

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0260.10
(751 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
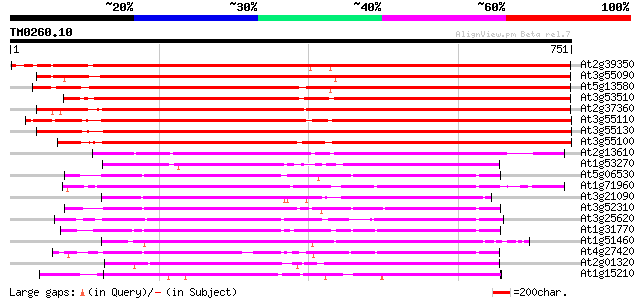
Sequences producing significant alignments: (bits) Value
At2g39350 ABC transporter like protein 1053 0.0
At3g55090 ABC transporter - like protein 1037 0.0
At5g13580 ABC transporter-like protein 1036 0.0
At3g53510 ABC transporter -like protein 991 0.0
At2g37360 ABC transporter like protein 979 0.0
At3g55110 ABC transporter - like protein 892 0.0
At3g55130 ABC transporter - like protein 866 0.0
At3g55100 ABC transporter - like protein 853 0.0
At2g13610 putative ABC transporter 367 e-101
At1g53270 putative ABC transporter emb|AAD22683.1 350 2e-96
At5g06530 ABC transporter like protein 270 2e-72
At1g71960 putative ABC transporter 267 2e-71
At3g21090 ABC transporter, putative 266 3e-71
At3g52310 ABC transporter-like protein 259 4e-69
At3g25620 membrane transporter, putative 258 1e-68
At1g31770 unknown protein 258 1e-68
At1g51460 ATP-dependent transmembrane transporter like protein 256 2e-68
At4g27420 putative protein 254 1e-67
At2g01320 putative ABC transporter 249 5e-66
At1g15210 putative ABC transporter 210 3e-54
>At2g39350 ABC transporter like protein
Length = 740
Score = 1053 bits (2724), Expect = 0.0
Identities = 551/758 (72%), Positives = 627/758 (82%), Gaps = 30/758 (3%)
Query: 3 SRIVAENVTDDATSYLDLMELSDLTRRPASGDLPTLGQLLKHVGDARKEAAGDGSETPLH 62
+RIVA N DD D MEL+ ++ S TLGQLLK+V D RK A GD ETP+H
Sbjct: 2 ARIVAAN--DD-----DSMELNTISSIHDS----TLGQLLKNVSDVRKMAIGD--ETPVH 48
Query: 63 HAL--DVVGMEPRSLPFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESV 120
+L D R++PFVLSF NLTY+V V+ KL F ++FPRRR + P + ++
Sbjct: 49 ESLNQDYNDGYMRTVPFVLSFDNLTYNVSVRPKLDFRNLFPRRRT------EDPEIAQTA 102
Query: 121 FSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALES 180
+TKTLLN+ISGE RDGEIMAVLGASGSGKSTLIDALANRIAKG LKGT+ LNGE L+S
Sbjct: 103 RPKTKTLLNNISGETRDGEIMAVLGASGSGKSTLIDALANRIAKGSLKGTVKLNGETLQS 162
Query: 181 RLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAA 240
R+LKVISAYVMQDDLLFPMLTVEETL FAAEFRLPR+L KSKKK RVQALIDQLG+RNAA
Sbjct: 163 RMLKVISAYVMQDDLLFPMLTVEETLMFAAEFRLPRSLPKSKKKLRVQALIDQLGIRNAA 222
Query: 241 KTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQS 300
KT+IGDEGHRG+SGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVL+RIAQS
Sbjct: 223 KTIIGDEGHRGISGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLKRIAQS 282
Query: 301 GSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFA 360
GSIVIMSIHQPS+R+LGLLDR+IFLSRG TVYSGSP LP FF EFG P+P+++NRTEFA
Sbjct: 283 GSIVIMSIHQPSHRVLGLLDRLIFLSRGHTVYSGSPASLPRFFTEFGSPIPENENRTEFA 342
Query: 361 LDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPER---PNGMSLKEAISASISR 417
LDLIR+LEGS GGT+ L+EFNK WQ M K + P PN ++LKEAI+ASISR
Sbjct: 343 LDLIRELEGSAGGTRGLIEFNKKWQEMKKQSNRQPPLTPPSSPYPN-LTLKEAIAASISR 401
Query: 418 GKLVSGATNTA-----TTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAV 472
GKLVSG + A T T + VP FANP WIE+ TLSKRS +SRR+PELFGIR+ +V
Sbjct: 402 GKLVSGGESVAHGGATTNTTTLAVPAFANPMWIEIKTLSKRSMLNSRRQPELFGIRIASV 461
Query: 473 MVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYN 532
++TGFILAT+FW LDNSPKGVQERLGFFAFAMST FYT ADALPVFLQERYIFMRETAYN
Sbjct: 462 VITGFILATVFWRLDNSPKGVQERLGFFAFAMSTMFYTCADALPVFLQERYIFMRETAYN 521
Query: 533 AYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSF 592
AYRRSSY++SHA+VS P L FLS+AFA T+WAVGLDGG +G LFY LII ASFW+G+SF
Sbjct: 522 AYRRSSYVLSHAIVSFPSLIFLSVAFAATTYWAVGLDGGLTGLLFYCLIILASFWSGSSF 581
Query: 593 VTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQ 652
VTFLSGVVP VMLGYTIVVAILAYFLL SGFFI+R+RIP YWIWFHY+SLVKYPYEAVLQ
Sbjct: 582 VTFLSGVVPSVMLGYTIVVAILAYFLLFSGFFINRNRIPDYWIWFHYMSLVKYPYEAVLQ 641
Query: 653 NEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLLGSMSGTLGTNITATTCLTTGADILQQ 712
NEF D KCFVRGVQIFDN+PL +P +KLKLLG++S +LG I++TTCLTTG+DIL+Q
Sbjct: 642 NEFSDATKCFVRGVQIFDNTPLGELPEVMKLKLLGTVSKSLGVTISSTTCLTTGSDILRQ 701
Query: 713 NGVTELSKWNCLWITVAWGFFFRFLFYLALLVGSKNKR 750
GV +LSKWNCL+ITVA+GFFFR LFY LL+GSKNKR
Sbjct: 702 QGVVQLSKWNCLFITVAFGFFFRILFYFTLLLGSKNKR 739
>At3g55090 ABC transporter - like protein
Length = 720
Score = 1037 bits (2681), Expect = 0.0
Identities = 532/727 (73%), Positives = 604/727 (82%), Gaps = 29/727 (3%)
Query: 37 TLGQLLKHVGDARKEAAGDGSETPLHHALDVVGME--------PRSLPFVLSFSNLTYSV 88
TLGQLLK+V D RK GD ETP+H D G R +PFVLSF+NLTY+V
Sbjct: 9 TLGQLLKNVSDVRKVEVGD--ETPVHEFFDRDGSSLDGDNDHLMRPVPFVLSFNNLTYNV 66
Query: 89 KVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLLNDISGEARDGEIMAVLGASG 148
V+RKL F + P RR FS+TKTLL++ISGE RDGEI+AVLGASG
Sbjct: 67 SVRRKLDFHDLVPWRRTS--------------FSKTKTLLDNISGETRDGEILAVLGASG 112
Query: 149 SGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISAYVMQDDLLFPMLTVEETLTF 208
SGKSTLIDALANRIAKG LKGT+ LNGEAL+SR+LKVISAYVMQDDLLFPMLTVEETL F
Sbjct: 113 SGKSTLIDALANRIAKGSLKGTVTLNGEALQSRMLKVISAYVMQDDLLFPMLTVEETLMF 172
Query: 209 AAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIH 268
AAEFRLPR+L KSKKK RVQALIDQLG+RNAAKT+IGDEGHRG+SGGERRRVSIGIDIIH
Sbjct: 173 AAEFRLPRSLPKSKKKLRVQALIDQLGIRNAAKTIIGDEGHRGISGGERRRVSIGIDIIH 232
Query: 269 DPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFLSRG 328
DPI+LFLDEPTSGLDSTSAFMVVKVL+RIA+SGSI+IMSIHQPS+R+L LLDR+IFLSRG
Sbjct: 233 DPIVLFLDEPTSGLDSTSAFMVVKVLKRIAESGSIIIMSIHQPSHRVLSLLDRLIFLSRG 292
Query: 329 QTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQSMT 388
TV+SGSP LPSFFA FG+P+P+++N+TEFALDLIR+LEGS GGT+ LVEFNK WQ M
Sbjct: 293 HTVFSGSPASLPSFFAGFGNPIPENENQTEFALDLIRELEGSAGGTRGLVEFNKKWQEMK 352
Query: 389 KVHSHSVSSQPERPN-GMSLKEAISASISRGKLVSGATNTATTTPSS----MVPTFANPF 443
K + + P PN ++LKEAISASISRGKLVSG ++ VP FANPF
Sbjct: 353 KQSNPQTLTPPASPNPNLTLKEAISASISRGKLVSGGGGGSSVINHGGGTLAVPAFANPF 412
Query: 444 WIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFA 503
WIE+ TL++RS +SRR+PEL G+RL V+VTGFILAT+FW LDNSPKGVQERLGFFAFA
Sbjct: 413 WIEIKTLTRRSILNSRRQPELLGMRLATVIVTGFILATVFWRLDNSPKGVQERLGFFAFA 472
Query: 504 MSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITF 563
MST FYT ADALPVFLQERYIFMRETAYNAYRRSSY++SHA+V+ P L FLSLAFAV TF
Sbjct: 473 MSTMFYTCADALPVFLQERYIFMRETAYNAYRRSSYVLSHAIVTFPSLIFLSLAFAVTTF 532
Query: 564 WAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGF 623
WAVGL+GG GFLFY LII ASFW+G+SFVTFLSGVVPHVMLGYTIVVAILAYFLL SGF
Sbjct: 533 WAVGLEGGLMGFLFYCLIILASFWSGSSFVTFLSGVVPHVMLGYTIVVAILAYFLLFSGF 592
Query: 624 FIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCFVRGVQIFDNSPLRSVPYELKL 683
