

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0252a.5
(150 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
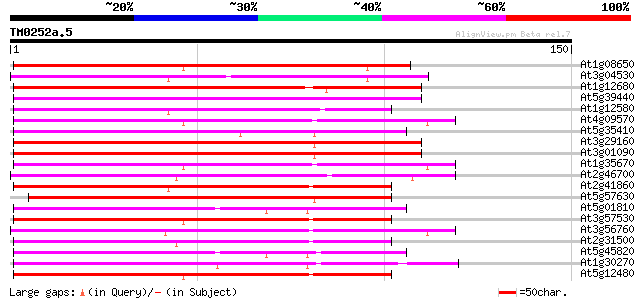
Score E
Sequences producing significant alignments: (bits) Value
At1g08650 putative calcium-dependent protein kinase (U90439) 129 8e-31
At3g04530 phosphoenolpyruvate carboxylase kinase 2 124 2e-29
At1g12680 Unknown protein (T12C24.22) 100 5e-22
At5g39440 serine/threonine-specific protein kinase -like protein 92 1e-19
At1g12580 unknown protein 90 4e-19
At4g09570 calmodulin-domain protein kinase CDPK isoform 4 (CPK4) 89 7e-19
At5g35410 serine/threonine protein kinase SOS2 (gb|AAF62923.1) 89 1e-18
At3g29160 SNF1-like protein kinase (AKin11) 88 2e-18
At3g01090 putative SNF1-related protein kinase 88 2e-18
At1g35670 calcium-dependent protein kinase 87 3e-18
At2g46700 calcium/calmodulin-dependent protein kinase CaMK4 87 3e-18
At2g41860 putative Ca2+ dependent protein kinase 86 7e-18
At5g57630 CBL-interacting protein kinase 21 (CIPK21) 85 1e-17
At5g01810 serine/threonine protein kinase ATPK10 85 1e-17
At3g57530 calcium-dependent protein kinase 85 2e-17
At3g56760 calcium-dependent protein kinase-like 85 2e-17
At2g31500 putative calcium-dependent protein kinase 84 2e-17
At5g45820 CBL-interacting protein kinase 20 (CIPK20) 84 4e-17
At1g30270 serine/threonine kinase, putative 84 4e-17
At5g12480 calcium-dependent protein kinase - like protein 83 5e-17
>At1g08650 putative calcium-dependent protein kinase (U90439)
Length = 284
Score = 129 bits (323), Expect = 8e-31
Identities = 65/126 (51%), Positives = 79/126 (62%), Gaps = 20/126 (15%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD-LKLADFGSAEWFGDG 60
G E + AS KQ+L+A++HCHR GV H D+KP+N+L D +K+ DFGS W G+G
Sbjct: 110 GTFFEPQTASFAKQILQALSHCHRYGVVHRDIKPENILVDLRNDTVKICDFGSGIWLGEG 169
Query: 61 RRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIW 101
GVVGTPYYVAPEVLMG+ GEKVD+WS GV IFEAV+
Sbjct: 170 ETTEGVVGTPYYVAPEVLMGYSYGEKVDLWSAGVVLYTMLAGTPPFYGETAEEIFEAVLR 229
Query: 102 GNLRFP 107
GNLRFP
Sbjct: 230 GNLRFP 235
>At3g04530 phosphoenolpyruvate carboxylase kinase 2
Length = 278
Score = 124 bits (310), Expect = 2e-29
Identities = 68/132 (51%), Positives = 79/132 (59%), Gaps = 21/132 (15%)
Query: 1 GGPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFG-AGGDLKLADFGSAEWFGD 59
GG + E E AS KQ+L A+ HCHR V H DVKPDNVL G +KL DFGSA W G
Sbjct: 106 GGRLSESESASYAKQILSALAHCHRCDVVHRDVKPDNVLVDLVSGGVKLCDFGSAVWLG- 164
Query: 60 GRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVI 100
G GVVGTPYYVAPEV+MG + EKVD+WS GV IFE+++
Sbjct: 165 GETAEGVVGTPYYVAPEVVMGRKYDEKVDIWSAGVVIYTMLAGEPPFNGETAEDIFESIL 224
Query: 101 WGNLRFPLESSG 112
