

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0258b.1
(207 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
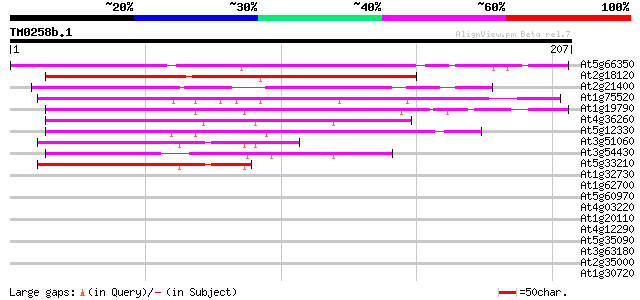
Score E
Sequences producing significant alignments: (bits) Value
At5g66350 zinc finger protein SHI-like 142 1e-34
At2g18120 unknown protein 140 6e-34
At2g21400 unknown protein 136 9e-33
At1g75520 lateral root primordium protein LRP1 like 133 8e-32
At1g19790 hypothetical protein 125 2e-29
At4g36260 STYLISH2 protein (STY2) 123 8e-29
At5g12330 lateral root primordia (LRP1) 111 3e-25
At3g51060 STYLISH 1 protein (STY1) 92 3e-19
At3g54430 putative protein 87 6e-18
At5g33210 putative protein 85 2e-17
At1g32730 unknown protein 39 0.001
At1g62700 unknown protein 36 0.016
At5g60970 DNA binding protein - like 33 0.14
At4g03220 hypothetical protein 31 0.40
At1g20110 unknown protein 31 0.53
At4g12290 copper amine oxidase -like protein 30 0.90
At5g35090 unknown protein 30 1.2
At3g63180 hypothetical protein 30 1.2
At2g35000 putative RING zinc finger protein 28 2.6
At1g30720 reticuline oxidase-like protein 28 3.4
>At5g66350 zinc finger protein SHI-like
Length = 331
Score = 142 bits (358), Expect = 1e-34
Identities = 87/234 (37%), Positives = 123/234 (52%), Gaps = 38/234 (16%)
Query: 1 MSMMLGQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLM 60
M M G G CQ+CGNQ+KK C++ RCRTCC ++G C THV+STW+P +RR R
Sbjct: 107 MMMRSGSGSGGPSCQDCGNQSKKDCSHMRCRTCCKSRGLDCPTHVKSTWVPAAKRRER-- 164
Query: 61 EHQPPPTTNNPHHLHEDIPQSHNQ----------------NPFTSLEELKFPEAMSSMAV 104
Q T P L +P+ + N + LE FP +SS AV
Sbjct: 165 -QQQLSTGQQPQQLGGSVPKRQRERIPARSTSMAYTRIPSNNTSGLEVGNFPPEVSSSAV 223
Query: 105 FSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIH 164
F VRV S+DD E AY+T+V+IGGH F G+LYDQGP ++S + Q LNL T
Sbjct: 224 FRCVRVSSVDDEEEEYAYKTAVSIGGHVFKGVLYDQGPAERSSSGG---GSQPLNLIT-- 278
Query: 165 SHDGATMAPPSSS-------ATAAH-----ELFFPQPRSLASFRSGVPYFSHTK 206
+ A+ + P+ S +T+ H L +P P + +F +G +FS+++
Sbjct: 279 AGPSASSSSPNVSCNNGVVGSTSDHYIDPASLNYPTP--INTFMTGTHFFSNSR 330
>At2g18120 unknown protein
Length = 222
Score = 140 bits (352), Expect = 6e-34
Identities = 68/144 (47%), Positives = 91/144 (62%), Gaps = 9/144 (6%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNPHH 73
CQECGNQAKKGC + RCRTCC + G C THVRSTWIP+ +RR R + Q P T+NP
Sbjct: 72 CQECGNQAKKGCTHGRCRTCCKSNGLHCPTHVRSTWIPIAKRRERQQQLQTP--TSNPTG 129
Query: 74 LHEDIPQSHNQNPFTSLE-------ELKFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSV 126
+ + + N +L+ E +FP+ +SS A+F VR+ DD + AYQT+V
Sbjct: 130 GSGRVGKYRDINQHATLDSSGLEMGETRFPDEVSSDALFRCVRMSGTDDGEGQYAYQTTV 189
Query: 127 NIGGHRFSGILYDQGPEQQSLNAS 150
I GH F GILY+QGPE +S+ ++
Sbjct: 190 GIAGHLFKGILYNQGPENKSMRST 213
>At2g21400 unknown protein
Length = 174
Score = 136 bits (342), Expect = 9e-33