FI+RDRIP YWIWFHYLSLVKYPYEAVLQNEF DP +CFVRGVQ+FDNSPL + Y +KL
Sbjct: 593 FINRDRIPQYWIWFHYLSLVKYPYEAVLQNEFSDPTECFVRGVQLFDNSPLGELTYGMKL 652
Query: 684 KLLGSMSGTLGTNITATTCLTTGADILQQNGVTELSKWNCLWITVAWGFFFRFLFYLALL 743
+LL S+S ++G I+++TCLTTGAD+L+Q GVT+LSKWNCL ITV +GF FR LFYL LL
Sbjct: 653 RLLDSVSRSIGMRISSSTCLTTGADVLKQQGVTQLSKWNCLLITVGFGFLFRILFYLCLL 712
Query: 744 VGSKNKR 750
+GSKNKR
Sbjct: 713 LGSKNKR 719
>At5g13580 ABC transporter-like protein
Length = 727
Score = 1036 bits (2680), Expect = 0.0
Identities = 529/724 (73%), Positives = 604/724 (83%), Gaps = 27/724 (3%)
Query: 31 ASGDLPTLGQLLKHVGDARKEAAGDGSETPLHHALDVVGMEPRSLPFVLSFSNLTYSVKV 90
AS T QLL++V D+ + + HH + +S+PFVLSF++LTYSVKV
Sbjct: 26 ASSSPTTFAQLLQNVDDSTRRSHHQ------HHVDVDLASPDQSVPFVLSFTDLTYSVKV 79
Query: 91 KRKLSFSSIFPRRRNRLGAVEDAPAVEESVFS-RTKTLLNDISGEARDGEIMAVLGASGS 149
+RK ++ R D A E +FS +TKTLLN I+GEARDGEI+AVLGASGS
Sbjct: 80 RRKFTW---------RRSVSSDPGAPSEGIFSSKTKTLLNGITGEARDGEILAVLGASGS 130
Query: 150 GKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISAYVMQDDLLFPMLTVEETLTFA 209
GKSTLIDALANRIAKG LKG + LNGE L S++ K ISAYVMQDDLLFPMLTVEETL FA
Sbjct: 131 GKSTLIDALANRIAKGSLKGNVTLNGEVLNSKMQKAISAYVMQDDLLFPMLTVEETLMFA 190
Query: 210 AEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHD 269
AEFRLPR+LSKSKK RVQALIDQLGLRNAA TVIGDEGHRG+SGGERRRVSIGIDIIHD
Sbjct: 191 AEFRLPRSLSKSKKSLRVQALIDQLGLRNAANTVIGDEGHRGISGGERRRVSIGIDIIHD 250
Query: 270 PILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFLSRGQ 329
PILLFLDEPTSGLDSTSA V+KVL+RIAQSGS+VIM++HQPSYR+L LLDR++FLSRGQ
Sbjct: 251 PILLFLDEPTSGLDSTSALSVIKVLKRIAQSGSMVIMTLHQPSYRLLRLLDRLLFLSRGQ 310
Query: 330 TVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTK 389
TV+SGSP LP FFAEFGHP+P+ +NRTEFALDLIR+LEGS GGT+SLVEFNK ++
Sbjct: 311 TVFSGSPAMLPRFFAEFGHPIPEHENRTEFALDLIRELEGSAGGTRSLVEFNKGFRQR-- 368
Query: 390 VHSHSVSSQPERPNGMSLKEAISASISRGKLVSGATNT---ATTTPSSMVPTFANPFWIE 446
++P G+SLKEAISASIS+GKLVSGAT T + ++P S +PTFANPFW+E
Sbjct: 369 ------KAEPRSQTGLSLKEAISASISKGKLVSGATTTTHSSGSSPVSTIPTFANPFWVE 422
Query: 447 MVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMST 506
+ L+KRS T+SRR+PELFGIRLGAV+VTGFILATMFW LDNSPKGVQERLG FAFAMST
Sbjct: 423 LAVLAKRSMTNSRRQPELFGIRLGAVLVTGFILATMFWQLDNSPKGVQERLGCFAFAMST 482
Query: 507 TFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAV 566
TFYT ADALPVFLQER+IFMRETAYNAYRRSSY++SH+LV+LP L LSLAFA ITFW V
Sbjct: 483 TFYTCADALPVFLQERFIFMRETAYNAYRRSSYVLSHSLVALPSLIILSLAFAAITFWGV 542
Query: 567 GLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFID 626
GLDGG GFLFYFL+I ASFWAG+SFVTFLSGVVPHVMLGYTIVVAILAYFLL SGFFI+
Sbjct: 543 GLDGGLMGFLFYFLVILASFWAGSSFVTFLSGVVPHVMLGYTIVVAILAYFLLFSGFFIN 602
Query: 627 RDRIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLL 686
RDRIP YWIWFHY+SLVKYPYEAVL NEF DP KCFVRGVQIFDN+PL +VP +K++LL
Sbjct: 603 RDRIPGYWIWFHYISLVKYPYEAVLLNEFGDPTKCFVRGVQIFDNTPLVAVPQGMKVRLL 662
Query: 687 GSMSGTLGTNITATTCLTTGADILQQNGVTELSKWNCLWITVAWGFFFRFLFYLALLVGS 746
+MS +LG IT++TCLTTG DILQQ GVT+L+KWNCLW+TVAWGFFFR LFY +LL+GS
Sbjct: 663 ATMSKSLGMRITSSTCLTTGYDILQQQGVTDLTKWNCLWVTVAWGFFFRILFYFSLLLGS 722
Query: 747 KNKR 750
KNKR
Sbjct: 723 KNKR 726
>At3g53510 ABC transporter -like protein
Length = 739
Score = 991 bits (2562), Expect = 0.0
Identities = 504/679 (74%), Positives = 566/679 (83%), Gaps = 21/679 (3%)
Query: 72 PRSLPFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLLNDI 131
P S PFVLSF +LTYSVK+K+K FP N +P + TK LLN I
Sbjct: 81 PSSSPFVLSFKDLTYSVKIKKKFK---PFPCCGN-------SPFDGNDMEMNTKVLLNGI 130
Query: 132 SGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISAYVM 191
SGEAR+GE+MAVLGASGSGKSTLIDALANRI+K L+G + LNGE LES L KVISAYVM
Sbjct: 131 SGEAREGEMMAVLGASGSGKSTLIDALANRISKESLRGDITLNGEVLESSLHKVISAYVM 190
Query: 192 QDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRG 251
QDDLLFPMLTVEETL F+AEFRLP +LSK KKKARVQALIDQLGLRNAAKTVIGDEGHRG
Sbjct: 191 QDDLLFPMLTVEETLMFSAEFRLPSSLSKKKKKARVQALIDQLGLRNAAKTVIGDEGHRG 250
Query: 252 VSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQP 311
VSGGERRRVSIG DIIHDPI+LFLDEPTSGLDSTSA+MVVKVLQRIAQSGSIVIMSIHQP
Sbjct: 251 VSGGERRRVSIGTDIIHDPIILFLDEPTSGLDSTSAYMVVKVLQRIAQSGSIVIMSIHQP 310
Query: 312 SYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSP 371
SYRILGLLD++IFLSRG TVYSGSPT LP FF+EFGHP+P+++N+ EFALDLIR+LE SP
Sbjct: 311 SYRILGLLDKLIFLSRGNTVYSGSPTHLPQFFSEFGHPIPENENKPEFALDLIRELEDSP 370
Query: 372 GGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKLVSGATNTATTT 431
GTKSLVEF+K W++ SSQ R +SLK+AISASISRGKLVSGATN
Sbjct: 371 EGTKSLVEFHKQWRAK------QTSSQSRRNTNVSLKDAISASISRGKLVSGATNLR--- 421
Query: 432 PSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPK 491
S TFANPFW EM+ + KRS +SRR+PELFGIRLGAV+VTG ILAT+FW LDNSP+
Sbjct: 422 --SSFQTFANPFWTEMLVIGKRSILNSRRQPELFGIRLGAVLVTGMILATIFWKLDNSPR 479
Query: 492 GVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPL 551
G+QERLGFFAFAMSTTFYT A+A+PVFLQERYIFMRETAYNAYRRSSY+++H ++S+P L
Sbjct: 480 GIQERLGFFAFAMSTTFYTCAEAIPVFLQERYIFMRETAYNAYRRSSYVLAHTIISIPAL 539
Query: 552 AFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVV 611
LS AFA TF AVGL GG+ GFLF+F I +FWAG+SFVTFLSGVV HVM+G+T+VV
Sbjct: 540 IILSAAFAASTFSAVGLAGGSEGFLFFFFTILTAFWAGSSFVTFLSGVVSHVMIGFTVVV 599
Query: 612 AILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCFVRGVQIFDN 671
AILAYFLL SGFFI RDRIP YWIWFHYLSLVKYPYE VLQNEF+DP KCFVRG+Q+FDN
Sbjct: 600 AILAYFLLFSGFFISRDRIPLYWIWFHYLSLVKYPYEGVLQNEFEDPTKCFVRGIQMFDN 659
Query: 672 SPLRSVPYELKLKLLGSMSGTLGTNITATTCLTTGADILQQNGVTELSKWNCLWITVAWG 731
SPL VP +K+ LL SMSG LG N+TA TC+TTG DIL+Q G+TE+SKWNCLWITVAWG
Sbjct: 660 SPLGQVPTAVKISLLKSMSGVLGINVTAETCVTTGIDILKQQGITEISKWNCLWITVAWG 719
Query: 732 FFFRFLFYLALLVGSKNKR 750
FFFR LFY LL+GSKNKR
Sbjct: 720 FFFRVLFYFTLLIGSKNKR 738
>At2g37360 ABC transporter like protein
Length = 755
Score = 979 bits (2531), Expect = 0.0
Identities = 501/725 (69%), Positives = 589/725 (81%), Gaps = 26/725 (3%)
Query: 37 TLGQLLKHVGDARKEAAGD----GSETPLHHALD-------VVGMEPRSLPFVLSFSNLT 85
T + L +V DAR + + G +P++ A S PFVLSF++LT
Sbjct: 45 TFAEHLMNVEDARNDESASSRALGIASPINSAASSFNSWASAPASSISSSPFVLSFTDLT 104
Query: 86 YSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLLNDISGEARDGEIMAVLG 145
YSVK+++K + + R N + SV TK LLN ISGEAR+GE+MAVLG
Sbjct: 105 YSVKIQKKFNPLACCRRSGN-----------DSSV--NTKILLNGISGEAREGEMMAVLG 151
Query: 146 ASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISAYVMQDDLLFPMLTVEET 205
ASGSGKSTLIDALANRIAK L+G++ LNGE LES + KVISAYVMQDDLLFPMLTVEET