GNLRFP + G
Sbjct: 225 RGNLRFPPKKFG 236
>At1g12680 Unknown protein (T12C24.22)
Length = 470
Score = 99.8 bits (247), Expect = 5e-22
Identities = 52/110 (47%), Positives = 70/110 (63%), Gaps = 3/110 (2%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G E A++ K L+ I +CH +GV H D+KP+N+L A G ++LADFG A G+
Sbjct: 194 GRYSEQRAANIFKDLMLVINYCHEMGVVHRDIKPENILLTAAGKIQLADFGLAMRIAKGQ 253
Query: 62 RMSGVVGTPYYVAPEVLMGWEN-GEKVDVWSCGVIFEAVIWGNLRFPLES 110
+SG+ G+P YVAPEVL EN EKVDVWS GV+ A++ G L F +S
Sbjct: 254 TLSGLAGSPAYVAPEVLS--ENYSEKVDVWSAGVLLYALLSGVLPFKGDS 301
>At5g39440 serine/threonine-specific protein kinase -like protein
Length = 494
Score = 92.0 bits (227), Expect = 1e-19
Identities = 43/109 (39%), Positives = 64/109 (58%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E E L +Q++ + +CHR + H D+KP+NVL + ++K+ DFG + DG
Sbjct: 112 GKLQEDEARHLFQQIISGVEYCHRNMIVHRDLKPENVLLDSQCNIKIVDFGLSNVMHDGH 171
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
+ G+P Y APEV+ G G VD+WSCGVI A++ G L F E+
Sbjct: 172 FLKTSCGSPNYAAPEVISGKPYGPDVDIWSCGVILYALLCGTLPFDDEN 220
>At1g12580 unknown protein
Length = 522
Score = 90.1 bits (222), Expect = 4e-19
Identities = 44/104 (42%), Positives = 62/104 (59%), Gaps = 4/104 (3%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFG---AGGDLKLADFGSAEWFG 58
G EV L K L++ + CH G+ H D+KP+N+L + +KLADFG A +
Sbjct: 138 GRYSEVRARVLFKHLMQVVKFCHDSGIVHRDLKPENILMATMSSSSPIKLADFGLATYIK 197
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G ++SG VG+P+Y+APEVL G N + DVWS GVI ++ G
Sbjct: 198 PGEKLSGTVGSPFYIAPEVLAGGYN-QAADVWSAGVILYILLSG 240
>At4g09570 calmodulin-domain protein kinase CDPK isoform 4 (CPK4)
Length = 501
Score = 89.4 bits (220), Expect = 7e-19
Identities = 53/122 (43%), Positives = 73/122 (59%), Gaps = 5/122 (4%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFG 58
G E E A L+K +L + CH LGV H D+KP+N LF + D LK DFG + ++
Sbjct: 119 GCFSEREAAKLIKTILGVVEACHSLGVMHRDLKPENFLFDSPSDDAKLKATDFGLSVFYK 178
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRFPLES-SGMFLLL 117
G+ + VVG+PYYVAPEVL G ++DVWS GVI ++ G F E+ SG+F +
Sbjct: 179 PGQYLYDVVGSPYYVAPEVLKKC-YGPEIDVWSAGVILYILLSGVPPFWAETESGIFRQI 237
Query: 118 LR 119
L+
Sbjct: 238 LQ 239
>At5g35410 serine/threonine protein kinase SOS2 (gb|AAF62923.1)
Length = 446
Score = 88.6 bits (218), Expect = 1e-18
Identities = 46/107 (42%), Positives = 64/107 (58%), Gaps = 2/107 (1%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDG- 60
G + E E +QL++A+ HCH GV H D+KP+N+L G+LK++DFG + +G
Sbjct: 104 GRLEESESRKYFQQLVDAVAHCHCKGVYHRDLKPENLLLDTNGNLKVSDFGLSALPQEGV 163
Query: 61 RRMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRF 106
+ GTP YVAPEVL G +G D+WSCGVI ++ G L F
