

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0253.9
(677 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
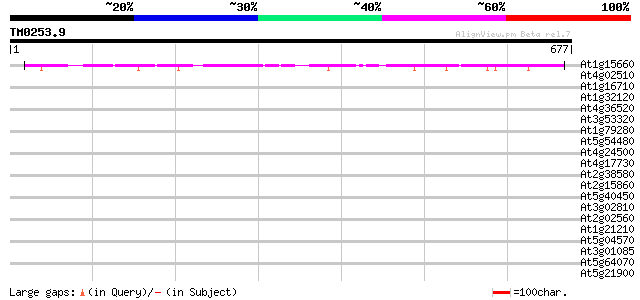
Score E
Sequences producing significant alignments: (bits) Value
At1g15660 unknown protein 215 7e-56
At4g02510 chloroplast protein import component Toc159-like 41 0.003
At1g16710 hypothetical protein 38 0.021
At1g32120 hypothetical protein 37 0.047
At4g36520 trichohyalin like protein 36 0.062
At3g53320 unknown protein 35 0.11
At1g79280 hypothetical protein 35 0.11
At5g54480 unknown protein 34 0.23
At4g24500 unknown protein 34 0.23
At4g17730 syntaxin 34 0.23
At2g38580 hypothetical protein 34 0.31
At2g15860 unknown protein 34 0.31
At5g40450 unknown protein 33 0.52
At3g02810 putative protein kinase 33 0.52
At2g02560 unknown protein 33 0.68
At1g21210 hypothetical protein 33 0.68
At5g04570 putative protein 32 0.89
At3g01085 putative protein 32 0.89
At5g64070 phosphatidylinositol 4-kinase (emb|CAB37928.1) 32 1.2
At5g21900 DNA excision repair protein 32 1.2
>At1g15660 unknown protein
Length = 710
Score = 215 bits (547), Expect = 7e-56
Identities = 213/717 (29%), Positives = 334/717 (45%), Gaps = 128/717 (17%)
Query: 18 DPLSNYRGLSLFPSTSSAL-----PLSVADDLDAIHNHLRSMALRSPAKLVEQTKAILEE 72
DPL Y GLSLFP T +L P ++DL H L+SM ++ EQ KAILE
Sbjct: 15 DPLQAYSGLSLFPRTLKSLSNPLPPSYQSEDLQQTHTLLQSMPFEIQSEHQEQAKAILE- 73
Query: 73 NPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTSVSLESLLPN 132
DVDVDV + R+RRPGL KR FSL +TS + P+
Sbjct: 74 ----------------DVDVDVQLNPIPNKRERRPGLDRKRKSFSLHLTTSQP-PPVAPS 116
Query: 133 LDIENLKDPVEFFKAHERMEN-----ARREIQKQMGVSVEQNQDDVSTRPRQRRPGLRGN 187
D +FF A+++ E A RE QKQ G SV Q++ +R R RRPG+ G
Sbjct: 117 FDPSKYPRSEDFFAAYDKFEPGFETVANREWQKQTGSSVIDIQENPPSR-RPRRPGIPGR 175
Query: 188 DQRSIK--YRHRYSTEI----SDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLENE 241
+R K + Y T++ + + + +AS+++ S + A +T+++ E
Sbjct: 176 KRRPFKESFTDSYFTDVINLEASEKEIPIASEQSLESATA-----------AHVTTVDRE 224
Query: 242 VTDSSATEGNKICEILDGLLRCNSEDLEGDGAMDLLQERLQIKPVVLEKVSVPDFPDNQP 301
V DS+ + +L LL C+ E+LEGDGA+ LL+ERLQIK +EK S+P+F D +
Sbjct: 225 VDDSTVDTDKDLNNVLKDLLACSREELEGDGAIKLLEERLQIKSFNIEKFSIPEFQDVRK 284
Query: 302 IDLKSLRGSLSKPRKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSL 361
++LK+ GS RK+LS+I N+L + N+ +R+++ SP+ Q
Sbjct: 285 MNLKA-SGSNPPNRKSLSDIQNIL-KGTNRVAVRKNSHSPSPQ----------------T 326
Query: 362 LNHISRLKPSVDPFSAHDIDHL-------STTKYSPVHK--MNQELNVVGSAKPSNELND 412
+ H S P VD FS DI +L S P+ K N VG+ ++ ND
Sbjct: 327 IKHFSSPNPPVDQFSFPDIHNLLPGDQQPSEVNVQPIAKDIPNTSPTNVGTVDVASPFND 386