Identities = 72/170 (42%), Positives = 98/170 (57%), Gaps = 23/170 (13%)
Query: 9 VKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTT 68
+ G +C++CGNQAKK C Y RCRTCC +K F CQTH++STW+P RR H + Q P +T
Sbjct: 4 IMGRKCEDCGNQAKKDCVYMRCRTCCKSKAFHCQTHIKSTWVPAYRRSHHKHQSQ-PLST 62
Query: 69 NNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSVNI 128
+ P + H FP +SS+A F V+V S+DD + AYQT+VNI
Sbjct: 63 SIPKGVQIHTTPGH------------FPAELSSLADFRCVKVSSIDDGKEQYAYQTTVNI 110
Query: 129 GGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSA 178
GGH F GIL+DQG L+ +D H N N S++ + PS+S+
Sbjct: 111 GGHVFRGILHDQG-----LHKVMVDHHYNKN-----SNNHQELLTPSTSS 150
>At1g75520 lateral root primordium protein LRP1 like
Length = 346
Score = 133 bits (334), Expect = 8e-32
Identities = 84/231 (36%), Positives = 116/231 (49%), Gaps = 48/231 (20%)
Query: 11 GSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRL-----MEHQPP 65
G CQ+CGNQAKK C + RCRTCC ++GF CQTHV+STW+P +RR RL ++H
Sbjct: 122 GMNCQDCGNQAKKDCPHMRCRTCCKSRGFHCQTHVKSTWVPAAKRRERLAQLASLQHHSA 181
Query: 66 PT--TNNPHHLHE------DIPQSH--------------NQNPFTSLE-ELKFPEAMSSM 102
+ T N L E + + H N N + LE P ++S
Sbjct: 182 SSRETQNAKRLREASGGDNNDDKDHSGGGGSALANTRVVNANSNSGLEVSQHLPPEVNSP 241
Query: 103 AVFSSVRVRSMDDSVNEM--AYQTSVNIGGHRFSGILYDQGPEQQ--------SLNASPL 152
A+F VRV S+++ ++ AYQT+VNIGGH F GILYDQGPE Q + A+
Sbjct: 242 AIFRCVRVSSIEEDEDDQAYAYQTAVNIGGHIFKGILYDQGPEHQDNHHLNLLASTATTT 301
Query: 153 DQHQNLNLTTIHSHDGATMAPPSSSATAAHELFFPQPRSLASFRSGVPYFS 203
+ + T +++ M PSS P L SF +G P+F+
Sbjct: 302 NVEETATKTVTGNNNNGLMLDPSSL----------YPAQLNSFIAGTPFFT 342
>At1g19790 hypothetical protein
Length = 345
Score = 125 bits (313), Expect = 2e-29
Identities = 84/235 (35%), Positives = 113/235 (47%), Gaps = 51/235 (21%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPT-----T 68
CQ+CGNQAKK C + RCRTCC ++GF CQTHV+STW+ +RR R + P
Sbjct: 119 CQDCGNQAKKDCPHMRCRTCCKSRGFDCQTHVKSTWVSAAKRRERQAQLAVLPAKRIRDA 178
Query: 69 NNPHHLHEDIPQSHNQN------------------PFTSLEELKFPEAMSSMAVFSSVRV 110
N+ +D ++ + LE P +SS AVF +RV
Sbjct: 179 NSRGGGDDDDDDKEDEKNDSCGGGSALACTRVVNASSSGLETSHLPPEISSPAVFRCMRV 238
Query: 111 RSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPE------QQSLNASPLDQHQNLNL---- 160
S+DD E AYQT+V+IGGH F GILYDQGP SLN QH +LNL
Sbjct: 239 SSIDDEDEEYAYQTAVSIGGHVFKGILYDQGPSSDHHRYSSSLNGETSHQH-HLNLMDST 297
Query: 161 ---------TTIHSHDGATMAPPSSSATAAHELFFPQPRSLASFRSGVPYFSHTK 206
T +++++G+ PSS TA F A G P+F+ ++
Sbjct: 298 PSAATTNAVTAVNTNNGS--IDPSSLYTAVATPF------NAFVAGGTPFFASSR 344
>At4g36260 STYLISH2 protein (STY2)
Length = 322
Score = 123 bits (308), Expect = 8e-29
Identities = 65/158 (41%), Positives = 87/158 (54%), Gaps = 23/158 (14%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNN--- 70
C++CGNQAKK C + RCRTCC ++GF C THVRSTWIPV RRR R + +
Sbjct: 94 CRDCGNQAKKDCTHMRCRTCCKSRGFDCSTHVRSTWIPVARRRERQQQLHMSTSGGGGGS 153