Sbjct: 164 ELLRTTCGTPNYVAPEVLSGQGYDGSAADIWSCGVILFVILAGYLPF 210
>At3g29160 SNF1-like protein kinase (AKin11)
Length = 512
Score = 88.2 bits (217), Expect = 2e-18
Identities = 44/110 (40%), Positives = 66/110 (60%), Gaps = 1/110 (0%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E E + +Q++ + +CHR V H D+KP+N+L + ++K+ADFG + DG
Sbjct: 113 GRLQEDEARNFFQQIISGVEYCHRNMVVHRDLKPENLLLDSRCNIKIADFGLSNVMRDGH 172
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
+ G+P Y APEV+ G G +VDVWSCGVI A++ G L F E+
Sbjct: 173 FLKTSCGSPNYAAPEVISGKLYAGPEVDVWSCGVILYALLCGTLPFDDEN 222
>At3g01090 putative SNF1-related protein kinase
Length = 512
Score = 87.8 bits (216), Expect = 2e-18
Identities = 44/110 (40%), Positives = 66/110 (60%), Gaps = 1/110 (0%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E E + +Q++ + +CHR V H D+KP+N+L + ++K+ADFG + DG
Sbjct: 112 GRLQEDEARNFFQQIISGVEYCHRNMVVHRDLKPENLLLDSKCNVKIADFGLSNIMRDGH 171
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
+ G+P Y APEV+ G G +VDVWSCGVI A++ G L F E+
Sbjct: 172 FLKTSCGSPNYAAPEVISGKLYAGPEVDVWSCGVILYALLCGTLPFDDEN 221
>At1g35670 calcium-dependent protein kinase
Length = 495
Score = 87.4 bits (215), Expect = 3e-18
Identities = 52/122 (42%), Positives = 72/122 (58%), Gaps = 5/122 (4%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFG 58
G E E L+K +L + CH LGV H D+KP+N LF + D LK DFG + ++
Sbjct: 120 GHFSEREAVKLIKTILGVVEACHSLGVMHRDLKPENFLFDSPKDDAKLKATDFGLSVFYK 179
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRFPLES-SGMFLLL 117
G+ + VVG+PYYVAPEVL G ++DVWS GVI ++ G F E+ SG+F +
Sbjct: 180 PGQYLYDVVGSPYYVAPEVLKKC-YGPEIDVWSAGVILYILLSGVPPFWAETESGIFRQI 238
Query: 118 LR 119
L+
Sbjct: 239 LQ 240
>At2g46700 calcium/calmodulin-dependent protein kinase CaMK4
Length = 595
Score = 87.0 bits (214), Expect = 3e-18
Identities = 49/123 (39%), Positives = 73/123 (58%), Gaps = 5/123 (4%)
Query: 1 GGPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAG---GDLKLADFGSAEWF 57
GG PE + +++ Q+L ++ CH GV H D+KP+N LF + DLKL DFG +++
Sbjct: 240 GGKYPEDDAKAIVVQILTVVSFCHLQGVVHRDLKPENFLFTSSREDSDLKLIDFGLSDFI 299
Query: 58 GDGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRF-PLESSGMFLL 116
R++ +VG+ YYVAPEVL + E D+WS GVI ++ G+ F SG+F
Sbjct: 300 RPDERLNDIVGSAYYVAPEVLHRSYSLE-ADIWSIGVITYILLCGSRPFWARTESGIFRT 358
Query: 117 LLR 119
+LR
Sbjct: 359 VLR 361
>At2g41860 putative Ca2+ dependent protein kinase
Length = 530
Score = 85.9 bits (211), Expect = 7e-18
Identities = 43/104 (41%), Positives = 63/104 (60%), Gaps = 4/104 (3%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFG---AGGDLKLADFGSAEWFG 58
G E AS++K ++E + CH+ GV H D+KP+N LF LK DFG + +F
Sbjct: 148 GHYTERAAASVIKTIIEVVQMCHKHGVMHRDLKPENFLFANKKETASLKAIDFGLSVFFK 207
