

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0263.5
(295 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
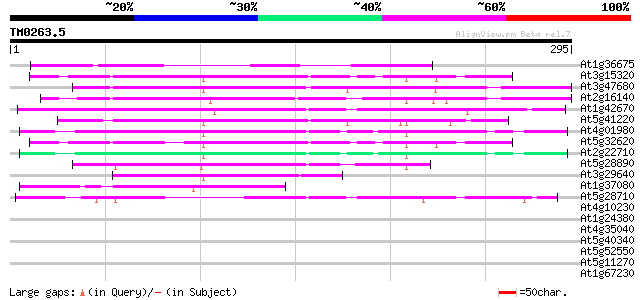
Sequences producing significant alignments: (bits) Value
At1g36675 putative protein 126 2e-29
At3g15320 hypothetical protein 98 6e-21
At3g47680 unknown protein 97 1e-20
At2g16140 pseudogene; similar to MURA transposase of maize Muta... 91 7e-19
At1g42670 unknown protein 91 7e-19
At5g41220 glutathione transferase-like 90 2e-18
At4g01980 hypothetical protein 79 4e-15
At5g32620 putative protein 77 1e-14
At2g22710 hypothetical protein 74 7e-14
At5g28890 putative protein 66 3e-11
At3g29640 hypothetical protein 65 4e-11
At1g37080 hypothetical protein 50 1e-06
At5g28710 putative protein 48 7e-06
At4g10230 hypothetical protein 39 0.003
At1g24380 hypothetical protein 39 0.003
At4g35040 bZip transcription factor AtbZip19 35 0.037
At5g40340 unknown protein 35 0.048
At5g52550 unknown protein 34 0.082
At5g11270 unknown protein 34 0.11
At1g67230 unknown protein 33 0.14
>At1g36675 putative protein
Length = 268
Score = 126 bits (316), Expect = 2e-29
Identities = 80/211 (37%), Positives = 100/211 (46%), Gaps = 73/211 (34%)
Query: 12 NVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYG 71
NVG T+FPEFST M LG M+ V+E PNEED T ++S W T+QNLVL+S WIKY
Sbjct: 37 NVGATEFPEFSTQMALGVMSGVHETIPNEEDLT--CNHSRSSKWTTDQNLVLLSRWIKYE 94
Query: 72 TSSVVGRNQKGETYWSQIAEYCHEHCSFDPPREGGACRNCFNYMSNQLNKWIGAYDGAKR 131
+ +K D KR
Sbjct: 95 QIVSLVETKK---------------------------------------------DNVKR 109
Query: 132 VQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSGSKR 191
+Q SGWS+D +LAKA ++Y+N GN GSGSSGSKR
Sbjct: 110 MQQSGWSKDGVLAKAHELYSNA--------------------------GNTGSGSSGSKR 143
Query: 192 SHESDACGSNSIGSIPRPMGREAAKKKNKKK 222
+HESD +NS+GS RP+G + AKKK KKK
Sbjct: 144 THESDVRDANSVGSSARPIGSDIAKKKTKKK 174
>At3g15320 hypothetical protein
Length = 287
Score = 97.8 bits (242), Expect = 6e-21
Identities = 74/280 (26%), Positives = 127/280 (44%), Gaps = 43/280 (15%)
Query: 11 FNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKY 70
F G T+ P FS+ ++ +++ +E++ P+ ++ W ++++VL+S W+
Sbjct: 4 FASGSTEVPAFSSQLS----EEGSDLEGSEDEVKPKQSISRKK-WTAKEDIVLVSAWLNT 58
Query: 71 GTSSVVGRNQKGETYWSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWIGAYDG 128
V+G +Q+G+++W +IA Y S D P RE C++ + ++ + K++G Y
Sbjct: 59 SKDPVIGNDQQGQSFWKRIAAYVAASPSLDGLPKREHAKCKHRWGKVNKSVTKFVGCYKT 118
Query: 129 AKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSG 188
+ SG SEDD++ A +IY N + FTL W L + +C G
Sbjct: 119 TTTHKTSGQSEDDVMKLAYEIYFNDTKKN-FTLDHAWRELRYDQKWCEATS---RKGDKN 174
Query: 189 SKRSHESDACGSNSIGSIP----------RPMGREAAKKKNKKKS--------------R 224
