sbjct: 603 IRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVICTEMLVKMLQSAEQKN 662
Query: 564 LFTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA 623
TG +V IS L FADD++F+ + + + + + I+ + SG +VN+ KS +
Sbjct: 663 QITGLKVARGAPPISHLLFADDSMFYCKVNDEALGQIIRIIEEYSLASGQRVNYLKSSIY 722
Query: 624 GISMVSRETQ-IFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHK 682
+S E + + L + G YLG+P K AT + ++ +K+ +
Sbjct: 723 FGKHISEERRCLVKRKLGIEREGGEGVYLGLPESFQGSKVATLSYLKDRLGKKVLGWQSN 782
Query: 683 SLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWE 741
LS GG+ L+K+V +LP + S FK PK I Q + +F W +EGRG+ W W
Sbjct: 783 FLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKEGRGLHWKAWC 842
Query: 742 EICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+ + K GGLG K++E+FN ALL K WR++ E++ L +V +S+
Sbjct: 843 HLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAKVFKSR 888
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 257 bits (656), Expect = 2e-68
Identities = 150/407 (36%), Positives = 227/407 (54%), Gaps = 12/407 (2%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + + LI KV P+ + +RP+SL ++KI+ K++ RL+ V+SK+I Q +FI GR
Sbjct: 349 NDALLVLIAKVAKPERIQQFRPVSLCNVLFKIITKMMVTRLKNVISKLIGPAQASFIPGR 408
Query: 449 FMLDSVVVANEVVEEAKRRK--KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHW 506
+D++V+ E V +R+K K + K+D EKAYD V W+FL + G S W
Sbjct: 409 LSIDNIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRVRWDFLQETLEAAGLSEGWTSR 468
Query: 507 IKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFT 566
I + S SVL NG T+ F RGLRQGDPL+P+LF++ E L L+ +V ++ +
Sbjct: 469 IMAGVTDPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGKREWK 528
Query: 567 GYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGIS 626
V ++S + FADD + F EAS I +++ +L F SG KV+ KSK+
Sbjct: 529 PIAVSCGGSKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSH 588
Query: 627 MVSRETQIFAAMLNCKVMGVPFT-----YLGIPVGANPRKAATWEPVIKKMQRKLSL*KH 681
VSRE + L + G+ T YLG+P+ T+ V++++ +L+ K
Sbjct: 589 NVSREME----QLISEESGIGCTKELGKYLGMPILQKRMNKETFGEVLERVSARLAGWKG 644
Query: 682 KSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKW 740
+SLS GRI L K+VLSS+P+ S P +D ++ R FLWG E + + W
Sbjct: 645 RSLSLAGRITLTKAVLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSW 704
Query: 741 EEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+ICK K EGG+G++ NKAL+ K WRLL+++E LW +V+ K
Sbjct: 705 RKICKPKAEGGIGLRSARDMNKALVAKVGWRLLQDKESLWARVVRKK 751
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 256 bits (654), Expect = 3e-68
Identities = 148/390 (37%), Positives = 221/390 (55%), Gaps = 23/390 (5%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I LIPK+ +P+ + +YRPISL YKI++K+L RL++ L VI D Q AF+ G+
Sbjct: 847 NQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKILIKRLKQCLGDVISDSQAAFVPGQ 906
Query: 449 FMLDSVVVANEVVEEAKRRKKEC----FVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWI 504
+ D+V+VA+E++ K R+ EC K D KAYD V+WNFL +M ++GF+ +W+
Sbjct: 907 NISDNVLVAHELLHSLKSRR-ECQSGYVAVKTDISKAYDRVEWNFLEKVMIQLGFAPRWV 965