Query: 71 -------------PHHLHEDIPQSHNQNPFTS------LEELKFPEAMSSMAVFSSVRVR 111
H +P + + + S + E FP +SS A+F V++
Sbjct: 154 GSGGAGGGGSSIPKRHRDPTLPGTSSSSRLPSHSAGLEMGEASFPGEVSSDALFRCVKMS 213
Query: 112 SMDDSVN-EMAYQTSVNIGGHRFSGILYDQGPEQQSLN 148
+DD + + AYQT+VNIGGH F GILYDQGPE ++
Sbjct: 214 GVDDGGDGQYAYQTTVNIGGHLFKGILYDQGPESSYMS 251
>At5g12330 lateral root primordia (LRP1)
Length = 320
Score = 111 bits (277), Expect = 3e-25
Identities = 68/191 (35%), Positives = 89/191 (45%), Gaps = 33/191 (17%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHR-LMEHQPPPT----- 67
CQ+CGNQAKK C RCRTCC ++GF C THV+STW+ RRR R +M PT
Sbjct: 112 CQDCGNQAKKECKQRRCRTCCKSRGFDCSTHVKSTWVSAARRRERQVMPTGANPTAGSSL 171
Query: 68 -----TNNPHHLHEDIPQSHNQNPFTSLEEL-------------------KFPEAMSSMA 103
T P + Q TS +P + + A
Sbjct: 172 STSSGTKKPRIVGSQQQQQQQATSHTSTSNTPPQSFETSSSRQDGGGSREAWPGQVRAAA 231
Query: 104 VFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTI 163
VF VRV +++D +E AYQ V IGGH F G LYDQG E + S D H +
Sbjct: 232 VFKCVRVTAVEDGDDEYAYQAVVKIGGHVFKGFLYDQGLEPKEGFPSMSDLHLG---GSA 288
Query: 164 HSHDGATMAPP 174
++H+G + + P
Sbjct: 289 NNHNGVSASAP 299
>At3g51060 STYLISH 1 protein (STY1)
Length = 252
Score = 91.7 bits (226), Expect = 3e-19
Identities = 49/112 (43%), Positives = 63/112 (55%), Gaps = 17/112 (15%)
Query: 11 GSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLME-HQPPPTTN 69
G CQ+CGNQAKK C++ RCRTCC ++GF+C THVRSTW+P +RR R + P T
Sbjct: 141 GVSCQDCGNQAKKDCSHMRCRTCCKSRGFECSTHVRSTWVPAAKRRERQQQLATVQPQTQ 200
Query: 70 NPHHLHEDIPQSHNQN-PFTS-------------LEELKFPEAMSSMAVFSS 107
P E +P+ H +N P TS LE FP +SS AV +
Sbjct: 201 LPR--GESVPKRHRENLPATSSSLVCTRIPSHSGLEVGNFPAEVSSSAVLGA 250
>At3g54430 putative protein
Length = 183
Score = 87.0 bits (214), Expect = 6e-18
Identities = 48/140 (34%), Positives = 73/140 (51%), Gaps = 22/140 (15%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNPHH 73
C++CGN+AKK C + RCRTCC ++G+ C THV+STWIP R ++++P
Sbjct: 41 CRDCGNRAKKECLFERCRTCCKSRGYNCVTHVKSTWIPSSATR----------SSSSPSE 90
Query: 74 LHEDIPQSHNQNP-------FTSLEELKF----PEAMSSMAVFSSVRVRSMDDSVN-EMA 121
+ + +P TS +E F P + + AVF RV ++ ++ E+
Sbjct: 91 RKKKLKIDKQSSPNVSLLPTTTSRQERGFREGLPGKIEAPAVFKRTRVTAISNNEQAEIG 150
Query: 122 YQTSVNIGGHRFSGILYDQG 141
YQ +V I GH F G L+ G
Sbjct: 151 YQATVTISGHIFKGFLHYYG 170
>At5g33210 putative protein
Length = 173
Score = 85.1 bits (209), Expect = 2e-17
Identities = 40/81 (49%), Positives = 52/81 (63%), Gaps = 4/81 (4%)
Query: 11 GSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLME-HQPPPTTN 69
G CQ+ GNQAKK C++ RCRTCC ++GF+C THVRSTW+P +RR R + P T
Sbjct: 49 GVSCQDFGNQAKKDCSHMRCRTCCKSRGFECSTHVRSTWVPATKRRERQQQLATVQPQTQ 108
Query: 70 NPHHLHEDIPQSHNQN-PFTS 89
P E +P+ H +N P TS
Sbjct: 109 LPR--GESVPKRHRENLPATS 127
>At1g32730 unknown protein