+KR CG + S P RP G +AAK K +K +
Sbjct: 175 AKRK----KCGDGNASSQPIHVEDDSVMSRPPGVKAAKAKARKSATIKEGKKPATVKDDS 230
Query: 225 DAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEK 264
++E N W +L+E++ +R EK S QQ L K
Sbjct: 231 GQSVEHFQNLW----ELKEKDWDRKEKQSKHQQLERLLSK 266
>At3g47680 unknown protein
Length = 302
Score = 96.7 bits (239), Expect = 1e-20
Identities = 75/282 (26%), Positives = 130/282 (45%), Gaps = 30/282 (10%)
Query: 34 NEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEYC 93
++V+PN+ TP+ ++ + W+ ++LVL+S W+ +V+G QKG +WS+IA Y
Sbjct: 30 DDVSPNQAAHTPKVKRERRK-WSAGEDLVLVSAWLNTSKDAVIGNEQKGYAFWSRIAAYY 88
Query: 94 HEHCSFD--PPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYT 151
+ RE G + + ++ + K++G+Y+ A + + SG ++DD++A A +IY
Sbjct: 89 GASPKLNGVEKRETGHIKQRWTKINEGVGKFVGSYEAATKQKSSGQNDDDVVALAHEIYN 148
Query: 152 NGKNSHPFTLMKEWLALCNEPHYCS--QVGGNLGSGSSGSKRSHESDACGSNSIGSIPRP 209
+ FTL W L E + S + S S ++ S + ++ RP
Sbjct: 149 SEHGK--FTLEHAWRVLRFEQKWLSAPSTKATVMSKRRKSDKASTSQPQTHEAEEAMSRP 206
Query: 210 MGREAAKKKNKKK---------------SRDAALEVVNNEWSEYKQLREQELERLEKISS 254
+G +AAK K KK + E+ +W ++ REQE E EK
Sbjct: 207 IGVKAAKAKAKKAVSKTTTVEDKGNVMLEIQSIWEIKQKDWELRQKDREQEKEDFEK--- 263
Query: 255 VQQEANRLEKMKMFLQL-SFEEHLDDRRKELLKQLSQELFEN 295
RL + K+ L + +E L D L +L E+ N
Sbjct: 264 ----QERLSRTKLLESLFTKKEPLTDIEVALKNKLINEMLSN 301
>At2g16140 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 311
Score = 90.9 bits (224), Expect = 7e-19
Identities = 78/306 (25%), Positives = 139/306 (44%), Gaps = 53/306 (17%)
Query: 17 DFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVV 76
D P FST T +P+ E+ P+ ++ + W+ ++++VL+S W+ +V+
Sbjct: 32 DVPIFSTQCT---------DSPSLEEHVPKVKRERMK-WSAKEDMVLVSSWLNTSKDAVI 81
Query: 77 GRNQKGETYWSQIAEYCHEHCSFDPPRE--GGACRNCFNYMSNQLNKWIGAYDGAKRVQG 134
G QK T+WS+IA Y + R+ G + + +++ + K++G+Y+ A R +
Sbjct: 82 GNEQKANTFWSRIAAYYDASPQLNGLRKRMQGNIKQRWAKINDGVCKFVGSYEAASREKS 141
Query: 135 SGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSGSKRSHE 194
SG +++D+++ A +I+ N + F L W L ++ +CSQ +S + +
Sbjct: 142 SGQNDNDVISLAHEIF-NNDYGYKFPLEHAWRVLRHDQKWCSQ--------ASVMSKRRK 192
Query: 195 SDACGSNSIGSIP---------RPMGREAAKKKNKK--------KSRDAAL-------EV 230
D S P RP+G +AAK K KK + + A+ E+
Sbjct: 193 CDKAAQPSTSQPPSHGVEEAMSRPIGVKAAKAKAKKTVTKTTTVEDKGNAMLEIQSIWEI 252
Query: 231 VNNEWSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQL-SFEEHLDDRRKELLKQLS 289
+W ++ REQE E EK +RL K + L + +E L D L +L
Sbjct: 253 KQKDWELRQKDREQEKEDFEK-------KDRLSKTTLLESLIAKKEPLTDNEVTLKNKLI 305
Query: 290 QELFEN 295
L E+
Sbjct: 306 DFLLEH 311
>At1g42670 unknown protein
Length = 579
Score = 90.9 bits (224), Expect = 7e-19
