

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0248.6
(573 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
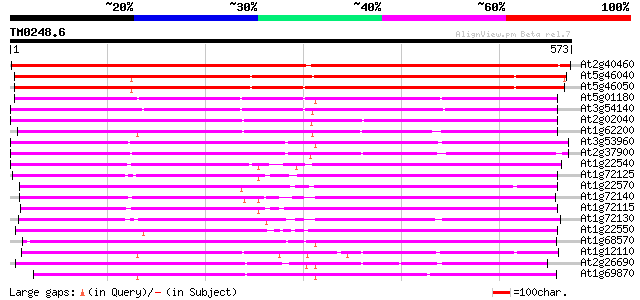
Score E
Sequences producing significant alignments: (bits) Value
At2g40460 putative PTR2 family peptide transporter 693 0.0
At5g46040 peptide transporter 486 e-137
At5g46050 peptide transporter 484 e-137
At5g01180 oligopeptide transporter - like protein 442 e-124
At3g54140 peptide transport - like protein 438 e-123
At2g02040 histidine transport protein (PTR2-B) 417 e-116
At1g62200 unknown protein (At1g62200) 407 e-113
At3g53960 transporter like protein 384 e-106
At2g37900 putative peptide/amino acid transporter 381 e-106
At1g22540 Similar to peptide transporter 374 e-103
At1g72125 oligopeptide like transporter 369 e-102
At1g22570 Similar to LeOPT1 [Lycopersicon esculentum 368 e-102
At1g72140 peptide transporter like protein (PTR2-B) 362 e-100
At1g72115 oligopeptide like transporter 362 e-100
At1g72130 peptide transporter PTR2-B -like protein 360 1e-99
At1g22550 Similar to LeOPT1 [Lycopersicon esculentum 353 1e-97
At1g68570 peptide transporter like 340 1e-93
At1g12110 nitrate/chlorate transporter CHL1 334 7e-92
At2g26690 putative nitrate transporter 331 6e-91
At1g69870 unknown protein 324 9e-89
>At2g40460 putative PTR2 family peptide transporter
Length = 583
Score = 693 bits (1788), Expect = 0.0
Identities = 324/570 (56%), Positives = 430/570 (74%), Gaps = 5/570 (0%)
Query: 3 AQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKN 62
A+ +T DGTVDL GRPV++S TG+W+AC+F+L Y+ FER A+YGI ++LVNY+T LH++
Sbjct: 4 AKVYTQDGTVDLQGRPVLASKTGRWRACSFLLGYEAFERMAFYGIASNLVNYLTKRLHED 63
Query: 63 VVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLG 122
++S+ +V+NWSG WITPI GAYIADSY+GRFWT T S LIY +G+ LL + ++K L
Sbjct: 64 TISSVRNVNNWSGAVWITPIAGAYIADSYIGRFWTFTASSLIYVLGMILLTMAVTVKSLR 123
Query: 123 PTCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFF 182
PTCENGVC +AS+ Q+ FFY+S+YTIAIG+G KPN+STFGADQFD + EK+ KVSFF
Sbjct: 124 PTCENGVCNKASSLQVTFFYISLYTIAIGAGGTKPNISTFGADQFDSYSIEEKKQKVSFF 183
Query: 183 NWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKAR 242
NWW+F++ G L +TL +VYI+E GW LGY I +G LV+ + F++GTP YRHK K
Sbjct: 184 NWWMFSSFLGALFATLGLVYIQENLGWGLGYGIPTVGLLVSLVVFYIGTPFYRHKVIKTD 243
Query: 243 SSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAITE 302
+ AK ++VP+ AF+NRKLQ P + EL+E + +Y S+G+ ++HHTP FRFLDKAAI
Sbjct: 244 NLAKDLVQVPIAAFKNRKLQCPDDHLELYELDSHYYKSNGKHQVHHTPVFRFLDKAAIKT 303
Query: 303 SNTDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHF 362