Length = 327
Score = 39.3 bits (90), Expect = 0.001
Identities = 36/167 (21%), Positives = 67/167 (39%), Gaps = 20/167 (11%)
Query: 13 RCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHV--------RSTWIP-VDRRRHRLMEHQ 63
+C +CGN A+ C + C+ CC+ C HV T P + E
Sbjct: 30 KCIQCGNVARSRCPFQSCKGCCSRAENPCPIHVLKVASTSGEKTQAPSTPSSEQKATEGT 89
Query: 64 PPPTT--NNPHHLHEDIPQSHNQNPFTSLEE---LKFPEAMSSMAVFSSVRVRSMDDSVN 118
P TT ++ L + Q +N N + + +K +A++ R + V
Sbjct: 90 PGSTTRVSSIRQLSSNFAQFNNLNASSRQRKPLTIKDAQALNEWRFTKLKEYRDRNIEVE 149
Query: 119 EMA---YQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTT 162
A Y ++VN+ FS + P+++S + +Q++ N+ +
Sbjct: 150 NEAFDRYMSNVNLLEEAFS---FTSVPDEESHGTAAPEQNKEENIVS 193
>At1g62700 unknown protein
Length = 394
Score = 35.8 bits (81), Expect = 0.016
Identities = 34/144 (23%), Positives = 61/144 (41%), Gaps = 24/144 (16%)
Query: 35 NNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELK 94
NN QC+ + + +++++HQ HHL E + F L +L+
Sbjct: 221 NNPNLQCKQELELHY-------NQMVQHQ-----QQNHHLRESM--------FLQLPQLE 260
Query: 95 FPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQ 154
P + + ++ R S + ++++ + G FS + YDQG EQ + + LD+
Sbjct: 261 SPTSNCNSDNNNNTRNISNLQKSSNISHEEQLQQGNQSFSSLYYDQGVEQMTTDWRVLDK 320
Query: 155 HQNLNLTTIHSHDGATMAPPSSSA 178
L S+D A SSS+
Sbjct: 321 FVASQL----SNDEEAAAVVSSSS 340
>At5g60970 DNA binding protein - like
Length = 360
Score = 32.7 bits (73), Expect = 0.14
Identities = 21/66 (31%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Query: 138 YDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATAAHELFF-PQPRSLASFR 196
Y+ G QQSL+ S + ++ ++++ + PPSSSA +LFF P P +++S
Sbjct: 232 YNLGHLQQSLDQSGNNVTVAISNVAANNNNNLNLHPPSSSAGDGSQLFFGPTPPAMSSLF 291
Query: 197 SGVPYF 202
P F
Sbjct: 292 PTYPSF 297
>At4g03220 hypothetical protein
Length = 498
Score = 31.2 bits (69), Expect = 0.40
Identities = 28/104 (26%), Positives = 47/104 (44%), Gaps = 12/104 (11%)
Query: 87 FTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQ-Q 145
FTS+ +LK P++ SS +++ + +RS DS N + + V + + ++ Q Q
Sbjct: 68 FTSINDLKNPKSFSSNSIYKVLSLRSHRDSNNLRSLRFRVPVTFTSLNSLIRLAVTHQVQ 127
Query: 146 SLNASPLDQ-----------HQNLNLTTIHSHDGATMAPPSSSA 178
L+ + QNL T+ S + PPSSSA
Sbjct: 128 DLDIEVTTKDYFNFPRWIVTSQNLRALTLKSANLGFRLPPSSSA 171
>At1g20110 unknown protein
Length = 601
Score = 30.8 bits (68), Expect = 0.53
Identities = 25/64 (39%), Positives = 29/64 (45%), Gaps = 12/64 (18%)
Query: 140 QGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATAAHELFFPQPRSLASFRSGV 199
Q P Q + SP DQHQ G + APP SSA A + P P S +S S
Sbjct: 138 QQPTSQHMYYSPYDQHQT---------SGYSSAPPPSSAPAPNP--NPAPYS-SSLYSAP 185
Query: 200 PYFS 203
PY S
Sbjct: 186 PYSS 189
>At4g12290 copper amine oxidase -like protein
Length = 756
Score = 30.0 bits (66), Expect = 0.90