Query: 505 HWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKL 564
WI C+ S S VL+NGSP + RG+RQGDPL+P+LFL AE L+ ++R+A K
Sbjct: 966 KWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSPYLFLFCAEVLSNMLRKAEVNKQ 1025
Query: 565 FTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA- 623
G ++ D + IS L FADD+LFF AS QNI + I + +E SG K+N+ KS +
Sbjct: 1026 IHGMKITKDCLAISHLLFADDSLFFCRASNQNIEQLALIFKKYEEASGQKINYAKSSIIF 1085
Query: 624 GISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKS 683
G + + Q +L + YLG+P RK +E ++ K++ + +
Sbjct: 1086 GQKIPTMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRKVELFEYIVTKVKERTEGWAYNY 1145
Query: 684 LSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEGRGVAWVKWEEI 743
LS G+ +IK++ +LP++ + F P I ++ N + F WG
Sbjct: 1146 LSPAGKEIVIKAIAMALPVYSMNCFLLPTLICNEINSLITAFWWG--------------- 1190
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLL 773
K+ EG LG KDL FN+ALL K WR+L
Sbjct: 1191 --KENEGDLGFKDLHQFNRALLAKQAWRIL 1218
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 254 bits (650), Expect = 1e-67
Identities = 147/406 (36%), Positives = 222/406 (54%), Gaps = 10/406 (2%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + LI KV P+ + +RPISL ++K + K++ RL+ V++K+I QT+FI GR
Sbjct: 1025 NDVLVVLIAKVLKPEKITQFRPISLCNVLFKTITKVMVGRLKGVINKLIGPAQTSFIPGR 1084
Query: 449 FMLDSVVVANEVVEEAKRRK--KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHW 506
D++VV EVV +R+K K + K+D EKAYD + W+ L + G W+ W
Sbjct: 1085 LSTDNIVVVQEVVHSMRRKKGVKGWMLLKLDLEKAYDRIRWDLLEDTLKAAGLPGTWVQW 1144
Query: 507 IKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFT 566
I C++ S +L NG T+ F RGLRQGDPL+P+LF++ E L L+ ++ K +
Sbjct: 1145 IMKCVEGPSMRLLWNGEKTDAFKPLRGLRQGDPLSPYLFVLCIERLCHLIESSIAAKKWK 1204
Query: 567 GYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGIS 626
++ +S + FADD + F EAS I V++ +L F SG KV+ KSK+
Sbjct: 1205 PIKISQSGPRLSHICFADDLILFAEASIDQIRVLRGVLEKFCGASGQKVSLEKSKIYFSK 1264
Query: 627 MVSRE----TQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHK 682
V R+ + M K +G YLG+P+ T+ VIK+ +L+ K +
Sbjct: 1265 NVLRDLGKRISEESGMKATKDLG---KYLGVPILQKRINKETFGEVIKRFSSRLAGWKGR 1321
Query: 683 SLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKWE 741
LSF GR+ L K+VL+S+ + S K P+ +D +K+ R FLWG E + V W
Sbjct: 1322 MLSFAGRLTLTKAVLTSILVHTMSTIKLPQSTLDGLDKVSRAFLWGSTLEKKKQHLVAWT 1381
Query: 742 EICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+C + EGGLG++ + NKAL+ K WR+L + LW QV+ SK
Sbjct: 1382 RVCLPRREGGLGIRSATAMNKALIAKVGWRVLNDGSSLWAQVVRSK 1427
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 254 bits (650), Expect = 1e-67
Identities = 145/404 (35%), Positives = 225/404 (54%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I LIPK P+ + +YRPI+L YKIVAK+++ R++ +L K+I + Q+AF+ GR
Sbjct: 380 NETHIRLIPKDLGPRKVADYRPIALCNIFYKIVAKIMTKRMQLILPKLISENQSAFVPGR 439
Query: 449 FMLDSVVVANEVVE--EAKRRKKEC-FVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++ +EV+ KK C K D KAYD V+W+FL ++ R GF WI
Sbjct: 440 VISDNVLITHEVLHFLRTSSAKKHCSMAVKTDMSKAYDRVEWDFLKKVLQRFGFHSIWID 499