Identities = 71/296 (23%), Positives = 132/296 (43%), Gaps = 16/296 (5%)
Query: 5 PSQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLI 64
P+ F +G + P +ST + D + ++ + K W++ +++VLI
Sbjct: 290 PNPYECFELGSANVPVYSTEWSDDDPSEDELPIAGKKGKKGKKAKKPRRNWSSTEDVVLI 349
Query: 65 SGWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFDPPREGG--ACRNCFNYMSNQLNKW 122
SGW+ VVG QKG +W +IA Y + + G C+ ++ +++ +NK+
Sbjct: 350 SGWLNTSKDPVVGNEQKGAAFWERIAVYYNSSPKLKGVEKRGHICCKQRWSKVNDAVNKF 409
Query: 123 IGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNL 182
+G+Y A + Q SG ++DD+++ A I++ FT W + + +Q
Sbjct: 410 VGSYLAASKQQTSGQNDDDVVSLAHQIFSKDYGC-KFTCEHAWRERRYDQKWIAQ----- 463
Query: 183 GSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAALEVVNNEWSEYK--- 239
+ +R E+D+ RP+G +AAK K K + V E + K
Sbjct: 464 STHGKAKRRKCEADSEPVGVEDKEARPIGVKAAKAAAKAKGKAKLSPVEGEETNALKEIQ 523
Query: 240 ---QLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQLSQEL 292
+++E++ EK+ ++ + NR + + L + E L D EL +L EL
Sbjct: 524 SIWEIKEKDHAAKEKLIIIKDKKNRTKFFERLLGKT--EPLSDIEIELKNKLINEL 577
>At5g41220 glutathione transferase-like
Length = 590
Score = 89.7 bits (221), Expect = 2e-18
Identities = 68/262 (25%), Positives = 120/262 (44%), Gaps = 33/262 (12%)
Query: 26 TLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVVGRNQKGETY 85
T+G+ + + N D RK W+ ++ +LIS W+ +V K +
Sbjct: 249 TIGEKSDDPNLVQNTTDRRKHRRK-----WSRAEDAILISAWLNTSKDPIVDNEHKACAF 303
Query: 86 WSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDIL 143
W +I Y + S P RE C+ ++ +++++ K++G YD A + SG SEDD+
Sbjct: 304 WKRIGAYFNNSASLANLPKREPSHCKQRWSKLNDKVCKFVGCYDQALNQRSSGQSEDDVF 363
Query: 144 AKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCS--QVGGNLGSGSSGSKRSHESDACGSN 201
A +YTN S+ FTL W L + +CS + G GSS + + D S+
Sbjct: 364 QVAYQVYTNNYKSN-FTLEHAWRELRHSKKWCSLYPFENSKGGGSSKRTKLNNGDRVYSS 422
Query: 202 SIG--SIP-----------RPMGREAAKKKNKKKSRDAALEV--------VNNEWSEYKQ 240
S S+P P+G +++K+K KK + +E + N W ++
Sbjct: 423 SSNPESVPIALDEEEQVMDLPLGVKSSKQKEKKVATIITIEEREADSGSRLENLWVLDEE 482
Query: 241 LREQELERLEKISSVQQEANRL 262
EQ ++R + S++Q+ N++
Sbjct: 483 --EQVMDRPLGVKSLEQKENKV 502
>At4g01980 hypothetical protein
Length = 302
Score = 78.6 bits (192), Expect = 4e-15
Identities = 76/301 (25%), Positives = 126/301 (41%), Gaps = 38/301 (12%)
Query: 6 SQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLIS 65
++T + VG ++ P F T + E+ W ++++VLIS
Sbjct: 26 NETNQYEVGSSEIPVFGTQ----------RCQESHEERYASRVAIARKKWAAKEDIVLIS 75
Query: 66 GWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWI 123
W+ VVG QK +W +IA Y + D P R C+ + M+ + K++
Sbjct: 76 AWLNTSKDPVVGNEQKAPAFWKRIASYVAANPDLDGVPKRASAQCKQRWAKMNELVMKFV 135