Query: 413 HITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDILAVSESCLIED 472
+ K + GE ++ + + S S++ N D +++ I S + L ++
Sbjct: 387 SVVKRS---GEDDS---HIHSGIHRSHLSRDGNP-------DICVMDSISNRSSAMLQKN 433
Query: 473 AVMNSTSTS-QMPMED-----NSMEPEFDANVDRNEPHADMDVDNGGSGMGE--TVMDDT 524
M + +PM + N+ + E DA ++ + + + + TV +D+
Sbjct: 434 VDMRTKGKEVDVPMSESGANRNTGDRENDAEINEETDNLERLAECASKEVTRPFTVEEDS 493
Query: 525 V-------AKPNIEVNEQCQFEGKILTENTDAFTASMPTDDTDIDVVNPLADQSSPGV-- 575
+ +K EQ G L E+ + ++ + + L +++P V
Sbjct: 494 IPYQQGASSKSPNRAPEQYNTMGGSL-EHAEHNQGLHEEENVNTGSASGLQVENAPEVHK 552
Query: 576 ---IQANSIDKR------------TTCTNEGSEQCLQEERDGSRAPVEQKRVKSRSQKDT 620
Q N KR T G ++ ++ SRA +Q + KS +++
Sbjct: 553 YSHKQTNKRRKRGSSDSNVKKRSKTVHGETGGDKQMKTLPHESRAK-KQTKGKSNEREEK 611
Query: 621 KSKR--------LAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
K K+ + R+SLAAAGT G+RRSTRI++RPLEYW+GER +YGRIH+S
Sbjct: 612 KPKKTLTHEGKLFSCRKSLAAAGTKIEGGVRRSTRIKSRPLEYWRGERFLYGRIHES 668
>At4g02510 chloroplast protein import component Toc159-like
Length = 1503
Score = 40.8 bits (94), Expect = 0.003
Identities = 49/249 (19%), Positives = 105/249 (41%), Gaps = 19/249 (7%)
Query: 399 NVVGSAKPSNELNDHITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLI 458
N+VG K S+E++ + +D+ E P+ + + E D+ S K +A +
Sbjct: 79 NLVGDGKVSDEVDGSLKEDSTT-PEATPKPEVV-----SGETIGVDDVSSLSPKPEA-VS 131
Query: 459 EDILAVSESCLIEDAVMNSTSTSQMPMEDNSMEPEF-DANVDRNEPHADMDVDNGGSGMG 517
+ + V E+ +++ V + + +E+ S++ + A+ D DV+ G
Sbjct: 132 DGVGVVEENKKVKEDVEDIKDDGESKIENGSVDVDVKQASTDGESESKVKDVEEEDVGTK 191
Query: 518 ETVMDDTVAKPNIEVNEQC----QFEGKILTENTDAFTASMPTDDTDIDVVNP----LAD 569
+ ++ ++V+++ + EG LT+ D S P + +DV P + D
Sbjct: 192 KDDEGESELGGKVDVDDKSDNVIEEEGVELTDKGDVIVNSSPVESVHVDVAKPGVVVVGD 251
Query: 570 QSSPGVIQANSIDKRTTCTNEGSEQCLQEERDGSRAPVEQKRVKSRSQKDT-KSKRLAKR 628
++ N+ + N+ + + ++ G PV K ++ +K T +S +A
Sbjct: 252 AEGSEELKINADAETLEVANKFDQ--IGDDDSGEFEPVSDKAIEEVEEKFTSESDSIADS 309
Query: 629 QSLAAAGTS 637
L + TS
Sbjct: 310 SKLESVDTS 318
>At1g16710 hypothetical protein
Length = 1706
Score = 37.7 bits (86), Expect = 0.021
Identities = 37/184 (20%), Positives = 76/184 (41%), Gaps = 8/184 (4%)
Query: 360 SLLNHISRLKPSVDPFSAHDIDHLSTTKYSPVHKMNQELNVVGSAKPSNELNDHITKDAV 419
+L+ H K P + + ++ E + VGS S ND ++ +A
Sbjct: 697 ALIGHYKNCKDPRCPVCVPVKTYQQQANVRALARLKNESSAVGSVNRSVVSNDSLSANAG 756
Query: 420 AIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDILAVSESCLIEDA------ 473
A+ T DTL N + + K + S + I++ +E+ L +DA
Sbjct: 757 AVSGTPRCADTLDNLQPSLKRLKVEQSFQPVVPKTESCKSSIVSTTEADLSQDAERKDHR 816