Identities = 18/47 (38%), Positives = 25/47 (52%), Gaps = 2/47 (4%)
Query: 61 EHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSS 107
+H P P T N H+ D P +H +P T E K +SS A+F+S
Sbjct: 78 KHDPKPNTKNHDHV-SDTP-NHPLDPLTVSEINKIRSILSSHALFTS 122
>At5g35090 unknown protein
Length = 159
Score = 29.6 bits (65), Expect = 1.2
Identities = 32/133 (24%), Positives = 51/133 (38%), Gaps = 15/133 (11%)
Query: 67 TTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDS-------VNE 119
TTN HH H++IP +P T+ + + P M S R++ S ++
Sbjct: 27 TTN--HHHHDNIPDDLPSSPVTTEKSTQSPHG-GGMKTKSPRLTRTLSKSHEKCKSLIHR 83
Query: 120 MAYQTSVNIGGH----RFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPS 175
M +GGH R + P +LN D+ N++ + + P
Sbjct: 84 MGGGVG-GVGGHGKHIRRHTADFHYDPSSYALNFDKGDEDDNIDRFPLRNFSARLPHSPP 142
Query: 176 SSATAAHELFFPQ 188
SSA AA + Q
Sbjct: 143 SSAKAADSSYTVQ 155
>At3g63180 hypothetical protein
Length = 975
Score = 29.6 bits (65), Expect = 1.2
Identities = 19/65 (29%), Positives = 30/65 (45%), Gaps = 12/65 (18%)
Query: 140 QGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATAAHELFFPQPRSLASFRSGV 199
+GP Q++ D+ NL L + + P + +A H P P ++AS+ SGV
Sbjct: 348 EGPNQET------DRDSNLRLQNLD------WSQPQQAKSAPHSSVLPLPVAVASWPSGV 395
Query: 200 PYFSH 204
P H
Sbjct: 396 PPQGH 400
>At2g35000 putative RING zinc finger protein
Length = 378
Score = 28.5 bits (62), Expect = 2.6
Identities = 22/75 (29%), Positives = 31/75 (41%), Gaps = 4/75 (5%)
Query: 125 SVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATAAHEL 184
S + + +GIL+ P S S + NL+ T+ D +S T H
Sbjct: 239 STGLSSWQITGILF---PRSHSTGHSLIQPAGNLDRFTLRLPDDVRRQLMKTSRTMGHVA 295
Query: 185 FFPQPRSLAS-FRSG 198
PQ RS S +RSG
Sbjct: 296 LLPQARSSRSGYRSG 310
>At1g30720 reticuline oxidase-like protein
Length = 527
Score = 28.1 bits (61), Expect = 3.4
Identities = 20/87 (22%), Positives = 38/87 (42%), Gaps = 6/87 (6%)
Query: 58 RLMEHQPP-PTTNNPHHLHEDIPQSHNQNPFTSLEELKF-----PEAMSSMAVFSSVRVR 111
R ++ QP PT+ N + S + + L+F P+ +S +A + ++
Sbjct: 31 RCLDRQPTDPTSPNSAVAYIPTNSSFTTVLRSRIPNLRFDKPTTPKPISVVAAATWTHIQ 90
Query: 112 SMDDSVNEMAYQTSVNIGGHRFSGILY 138
+ E++ Q + GGH F G+ Y
Sbjct: 91 AAVGCARELSLQVRIRSGGHDFEGLSY 117
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.129 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,049,781
Number of Sequences: 26719
Number of extensions: 213061
Number of successful extensions: 683
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 21
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 648
Number of HSP's gapped (non-prelim): 40
length of query: 207
length of database: 11,318,596
effective HSP length: 95
effective length of query: 112
effective length of database: 8,780,291
effective search space: 983392592
effective search space used: 983392592
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0258b.1