Query: 124 GAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLG 183
G Y + SG +E+D++ A +++ N F+L W L ++ + + N
Sbjct: 136 GCYATTTNQKASGQTENDVMLFANELFENDMKK-KFSLDHAWRLLRHDQKW---LISNAP 191
Query: 184 SGSSGSKRSHESDACGSNSIGSIP----------RPMGREAAKKKNKKKSRDAALEVVNN 233
G +KR G+ + S P RP G +AAK K KK EV +
Sbjct: 192 KGKGIAKR--RKVRVGTQAASSQPIDLEDDDVMRRPPGVKAAKAKAKKTPTVKGEEVNSV 249
Query: 234 E-WSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQLSQEL 292
E + L+E++ +R +K Q + LE + LS E L + L K+L +EL
Sbjct: 250 EHLTSLWDLKERDWDRKDK----QSKNQMLESL-----LSRTEPLSECDLVLKKKLIEEL 300
Query: 293 F 293
F
Sbjct: 301 F 301
>At5g32620 putative protein
Length = 301
Score = 76.6 bits (187), Expect = 1e-14
Identities = 70/280 (25%), Positives = 117/280 (41%), Gaps = 52/280 (18%)
Query: 11 FNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKY 70
F G T+ P FS+ ++ +E+ +E++ P+ ++ W ++++VL+S W+
Sbjct: 27 FASGSTEVPAFSSQLS----EEGSELEGSEDEVKPKQSISRKK-WTAKEDIVLVSAWLNP 81
Query: 71 GTSSVVGRNQKGETYWSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWIGAYDG 128
V+G +Q+ Y S D P RE C+ + ++ + K++ Y
Sbjct: 82 SKDPVIGNDQQA---------YVAASPSLDGLPKREHAKCKQRWGKVNKSVTKFVACYKT 132
Query: 129 AKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSG 188
+ SG SEDD++ A +IY N + FTL W L + +C G
Sbjct: 133 TTTHKTSGQSEDDVMKLAYEIYFNDTKKN-FTLDHAWRELRYDQKWCEATS---RKGDEN 188
Query: 189 SKRSHESDACGSNSIGSIP----------RPMGREAAKKKNKKKS--------------R 224
+KR CG + S P RP G +AAK K +K +
Sbjct: 189 AKRR----KCGDGNASSHPIHVEDDSIMSRPPGVKAAKAKGRKSAIVKEGKKPATVKDDS 244
Query: 225 DAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEK 264
++E N W +L+E++ +R EK S QQ L K
Sbjct: 245 GQSVEHFQNLW----ELKEKDWDRKEKQSKHQQLERLLSK 280
>At2g22710 hypothetical protein
Length = 300
Score = 74.3 bits (181), Expect = 7e-14
Identities = 75/301 (24%), Positives = 123/301 (39%), Gaps = 38/301 (12%)
Query: 6 SQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLIS 65
++T + VG ++ P F T + E W ++++VLIS
Sbjct: 24 NETNQYEVGSSEIPVFGTQ----------RCQESHEKRYASRVAIARKKWAAKEDIVLIS 73
Query: 66 GWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFD--PPREGGACRNCFNYMSNQLNKWI 123
W+ VVG QK +W +IA Y D P R C+ + M+ + K++
Sbjct: 74 AWLNTSKDPVVGNEQKAPAFWKRIASYVAASPDLDGVPKRASAQCKQRWAKMNELVMKFV 133
Query: 124 GAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLG 183
G Y + SG +E+D++ A +++ N F+L W L ++ + + N
Sbjct: 134 GCYATTTNQKASGQTENDVMLFANELFKNDMKK-KFSLDHAWRLLRHDQKW---LISNAP 189
Query: 184 SGSSGSKRSHESDACGSNSIGSIP----------RPMGREAAKKKNKKKSRDAALEVVNN 233
G +KR G+ + S P RP G +AAK K K EV +
Sbjct: 190 KGKGIAKR--RKVRVGTQAASSQPIDLEDDDVMRRPPGVKAAKAKAKMTPTVKGEEVNSV 247
Query: 234 E-WSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQLSQEL 292