S S PCTVT+VE AK V+G+ IWL+ LIPS WA T+FVKQGTT+DR +G +F
Sbjct: 304 S----SRVPCTVTKVEVAKRVLGLIFIWLVTLIPSTLWAQVNTLFVKQGTTLDRKIGSNF 359
Query: 363 HFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVT 422
PAASL SF +M++S+P+YD F+PFMR+ TGN RG+ LLQR+G+G AIQI+ IA+
Sbjct: 360 QIPAASLGSFVTLSMLLSVPMYDQSFVPFMRKKTGNPRGITLLQRLGVGFAIQIVAIAIA 419
Query: 423 YAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEM 482
AVE++RM VI++ +I P +VVPMSIFWLLPQ + G+ + F A GLLEFFYDQSPEEM
Sbjct: 420 SAVEVKRMRVIKEFHITSPTQVVPMSIFWLLPQYSLLGIGDVFNAIGLLEFFYDQSPEEM 479
Query: 483 KVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVIAI 542
+ LGTTF+TS + GN+ NSFLVT++DK+T K SWIG+NLN+S LD+YY FL VI+I
Sbjct: 480 QSLGTTFFTSGIGLGNFLNSFLVTMIDKITSKGGGKSWIGNNLNDSRLDYYYGFLVVISI 539
Query: 543 FNFGVFLWVSNGYIYKKECTSTTEPNDGFC 572
N G+F+W ++ Y+YK + T E + G C
Sbjct: 540 VNMGLFVWAASKYVYKSD-DDTKEFSGGGC 568
>At5g46040 peptide transporter
Length = 586
Score = 486 bits (1252), Expect = e-137
Identities = 252/573 (43%), Positives = 362/573 (62%), Gaps = 13/573 (2%)
Query: 6 HTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVT 65
+T DGTVDL G V S TG+WKAC+F++VY+VFER AYYGI ++LV YMT LH+ V
Sbjct: 10 YTKDGTVDLRGNRVRRSQTGRWKACSFVVVYEVFERMAYYGISSNLVIYMTTKLHQGTVK 69
Query: 66 SITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGP-- 123
S +V+NW G SW+TPILGAY+AD++ GR+ T +S IY +G+ LL L+ SL GL P
Sbjct: 70 SSNNVTNWVGTSWLTPILGAYVADAHFGRYITFVISSAIYLLGMALLTLSVSLPGLKPPK 129
Query: 124 --TCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSF 181
T C++AS QLA F+ ++YT+AIG+G KPN+ST GADQFD+F P +K HK SF
Sbjct: 130 CSTANVENCEKASVIQLAVFFGALYTLAIGTGGTKPNISTIGADQFDEFDPKDKIHKHSF 189
Query: 182 FNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKA 241
FNWW+F+ GT +T +VY+++ GWA+GY +S +G + F LGT +YRHK
Sbjct: 190 FNWWMFSIFFGTFFATTVLVYVQDNVGWAIGYGLSTLGLAFSIFIFLLGTRLYRHKLPMG 249
Query: 242 RSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAIT 301
K RV V + R + + S+ + +E Y S +H T RFL++A++
Sbjct: 250 SPFTK-MARVIVASLRKAREPMSSDSTRFYELPPMEYASKRAFPIHSTSSLRFLNRASLK 308
Query: 302 ESNTDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPH 361
+T CT+T+VE+ K ++ M + + +PS A +T+F+KQGTT+DR L +
Sbjct: 309 TGSTHKWR-LCTITEVEETKQMLKMLPVLFVTFVPSMMLAQIMTLFIKQGTTLDRRLTNN 367
Query: 362 FHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAV 421
F P ASL F F+M++S+ IYD F+ FMR+ TGN RG+ LLQR+GIG+ + I+ + +
Sbjct: 368 FSIPPASLLGFTTFSMLVSIVIYDRVFVKFMRKLTGNPRGITLLQRMGIGMILHILIMII 427
Query: 422 TYAVEIQRMHVIRKHNIVGPKEV-VPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPE 480