Sbjct: 152 ASGSGKSTLIDALANRIAKDSLRGSITLNGEVLESSMQKVISAYVMQDDLLFPMLTVEET 211
Query: 206 LTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGID 265
L F+AEFRLPR+LSK KKKARVQALIDQLGLR+AAKTVIGDEGHRGVSGGERRRVSIG D
Sbjct: 212 LMFSAEFRLPRSLSKKKKKARVQALIDQLGLRSAAKTVIGDEGHRGVSGGERRRVSIGND 271
Query: 266 IIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFL 325
IIHDPI+LFLDEPTSGLDSTSA+MV+KVLQRIAQSGSIVIMSIHQPSYRI+GLLD++IFL
Sbjct: 272 IIHDPIILFLDEPTSGLDSTSAYMVIKVLQRIAQSGSIVIMSIHQPSYRIMGLLDQLIFL 331
Query: 326 SRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQ 385
S+G TVYSGSPT LP FF+EF HP+P+++N+TEFALDLIR+LE S GTK LVEF+K W+
Sbjct: 332 SKGNTVYSGSPTHLPQFFSEFKHPIPENENKTEFALDLIRELEYSTEGTKPLVEFHKQWR 391
Query: 386 SMTKVHSHSVSSQPERPNGMSLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWI 445
+ + S++ ++ N SLKEAI+ASISRGKLVSGATN ++ + TFANPFWI
Sbjct: 392 AK-QAPSYN-NNNKRNTNVSSLKEAITASISRGKLVSGATNNNSSNLTPSFQTFANPFWI 449
Query: 446 EMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMS 505
EM+ + KR+ +SRR+PEL G+RLGAVMVTG ILATMF NLDNSPKG QERLGFFAFAMS
Sbjct: 450 EMIVIGKRAILNSRRQPELLGMRLGAVMVTGIILATMFTNLDNSPKGAQERLGFFAFAMS 509
Query: 506 TTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWA 565
TTFYT A+A+PVFLQERYIFMRETAYNAYRRSSY++S +++S+P L LS +FA TFWA
Sbjct: 510 TTFYTCAEAIPVFLQERYIFMRETAYNAYRRSSYVLSQSIISIPALIVLSASFAATTFWA 569
Query: 566 VGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFI 625
VGLDGGA+GF F++ I ASFWAG+SFVTFLSGV+P+VMLG+T+VVAILAYFLL SGFFI
Sbjct: 570 VGLDGGANGFFFFYFTILASFWAGSSFVTFLSGVIPNVMLGFTVVVAILAYFLLFSGFFI 629
Query: 626 DRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKL 685
RDRIP YW+WFHY+SLVKYPYE VLQNEF +P +CF RGVQ+FDNSPL P ++K+ L
Sbjct: 630 SRDRIPVYWLWFHYISLVKYPYEGVLQNEFQNPTRCFARGVQLFDNSPLGEFPNDVKVNL 689
Query: 686 LGSMSGTLGTNITATTCLTTGADILQQNGVTELSKWNCLWITVAWGFFFRFLFYLALLVG 745
L SMSG LGTN+TA TC+TTG DIL+Q G+T++SKWNCLWITVAWGFFFR LFY LL+G
Sbjct: 690 LKSMSGVLGTNVTAETCVTTGIDILKQQGITDISKWNCLWITVAWGFFFRVLFYFTLLIG 749
Query: 746 SKNKR 750
SKNKR
Sbjct: 750 SKNKR 754
>At3g55110 ABC transporter - like protein
Length = 708
Score = 892 bits (2305), Expect = 0.0
Identities = 460/735 (62%), Positives = 570/735 (76%), Gaps = 38/735 (5%)
Query: 22 ELSDLTRRPASGDLPTLGQLLKHVGDA-RKEAAGDGSETPLHHALDVV-GMEPRSLPFVL 79
E+ D++ + PTLG+LLK D+ RK+ G+ + P H LD+ E RS+PF+L
Sbjct: 7 EILDISHT-GGNESPTLGELLKDFNDSGRKKYPGENA--PTQHILDLAPAAETRSVPFLL 63
Query: 80 SFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLLNDISGEARDGE 139
SF+NL+Y+V ++R+ FS RR+ + KTLL+DI+GEARDGE
Sbjct: 64 SFNNLSYNVVLRRRFDFS----RRKT----------------ASVKTLLDDITGEARDGE 103
Query: 140 IMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGE-ALESRLLKVISAYVMQDDLLFP 198
I+AVLG SG+GKSTLIDALA R+A+ LKGT+ LNGE L+SRLLKVISAYVMQDDLLFP
Sbjct: 104 ILAVLGGSGAGKSTLIDALAGRVAEDSLKGTVTLNGEKVLQSRLLKVISAYVMQDDLLFP 163
Query: 199 MLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERR 258
MLTV+ETL FA+EFRLPR+L KSKK RV+ LIDQLGLRNAA TVIGDEGHRGVSGGERR
Sbjct: 164 MLTVKETLMFASEFRLPRSLPKSKKMERVETLIDQLGLRNAADTVIGDEGHRGVSGGERR 223
Query: 259 RVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSYRILGL 318
RVSIGIDIIHDPILLFLDEPTSGLDST+AFMVV+VL+RIAQSGS+VIMSIHQPS RI+GL
Sbjct: 224 RVSIGIDIIHDPILLFLDEPTSGLDSTNAFMVVQVLKRIAQSGSVVIMSIHQPSARIIGL 283
Query: 319 LDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGGTKSLV 378
LDR+I LS G++V++GSP LPSFF+ FG P+P+ +N TEFALD+IR+LEGS GT+ LV
Sbjct: 284 LDRLIILSHGKSVFNGSPVSLPSFFSSFGRPIPEKENITEFALDVIRELEGSSEGTRDLV 343
Query: 379 EFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKLVSGATNTATTTPSSM--V 436
EFN+ WQ + + S +SLKEAI+AS+SRGKLVSG++ P SM V
Sbjct: 344 EFNEKWQQNQTARATTQSR-------VSLKEAIAASVSRGKLVSGSSGA---NPISMETV 393
Query: 437 PTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQER 496
++ANP E L+KR + R PEL G+R+G VMVTG +LAT++W LDN+P+G QER
Sbjct: 394 SSYANPPLAETFILAKRYIKNWIRTPELIGMRIGTVMVTGLLLATVYWRLDNTPRGAQER 453
Query: 497 LGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSL 556
+GFFAF MST FY AD +PVF+QERYIF+RET +NAYR SSY+ISHALVSLP L LS+
Sbjct: 454 MGFFAFGMSTMFYCCADNIPVFIQERYIFLRETTHNAYRTSSYVISHALVSLPQLLALSI 513
Query: 557 AFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAY 616
AFA TFW VGL GG F +Y LII+A+FW+G+S VTF+SG++P+VM+ Y + +A L+Y
Sbjct: 514 AFAATTFWTVGLSGGLESFFYYCLIIYAAFWSGSSIVTFISGLIPNVMMSYMVTIAYLSY 573
Query: 617 FLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCFVRGVQIFDNSPLRS 676
LLL GF+I+RDRIP YWIWFHY+SL+KYPYEAVL NEFDDP +CFV+GVQ+FD + L
Sbjct: 574 CLLLGGFYINRDRIPLYWIWFHYISLLKYPYEAVLINEFDDPSRCFVKGVQVFDGTLLAE 633
Query: 677 VPYELKLKLLGSMSGTLGTNITATTCLTTGADILQQNGVTELSKWNCLWITVAWGFFFRF 736
V + +K+KLL ++SG+LGT IT +TCL TG D+L Q G+T+LSKW+CLWIT+AWG FFR
Sbjct: 634 VSHVMKVKLLDTLSGSLGTKITESTCLRTGPDLLMQQGITQLSKWDCLWITLAWGLFFRI 693
Query: 737 LFYLALLVGSKNKRS 751
LFYL+LL GSKNKR+
Sbjct: 694 LFYLSLLFGSKNKRT 708
>At3g55130 ABC transporter - like protein
Length = 725
Score = 866 bits (2237), Expect = 0.0
Identities = 443/721 (61%), Positives = 561/721 (77%), Gaps = 28/721 (3%)
Query: 36 PTLGQLLKHVGDARKEAAGDG--SETPLHHALDVVGMEPRSLPFVLSFSNLTYSVKVKRK 93
PTL +LLK RK +GDG S+ P HH +DV + + +P+VL+F+NL Y V ++R+
Sbjct: 28 PTLDELLKDCDSFRKGDSGDGVKSDDPAHHIIDVEALYVKPVPYVLNFNNLQYDVTLRRR 87
Query: 94 LSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLLNDISGEARDGEIMAVLGASGSGKST 153
FS R+N + KTLL+D+SGEA DG+I+AVLGASG+GKST
Sbjct: 88 FGFS-----RQNGV-----------------KTLLDDVSGEASDGDILAVLGASGAGKST 125
Query: 154 LIDALANRIAKGRLKGTLALNGE-ALESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEF 212
LIDALA R+A+G L+G++ LNGE L+SRLLKVISAYVMQDDLLFPMLTV+ETL FA+EF
Sbjct: 126 LIDALAGRVAEGSLRGSVTLNGEKVLQSRLLKVISAYVMQDDLLFPMLTVKETLMFASEF 185
Query: 213 RLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPIL 272
RLPR+LSKSKK RV+ALIDQLGLRNAA TVIGDEGHRGVSGGERRRVSIGIDIIHDPI+
Sbjct: 186 RLPRSLSKSKKMERVEALIDQLGLRNAANTVIGDEGHRGVSGGERRRVSIGIDIIHDPIV 245
Query: 273 LFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVY 332
LFLDEPTSGLDST+AFMVV+VL+RIAQSGSIVIMSIHQPS RI+ LLDR+I LSRG++V+
Sbjct: 246 LFLDEPTSGLDSTNAFMVVQVLKRIAQSGSIVIMSIHQPSARIVELLDRLIILSRGKSVF 305
Query: 333 SGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQS--MTKV 390
+GSP LP FF++FG P+P+ +N +EFALDL+R+LEGS GTK+LV+FN+ WQ ++ +
Sbjct: 306 NGSPASLPGFFSDFGRPIPEKENISEFALDLVRELEGSNEGTKALVDFNEKWQQNKISLI 365
Query: 391 HSHSVSSQPERPNGMSLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTL 450
S +++ ++ +SLKEAI+AS+SRGKLVSG++ + T+ + V ++ANP E L
Sbjct: 366 QSAPQTNKLDQDRSLSLKEAINASVSRGKLVSGSSRSNPTSMET-VSSYANPSLFETFIL 424