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
W+ C+ S S S L+NG+P + RGLRQGDPL+P LF++ E L+GL +A + +
Sbjct: 500 WVLECVTSVSYSFLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLSGLCTRAQRLRQL 559
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G RV + ++ L FADDT+FF ++ ++ + IL + SG +NF KS +
Sbjct: 560 PGVRVSINGPRVNHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQSINFHKSSVTFS 619
Query: 626 SMVSRETQ-IFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S R + +L + G YLG+P RK + +I K+++K + L
Sbjct: 620 SKTPRSVKGQVKRILKIRKEGGTGKYLGLPEHFGRRKRDIFGAIIDKIRQKSHSWASRFL 679
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKWEEI 743
S G+ ++K+VL+S+PL+ S FK P + + + +F W + + R +WV W ++
Sbjct: 680 SQAGKQVMLKAVLASMPLYSMSCFKLPSALCRKIQSLLTRFWWDTKPDVRKTSWVAWSKL 739
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K GGLG +D+E N +LL K WRLL E L ++L K
Sbjct: 740 TNPKNAGGLGFRDIERCNDSLLAKLGWRLLNSPESLLSRILLGK 783
>At1g24640 hypothetical protein
Length = 1270
Score = 252 bits (644), Expect = 5e-67
Identities = 141/404 (34%), Positives = 225/404 (54%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + + LIPK ++P + + RPISL +YKI++K+++ RL+ L +++ D Q+AF+ R
Sbjct: 448 NYTHLCLIPKTQHPTEMVDLRPISLCSVLYKIISKIMAKRLQPWLPEIVSDTQSAFVSER 507
Query: 449 FMLDSVVVANEVVEEAK---RRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+++VA+E+V K R E K D KAYD V+W++L ++ +GF KW++
Sbjct: 508 LITDNILVAHELVHSLKVHPRISSEFMAVKSDMSKAYDRVEWSYLRSLLLSLGFHLKWVN 567
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI C+ S + SVL+N P + RGLRQGDPL+PFLF++ EGL L+ +A +
Sbjct: 568 WIMVCVSSVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTHLLNKAQWEGAL 627
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G + + + L FADD+LF +AS + LV++ IL+ + +G +N KS +
Sbjct: 628 EGIQFSENGPMVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQTINLNKSSITFG 687
Query: 626 SMVSRETQ-IFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
V + + L G TYLG+P + K + +++ KL + + L
Sbjct: 688 EKVDEQLKGTIRTCLGIFTEGGAGTYLGLPECFSGSKVDMLHYLKDRLKEKLDVWFTRCL 747
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGG-EEGRGVAWVKWEEI 743
S GG+ L+KSV ++P+F S FK P + F W + R + W WE +
Sbjct: 748 SQGGKEVLLKSVALAMPVFAMSCFKLPITTCENLESAMASFWWDSCDHSRKIHWQSWERL 807
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
C K+ GGLG +D++SFN+ALL K WRLL + L ++L+S+
Sbjct: 808 CLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRLLKSR 851
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 250 bits (638), Expect = 2e-66
Identities = 142/404 (35%), Positives = 227/404 (56%), Gaps = 12/404 (2%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + +TLIPK+ P+ + +YRPI+L YKIVAK+L+ RL+ LS++I Q+AF+ GR
Sbjct: 426 NETHVTLIPKISAPRKVSDYRPIALCNVQYKIVAKILTRRLQPWLSELISLHQSAFVPGR 485
Query: 449 FMLDSVVVANEVVE--EAKRRKKEC-FVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++ +E++ KK C K D KAYD + WNFL ++ R+GF KWI
Sbjct: 486 AIADNVLITHEILHFLRVSGAKKYCSMAIKTDMSKAYDRIKWNFLQEVLMRLGFHDKWIR 545
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