E R+ V +H + V +P+SIF LLPQ V+ G+A+ F+ LEFFYDQ+PE
Sbjct: 428 ASITERYRLKVAAEHGLTHQTAVPIPLSIFTLLPQYVLMGLADAFIEIAKLEFFYDQAPE 487
Query: 481 EMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVI 540
MK LGT++ +++MA G + +S L++ V ++T+K WI +NLNES LD+YY F V+
Sbjct: 488 SMKSLGTSYTSTSMAVGYFMSSILLSSVSQITKKQ-GRGWIQNNLNESRLDNYYMFFAVL 546
Query: 541 AIFNFGVFLWVSNGYIYKKECTSTT-----EPN 568
+ NF +FL V Y Y+ + T + EPN
Sbjct: 547 NLLNFILFLVVIRFYEYRADVTQSANVEQKEPN 579
>At5g46050 peptide transporter
Length = 582
Score = 484 bits (1247), Expect = e-137
Identities = 248/566 (43%), Positives = 356/566 (62%), Gaps = 8/566 (1%)
Query: 6 HTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVT 65
+T DGTVDL G PV S+ G+WKAC+F++VY+VFER AYYGI ++L YMT LH+ V
Sbjct: 10 YTKDGTVDLQGNPVRRSIRGRWKACSFVVVYEVFERMAYYGISSNLFIYMTTKLHQGTVK 69
Query: 66 SITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGP-- 123
S +V+NW G SW+TPILGAY+ D+ LGR+ T +S IY G+ +L L+ ++ G+ P
Sbjct: 70 SSNNVTNWVGTSWLTPILGAYVGDALLGRYITFVISCAIYFSGMMVLTLSVTIPGIKPPE 129
Query: 124 --TCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSF 181
T C++AS QLA F+ ++YT+AIG+G KPN+ST GADQFD F P EK K+SF
Sbjct: 130 CSTTNVENCEKASVLQLAVFFGALYTLAIGTGGTKPNISTIGADQFDVFDPKEKTQKLSF 189
Query: 182 FNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKA 241
FNWW+F+ GTL + +VY+++ GW LGY + +G ++ F LGTP YRHK
Sbjct: 190 FNWWMFSIFFGTLFANTVLVYVQDNVGWTLGYGLPTLGLAISITIFLLGTPFYRHKLPTG 249
Query: 242 RSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAIT 301
K RV V +FR + + + HE Y G +H TP RFLD+A++
Sbjct: 250 SPFTK-MARVIVASFRKANAPMTHDITSFHELPSLEYERKGAFPIHPTPSLRFLDRASL- 307
Query: 302 ESNTDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPH 361
++ T++ CT T+VE+ K ++ M + + +PS A T+FVKQGTT+DR +
Sbjct: 308 KTGTNHKWNLCTTTEVEETKQMLRMLPVLFITFVPSMMLAQINTLFVKQGTTLDRKVTGS 367
Query: 362 FHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAV 421
F P ASL F +M+IS+ +YD F+ R+ TGN RG+ LLQR+GIG+ I+ + V
Sbjct: 368 FSIPPASLSGFVTLSMLISIVLYDRVFVKITRKFTGNPRGITLLQRMGIGLIFHILIMIV 427
Query: 422 TYAVEIQRMHVIRKHNIVGPKEV-VPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPE 480
E R+ V H ++ V +P++IF LLPQ V+ G+A++FL LEFFYDQ+PE
Sbjct: 428 ASVTERYRLKVAADHGLIHQTGVKLPLTIFALLPQFVLMGMADSFLEVAKLEFFYDQAPE 487
Query: 481 EMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVI 540
MK LGT++ T+++A GN+ +SFL++ V ++T+K WI +NLNES LD+YY F V+
Sbjct: 488 SMKSLGTSYSTTSLAIGNFMSSFLLSTVSEITKKR-GRGWILNNLNESRLDYYYLFFAVL 546
Query: 541 AIFNFGVFLWVSNGYIYKKECTSTTE 566
+ NF +FL V Y+Y+ E T + +
Sbjct: 547 NLVNFVLFLVVVKFYVYRAEVTDSVD 572