Query: 474 -VMNSTSTSQMPMEDNSMEPEFDANVDRNEPHADMDVDNGGSGMGE-TVMDDTVAKPNIE 531
+ + T ++ + DNS++ F ++EP ++ S G+ + D+ + NI+
Sbjct: 817 PLKSETMEVKVEIPDNSVQAGFGIKETKSEPFENVPKPKPVSEPGKHGLSGDSPKQENIK 876
Query: 532 VNEQ 535
+ ++
Sbjct: 877 MKKE 880
>At1g32120 hypothetical protein
Length = 1206
Score = 36.6 bits (83), Expect = 0.047
Identities = 37/159 (23%), Positives = 60/159 (37%), Gaps = 12/159 (7%)
Query: 328 RKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHISRLKPSVDPFSAHDIDHLSTTK 387
R +K P + S + PPRS A S NH L+ S +P+ + + S+T+
Sbjct: 770 RVSKEPYMSSSLSVSSSSTRKKPPRSHEAANSRGYNHTPSLRASKEPYKSSSLSGSSSTR 829
Query: 388 YSPVH----KMNQELNVVGSAKPSNELNDHITKDAVAIGETNAVPDTLRNCASASENSKE 443
P ++ N S + S ELN + N P + N S +S
Sbjct: 830 KKPPRSHEASSSRGYNHPPSPRVSKELNKTPSISGSPSATRNKSPRSSENVNSCGNSSSG 889
Query: 444 DNSGKSSN--------KLDAPLIEDILAVSESCLIEDAV 474
NS N K ++P+ + + L+E+ V
Sbjct: 890 GNSVVKRNTDTSIIVSKQESPIKTQLDISKANKLLEEVV 928
>At4g36520 trichohyalin like protein
Length = 1432
Score = 36.2 bits (82), Expect = 0.062
Identities = 46/206 (22%), Positives = 86/206 (41%), Gaps = 14/206 (6%)
Query: 49 NHLRSMALRSPAKLVEQTKAILEENPGLFNSQD-DERNDVVDVDVDVAEDGQDFPRKRRP 107
N R R A+L ++ KA LE+ ++ ER + +V E ++ RK +
Sbjct: 749 NERRIKEAREKAELEQRLKATLEQEEKERQIKERQEREENERRAKEVLEQAEN-ERKLKE 807
Query: 108 GLGLKRARFSLKPSTSVSLESLLPNLDIENLKDPVEFFKAHERMENA--RREIQKQMGVS 165
L K LK E+ + + L++ +E + +R+ A R EI++++
Sbjct: 808 ALEQKENERRLK-------ETREKEENKKKLREAIELEEKEKRLIEAFERAEIERRLKED 860
Query: 166 VEQNQDDVSTRPRQRRPGLRGNDQR---SIKYRHRYSTEISDKNDHAMASQEAFGSDSLD 222
+EQ + + + + R L +Q + + +H YS E SD+ + E + +
Sbjct: 861 LEQEEMRMRLQEAKERERLHRENQEHQENERKQHEYSGEESDEKERDACEMEKTCETTKE 920
Query: 223 PDTEKIENGEACLTSLENEVTDSSAT 248
E+ N T ENE D+ +
Sbjct: 921 AHGEQSSNESLSDTLEENESIDNDVS 946
>At3g53320 unknown protein
Length = 553
Score = 35.4 bits (80), Expect = 0.11
Identities = 65/322 (20%), Positives = 111/322 (34%), Gaps = 65/322 (20%)
Query: 290 KVSVPDFPDNQPIDLKSLRGSLSKPRKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPT 349
+V P QP+ + L S+SKP LS + L N++ L D ++ S
Sbjct: 221 RVQGPGKATKQPVATRGLSTSISKPPNGLSKVRPLSTTSTNRSSL--DISKTQQEKNSKL 278
Query: 350 PP-RSPFAPLSSLLNHISRL--KPSVDPFSAHDIDHLSTTKYSPVHKMNQELNVVGSAKP 406
P + P P S+ + KP V PF + S++
Sbjct: 279 PAGKEPLGPRISMSRRAKPVLPKPGV-PFKS-------------------------SSRS 312
Query: 407 SNELNDHITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDILAVSE 466
S+ + +T +L +CASAS ++ S S K +
Sbjct: 313 SDASKNEMTSSC----------SSLESCASASSSASHKPSIDSIKKKN----------DS 352
Query: 467 SCLIEDAVMNSTSTSQMPMEDNSMEPEFDANVDRNEPHADMDVDNGGSGMGETVMDDTVA 526