W+ C+ + S S L+NGSP RGLRQGDPL+P+LF++ E L+GL R+A ++ +
Sbjct: 546 WVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGDPLSPYLFILCTEVLSGLCRKAQEKGVM 605
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G RV +++ L FADDT+FF + + + +IL+ +E SG +N KS +
Sbjct: 606 VGIRVARGSPQVNHLLFADDTMFFCKTNPTCCGALSNILKKYELASGQSINLAKSAITFS 665
Query: 626 SMVSRETQIFAAM-LNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S ++ + + L G YLG+P RK + ++ +++++ + L
Sbjct: 666 SKTPQDIKRRVKLSLRIDNEGGIGKYLGLPEHFGRRKRDIFSSIVDRIRQRSHSWSIRFL 725
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKWEEI 743
S G+ L+K+VLSS+P + FK P + Q + +F W + + R +AWV W+++
Sbjct: 726 SSAGKQILLKAVLSSMPSYAMMCFKLPASLCKQIQSVLTRFWWDSKPDKRKMAWVSWDKL 785
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
EGGLG +++E+ K WR+L+E L +VL K
Sbjct: 786 TLPINEGGLGFREIEA-------KLSWRILKEPHSLLSRVLLGK 822
>At2g11240 pseudogene
Length = 1044
Score = 246 bits (628), Expect = 4e-65
Identities = 142/411 (34%), Positives = 230/411 (55%), Gaps = 19/411 (4%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + ITLIPK+++ + + +YRPI+L YKI++KLLS RL+ +L ++I + Q+AF+ R
Sbjct: 311 NKTHITLIPKIQSLKRMVDYRPIALCTVFYKIISKLLSRRLQPILQEIISENQSAFVPKR 370
Query: 449 FMLDSVVVANEVVEEAKR--RKKECFV-FKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
D+V++ +E + K +K CF+ K + KAYD ++W+F+ +M +GF WI
Sbjct: 371 ASNDNVLITHEALHYLKSLGAEKRCFMAVKTNMSKAYDRIEWDFIKLVMQEMGFHQTWIS 430
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI CI + S S L+NGS RGLRQGDPL+PFLF+I +E L+GL R+A
Sbjct: 431 WILQCITTVSYSFLLNGSAQGAVTPERGLRQGDPLSPFLFIICSEVLSGLCRKAQLDGSL 490
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G RV ++ L FADDT+FF + ++ IL+ +E SG +N KS +
Sbjct: 491 LGLRVSKGNPRVNHLLFADDTIFFCRSDLKSCKTFLCILKKYEEASGQMINKSKSAIT-- 548
Query: 626 SMVSRETQIFAAMLNCKVMGVPFT-----YLGIPVGANPRKAATWEPVIKKMQRKLSL*K 680
SR+T +++G+ YLG+P +K + ++ +++++
Sbjct: 549 --FSRKTPDHIKTEAQQILGIQLVGGLGKYLGLPKMFGRKKRDLFNQIVDRIRQRSLSWS 606
Query: 681 HKSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQ---RKFLW-GGEEGRGVA 736
+ LS G+ ++KSVL+S+P + S F K ++ C +IQ F W + + +
Sbjct: 607 SRFLSTAGKTTMLKSVLASMPTYTMSCF---KLLVSLCKRIQSALTHFWWDSSADKKKMC 663
Query: 737 WVKWEEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
W+ W ++ K K+EGGLG KD+ +FN ALL K WR+++ + ++L K
Sbjct: 664 WIAWSKMAKNKKEGGLGFKDITNFNDALLAKLSWRIVQSPSCVLVRILLGK 714
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 243 bits (619), Expect = 4e-64
Identities = 139/400 (34%), Positives = 235/400 (58%), Gaps = 9/400 (2%)
Query: 386 RGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFI 445
+G N++ + LIPK + + + +YRPIS +YK ++K+L+ RL+ +L K I Q+AF+
Sbjct: 228 KGVNSTILALIPKKKEAREIKDYRPISCCNVLYKAISKILANRLKRILPKFIVGNQSAFV 287
Query: 446 GGRFMLDSVVVANEVVEEAKRRK--KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKW 503
R ++++V++A E+V++ + C K+D KA+DS+ W+FL ++++ + F ++
Sbjct: 288 KDRLLIENVLLATELVKDYHKDSISTRC-AMKIDISKAFDSLQWSFLTHVLAAMNFPGEF 346
Query: 504 IHWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQK 563