E + L+E++ +R +K Q + LE + LS E L + L K+L +EL
Sbjct: 248 EHLTSLWDLKEKDWDRKDK----QSKNQMLESL-----LSRTEPLSECDLVLKKKLIEEL 298
Query: 293 F 293
F
Sbjct: 299 F 299
>At5g28890 putative protein
Length = 232
Score = 65.9 bits (159), Expect = 3e-11
Identities = 51/205 (24%), Positives = 84/205 (40%), Gaps = 25/205 (12%)
Query: 34 NEVTPNEEDSTPRSRKTQSPA-----WNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQ 88
NE P D T A W+ +++++LIS W+ VVG QK +W +
Sbjct: 21 NENNPQLSDEDHEDVNTPKVAIPKKKWSAKEDVILISAWLNTSKDPVVGNEQKAPAFWKR 80
Query: 89 IAEYCHEHCSF--DPPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKA 146
IA Y P RE C+ + M+ + K++G Y + SG +E+D++ A
Sbjct: 81 IATYVAASPDLVGFPKRESAQCKQRWAKMNELVMKFVGCYATETNQKASGQTENDVMLFA 140
Query: 147 QDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSGSKRSHESDACGSNSIGSI 206
+++ N F+L W + +E + + S + KR + GS + S
Sbjct: 141 NELFENDMKK-KFSLDHAWRLVRHEQKW-------IISNTPKEKRMSKRRKVGSQAKSSQ 192
Query: 207 P----------RPMGREAAKKKNKK 221
P RP+ + K K KK
Sbjct: 193 PINLEDYDVMARPLRVKVPKAKTKK 217
>At3g29640 hypothetical protein
Length = 171
Score = 65.1 bits (157), Expect = 4e-11
Identities = 35/123 (28%), Positives = 61/123 (49%), Gaps = 3/123 (2%)
Query: 55 WNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFD--PPREGGACRNCF 112
W ++++VL+ W+ V+G +Q+ +++ QIA Y D P RE C+ +
Sbjct: 31 WFPKEDIVLVRAWLNTNKDHVIGNDQQCQSFLKQIASYVAISPQLDYLPKREHAKCKQRW 90
Query: 113 NYMSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEP 172
+ ++ + K++G Y A + SG SEDD++ A +IY N F L W L +
Sbjct: 91 SKVNKSVTKFVGCYKTATTHKTSGQSEDDVMKLAYEIYYN-DTKKKFNLEHTWRKLRYDQ 149
Query: 173 HYC 175
+C
Sbjct: 150 KWC 152
>At1g37080 hypothetical protein
Length = 224
Score = 50.4 bits (119), Expect = 1e-06
Identities = 32/142 (22%), Positives = 67/142 (46%), Gaps = 10/142 (7%)
Query: 6 SQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLIS 65
+ +P +G ++F F+T + N T + +T R++ W +++++VLIS
Sbjct: 30 THSPSIKLGESEFLVFNTQWHESHLQGGNTTT--KRTTTTRNK------WTSKEDIVLIS 81
Query: 66 GWIKYGTSSVVGRNQKGETYWSQIAEYCHE--HCSFDPPREGGACRNCFNYMSNQLNKWI 123
W+ SVVG Q+ + +W ++ Y + S RE + + ++ + +
Sbjct: 82 TWLNTSKDSVVGNEQRADAFWKRVVVYFASTLNVSSQIKREPSHSKQRWGKINKIVCMFG 141
Query: 124 GAYDGAKRVQGSGWSEDDILAK 145
G+++ A SG +ED ++ K
Sbjct: 142 GSHEAANTQMASGMNEDVLMNK 163
>At5g28710 putative protein
Length = 264
Score = 47.8 bits (112), Expect = 7e-06
Identities = 62/297 (20%), Positives = 122/297 (40%), Gaps = 73/297 (24%)
Query: 4 SPSQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDST---PRSRKTQSPA--WNTE 58
SP+ +G T+ P FST + D P+E+++ + +K + P W++
Sbjct: 27 SPNPYECLELGSTNVPVFSTEWSDDD--------PSEDEALIAGKKGKKAKKPRRNWSSI 78
Query: 59 QNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFDPPREGGACRNCFNYMSNQ 118