S + + + STS+ M + P+ + P V GS + D +
Sbjct: 353 SSRLSSQPLANRSTSRGIMGQPRIPPQQTNKTSK--PKLSSSVPTAGS------ISDYSS 404
Query: 527 KPNIEVNEQCQFEGKILTENTDAFTASMPTDDTDIDVVNPLADQSSPGVIQANSID--KR 584
+ + G T + + +P +D + V PL + V+QA++ + KR
Sbjct: 405 ESSRASETSKMANGNQKTVSRE----KVPANDNTVQTVKPLKNSKDTSVVQADAKEGTKR 460
Query: 585 TTCTNEGSEQCLQEERDGSRAP 606
+ N G + G R P
Sbjct: 461 VSAINGGLVPSASAKPSGLRVP 482
>At1g79280 hypothetical protein
Length = 2111
Score = 35.4 bits (80), Expect = 0.11
Identities = 75/415 (18%), Positives = 145/415 (34%), Gaps = 64/415 (15%)
Query: 149 ERMENARREIQKQMGVS---VEQNQDDVSTRPRQRRPGLRGNDQRSIKYRHRYSTEISDK 205
E+ R E +K++ + V Q +D+V R++ L+ D+ K R + +
Sbjct: 1507 EQSVKEREEKEKRIQILDKYVHQLKDEV----RKKTEDLKKKDEELTKERSERKSVEKEV 1562
Query: 206 NDHAMASQEAFGSDSLDPDTEKIENGEACLTSLENEV-----TDSSATEGNKICEILDGL 260
D ++ +D + K+E + LT L E+ D + EG ++L G
Sbjct: 1563 GDSLTKIKKE--KTKVDEELAKLERYQTALTHLSEELEKLKHADGNLPEGTSAVQVLSGS 1620
Query: 261 LRCNSEDLEGDGAMDLLQE---------RLQIKPVVL-----------EKVSVPDFPDNQ 300
+ N + A++ + ++ KP + E ++ P +
Sbjct: 1621 I-LNDQAAAYVSAVEYFERVARSIASNSQVSTKPTDMVTEPSSGIPAAEPSTMTRVPSST 1679
Query: 301 PIDLKSLRGSLSKPRKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSS 360
P+ + + P+ A N + L +K T R+ +G + P SP +
Sbjct: 1680 PLIKSPVATTQQLPKVASDNKEKRLISQKPSTEFRRPSGRRIVRPQLVKPEESPKVDVD- 1738
Query: 361 LLNHISRLKPSVDPFSAHDIDHLSTTKYSPVHKM-----------NQELNVVGSAKPSNE 409
+ +AH+ + TT PV + + + + + S+E
Sbjct: 1739 -MPEAEGTGDEGKQPAAHEPESQVTTSVRPVQTLVRKRQADSLVSEPQQDSLTQGETSSE 1797
Query: 410 LNDHITKDA--------VAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDI 461
+ +K A + GE A + A+ + DN + + E +
Sbjct: 1798 IAPPASKKAKGSESHPDTSEGENLAKEPAIDELMDATTTTDGDNEETEAENAEEKTEEYV 1857
Query: 462 LAVSESCLIEDAVMNSTSTSQMPMEDNS--------MEPEFDANVDRNEPHADMD 508
A ++ E + T T +P E+ S EP D D+ E D+D
Sbjct: 1858 EAQQDNEADEPVEESPTETETIPTEEESRDQTEEENQEPLTDMESDKEEGELDLD 1912
>At5g54480 unknown protein
Length = 720
Score = 34.3 bits (77), Expect = 0.23
Identities = 42/199 (21%), Positives = 75/199 (37%), Gaps = 32/199 (16%)
Query: 78 NSQDDERNDVV-DVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTSVSLESLLPNLDIE 136
N +R VV D + +GQ+ K R GLG + +F PS S P
Sbjct: 111 NDDQSQRKPVVGDGTCGASNNGQEGMEKSREGLGFEGTQFVSDPSEDEMKSSQHP----- 165
Query: 137 NLKDPVEFFKAHERMENAR----------------REIQKQMGVSVEQNQDDVSTRPRQR 180
D + F+K+ M+ REI+++ G+ + + D ++ R+
Sbjct: 166 --YDILGFYKSVYNMQQCEIIELDHRDDEVESDGFREIREREGIPDLEPESDYNSLIRKN 223
Query: 181 RPGLRGNDQRSIKYRHRYSTEISDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLEN 240