IHWI C+ +AS S+ VNG F RGLRQG L+P+LF+I + L+ ++ +A +
Sbjct: 347 IHWISLCMSTASFSIQVNGELAGYFRSARGLRQGCSLSPYLFVISMDVLSRMLDKAAGAR 406
Query: 564 LFTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKL- 622
F GY + ++ L FADD + + +++ + +L F A GLK+ K+ L
Sbjct: 407 EF-GYHPRCKTLGLTHLCFADDLMILTDGKIRSVDGIVKVLNQFAAKLGLKICMEKTTLY 465
Query: 623 -AGISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KH 681
AG+S SR Q+ ++ + V +P YLG+P+ + + P+I +++R++ +
Sbjct: 466 LAGVSDHSR--QLMSSRYSFGVGKLPVRYLGLPLVTKRLTTSDYSPLIDQIRRRIGMWTS 523
Query: 682 KSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEE-GRGVAWVKW 740
+ LSF GR+ LI SVL S+ F+ + F+ P+ I++ N+I LW G E A V W
Sbjct: 524 RYLSFAGRLSLINSVLWSITNFWMNAFRLPRECINEINRISSALLWSGPELNPKKAKVSW 583
Query: 741 EEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLW 780
+EICK K+EGGLG++ L NK +K WRLL ++ LW
Sbjct: 584 DEICKPKKEGGLGLQSLREANKVSSLKLIWRLLSCQDSLW 623
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 243 bits (619), Expect = 4e-64
Identities = 142/399 (35%), Positives = 232/399 (57%), Gaps = 7/399 (1%)
Query: 386 RGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFI 445
+G N + + LI K G+ +YRPIS +YKIV+KL++ RL+E+L I Q+AFI
Sbjct: 654 KGINTTILALISKKHEVSGMKDYRPISCCNVLYKIVSKLMANRLKEILPASIAPNQSAFI 713
Query: 446 GGRFMLDSVVVANEVVEEAKRRK-KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWI 504
R M++++++A+E+V++ + K+D KA+D V W FL ++ I +I
Sbjct: 714 KDRLMMENLLLASELVKDYHKESISSRSALKIDISKAFDFVQWPFLINVLKAIHLPEMFI 773
Query: 505 HWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKL 564
HWI+ CI +AS SV VNG + F RGLRQG L+P+L++I L+ ++ +A +K
Sbjct: 774 HWIELCIGTASFSVQVNGELSGFFRSERGLRQGCSLSPYLYVICMNVLSCMLDKAAVEKK 833
Query: 565 FTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSK--L 622
+ Y + ++ L FADD + F + ++++I +I F AMS LK++ KS +
Sbjct: 834 IS-YHPRCRNMNLTHLCFADDIMVFSDGTSKSIQGTLAIFEKFAAMSWLKISLEKSTIFM 892
Query: 623 AGISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHK 682
AGIS ++ + ++ +P YLG+P+ + + P+++K++ +++ ++
Sbjct: 893 AGISPNAKTS--ILQQFPFELGTLPVKYLGLPLLTKRMTQSDYLPLVEKIRARITSWTNR 950
Query: 683 SLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEE-GRGVAWVKWE 741
LSF GR+ LIKSVLSS+ F+ S F+ PK + + K+ FLW G + A + W
Sbjct: 951 FLSFAGRLQLIKSVLSSITNFWLSVFRLPKACLQEIEKMFSAFLWSGPDLNTKKAKIAWS 1010
Query: 742 EICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLW 780
E+CK KEEGGLG+K L+ N+ L+K WR+L R+ LW
Sbjct: 1011 EVCKLKEEGGLGLKPLKEANEVSLLKLIWRILSARDSLW 1049
>At3g25485 unknown protein
Length = 979
Score = 242 bits (618), Expect = 5e-64
Identities = 141/405 (34%), Positives = 233/405 (56%), Gaps = 9/405 (2%)
Query: 386 RGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFI 445
+G N++ + LIPK + + +YRPIS +YK+++KL++ RL+E+L + I Q+AF+
Sbjct: 423 KGVNSTILALIPKKFETKEMKDYRPISCCNVIYKVISKLIAKRLKEILPQFIAGNQSAFV 482
Query: 446 GGRFMLDSVVVANEVVEEA-KRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWI 504
R ++ ++++A E+V++ K + K+D KA+DSV W+FL ++ + FS ++I
Sbjct: 483 KDRLLIQNLLLATEIVKDYHKESVSDRCAIKIDISKAFDSVQWSFLENVLHSLNFSQEFI 542