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G R + +VG+PYY+APEVL G+++D+WS GVI ++ G
Sbjct: 208 PGERFNEIVGSPYYMAPEVLRR-SYGQEIDIWSAGVILYILLCG 250
>At5g57630 CBL-interacting protein kinase 21 (CIPK21)
Length = 416
Score = 85.1 bits (209), Expect = 1e-17
Identities = 41/98 (41%), Positives = 62/98 (62%), Gaps = 1/98 (1%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSG 65
E + L +QL++A+ +CH GV H D+KP N+L + G+LK++DFG + G +S
Sbjct: 108 ESDARKLFQQLIDAVDYCHNRGVYHRDLKPQNLLLDSKGNLKVSDFGLSAVPKSGDMLST 167
Query: 66 VVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWG 102
G+P Y+APE++M +G VDVWSCGVI ++ G
Sbjct: 168 ACGSPCYIAPELIMNKGYSGAAVDVWSCGVILFELLAG 205
>At5g01810 serine/threonine protein kinase ATPK10
Length = 421
Score = 85.1 bits (209), Expect = 1e-17
Identities = 47/110 (42%), Positives = 64/110 (57%), Gaps = 6/110 (5%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E +QL+ A+ CH GV H D+KP+N+L G+LK++DFG + D R
Sbjct: 104 GKLREDVARKYFQQLVRAVDFCHSRGVCHRDLKPENLLLDEHGNLKISDFGLSA-LSDSR 162
Query: 62 RMSGVV----GTPYYVAPEVL-MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
R G++ GTP YVAPEV+ +G K DVWSCGVI ++ G L F
Sbjct: 163 RQDGLLHTTCGTPAYVAPEVISRNGYDGFKADVWSCGVILFVLLAGYLPF 212
>At3g57530 calcium-dependent protein kinase
Length = 538
Score = 84.7 bits (208), Expect = 2e-17
Identities = 44/104 (42%), Positives = 63/104 (60%), Gaps = 4/104 (3%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFG 58
G E A++ K ++E + CH+ GV H D+KP+N LFG + LK DFG + +F
Sbjct: 157 GHYTERAAAAVTKTIMEVVQVCHKHGVMHRDLKPENFLFGNKKETAPLKAIDFGLSVFFK 216
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G R + +VG+PYY+APEVL G +VD+WS GVI ++ G
Sbjct: 217 PGERFNEIVGSPYYMAPEVLKR-NYGPEVDIWSAGVILYILLCG 259
>At3g56760 calcium-dependent protein kinase-like
Length = 577
Score = 84.7 bits (208), Expect = 2e-17
Identities = 48/123 (39%), Positives = 70/123 (56%), Gaps = 5/123 (4%)
Query: 1 GGPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLF---GAGGDLKLADFGSAEWF 57
GG EV+ +M Q+L + +CH GV H D+KP+N LF LK DFG +++
Sbjct: 221 GGKYSEVDAKKVMIQILSVVAYCHLQGVVHRDLKPENFLFTTKDESSPLKAIDFGLSDYV 280
Query: 58 GDGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRFPLES-SGMFLL 116
R++ +VG+ YYVAPEVL G + D+WS GVI ++ G+ F S SG+F
Sbjct: 281 RPDERLNDIVGSAYYVAPEVLHR-TYGTEADMWSIGVIAYILLCGSRPFWARSESGIFRA 339
Query: 117 LLR 119
+L+
Sbjct: 340 VLK 342
>At2g31500 putative calcium-dependent protein kinase
Length = 582
Score = 84.3 bits (207), Expect = 2e-17
Identities = 45/104 (43%), Positives = 61/104 (58%), Gaps = 4/104 (3%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAG---GDLKLADFGSAEWFG 58
G E AS+ K +LE + CH GV H D+KP+N LF G LK DFG + +F