+ + N ++ K R S+ ++ + D D E + EAC EN
Sbjct: 224 K---KKNKKKKKKKNVRESSSVASEIDKRDVEANTCNGQVAD---ETDNSNEACAQETEN 277
Query: 241 --EVTDSSATEGNKICEIL 257
E+ T ++ E++
Sbjct: 278 TPEIVTEEVTRSDEEVEVV 296
>At4g24500 unknown protein
Length = 319
Score = 34.3 bits (77), Expect = 0.23
Identities = 24/81 (29%), Positives = 36/81 (43%), Gaps = 16/81 (19%)
Query: 328 RKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHISRLKPSVDPFSAHDIDHLSTTK 387
+KNKTP +Q SP+ Q +SP PP+ P PSV P S ++ + T
Sbjct: 73 KKNKTPKQQYISSPSHQGSSPVPPQFP---------------PSVPPGSLCS-EYQAQTN 116
Query: 388 YSPVHKMNQELNVVGSAKPSN 408
+ H + E + PS+
Sbjct: 117 HGGFHAAHYEPRGMAHLSPSH 137
>At4g17730 syntaxin
Length = 255
Score = 34.3 bits (77), Expect = 0.23
Identities = 35/158 (22%), Positives = 64/158 (40%), Gaps = 13/158 (8%)
Query: 386 TKYSPVHKMNQELNVVGSAKPSNELNDHITKDAVAIGETNAVPDTLRNCASASENSKEDN 445
T S H++ +N +G+ K + EL + + K + IG+ V DT ASE +
Sbjct: 42 TSVSTFHRL---VNTLGTPKDTPELREKLHKTRLYIGQL--VKDTSAKLKEASETDHQRG 96
Query: 446 SGKSSNKLDAPLIEDILAVSESCLIEDAVMNSTSTSQMPMEDNSMEP------EFDANVD 499
+ +DA L +D AV + + T P+ P E D N D
Sbjct: 97 VNQKKKIVDAKLAKDFQAVLKEFQKAQRLAAERETVYAPLVHKPSLPSSYTSSEIDVNGD 156
Query: 500 RNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQ 537
++ + V++ + ++D+ +A + E+ Q
Sbjct: 157 KHPEQRALLVESKRQEL--VLLDNEIAFNEAVIEEREQ 192
>At2g38580 hypothetical protein
Length = 377
Score = 33.9 bits (76), Expect = 0.31
Identities = 37/205 (18%), Positives = 78/205 (38%), Gaps = 16/205 (7%)
Query: 439 ENSKEDNSGKSSNKLDAPLIEDILAVSESCLIEDAVMNSTSTSQMPMEDNSMEPEFDANV 498
++ K+ + K NK + ++D LA + +++ + Q+ ++ E + +
Sbjct: 3 DDKKKKRNKKKKNKQNNKRVDDALASGATTSVDENHIGDGDVPQISGGPDADESQSSHQI 62
Query: 499 DRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKI------LTENTDAFTA 552
D D V+N G E ++++T+ + E Q E K+ D
Sbjct: 63 DVVATEDDSGVENKSQG-SEVLLEETIKQLREENGSYLQKEEKLEERLVQYKNKNDMLLR 121
Query: 553 SMPTDDTDIDVVNPLADQSSPGVIQANSIDKRTTCTNEGSEQCLQEERDGSRA------P 606
M + + + L D+ S + S++K+ E + EE+
Sbjct: 122 EMSSTEAQ---MRQLLDERSTFTQKEASLEKKVQQLQHDEESLVAEEKSSREMISSLNNE 178
Query: 607 VEQKRVKSRSQKDTKSKRLAKRQSL 631
+ + R + + +KS L + QSL
Sbjct: 179 IARLRAQVTELEKSKSNLLEQNQSL 203
>At2g15860 unknown protein
Length = 512
Score = 33.9 bits (76), Expect = 0.31
Identities = 63/290 (21%), Positives = 111/290 (37%), Gaps = 47/290 (16%)
Query: 64 EQTKAILEENPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTS 123
E+ ++++E P + E+ V DVD P K G G + FS+
Sbjct: 9 EEKSSLVKEEPVRGEEPEIEKLTVADVDA---------PPKSTGGWGWGFSGFSVLSDLQ 59
Query: 124 VSLESLLPNLDI---ENLKDPVEFFKAHERMENARREIQKQMGVSVEQNQDDVSTR-PRQ 179