+++VLI GW+ +VG QKG
Sbjct: 79 EDIVLICGWLNTSKDPMVGNEQKG------------------------------------ 102
Query: 119 LNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQV 178
AY A + Q SG ++D +++ A I++ FT W L + + +Q
Sbjct: 103 -----AAYLTASKQQTSGQNDDFVVSLAHQIFSKDYGC-KFTCEHAWRELRYDQKWIAQ- 155
Query: 179 GGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAK--KKNKKKSRDA-ALEVVNNEW 235
+ +R E+D+ RP+ +AAK K++ + + AL+ + + W
Sbjct: 156 ----STHGKAKRRKCEADSDSVGVEDKEARPISVKAAKAAKQSPVEGEETNALKEIQSIW 211
Query: 236 SEYKQLREQELERLEKISSVQQEANRLEKMKMFL----QLSFEEHLDDRRKELLKQL 288
++++++ EK+ ++++ NR + ++ L LS E D +K+L+ +L
Sbjct: 212 ----EIKDKDHAAKEKLIIIKEKKNRTKLLERLLGKTKPLSIIE--IDLKKKLINEL 262
>At4g10230 hypothetical protein
Length = 273
Score = 39.3 bits (90), Expect = 0.003
Identities = 44/180 (24%), Positives = 76/180 (41%), Gaps = 35/180 (19%)
Query: 119 LNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNG-------KNSHPFTLMKEWLALCNE 171
++K G + + SG SE+DIL +A+ + T K H + ++K N+
Sbjct: 8 VSKLRGCVNQIENKNPSGASEEDILNQAKMLLTQYEKYKRGFKFDHVWPILKGIEKFAND 67
Query: 172 ----PHYCSQVGGNLGSGSSGSKRSHESDACGSNSI-----------GSIPRPMGREAAK 216
P G ++ S SS S + S + G NSI RPMG + AK
Sbjct: 68 NMKTPPAFQGEGRDVTSSSSFSINTESSPSPGMNSIDLNMDSEDANFSLSSRPMGLKKAK 127
Query: 217 KKNKKKSRDAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEH 276
+K + + ++KQL EQ + ++ I+ E N +++ K+ + EE+
Sbjct: 128 RKQQSEE-------------QFKQLLEQNDKLIKAITKGTSERNEIQRQKIEVARMKEEN 174
>At1g24380 hypothetical protein
Length = 273
Score = 39.3 bits (90), Expect = 0.003
Identities = 44/180 (24%), Positives = 76/180 (41%), Gaps = 35/180 (19%)
Query: 119 LNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNG-------KNSHPFTLMKEWLALCNE 171
++K G + + SG SE+DIL +A+ + T K H + ++K N+
Sbjct: 8 VSKLRGCVNQIENKNPSGASEEDILNQAKMLLTQYEKYKRGFKFDHVWPILKGIEKFAND 67
Query: 172 ----PHYCSQVGGNLGSGSSGSKRSHESDACGSNSI-----------GSIPRPMGREAAK 216
P G ++ S SS S + S + G NSI RPMG + AK
Sbjct: 68 NMKTPAAFQGEGRDVTSSSSFSINTESSPSPGMNSIDLNMDSEDANFSLSSRPMGLKKAK 127
Query: 217 KKNKKKSRDAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEH 276
+K + + ++KQL EQ + ++ I+ E N +++ K+ + EE+
Sbjct: 128 RKQQSEE-------------QFKQLLEQNDKLIKAITKGTSERNEIQRQKIEVARMKEEN 174
>At4g35040 bZip transcription factor AtbZip19
Length = 261
Score = 35.4 bits (80), Expect = 0.037
Identities = 28/93 (30%), Positives = 50/93 (53%), Gaps = 6/93 (6%)
Query: 173 HYCSQVGGNLGSGSSGSKRSHESDACGSNSIGSIPRPMG-REAAKK-KNKKKSRDAALEV 230
H C V + S K S + A G RP+G REA +K + KKK++ A+LE
Sbjct: 61 HTCFHVHTKILPDESDEKVSTDDTAESCGKKGE-KRPLGNREAVRKYREKKKAKAASLE- 118
Query: 231 VNNEWSEYKQLREQELERLEKISSVQQEANRLE 263
+E + + + +Q ++RL+ ++++ E +RL+