Query: 451 SKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYT 510
+KR + R PEL G R+ VMVTG +LAT++W LD++P+G QERL FAF + T FY
Sbjct: 425 AKRYMKNWIRMPELVGTRIATVMVTGCLLATVYWKLDHTPRGAQERLTLFAFVVPTMFYC 484
Query: 511 TADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDG 570
D +PVF+QERYIF+RET +NAYR SSY+ISH+LVSLP L SL F+ ITFW VGL G
Sbjct: 485 CLDNVPVFIQERYIFLRETTHNAYRTSSYVISHSLVSLPQLLAPSLVFSAITFWTVGLSG 544
Query: 571 GASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRI 630
G GF+FY L+I+ASFW+G+S VTF+SGVVP++ML Y + + LAY LLLSGF+++RDRI
Sbjct: 545 GLEGFVFYCLLIYASFWSGSSVVTFISGVVPNIMLCYMVSITYLAYCLLLSGFYVNRDRI 604
Query: 631 PSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLLGSMS 690
P YW WFHY+S++KYPYEAVL NEFDDP +CFVRGVQ+FD++ L V K+KLL ++S
Sbjct: 605 PFYWTWFHYISILKYPYEAVLINEFDDPSRCFVRGVQVFDSTLLGGVSDSGKVKLLETLS 664
Query: 691 GTLGTNITATTCLTTGADILQQNGVTELSKWNCLWITVAWGFFFRFLFYLALLVGSKNKR 750
+L T IT +TCL TG+D+L Q G+T+LSKW+CLWIT A G FFR LFY ALL GS+NKR
Sbjct: 665 KSLRTKITESTCLRTGSDLLAQQGITQLSKWDCLWITFASGLFFRILFYFALLFGSRNKR 724
Query: 751 S 751
+
Sbjct: 725 T 725
>At3g55100 ABC transporter - like protein
Length = 662
Score = 853 bits (2203), Expect = 0.0
Identities = 431/687 (62%), Positives = 535/687 (77%), Gaps = 32/687 (4%)
Query: 65 LDVVGMEPRSLPFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRT 124
+DV E +PFVL+F++LTY+V ++++ R G +PA +
Sbjct: 8 IDVDESEIPPIPFVLAFNDLTYNVTLQQRFGL---------RFG---HSPA-------KI 48
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLK 184
KTLLN I+GEA++GEI+A+LGASG+GKSTLIDALA +IA+G LKGT+ LNGEAL+SRLL+
Sbjct: 49 KTLLNGITGEAKEGEILAILGASGAGKSTLIDALAGQIAEGSLKGTVTLNGEALQSRLLR 108
Query: 185 VISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVI 244
VISAYVMQ+DLLFPMLTVEETL FAAEFRLPR+LSKSKK+ RV+ LIDQLGL TVI
Sbjct: 109 VISAYVMQEDLLFPMLTVEETLMFAAEFRLPRSLSKSKKRNRVETLIDQLGLTTVKNTVI 168
Query: 245 GDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIV 304
GDEGHRGVSGGERRRVSIG DIIHDPI+LFLDEPTSGLDSTSAFMVV+VL++IA+SGSIV
Sbjct: 169 GDEGHRGVSGGERRRVSIGTDIIHDPIVLFLDEPTSGLDSTSAFMVVQVLKKIARSGSIV 228
Query: 305 IMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLI 364
IMSIHQPS RI+ LDR+I LS GQ V+S SP LP FF+EFG P+P+ +N EF LDLI
Sbjct: 229 IMSIHQPSGRIMEFLDRVIVLSSGQIVFSDSPATLPLFFSEFGSPIPEKENIAEFTLDLI 288
Query: 365 RDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKLVSGA 424
+DLEGSP GT+ LVEFN++WQ H SQ N SL EAI+ASISRGKLVS
Sbjct: 289 KDLEGSPEGTRGLVEFNRNWQ-----HRKLRVSQEPHHNSSSLGEAINASISRGKLVS-- 341
Query: 425 TNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFW 484
T +P++ NP+W+E V L+KR + R PEL G R+ VM+TGF+LAT++W
Sbjct: 342 ------TSYRSIPSYVNPWWVETVILAKRYMINWTRTPELIGTRVFIVMMTGFLLATVYW 395
Query: 485 NLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHA 544
+D+SP+GVQERL FF+FAM+T FY+ AD LP F+QERYIF+RETA+NAYRRSSY+ISH+
Sbjct: 396 KVDDSPRGVQERLSFFSFAMATMFYSCADGLPAFIQERYIFLRETAHNAYRRSSYVISHS 455
Query: 545 LVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVM 604
LV+LP L LS+ FA TFW VGL+GG +GF++Y +IIFASFW+G SFVTF+SGV+P+VM
Sbjct: 456 LVTLPHLFALSIGFAATTFWFVGLNGGLAGFIYYLMIIFASFWSGCSFVTFVSGVIPNVM 515
Query: 605 LGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCFVR 664
+ Y + L+Y LL SGF+++RDRI YWIW HY+SL+KYPYEAVL NEFDDP +CFVR
Sbjct: 516 MSYMVTFGYLSYCLLFSGFYVNRDRIHLYWIWIHYISLLKYPYEAVLHNEFDDPSRCFVR 575
Query: 665 GVQIFDNSPLRSVPYELKLKLLGSMSGTLGTNITATTCLTTGADILQQNGVTELSKWNCL 724
G Q+FDN+ + V K KLL +MSG LG +T +TCLTTG+D+L+Q+G+ +L KW CL
Sbjct: 576 GNQVFDNTIMEGVSETTKAKLLETMSGYLGMELTESTCLTTGSDLLKQHGIEQLDKWGCL 635
Query: 725 WITVAWGFFFRFLFYLALLVGSKNKRS 751
W+T+AWGFFFR LFY +LL+GSKNKR+
Sbjct: 636 WVTLAWGFFFRILFYFSLLLGSKNKRA 662
>At2g13610 putative ABC transporter
Length = 649
Score = 367 bits (942), Expect = e-101
Identities = 221/634 (34%), Positives = 345/634 (53%), Gaps = 46/634 (7%)
Query: 111 EDAPAVEESVFSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGT 170
E++ +E+ ++ K +L ++ A+ EI+A++G SG+GKS+L++ LA R+ G+
Sbjct: 46 EESLKLEDETGNKVKHVLKGVTCRAKPWEILAIVGPSGAGKSSLLEILAARLIPQT--GS 103
Query: 171 LALNGEALESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQAL 230
+ +N ++ K IS YV Q D LFP+LTVEETL F+A+ RL L + ++RV++L
Sbjct: 104 VYVNKRPVDRANFKKISGYVTQKDTLFPLLTVEETLLFSAKLRLK--LPADELRSRVKSL 161
Query: 231 IDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMV 290
+ +LGL A +GD+ RG+SGGERRRVSIG+++IHDP +L LDEPTSGLDSTSA ++
Sbjct: 162 VHELGLEAVATARVGDDSVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALLI 221
Query: 291 VKVLQRIAQS-GSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHP 349
+ +L+ +A++ G +I++IHQP +RI+ + ++ L+ G T+ GS QL + G
Sbjct: 222 IDMLKHMAETRGRTIILTIHQPGFRIVKQFNSVLLLANGSTLKQGSVDQLGVYLRSNGLH 281
Query: 350 LPDSDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKE 409
P +N EFA++ I + +S + T S SQ E +G K
Sbjct: 282 PPLHENIVEFAIESIESITKQQRLQESRRAAHVLTPQTTLQEKRSEDSQGESKSG---KF 338
Query: 410 AISASISRGKLVS-GATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIR 468
+ + ++ G N AT FAN E + L+ R + R ELF R
Sbjct: 339 TLQQLFQQTRVADVGTMNIAT----EFTRDFANSRLEETMILTHRFSKNIFRTKELFACR 394
Query: 469 LGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRE 528
++ +G +L +F NL + KG +ER+G FAF ++ +T +ALP+FLQER I M+E
Sbjct: 395 TVQMLGSGIVLGLIFHNLKDDLKGARERVGLFAFILTFLLTSTIEALPIFLQEREILMKE 454
Query: 529 TAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWA 588
T+ +YR SSY +++ LV LP L L++ F+ +W VGL+ FL + L+I+ +
Sbjct: 455 TSSGSYRVSSYAVANGLVYLPFLLILAILFSTPVYWLVGLNPSFMAFLHFSLLIWLILYT 514
Query: 589 GNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYE 648
NS V S +VP+ ++G +++ ++ F L SG+FI IP YWI+ HY+SL KYP+E
Sbjct: 515 ANSVVVCFSALVPNFIVGNSVISGVMGSFFLFSGYFISNHEIPGYWIFMHYISLFKYPFE 574
Query: 649 AVLQNEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLLGSMSGTLGTNITATTCLTTGAD 708
L NEF KC G CL T D
Sbjct: 575 GFLINEFSKSNKCLEYGF---------------------------------GKCLVTEED 601
Query: 709 ILQQNGVTELSKWNCLWITVAWGFFFRFLFYLAL 742
+L++ E S+W + I + + +RF+ Y+ L
Sbjct: 602 LLKEERYGEESRWRNVVIMLCFVLLYRFISYVIL 635
>At1g53270 putative ABC transporter emb|AAD22683.1
Length = 590
Score = 350 bits (898), Expect = 2e-96
Identities = 203/536 (37%), Positives = 313/536 (57%), Gaps = 45/536 (8%)
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLK 184
K +L D+S +AR EI A+ G SG+GK+TL++ LA +++ G++ G + +NG ++ +
Sbjct: 48 KVILKDVSCDARSAEITAIAGPSGAGKTTLLEILAGKVSHGKVSGQVLVNGRPMDGPEYR 107
Query: 185 VISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKK---ARVQALIDQLGLRNAAK 241