+ E + N + K E + E E++ +E +K EQ+ DD + + +
Sbjct: 60 KAAEDISRNAAAVAEKAAKSIAEMGEVDEDSESSAKEEEKTEEADTEQDSDDENAKLKKS 119
Query: 180 RRPGLRGNDQRSIKYRHRYSTEISDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLE 239
L G + S+ + ++ D S E+F S + ++ G + + LE
Sbjct: 120 ALERLEGASEESLLSQ---GLKVFDD------SVESFTSGAWQAFGNALKGGTSLVQKLE 170
Query: 240 NEVTDSSATE--GNKICEILDGLLRCNSEDLE-----GDGAMDLL---------QERLQI 283
N V S+ G+ +L+ ++ ++ G MDLL ++R+
Sbjct: 171 NSVQQGSSPREAGSGAPSLLETGKALTAKGMQVLEFVGKETMDLLITETGIGAEKDRVDF 230
Query: 284 KPVVLEKVSVPD--FPDNQPIDLKSLRGSLSKPRKALSNIDNLLNRRKNK 331
K VLE+V+ + P L+ L S+ L NRRK K
Sbjct: 231 KDQVLEEVTFDRCFYIYGGPEQLEELEA-------LASHYTLLFNRRKGK 273
>At5g40450 unknown protein
Length = 2910
Score = 33.1 bits (74), Expect = 0.52
Identities = 43/171 (25%), Positives = 70/171 (40%), Gaps = 26/171 (15%)
Query: 439 ENSKEDNSGKSSNKLDAPLIEDILAVSESCL-IEDAVMNSTSTSQMPMEDNSMEP----- 492
E SKED S + S IE+I A S+ L IE + ++T +S++ E + P
Sbjct: 1805 EESKEDKSQEISET-----IEEIEATSDQTLPIETSHTDNTLSSELVSEQDDQSPKKVEE 1859
Query: 493 ----------EFDANVDRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKI 542
+ +A +RN P D DN + ++ +T + ++ E E I
Sbjct: 1860 IHEEEPKEAHDVEATSERNLPVETSDADN---TLSSQLVSETKEEHKLQAGEILPTE-II 1915
Query: 543 LTENTDAFTASMPTDDTDIDVVNPLADQSSPGVIQANSIDKRTTCTNEGSE 593
E++D SM D D V D + V + N I + + + E E
Sbjct: 1916 PRESSDEALVSMLASRED-DKVALQEDNCADDVRETNDIQEERSISVETEE 1965
Score = 30.4 bits (67), Expect = 3.4
Identities = 55/253 (21%), Positives = 99/253 (38%), Gaps = 52/253 (20%)
Query: 381 DHLSTTKYSPVHKMNQELNVVGSAKPSNELNDHITKDAVAIGETNAVPDTLRNC------ 434
+ +S + SPV +++ E+ V SA S +A IGE V D ++
Sbjct: 1605 EEVSNDQQSPVEEISNEVIQVSSASLSEGPEYETVVEAEKIGE-EQVADKIQKSFETGEI 1663
Query: 435 ----ASASENSKEDNSGKSSNKLDAPLIEDILAVSESCLIEDAVMNSTSTSQMPMEDNSM 490
+S +S+E S K D ++D + + + + ++ T+ + +
Sbjct: 1664 VEAHSSLPSSSEEKEHETVSEKTDDEKVKDAEPIGD---MRERGLDIAETTHLSL----- 1715
Query: 491 EPEFDANVDRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILTENTDAF 550
P D D +E H +VA P ++EQ E TE +
Sbjct: 1716 -PSVDQKEDVDEIHI-----------------PSVALP---LDEQ---EKVTSTEKGETK 1751
Query: 551 TASMPTDDTDIDVVNPLADQSSPGVIQANSIDKRTTCTNEGSEQCLQEERDGS---RAPV 607
++ D D V +SP + + N + +T+ T+E + C+Q+E G+ P
Sbjct: 1752 SSEAEDDKPDEHV----DSSTSPMLSEKNDNETQTSKTSE--DVCMQQEESGTLEVPKPE 1805
Query: 608 EQKRVKSRSQKDT 620
E K KS+ +T
Sbjct: 1806 ESKEDKSQEISET 1818
>At3g02810 putative protein kinase
Length = 558
Score = 33.1 bits (74), Expect = 0.52
Identities = 35/123 (28%), Positives = 57/123 (45%), Gaps = 21/123 (17%)