Sbjct: 119 --DEVARLRAVNQQLVKRLQNQATLEAEVSRLK 149
>At5g40340 unknown protein
Length = 1008
Score = 35.0 bits (79), Expect = 0.048
Identities = 29/110 (26%), Positives = 44/110 (39%), Gaps = 6/110 (5%)
Query: 182 LGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAALEVVNNEWSEYKQL 241
LGS S K E G P G+E KK K K + +EV E +E +
Sbjct: 672 LGSYDSSDKEKEELSEMGK------PVTKGKEKKDKKGKAKQKAEEIEVTGKEENETDKH 725
Query: 242 REQELERLEKISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQLSQE 291
+ + ER K S ++E E+ + S ++ ++ E KQ E
Sbjct: 726 GKMKKERKRKKSESKKEGGEGEETQKEANESTKKERKRKKSESKKQSDGE 775
>At5g52550 unknown protein
Length = 360
Score = 34.3 bits (77), Expect = 0.082
Identities = 17/51 (33%), Positives = 28/51 (54%)
Query: 214 AAKKKNKKKSRDAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEK 264
A +K K+ ++ LE + E S ++QE + LE+I + + NRLEK
Sbjct: 52 AELEKKKQMKKEGQLEAADEEDSADAAKKKQERDELERIKQAENKKNRLEK 102
Score = 33.9 bits (76), Expect = 0.11
Identities = 16/51 (31%), Positives = 28/51 (54%)
Query: 214 AAKKKNKKKSRDAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEK 264
A +K K+ ++ L+ E S Y ++QE E LE+I +++ R+EK
Sbjct: 155 AELEKKKQMKKEGQLDAAVEEDSAYAAKKKQEREELERIKQAERKKRRIEK 205
>At5g11270 unknown protein
Length = 354
Score = 33.9 bits (76), Expect = 0.11
Identities = 22/76 (28%), Positives = 37/76 (47%), Gaps = 14/76 (18%)
Query: 220 KKKSRDAALEVVNNEWSEYKQLREQELERLEK------------ISSVQQEAN--RLEKM 265
K+K +D A+ V+ WS K++++ +E LEK +SS+ Q N R +
Sbjct: 272 KEKVKDEAVHVMQQRWSAQKRVKKAHIETLEKVYRRSKRPTNAVVSSIVQVTNLPRKRVL 331
Query: 266 KMFLQLSFEEHLDDRR 281
K F E+ + D+R
Sbjct: 332 KWFEDKRAEDGVPDKR 347
>At1g67230 unknown protein
Length = 1132
Score = 33.5 bits (75), Expect = 0.14
Identities = 25/87 (28%), Positives = 45/87 (50%), Gaps = 6/87 (6%)
Query: 206 IPRPMGREAAKKKNKKKSRDAALEVVNNEWSEYKQLREQELERLEKISS----VQQEANR 261
+ + G A+ K +D L V E SEY +L+ + E++EK S +Q+EA
Sbjct: 455 VEKVSGENQAQLSEINKEKDE-LRVTEEERSEYLRLQTELKEQIEKCRSQQELLQKEAED 513
Query: 262 LEKMKMFLQLSFEEHLDDRRKELLKQL 288
L+ + + +EE LD+R+ ++ +L
Sbjct: 514 LKAQRESFEKEWEE-LDERKAKIGNEL 539
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,448,233
Number of Sequences: 26719
Number of extensions: 341140
Number of successful extensions: 1328
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 55
Number of HSP's that attempted gapping in prelim test: 1234
Number of HSP's gapped (non-prelim): 127
length of query: 295
length of database: 11,318,596
effective HSP length: 99
effective length of query: 196
effective length of database: 8,673,415
effective search space: 1699989340
effective search space used: 1699989340
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0263.5