+S +V Q+D LFP LTV+ETLT++A RL K+K+K A+V+ LI +LGL + A
Sbjct: 108 RVSGFVPQEDALFPFLTVQETLTYSALLRL-----KTKRKDAAAKVKRLIQELGLEHVAD 162
Query: 242 TVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIA-QS 300
+ IG G+SGGERRRVSIG++++HDP ++ +DEPTSGLDS SA VV +L+ + +
Sbjct: 163 SRIGQGSRSGISGGERRRVSIGVELVHDPNVILIDEPTSGLDSASALQVVTLLKDMTIKQ 222
Query: 301 GSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFA 360
G ++++IHQP +RIL +DR++ LS G V +GS L GH +P N E+A
Sbjct: 223 GKTIVLTIHQPGFRILEQIDRIVLLSNGMVVQNGSVYSLHQKIKFSGHQIPRRVNVLEYA 282
Query: 361 LDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKL 420
+D+ LE P T+S E + + HS K S IS G
Sbjct: 283 IDIAGSLE--PIRTQSCREIS--------CYGHS-------------KTWKSCYISAG-- 317
Query: 421 VSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILA 480
G + + + +S++ E+ L +RS + R +LF R + G IL
Sbjct: 318 --GELHQSDSHSNSVLE--------EVQILGQRSCKNIFRTKQLFTTRALQASIAGLILG 367
Query: 481 TMFWNLDNSPKGVQE-RLGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSY 539
+++ N+ N K + R GFFAF ++ +T + LP+FLQ+R I MRET+ AYR SY
Sbjct: 368 SIYLNVGNQKKEAKVLRTGFFAFILTFLLSSTTEGLPIFLQDRRILMRETSRRAYRVLSY 427
Query: 540 LISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGV 599
+++ L+ +P L +S+ FA +W VGL GFL++ L+I+ NSFV S +
Sbjct: 428 VLADTLIFIPFLLIISMLFATPVYWLVGLRRELDGFLYFSLVIWIVLLMSNSFVACFSAL 487
Query: 600 VPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEF 655
VP+ ++G +++ ++ F L SG+FI +DRIP YW + HYLSL KYP+E ++ NE+
Sbjct: 488 VPNFIMGTSVISGLMGSFFLFSGYFIAKDRIPVYWEFMHYLSLFKYPFECLMINEY 543
>At5g06530 ABC transporter like protein
Length = 627
Score = 270 bits (691), Expect = 2e-72
Identities = 186/595 (31%), Positives = 291/595 (48%), Gaps = 48/595 (8%)
Query: 74 SLPFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLLNDISG 133
+LP L F ++TY V +K+ S S K +L ISG
Sbjct: 28 TLPIFLKFRDVTYKVVIKKLTS--------------------------SVEKEILTGISG 61
Query: 134 EARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISAYVMQD 193
GE++A++G SGSGK+TL+ LA RI++ G++ N + S+ LK +V QD
Sbjct: 62 SVNPGEVLALMGPSGSGKTTLLSLLAGRISQSSTGGSVTYNDKPY-SKYLKSKIGFVTQD 120
Query: 194 DLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVS 253
D+LFP LTV+ETLT+AA RLP+TL++ +KK R +I +LGL T+IG RGVS
Sbjct: 121 DVLFPHLTVKETLTYAARLRLPKTLTREQKKQRALDVIQELGLERCQDTMIGGAFVRGVS 180
Query: 254 GGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSY 313
GGER+RVSIG +II +P LL LDEPTSGLDST+A + +L IA++G VI +IHQPS
Sbjct: 181 GGERKRVSIGNEIIINPSLLLLDEPTSGLDSTTALRTILMLHDIAEAGKTVITTIHQPSS 240
Query: 314 RILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGG 373
R+ D++I L RG +Y G ++ +F+ G + N EF LDL G
Sbjct: 241 RLFHRFDKLILLGRGSLLYFGKSSEALDYFSSIGCSPLIAMNPAEFLLDL-------ANG 293
Query: 374 TKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAI-------SASISRGKLVSGA-- 424
+ + +V + +Q +P+ ++ E + A + KL+
Sbjct: 294 NINDISVPSELDDRVQVGNSGRETQTGKPSPAAVHEYLVEAYETRVAEQEKKKLLDPVPL 353
Query: 425 TNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFW 484
A + + + +W + L R + RR +R+ V+ T IL ++W
Sbjct: 354 DEEAKAKSTRLKRQWGTCWWEQYCILFCRGLKE-RRHEYFSWLRVTQVLSTAVILGLLWW 412
Query: 485 NLD-NSPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLIS 542
D +P G+Q++ G F A+ F+ A+ F QER + +E A + YR S+Y ++
Sbjct: 413 QSDIRTPMGLQDQAGLLFFIAVFWGFFPVFTAIFAFPQERAMLNKERAADMYRLSAYFLA 472
Query: 543 HALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPH 602
LP L F ++ ++ GL F L +F A + ++
Sbjct: 473 RTTSDLPLDFILPSLFLLVVYFMTGLRISPYPFFLSMLTVFLCIIAAQGLGLAIGAILMD 532
Query: 603 VMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDD 657
+ T+ + F+L GFF+ ++P + W YLS + Y+ +L+ ++ D
Sbjct: 533 LKKATTLASVTVMTFMLAGGFFV--KKVPVFISWIRYLSFNYHTYKLLLKVQYQD 585
>At1g71960 putative ABC transporter
Length = 662
Score = 267 bits (682), Expect = 2e-71
Identities = 201/681 (29%), Positives = 323/681 (46%), Gaps = 58/681 (8%)
Query: 71 EPRSL------PFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRT 124
EPRSL P L F ++ Y VK+ + S +L ++ P+ +E+ +
Sbjct: 26 EPRSLLSSSCFPITLKFVDVCYRVKIHGMSNDSCNI----KKLLGLKQKPS-DETRSTEE 80
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLK 184
+T+L+ ++G GE MAVLG SGSGKSTL++A+A R+ L G + +N + + LK
Sbjct: 81 RTILSGVTGMISPGEFMAVLGPSGSGKSTLLNAVAGRLHGSNLTGKILINDGKITKQTLK 140
Query: 185 VISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVI 244
+ +V QDDLL+P LTV ETL F A RLPR+L++ K +++I +LGL TV+
Sbjct: 141 R-TGFVAQDDLLYPHLTVRETLVFVALLRLPRSLTRDVKLRAAESVISELGLTKCENTVV 199
Query: 245 GDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQ-SGSI 303
G+ RG+SGGER+RVSI +++ +P LL LDEPTSGLD+T+A +V+ L +A G
Sbjct: 200 GNTFIRGISGGERKRVSIAHELLINPSLLVLDEPTSGLDATAALRLVQTLAGLAHGKGKT 259
Query: 304 VIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDL 363
V+ SIHQPS R+ + D ++ LS G+ ++ G ++F G N +F LDL
Sbjct: 260 VVTSIHQPSSRVFQMFDTVLLLSEGKCLFVGKGRDAMAYFESVGFSPAFPMNPADFLLDL 319
Query: 364 IRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKLVSG 423
+ + G T+ E Q++ + ++ Q + +S +A + ++ G
Sbjct: 320 ANGVCQTDGVTER--EKPNVRQTLVTAYDTLLAPQVKTCIEVSHFPQDNARFVKTRVNGG 377
Query: 424 ATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMF 483
T T W + + RR +R+ V+ + M+
Sbjct: 378 GITTCIAT------------WFSQLCILLHRLLKERRHESFDLLRIFQVVAASILCGLMW 425
Query: 484 WNLDNSPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLIS 542
W+ D + V +RLG F ++ + +A+ F QER IF RE A Y SSY ++
Sbjct: 426 WHSDY--RDVHDRLGLLFFISIFWGVLPSFNAVFTFPQERAIFTRERASGMYTLSSYFMA 483
Query: 543 HALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPH 602
H L SL L +F T+W V L G FL ++ A L +
Sbjct: 484 HVLGSLSMELVLPASFLTFTYWMVYLRPGIVPFLLTLSVLLLYVLASQGLGLALGAAIMD 543
Query: 603 VMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCF 662
TIV + F+L G+++ +++PS +W Y+S Y Y ++ ++
Sbjct: 544 AKKASTIVTVTMLAFVLTGGYYV--NKVPSGMVWMKYVSTTFYCYRLLVAIQYGS----- 596
Query: 663 VRGVQIFDNSPLRSVPYELKLKLLGSMS-GTLGTNITATTCLTTGADILQQNGVTELSKW 721
G +I L++LG S G G A+ + G +++ + ++ W
Sbjct: 597 --GEEI--------------LRMLGCDSKGKQG----ASAATSAGCRFVEEEVIGDVGMW 636
Query: 722 NCLWITVAWGFFFRFLFYLAL 742
+ + F +R L YLAL
Sbjct: 637 TSVGVLFLMFFGYRVLAYLAL 657
>At3g21090 ABC transporter, putative
Length = 680
Score = 266 bits (680), Expect = 3e-71
Identities = 183/541 (33%), Positives = 279/541 (50%), Gaps = 42/541 (7%)
Query: 124 TKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGR-LKGTLALNGEALESRL 182
T+ LL ++G A G IMA++G SGSGKSTL+D+LA R+A+ + G L LNG+ ++RL
Sbjct: 42 TRRLLQRLNGYAEPGRIMAIMGPSGSGKSTLLDSLAGRLARNVVMTGNLLLNGK--KARL 99