Query: 119 KPSTSVSLESLLPNLDIENLKDPV---EFFKAHERMENARREIQKQMGVSVEQNQDDVST 175
K STS ES L E+ K+ V E+ K HE E++ E + + E +D
Sbjct: 389 KSSTSSGEESSL-----ESEKESVSKNEYKKKHEE-EDSSMESDDESDSNSEHEKD---- 438
Query: 176 RPRQRRPGLRGNDQRSIKYRHRYSTEISDKNDHAMASQEAFGSDS------LDPDTEKIE 229
+ +P N +S+K ++RYS E D ND ++S+ + S+ D D ++ E
Sbjct: 439 --QPPKPIDEKNQAQSLKIKYRYSWEDIDVNDERLSSKSSQKSNDESTSSRYDSDRDQDE 496
Query: 230 NGE 232
G+
Sbjct: 497 KGK 499
>At2g02560 unknown protein
Length = 1217
Score = 32.7 bits (73), Expect = 0.68
Identities = 43/206 (20%), Positives = 83/206 (39%), Gaps = 27/206 (13%)
Query: 425 NAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDILAVSESCLIEDAVMNSTSTSQMP 484
N VP + C SASEN +E S L++ L+ +S C D ++N T
Sbjct: 250 NTVPVLINYCTSASENDEELRE-YSLQALESFLLRCPRDISPYC---DEILNLT------ 299
Query: 485 MEDNSMEPEFDANVDRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILT 544
+E S +P F N++ + + ++ + E D+ + +C G I++
Sbjct: 300 LEYISYDPNFTDNMEEDTDNETLEDEEDDESANEYTDDEDASWKVRRAAAKC-LAGLIVS 358
Query: 545 EN---TDAFTASMP---------TDDTDIDVVNPLADQSSPGVIQANSIDKRTTCTNEGS 592
+ T + + P ++ +DV N D + Q ++ K T T+E S
Sbjct: 359 RSEMLTKVYQEACPKLIDRFKEREENVKMDVFNTFIDL----LRQTGNVTKGQTDTDESS 414
Query: 593 EQCLQEERDGSRAPVEQKRVKSRSQK 618
+ L ++ ++++ +S K
Sbjct: 415 PKWLLKQEVSKIVKSINRQLREKSVK 440
>At1g21210 hypothetical protein
Length = 738
Score = 32.7 bits (73), Expect = 0.68
Identities = 39/151 (25%), Positives = 60/151 (38%), Gaps = 20/151 (13%)
Query: 60 AKLVEQTKAILEE---NPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARF 116
A +V+ T L+ N GL N E++DV V + E L ++A
Sbjct: 568 ATMVQGTLGYLDPEYYNTGLLN----EKSDVYSFGVVLMEL-----------LSGQKALC 612
Query: 117 SLKPSTSVSLESLLPNLDIENLKDPVEFFKAHERMENARREIQKQMGVSVEQNQDDVSTR 176
+P TS + S + EN E EN +REIQK ++VE + R
Sbjct: 613 FERPQTSKHIVSYFASATKENRLH--EIIDGQVMNENNQREIQKAARIAVECTRLTGEER 670
Query: 177 PRQRRPGLRGNDQRSIKYRHRYSTEISDKND 207
P + R K +H++S E ++ D
Sbjct: 671 PGMKEVAAELEALRVTKTKHKWSDEYPEQED 701
>At5g04570 putative protein
Length = 555
Score = 32.3 bits (72), Expect = 0.89
Identities = 51/216 (23%), Positives = 82/216 (37%), Gaps = 33/216 (15%)
Query: 396 QELNVVGSAKPSNELNDHITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDA 455
QE V KP E + KD+V G+ P N + +S E + + + LD
Sbjct: 178 QETTNVAQKKPDLEKTMN-WKDSVCFGQ----PRNDTNWQTTPSSSYEQCATRQPHVLD- 231
Query: 456 PLIEDILAVSESCLIE--------DAVMNSTSTSQMPMEDNSMEPEFDANVDRNEPHADM 507
IED E D V N + + S+ EF + + PH
Sbjct: 232 --IEDFGMQGEGLGYSWMSISPRVDRVKNKNVPRRFFRQGGSVPREFTGQIIPSTPHELP 289
Query: 508 DVD-NGGSGMGETVMDDTVAKPNIEVNEQCQFEGK---ILTENTDAFTAS---------- 553