Query: 505 HWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKL 564
HWI CI +AS SV VNG FN RGLRQG L+P+LF+I + L+ ++ +A K
Sbjct: 543 HWIMLCITTASFSVQVNGELVGFFNSSRGLRQGCSLSPYLFVIAMDVLSKMLDRAAGFKK 602
Query: 565 FTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSK--L 622
F GY + ++ L FADD + + +++ + S+ F SGLK++ KS L
Sbjct: 603 F-GYHPRCQTIGLTHLTFADDLMVLSDGKVRSVEGIVSVFDEFAKKSGLKISMEKSTIYL 661
Query: 623 AG-ISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KH 681
AG +M RE + V +P YLG+P+ +A + P++++++RK+
Sbjct: 662 AGSANMYRREIE---ERFKFAVGSLPVRYLGLPLVTKRFSSADYRPLVEQIKRKIGTWTA 718
Query: 682 KSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEE-GRGVAWVKW 740
+ LS+ GR+ LI SV+ S+ F+ + F+ P+ I + +K+ FLW G E A V W
Sbjct: 719 RYLSYAGRLNLISSVIWSICNFWIAAFRLPRECIREIDKLCSAFLWSGPELNSKRAKVNW 778
Query: 741 EEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLE 785
E ICK K EGGLG++ L+ N +K WR + + LW ++L+
Sbjct: 779 EAICKPKREGGLGLRSLKEANDVSCLKLIWRWVSRGDSLWRKLLK 823
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 242 bits (618), Expect = 5e-64
Identities = 134/404 (33%), Positives = 219/404 (54%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + + LIPK NPQ + + RPISL +YKI++K+LS +L++ L ++ Q+AF R
Sbjct: 437 NHTHLYLIPKFTNPQRMSDIRPISLCSVLYKIISKILSFKLKKHLPSIVSPSQSAFFAER 496
Query: 449 FMLDSVVVANEVVEEAKRR---KKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D++++A+E+V + KE VFK D KAYD V+W+FL ++ +GF+ KWI
Sbjct: 497 LISDNILIAHEIVHSLRTNDKISKEFMVFKTDMSKAYDRVEWSFLQEILVALGFNDKWIS 556
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI GC+ S + SVL+NG RG+RQGDP++PFLF++ E L +++QA K
Sbjct: 557 WIMGCVTSVTYSVLINGQHFGHITPERGIRQGDPISPFLFVLCTEALIHILQQAENSKKV 616
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA-G 624
+G + G ++ L F DDT A+ + + L + +SG +N KS + G
Sbjct: 617 SGIQFNGSGPSVNHLLFVDDTQLVCRATKSDCEQMMLCLSQYGHISGQLINVEKSSITFG 676
Query: 625 ISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
+ + + + G YLG+P + K + + +K+Q LS K+L
Sbjct: 677 VKVDEDTKRWIKNRSGIHLEGGTGKYLGLPENLSGSKQDLFGYIKEKLQSHLSGWYDKTL 736
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEE-GRGVAWVKWEEI 743
S GG+ L+KS+ +LP++ + F+ PK + + + F W E + W+ +++
Sbjct: 737 SQGGKEILLKSIALALPVYIMTCFRLPKGLCTKLTSVMMDFWWNSMEFSNKIHWIGGKKL 796
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K GG G KDL+ FN+ALL K WRL + + + Q+ +S+
Sbjct: 797 TLPKSLGGFGFKDLQCFNQALLAKQAWRLFSDSKSIVSQIFKSR 840
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 238 bits (606), Expect = 1e-62
Identities = 136/404 (33%), Positives = 222/404 (54%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + + LIPK+ P + + RPISL +YKI++K+LS RL++ L ++ Q+AF+ R
Sbjct: 214 NHTHLCLIPKITKPARMADIRPISLCSVMYKIISKILSARLKKYLPVIVSPTQSAFVAER 273
Query: 449 FMLDSVVVANEVVEEAKRRKK---ECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D++++A+E+V + +K + VFK D KAYD V+W FL ++ +GF+ WI+
Sbjct: 274 LVSDNIILAHEIVHNLRTNEKISKDFMVFKTDMSKAYDRVEWPFLKGILLALGFNSTWIN 333
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