Query: 183 LKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKT 242
+ AYV Q+D+L LTV ET+T++A RLP +SK + V+ I +LGL++ +
Sbjct: 100 DYGLVAYVTQEDVLLGTLTVRETITYSAHLRLPSDMSKEEVSDIVEGTIMELGLQDCSDR 159
Query: 243 VIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGS 302
VIG+ RGVSGGER+RVSI ++I+ P +LFLDEPTSGLDS SAF V++ L+ IA+ G
Sbjct: 160 VIGNWHARGVSGGERKRVSIALEILTRPQILFLDEPTSGLDSASAFFVIQALRNIARDGR 219
Query: 303 IVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALD 362
VI S+HQPS + L D + LS G++VY G FFAE G P P N ++ L
Sbjct: 220 TVISSVHQPSSEVFALFDDLFLLSSGESVYFGEAKSAVEFFAESGFPCPKKRNPSDHFLR 279
Query: 363 LIRD--------LEGS------PGGTKSLVEFNKSWQSMTKVHSHSVS----SQPERPNG 404
I L+GS P + L+ S V ++ S S R
Sbjct: 280 CINSDFDTVTATLKGSQRIQETPATSDPLMNLATSVIKARLVENYKRSKYAKSAKSRIRE 339
Query: 405 MSLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPEL 464
+S E + I +G S AT +W ++ TL+ RSF + R
Sbjct: 340 LSNIEGLEMEIRKG---SEAT-----------------WWKQLRTLTARSFINMCRDVGY 379
Query: 465 FGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYI 524
+ R+ + +V + T+F+++ S + R+ F + + P FL+E +
Sbjct: 380 YWTRIISYIVVSISVGTIFYDVGYSYTSILARVSCGGFITGFMTFMSIGGFPSFLEEMKV 439
Query: 525 FMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFA 584
F +E Y S Y++S+ + S P L +S+ IT+ V G S + F+ L IF
Sbjct: 440 FYKERLSGYYGVSVYILSNYISSFPFLVAISVITGTITYNLVKFRPGFSHYAFFCLNIFF 499
Query: 585 SFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVK 644
S S + ++ VVP+ ++G ++ ++ SGFF +P + W + +S +
Sbjct: 500 SVSVIESLMMVVASVVPNFLMGLITGAGLIGIIMMTSGFFRLLPDLPKIF-WRYPVSYIS 558
Query: 645 Y 645
Y
Sbjct: 559 Y 559
>At3g52310 ABC transporter-like protein
Length = 737
Score = 259 bits (662), Expect = 4e-69
Identities = 181/590 (30%), Positives = 294/590 (49%), Gaps = 42/590 (7%)
Query: 74 SLPFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLLNDISG 133
+ P L F ++TY V K S S K++LN ISG
Sbjct: 140 TFPIYLKFIDITYKVTTKGMTSSSE--------------------------KSILNGISG 173
Query: 134 EARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISAYVMQD 193
A GE++A++G SGSGK+TL++AL R + + G+++ N + S+ LK +V QD
Sbjct: 174 SAYPGELLALMGPSGSGKTTLLNALGGRFNQQNIGGSVSYNDKPY-SKHLKTRIGFVTQD 232
Query: 194 DLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVS 253
D+LFP LTV+ETLT+ A RLP+TL++ +K+ R ++I +LGL T+IG RGVS
Sbjct: 233 DVLFPHLTVKETLTYTALLRLPKTLTEQEKEQRAASVIQELGLERCQDTMIGGSFVRGVS 292
Query: 254 GGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSY 313
GGER+RV IG +I+ +P LL LDEPTS LDST+A +V++L IA++G ++ +IHQPS
Sbjct: 293 GGERKRVCIGNEIMTNPSLLLLDEPTSSLDSTTALKIVQMLHCIAKAGKTIVTTIHQPSS 352
Query: 314 RILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGG 373
R+ D+++ LSRG +Y G ++ S+F+ G + N EF LDL+
Sbjct: 353 RLFHRFDKLVVLSRGSLLYFGKASEAMSYFSSIGCSPLLAMNPAEFLLDLVNGNMNDISV 412
Query: 374 TKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASIS---RGKLVSGATNTATT 430
+L E K + +++ +V E L+EA I+ + KL++
Sbjct: 413 PSALKEKMKIIR--LELYVRNVKCDVET---QYLEEAYKTQIAVMEKMKLMAPVPLDEEV 467
Query: 431 TPSSMVP--TFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLD- 487
P + +W + LS R + RR +R+ V+ T IL ++W D
Sbjct: 468 KLMITCPKREWGLSWWEQYCLLSLRGIKE-RRHDYFSWLRVTQVLSTAIILGLLWWQSDI 526
Query: 488 NSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVS 547
S + + L FF A+ F+ A+ F QER + +E N YR S+Y ++
Sbjct: 527 TSQRPTRSGLLFF-IAVFWGFFPVFTAIFTFPQERAMLSKERESNMYRLSAYFVARTTSD 585
Query: 548 LPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGY 607
LP L + F V+ ++ GL A F L +F A + + +
Sbjct: 586 LPLDLILPVLFLVVVYFMAGLRLRAESFFLSVLTVFLCIVAAQGLGLAIGASLMDLKKAT 645
Query: 608 TIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDD 657
T+ + F+L G+F+ ++P + W ++S + Y+ +++ ++++
Sbjct: 646 TLASVTVMTFMLAGGYFV--KKVPFFIAWIRFMSFNYHTYKLLVKVQYEE 693
>At3g25620 membrane transporter, putative
Length = 646
Score = 258 bits (658), Expect = 1e-68
Identities = 184/610 (30%), Positives = 300/610 (49%), Gaps = 66/610 (10%)
Query: 60 PLHHALDVVGMEPRSLPFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEES 119
P H + + P +L F LTYS+K S + G+ E P
Sbjct: 48 PSHQSRQSSVLRQSLRPIILKFEELTYSIK--------SQTGKGSYWFGSQEPKP----- 94
Query: 120 VFSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALE 179
+ +L +SG + GE++A+LG SGSGK+TL+ ALA R+ +G+L GT++ NGE
Sbjct: 95 ----NRLVLKCVSGIVKPGELLAMLGPSGSGKTTLVTALAGRL-QGKLSGTVSYNGEPFT 149
Query: 180 SRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNA 239
S + K + +V QDD+L+P LTV ETLT+ A RLP+ L++ +K +V+ ++ LGL
Sbjct: 150 SSV-KRKTGFVTQDDVLYPHLTVMETLTYTALLRLPKELTRKEKLEQVEMVVSDLGLTRC 208
Query: 240 AKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQ 299
+VIG RG+SGGER+RVSIG +++ +P LL LDEPTSGLDST+A +V L+ +A+
Sbjct: 209 CNSVIGGGLIRGISGGERKRVSIGQEMLVNPSLLLLDEPTSGLDSTTAARIVATLRSLAR 268
Query: 300 SGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGH-PLPDSDNRTE 358
G V+ +IHQPS R+ + D+++ LS G +YSG ++ +F G+ P N +
Sbjct: 269 GGRTVVTTIHQPSSRLYRMFDKVLVLSEGCPIYSGDSGRVMEYFGSIGYQPGSSFVNPAD 328
Query: 359 FALDLIRDLEGSPGGTKSL--VEFNKSWQSMTKVHS--HSVSSQPERPNGMSLKEAISAS 414
F LDL G TK +E N + + +S S+ S ++ LKE +S +
Sbjct: 329 FVLDL---ANGITSDTKQYDQIETNGRLDRLEEQNSVKQSLISSYKKNLYPPLKEEVSRT 385
Query: 415 ISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMV 474
+ + A ++ + +W++ L KR
Sbjct: 386 FPQDQ------TNARLRKKAITNRWPTSWWMQFSVLLKRGL------------------- 420
Query: 475 TGFILATMFWNLDNSPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAYNA 533
++W+ + +Q+++G F F++ F+ +A+ F QER + ++E +
Sbjct: 421 -------LWWH--SRVAHLQDQVGLLFFFSIFWGFFPLFNAIFTFPQERPMLIKERSSGI 471
Query: 534 YRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFV 593
YR SSY I+ + LP L F IT+W GL + F+ +I+ +
Sbjct: 472 YRLSSYYIARTVGDLPMELILPTIFVTITYWMGGLKPSLTTFIMTLMIVLYNVLVAQGVG 531
Query: 594 TFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAV--L 651
L ++ T+ ++ FLL G++I IP + W Y+S Y Y+ + +
Sbjct: 532 LALGAILMDAKKAATLSSVLMLVFLLAGGYYI--QHIPGFIAWLKYVSFSHYCYKLLVGV 589
Query: 652 QNEFDDPVKC 661
Q +D+ +C
Sbjct: 590 QYTWDEVYEC 599
>At1g31770 unknown protein
Length = 648
Score = 258 bits (658), Expect = 1e-68
Identities = 176/590 (29%), Positives = 283/590 (47%), Gaps = 41/590 (6%)
Query: 69 GMEPRSLPFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLL 128
G++ P L F + Y VK+ E S S+ KT+L
Sbjct: 43 GLQMSMYPITLKFEEVVYKVKI--------------------EQTSQCMGSWKSKEKTIL 82
Query: 129 NDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISA 188
N I+G GE +A+LG SGSGK+TL+ AL R++K G + NG+ S +K +
Sbjct: 83 NGITGMVCPGEFLAMLGPSGSGKTTLLSALGGRLSK-TFSGKVMYNGQPF-SGCIKRRTG 140