+ +G S + DDT E+N+ + +L + + T
Sbjct: 290 GMGLSGSSSAVQEHQDDTQHNQQDEMNKASHLQKTFLDLLNSSEECLTRQSSTKQNITDG 349
Query: 554 -MPTDDTDIDVVNPLADQSSPG--VIQANSIDKRTT 586
+P D T DVV+PL++ SS ++++NS +K T
Sbjct: 350 CLPRDRTAEDVVDPLSNNSSLQNILVESNSSNKEQT 385
>At3g01085 putative protein
Length = 536
Score = 32.3 bits (72), Expect = 0.89
Identities = 26/123 (21%), Positives = 51/123 (41%), Gaps = 11/123 (8%)
Query: 213 QEAFGSDSLDPDTEKIE---NGEACLTSLENEVTDSSATEGNKICEILDGLLRCNSEDLE 269
+E + + L P T+ E CL +E ++ +L+ LL + E
Sbjct: 242 EEFWEKNKLHPQTKMFRPQHQYEGCLRERFDEFPKTAIN-------LLENLLSIDPEK-R 293
Query: 270 GDGAMDLLQERLQIKPVVLEKVSVPDFPDNQPIDLKSLRGSLSKPRKALSNIDNLLNRRK 329
G + L+ E +P + ++P +P N+ +D K + R ++ DNL ++
Sbjct: 294 GTASSALMSEYFNTQPYACDPSTLPKYPPNKEMDAKYREELQRRRRVSIKKRDNLATKKL 353
Query: 330 NKT 332
K+
Sbjct: 354 GKS 356
>At5g64070 phosphatidylinositol 4-kinase (emb|CAB37928.1)
Length = 1121
Score = 32.0 bits (71), Expect = 1.2
Identities = 26/107 (24%), Positives = 47/107 (43%), Gaps = 13/107 (12%)
Query: 77 FNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTSVSLESLLPNLDIE 136
F + D++ D+V VD DG + P L + F + P E P + E
Sbjct: 430 FKEKSDDKKDIVKVD-----DGNESEGDESPEFSLFKRLFRIHP------EDAKPTSENE 478
Query: 137 NLKDPVEFFKAHERMENARREIQKQMGVSVEQNQDDVSTRPRQRRPG 183
N + + ++ EN R++ + SVE ++ S + +++RPG
Sbjct: 479 NSSNGL--VESSPGTENFFRKLFRDRDQSVEDSELFGSKKHKEKRPG 523
>At5g21900 DNA excision repair protein
Length = 544
Score = 32.0 bits (71), Expect = 1.2
Identities = 25/97 (25%), Positives = 46/97 (46%), Gaps = 4/97 (4%)
Query: 77 FNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRAR-FSLKPSTSVSL-ESLLPNLD 134
F +QD E + + DV + G D+ R K A+ F+L+ + + + E ++
Sbjct: 42 FQNQDVEMEERKSTEADVVKTGSDYWRDYEVKRNFKAAKHFALELHSDLKIVEKTEEEVE 101
Query: 135 IENLK--DPVEFFKAHERMENARREIQKQMGVSVEQN 169
E ++ D EF KA E ++ R+EI + ++ N
Sbjct: 102 EEVIRDEDKSEFRKAKESIKKRRQEISTTSDMEIDLN 138
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.309 0.128 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,529,752
Number of Sequences: 26719
Number of extensions: 704348
Number of successful extensions: 2169
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 60
Number of HSP's that attempted gapping in prelim test: 2133
Number of HSP's gapped (non-prelim): 89
length of query: 677
length of database: 11,318,596
effective HSP length: 106
effective length of query: 571
effective length of database: 8,486,382
effective search space: 4845724122
effective search space used: 4845724122
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0253.9