W+ C+ S S SVL+NG P RGLRQGDPL+PFLF++ E L ++ QA +
Sbjct: 334 WMMACVSSVSYSVLINGQPFGHITPHRGLRQGDPLSPFLFVLCTEALIHILNQAEKIGKI 393
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
+G + G ++ L FADDTL +AS + L + +SG +N KS +
Sbjct: 394 SGIQFNGTGPSVNHLLFADDTLLICKASQLECAEIMHCLSQYGHISGQMINSEKSAITFG 453
Query: 626 SMVSRET-QIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
+ V+ ET Q + G YLG+P K + + +K+Q +LS K+L
Sbjct: 454 AKVNEETKQWIMNRSGIQTEGGTGKYLGLPECFQGSKQVLFGFIKEKLQSRLSGWYAKTL 513
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGG-EEGRGVAWVKWEEI 743
S GG+ L+KS+ + P++ + F+ K + + + F W ++ + + W+ +++
Sbjct: 514 SQGGKDILLKSIAMAFPVYAMTCFRLSKTLCTKLTSVMMDFWWNSVQDKKKIHWIGAQKL 573
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K GG G KDL+ FN+ALL K RL + + L Q+L+S+
Sbjct: 574 MLPKFLGGFGFKDLQCFNQALLAKQASRLHTDSDSLLSQILKSR 617
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 236 bits (603), Expect = 3e-62
Identities = 140/403 (34%), Positives = 215/403 (52%), Gaps = 21/403 (5%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + L+ KV P+ + +RP+SL ++KI+ K++ +RL+ V+SK+I Q +FI GR
Sbjct: 301 NDVLLVLLAKVAKPERITQFRPVSLCNVLFKIITKMMVIRLKNVISKLIGPAQASFIPGR 360
Query: 449 FMLDSVVVANEVVEEAKRRK--KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHW 506
D++VV E V +R+K K + K+D EKAYD + W+FL + G S WI
Sbjct: 361 LSFDNIVVVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRIRWDFLAETLEAAGLSEGWIKR 420
Query: 507 IKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFT 566
I C+ S+L NG T+ F RGLRQGDP++P+LF++ E L + AV + +
Sbjct: 421 IMECVAGPEMSLLWNGEKTDSFTPERGLRQGDPISPYLFVLCIERLCHQIETAVGRGDWK 480
Query: 567 GYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGIS 626
+ ++S + FADD + F EAS KV+ KSK+ +
Sbjct: 481 SISISQGGPKVSHVCFADDLILFAEASVAQ-----------------KVSLEKSKIFFSN 523
Query: 627 MVSRETQ-IFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLS 685
VSR+ + + A YLG+PV T+ V++++ +LS K +SLS
Sbjct: 524 NVSRDLEGLITAETGIGSTRELGKYLGMPVLQKRINKDTFGEVLERVSSRLSGWKSRSLS 583
Query: 686 FGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKWEEIC 744
GRI L K+VL S+P+ S P +++Q +K+ R FLWG E R + W+++C
Sbjct: 584 LAGRITLTKAVLMSIPIHTMSSILLPASLLEQLDKVSRNFLWGSTVEKRKQHLLSWKKVC 643
Query: 745 KKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+ K GGLG++ + N+ALL K WRLL ++ LW +VL K
Sbjct: 644 RPKAAGGLGLRASKDMNRALLAKVGWRLLNDKVSLWARVLRRK 686
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.341 0.150 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,355,066
Number of Sequences: 26719
Number of extensions: 660742
Number of successful extensions: 2124
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 65
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 1855
Number of HSP's gapped (non-prelim): 115
length of query: 787
length of database: 11,318,596
effective HSP length: 107
effective length of query: 680
effective length of database: 8,459,663
effective search space: 5752570840
effective search space used: 5752570840
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0263.7