Query: 189 YVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEG 248
+V QDD+L+P LTV ETL F A RLP +L++ +K V +I +LGL ++IG
Sbjct: 141 FVAQDDVLYPHLTVWETLFFTALLRLPSSLTRDEKAEHVDRVIAELGLNRCTNSMIGGPL 200
Query: 249 HRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSI 308
RG+SGGE++RVSIG +++ +P LL LDEPTSGLDST+A +V ++R+A G V+ +I
Sbjct: 201 FRGISGGEKKRVSIGQEMLINPSLLLLDEPTSGLDSTTAHRIVTTIKRLASGGRTVVTTI 260
Query: 309 HQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLE 368
HQPS RI + D+++ LS G +Y G+ + +F+ G + N + LDL
Sbjct: 261 HQPSSRIYHMFDKVVLLSEGSPIYYGAASSAVEYFSSLGFSTSLTVNPADLLLDL---AN 317
Query: 369 GSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKLVSGATNTA 428
G P T+ S Q V VS+ + + E +A + A
Sbjct: 318 GIPPDTQK----ETSEQEQKTVKETLVSAYEKNISTKLKAELCNAESHSYEYTKAAAKNL 373
Query: 429 TTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDN 488
+ + +W + L +R + RR +R+ V+ F+ ++W+
Sbjct: 374 KS------EQWCTTWWYQFTVLLQRGVRE-RRFESFNKLRIFQVISVAFLGGLLWWHTPK 426
Query: 489 SPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVS 547
S +Q+R F F++ FY +A+ F QE+ + ++E + YR SSY ++ +
Sbjct: 427 S--HIQDRTALLFFFSVFWGFYPLYNAVFTFPQEKRMLIKERSSGMYRLSSYFMARNVGD 484
Query: 548 LPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGY 607
LP L AF I +W GL + F+ L++ S ++ ++
Sbjct: 485 LPLELALPTAFVFIIYWMGGLKPDPTTFILSLLVVLYSVLVAQGLGLAFGALLMNIKQAT 544
Query: 608 TIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDD 657
T+ FL+ G+++ +IP + +W YLS Y Y+ +L ++ D
Sbjct: 545 TLASVTTLVFLIAGGYYV--QQIPPFIVWLKYLSYSYYCYKLLLGIQYTD 592
>At1g51460 ATP-dependent transmembrane transporter like protein
Length = 678
Score = 256 bits (655), Expect = 2e-68
Identities = 179/587 (30%), Positives = 295/587 (49%), Gaps = 32/587 (5%)
Query: 124 TKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEAL----E 179
TK LLN ++G I+A++G SGSGKSTL+DALA GRL G + ++G+ L +
Sbjct: 27 TKRLLNGVNGCGEPNRILAIMGPSGSGKSTLLDALA-----GRLAGNVVMSGKVLVNGKK 81
Query: 180 SRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNA 239
RL +AYV Q+D+L LTV E+++++A RLP L++ + V+A I +GL
Sbjct: 82 RRLDFGAAAYVTQEDVLLGTLTVRESISYSAHLRLPSKLTREEISDIVEATITDMGLEEC 141
Query: 240 AKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQ 299
+ IG+ RG+SGGE++R+SI ++++ P LLFLDEPTSGLDS SAF VV++L+ IA
Sbjct: 142 SDRTIGNWHLRGISGGEKKRLSIALEVLTKPSLLFLDEPTSGLDSASAFFVVQILRNIAS 201
Query: 300 SGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEF 359
SG V+ SIHQPS + L D ++ LS G+TVY G FF E G P P N ++
Sbjct: 202 SGKTVVSSIHQPSGEVFALFDDLLLLSGGETVYFGEAESATKFFGEAGFPCPSRRNPSDH 261
Query: 360 ALDLIR-DLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQP--ERPNG-------MSLKE 409
L + D + N S S+ ++H + + P + P K
Sbjct: 262 FLRCVNSDFDNVTAALVESRRINDSSFSLHQLHETTNTLDPLDDIPTAEIRTTLVRKFKC 321
Query: 410 AISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRL 469
++ A+ SR ++ A+ T +W ++ L++RSF + R + +R+
Sbjct: 322 SLYAAASRARIQEIASIVGIVTERK--KGSQTNWWKQLRILTQRSFINMSRDLGYYWMRI 379
Query: 470 GAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRET 529
+V + ++F+N+ + V F + + F++E +F RE
Sbjct: 380 AVYIVLSICVGSIFFNVGRNHTNVMSTAACGGFMAGFMTFMSIGGFQSFIEEMKVFSRER 439
Query: 530 AYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAG 589
Y + Y +S+ L SLP + + L+ + IT + V G S F + L + +
Sbjct: 440 LNGHYGVAVYTVSNLLSSLPFIILMCLSTSSITIYMVRFQSGGSHFFYNCLDLICAITTV 499
Query: 590 NSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEA 649
S + ++ VVP+ ++G + + +L +GFF +P + W + +S + Y A
Sbjct: 500 ESCMMMIASVVPNFLMGVMLGAGYIGIMVLSAGFFRFFPDLPMVF-WRYPVSYINYGAWA 558
Query: 650 VLQNEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLLGSMSGTLGTN 696
LQ + + + + + +SPL VP ++K +L+ + LG N
Sbjct: 559 -LQGAYKNEM------IGVEYDSPLPLVP-KMKGELI--LQTVLGIN 595
>At4g27420 putative protein
Length = 635
Score = 254 bits (649), Expect = 1e-67
Identities = 185/622 (29%), Positives = 292/622 (46%), Gaps = 84/622 (13%)
Query: 58 ETPLHHALDVVGMEPRSLPF----------VLSFSNLTYSVKVKRKLSFSSIFPRRRNRL 107
ETP+ D RSLPF L F NL Y+VK+K +
Sbjct: 11 ETPIAKTND-----DRSLPFSIFKKANNPVTLKFENLVYTVKLK-------------DSQ 52
Query: 108 GAVEDAPAVEESVFSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIA--KG 165
G EE +T+L ++G + GEI+A+LG SGSGK++L+ AL R+ KG
Sbjct: 53 GCFGKNDKTEE------RTILKGLTGIVKPGEILAMLGPSGSGKTSLLTALGGRVGEGKG 106
Query: 166 RLKGTLALNGEALESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKA 225
+L G ++ N + L S+ +K + +V QDD L+P LTV ETL F A RLP + K +K
Sbjct: 107 KLTGNISYNNKPL-SKAVKRTTGFVTQDDALYPNLTVTETLVFTALLRLPNSFKKQEKIK 165
Query: 226 RVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDST 285
+ +A++ +LGL T+IG RGVSGGER+RVSIG +I+ +P LLFLDEPTSGLDST
Sbjct: 166 QAKAVMTELGLDRCKDTIIGGPFLRGVSGGERKRVSIGQEILINPSLLFLDEPTSGLDST 225
Query: 286 SAFMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAE 345
+A +V +L +A+ G V+ +IHQP S+G VY G + +FA
Sbjct: 226 TAQRIVSILWELARGGRTVVTTIHQP--------------SKGNPVYFGLGSNAMDYFAS 271
Query: 346 FGH-PLPDSDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNG 404
G+ PL + N ++F LD+ G P + W S+ S + +RP
Sbjct: 272 VGYSPLVERINPSDFLLDI---ANGKP------LLVISCWPSVG-------SDESQRPEA 315
Query: 405 M----------SLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRS 454
M +L +++ + + ++ ++ + +W + L KR
Sbjct: 316 MKAALVAFYKTNLLDSVINEVKGQDDLCNKPRESSRVATNTYGDWPTTWWQQFCVLLKRG 375
Query: 455 FTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMS-TTFYTTAD 513
RR G+++ + + F+ ++W S +Q+++G F S F+
Sbjct: 376 L-KQRRHDSFSGMKVAQIFIVSFLCGLLWWQTKIS--RLQDQIGLLFFISSFWAFFPLFQ 432
Query: 514 ALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGAS 573
+ F QER + +E + YR S Y +S + LP L F VIT+W GL+ +
Sbjct: 433 QIFTFPQERAMLQKERSSGMYRLSPYFLSRVVGDLPMELILPTCFLVITYWMAGLNHNLA 492
Query: 574 GFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSY 633
F L++ L +V T+ I+ FLL G+++ +P +
Sbjct: 493 NFFVTLLVLLVHVLVSGGLGLALGALVMDQKSATTLGSVIMLTFLLAGGYYV--QHVPVF 550
Query: 634 WIWFHYLSLVKYPYEAVLQNEF 655
W Y+S+ Y Y+ ++ ++
Sbjct: 551 ISWIKYVSIGYYTYKLLILGQY 572
>At2g01320 putative ABC transporter
Length = 725
Score = 249 bits (635), Expect = 5e-66
Identities = 157/537 (29%), Positives = 273/537 (50%), Gaps = 31/537 (5%)
Query: 127 LLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKG---RLKGTLALNGEALESRLL 183
LL ++SGEA+ G ++A++G SGSGK+TL++ LA +++ L G L +NG+ S+
Sbjct: 90 LLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLSLSPRLHLSGLLEVNGKPSSSKAY 149
Query: 184 KVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTV 243
K+ A+V Q+DL F LTV ETL+FAAE +LP S ++ V L+ +LGL + A +