Sbjct: 160 GHYTERAAASVAKTILEVVKVCHEHGVIHRDLKPENFLFSNGTETAQLKAIDFGLSIFFK 219
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
+R + +VG+PYY+APEVL G ++DVWS GVI ++ G
Sbjct: 220 PAQRFNEIVGSPYYMAPEVLRR-NYGPEIDVWSAGVILYILLCG 262
>At5g45820 CBL-interacting protein kinase 20 (CIPK20)
Length = 439
Score = 83.6 bits (205), Expect = 4e-17
Identities = 46/111 (41%), Positives = 67/111 (59%), Gaps = 8/111 (7%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E +QL+ AI +CH GV H D+KP+N+L GDLK++DFG + + +
Sbjct: 104 GKLKENIARKYFQQLIGAIDYCHSRGVYHRDLKPENLLLDENGDLKISDFGLSA-LRESK 162
Query: 62 RMSGVV----GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
+ G++ GTP YVAPEV+ G++ G K DVWSCGV+ ++ G L F
Sbjct: 163 QQDGLLHTTCGTPAYVAPEVIGKKGYD-GAKADVWSCGVVLYVLLAGFLPF 212
>At1g30270 serine/threonine kinase, putative
Length = 482
Score = 83.6 bits (205), Expect = 4e-17
Identities = 49/124 (39%), Positives = 71/124 (56%), Gaps = 8/124 (6%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSA---EWFG 58
G + E E +QL+ A+ +CH GV H D+KP+N+L A G LK++DFG + +
Sbjct: 124 GRLKEDEARKYFQQLINAVDYCHSRGVYHRDLKPENLLLDANGALKVSDFGLSALPQQVR 183
Query: 59 DGRRMSGVVGTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLL 116
+ + GTP YVAPEV+ G++ G K D+WSCGVI ++ G L P E S + L
Sbjct: 184 EDGLLHTTCGTPNYVAPEVINNKGYD-GAKADLWSCGVILFVLMAGYL--PFEDSNLTSL 240
Query: 117 LLRI 120
+I
Sbjct: 241 YKKI 244
>At5g12480 calcium-dependent protein kinase - like protein
Length = 535
Score = 83.2 bits (204), Expect = 5e-17
Identities = 43/104 (41%), Positives = 63/104 (60%), Gaps = 4/104 (3%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFG 58
G E A++MK ++E + CH+ GV H D+KP+N LF + LK DFG + +F
Sbjct: 153 GHYTERAAAAVMKTIVEVVQICHKQGVMHRDLKPENFLFANKKETSALKAIDFGLSVFFK 212
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G + + +VG+PYY+APEVL G ++DVWS GVI ++ G
Sbjct: 213 PGEQFNEIVGSPYYMAPEVLRR-NYGPEIDVWSAGVILYILLCG 255
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.334 0.150 0.505
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,356,666
Number of Sequences: 26719
Number of extensions: 141576
Number of successful extensions: 2292
Number of sequences better than 10.0: 945
Number of HSP's better than 10.0 without gapping: 828
Number of HSP's successfully gapped in prelim test: 117
Number of HSP's that attempted gapping in prelim test: 552
Number of HSP's gapped (non-prelim): 1025
length of query: 150
length of database: 11,318,596
effective HSP length: 90
effective length of query: 60
effective length of database: 8,913,886
effective search space: 534833160
effective search space used: 534833160
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0252a.5