Sbjct: 150 KL--AFVRQEDLFFSQLTVRETLSFAAELQLPEISSAEERDEYVNNLLLKLGLVSCADSC 207
Query: 244 IGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSI 303
+GD RG+SGGE++R+S+ ++I P ++F DEPT+GLD+ A V++ LQ++AQ G
Sbjct: 208 VGDAKVRGISGGEKKRLSLACELIASPSVIFADEPTTGLDAFQAEKVMETLQKLAQDGHT 267
Query: 304 VIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLP-SFFAEFGHPLPDSDNRTEFALD 362
VI SIHQP + D ++ L+ G VY+G + P ++F FG P+ N EF D
Sbjct: 268 VICSIHQPRGSVYAKFDDIVLLTEGTLVYAGPAGKEPLTYFGNFGFLCPEHVNPAEFLAD 327
Query: 363 LIRDLEGSPGGTKSLVEFNKS---WQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGK 419
LI V+++ S + S +VH+ V + +R + + +S
Sbjct: 328 LIS------------VDYSSSETVYSSQKRVHA-LVDAFSQRSSSVLYATPLS------- 367
Query: 420 LVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFIL 479
+ T + + +W + L KR++ + R +R + + I
Sbjct: 368 -MKEETKNGMRPRRKAIVERTDGWWRQFFLLLKRAWMQASRDGPTNKVRARMSVASAVIF 426
Query: 480 ATMFWNLDNSPKGVQERLGFF-AFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSS 538
++FW + S +Q+R+G A++T + VF +ER I RE + +Y
Sbjct: 427 GSVFWRMGKSQTSIQDRMGLLQVAAINTAMAALTKTVGVFPKERAIVDRERSKGSYSLGP 486
Query: 539 YLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSG 598
YL+S + +P A L F + + L+ S F + I+ +A ++ +
Sbjct: 487 YLLSKTIAEIPIGAAFPLMFGAVLYPMARLNPTLSRFGKFCGIVTVESFAASAMGLTVGA 546
Query: 599 VVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEF 655
+VP + +++ F++ G++++ D P + W SL+++ ++ + NEF
Sbjct: 547 MVPSTEAAMAVGPSLMTVFIVFGGYYVNADNTPIIFRWIPRASLIRWAFQGLCINEF 603
>At1g15210 putative ABC transporter
Length = 1442
Score = 210 bits (534), Expect = 3e-54
Identities = 156/632 (24%), Positives = 278/632 (43%), Gaps = 67/632 (10%)
Query: 41 LLKHVGDARKEAAGDGSETPLHHALDVVGMEPRSLPFVLSFSNLTYSVKVKRKLSFSSIF 100
L K + K AG ET + GM P +SF ++ Y V + ++
Sbjct: 802 LPKEEDEEAKGKAGSNKETEMESVSAKKGMVLPFTPLAMSFDDVKYFVDMPAEMR----- 856
Query: 101 PRRRNRLGAVEDAPAVEESVFSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALAN 160
E+ V LL ++ R G + A++G SG+GK+TL+D LA
Sbjct: 857 ----------------EQGVQETRLQLLKGVTSAFRPGVLTALMGVSGAGKTTLMDVLAG 900
Query: 161 RIAKGRLKGTLALNGEALESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSK 220
R G ++G + ++G + IS Y Q D+ P +TV E+L F+A RL + +SK
Sbjct: 901 RKTGGYIEGDVRVSGFPKKQETFARISGYCEQTDIHSPQVTVRESLIFSAFLRLAKEVSK 960
Query: 221 SKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTS 280
K V +++ + L + ++G G G+S +R+R++I ++++ +P ++F+DEPTS
Sbjct: 961 EDKLMFVDQVMELVELVDLRDAIVGLPGVTGLSTEQRKRLTIAVELVANPSIIFMDEPTS 1020
Query: 281 GLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFLSRG-QTVYSGSPTQL 339
GLD+ +A +V++ ++ +G V+ +IHQPS I D ++ + RG +YSG
Sbjct: 1021 GLDARAAAIVMRAVRNTVDTGRTVVCTIHQPSIDIFEAFDELLLMKRGGHVIYSG----- 1075
Query: 340 PSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQP 399
PL + ++ ++ E PG K ++N + + + S
Sbjct: 1076 ---------PLGRNSHK------VVEYFESFPGVPKIPEKYNPATWML-----EASSLAA 1115
Query: 400 ERPNGMSLKEAISASI--SRGKLV--------SGATNTATTTPSSMVPTFANPFWIEMVT 449
E G+ E AS R K + GAT+ T F+ W + +
Sbjct: 1116 ELKLGVDFAELYKASALCQRNKALVQELSVPPQGATDLYFATQ------FSQNTWGQFKS 1169
Query: 450 LSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERL----GFFAFAMS 505
+ + R P+ +R + T ++ ++FW + VQ+ +A +
Sbjct: 1170 CLWKQWWTYWRSPDYNLVRFIFTLATSLMIGSVFWQIGGKRSNVQDLTMVIGAIYAAVVF 1229
Query: 506 TTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWA 565
+ P+ ER +F RE A Y Y IS LP + + +++I +
Sbjct: 1230 VGINNCSTVQPMVAVERTVFYREKAAGMYSAIPYAISQVTCELPYVLIQTTYYSLIIYSM 1289
Query: 566 VGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFI 625
VG + AS FL++ I + SF + + P+ + A F L SGFFI
Sbjct: 1290 VGFEWKASKFLWFIFINYFSFLYWTYYGMMTVSLTPNQQVASIFASAFYGIFNLFSGFFI 1349
Query: 626 DRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDD 657
R +IP +W+W++++ V + ++ +++ D
Sbjct: 1350 PRPKIPKWWVWYYWICPVAWTIYGLITSQYGD 1381
Score = 178 bits (451), Expect = 1e-44
Identities = 143/571 (25%), Positives = 259/571 (45%), Gaps = 55/571 (9%)
Query: 126 TLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGR-LKGTLALNGEALESRLLK 184
T+L D+SG + + +LG SGK+TL+ ALA ++ K + G + NG L +
Sbjct: 183 TILKDVSGIVKPSRMTLLLGPPSSGKTTLLLALAGKLDKSLDVSGEVTYNGYRLNEFVPI 242
Query: 185 VISAYVMQDDLLFPMLTVEETLTFAAE-------FRLPRTLSKSKKKARV--QALIDQ-- 233
SAY+ Q+DL ++TV+ETL F+A + L L++ +K A + +A +D
Sbjct: 243 KTSAYISQNDLHVGIMTVKETLDFSARCQGVGTRYDLLNELARREKDAGIFPEADVDLFM 302
Query: 234 --------------------LGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILL 273
LGL T++GD+ RG+SGG+++RV+ G I+ L
Sbjct: 303 KASAAQGVKSSLITDYTLKILGLDICKDTIVGDDMMRGISGGQKKRVTTGEMIVGPTKTL 362
Query: 274 FLDEPTSGLDSTSAFMVVKVLQRIAQ-SGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVY 332
F+DE ++GLDS++ F +VK LQ+I + + V++S+ QP+ L D +I LS GQ VY
Sbjct: 363 FMDEISTGLDSSTTFQIVKCLQQIVHLTEATVLISLLQPAPETFDLFDDIILLSEGQIVY 422
Query: 333 SGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHS 392
G + FF FG P+ +F ++++ + V+ N+ ++ + S
Sbjct: 423 QGPRDHILEFFESFGFKCPERKGTADF----LQEVTSKKDQEQYWVDPNRPYRYIPV--S 476
Query: 393 HSVSSQPERPNGMSLKEAISASISRGKLVSGAT--NTATTTPSSMVPTFANPFWIEMVTL 450
SS + G L +S + K A + + + ++ + + W+ M
Sbjct: 477 EFASSFKKFHVGSKLSNELSVPYDKSKSHKAALMFDKYSIKKTELLKSCWDKEWMLM--- 533
Query: 451 SKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQER---LGFFAFAMSTT 507
+R + + +++ I +T++ + + + +G FAM
Sbjct: 534 --------KRNSFFYVFKTVQIIIIAAITSTLYLRTEMHTRNEIDANIYVGSLLFAMIVN 585
Query: 508 FYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVG 567
+ + + +Q +F ++ + +Y + L+ +P F S A+ V+T++++G
Sbjct: 586 MFNGLAEMAMTIQRLPVFYKQRDLLFHPPWTYTLPTFLLGIPISIFESTAWMVVTYYSIG 645
Query: 568 LDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDR 627
A F FLIIF F++ + + T V +L L GF + R
Sbjct: 646 YAPDAERFFKQFLIIFLIQQMAAGIFRFIASTCRTMTIANTGGVLVLLVVFLTGGFLLPR 705
Query: 628 DRIPSYWIWFHYLSLVKYPYEAVLQNEFDDP 658
IP +W W +++S + Y + A+ NE P
Sbjct: 706 SEIPVWWRWAYWISPLSYAFNAITVNELFAP 736
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.137 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,996,762
Number of Sequences: 26719
Number of extensions: 688284
Number of successful extensions: 2265
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 111
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 1815
Number of HSP's gapped (non-prelim): 240
length of query: 751
length of database: 11,318,596
effective HSP length: 107
effective length of query: 644
effective length of database: 8,459,663
effective search space: 5448022972
effective search space used: 5448022972
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0260.10