>At5g01180 oligopeptide transporter - like protein
Length = 570
Score = 442 bits (1138), Expect = e-124
Identities = 223/561 (39%), Positives = 340/561 (59%), Gaps = 11/561 (1%)
Query: 6 HTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVT 65
+T DGT+D+ +P + TG WKAC FIL + ER AYYG+ +L+NY+ ++ V+
Sbjct: 8 YTKDGTLDIHKKPANKNKTGTWKACRFILGTECCERLAYYGMSTNLINYLEKQMNMENVS 67
Query: 66 SITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTC 125
+ SVSNWSG + TP++GA+IAD+YLGR+WTI ++IY G+ LL +++S+ GL PTC
Sbjct: 68 ASKSVSNWSGTCYATPLIGAFIADAYLGRYWTIASFVVIYIAGMTLLTISASVPGLTPTC 127
Query: 126 ENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWW 185
C A+ Q A ++++Y IA+G+G +KP +S+FGADQFDD EKE K SFFNW+
Sbjct: 128 SGETC-HATAGQTAITFIALYLIALGTGGIKPCVSSFGADQFDDTDEKEKESKSSFFNWF 186
Query: 186 VFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSSA 245
F G + ++ +V+I+ GW G + + +A + FF G+ YR + S
Sbjct: 187 YFVINVGAMIASSVLVWIQMNVGWGWGLGVPTVAMAIAVVFFFAGSNFYR-LQKPGGSPL 245
Query: 246 KGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAITESNT 305
++V V + R K+++P + S L+E + G RKL HT F DKAA+ E+ +
Sbjct: 246 TRMLQVIVASCRKSKVKIPEDESLLYENQDAESSIIGSRKLEHTKILTFFDKAAV-ETES 304
Query: 306 DNSNPP-------CTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSL 358
DN CTVTQVE+ K +I + IW ++ ++ ++ T+FV QG T+D+ +
Sbjct: 305 DNKGAAKSSSWKLCTVTQVEELKALIRLLPIWATGIVFASVYSQMGTVFVLQGNTLDQHM 364
Query: 359 GPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIG 418
GP+F P+ASL F +++ P+YD +PF R++TG+ RG LQR+GIG+ I I
Sbjct: 365 GPNFKIPSASLSLFDTLSVLFWAPVYDKLIVPFARKYTGHERGFTQLQRIGIGLVISIFS 424
Query: 419 IAVTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQS 478
+ +E+ R++ ++ HN+ +E +PM+IFW +PQ + G A F G LEFFYDQ+
Sbjct: 425 MVSAGILEVARLNYVQTHNLYN-EETIPMTIFWQVPQYFLVGCAEVFTFIGQLEFFYDQA 483
Query: 479 PEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLF 538
P+ M+ L + + +A GNY ++FLVT+V KVTR WI NLN HLD+++ L
Sbjct: 484 PDAMRSLCSALSLTAIAFGNYLSTFLVTLVTKVTRSGGRPGWIAKNLNNGHLDYFFWLLA 543
Query: 539 VIAIFNFGVFLWVSNGYIYKK 559
++ NF V+LW++ Y YKK
Sbjct: 544 GLSFLNFLVYLWIAKWYTYKK 564
>At3g54140 peptide transport - like protein
Length = 570
Score = 438 bits (1127), Expect = e-123
Identities = 232/566 (40%), Positives = 337/566 (58%), Gaps = 12/566 (2%)
Query: 2 EAQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHK 61
E +T DGTVD+ P TG WKAC FIL + ER AYYG+G +LVNY+ L++
Sbjct: 3 EKDVYTQDGTVDIHKNPANKEKTGNWKACRFILGNECCERLAYYGMGTNLVNYLESRLNQ 62
Query: 62 NVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL 121
T+ +V+NWSG +ITP++GA+IAD+YLGR+WTI + IY G+ LL L++S+ GL
Sbjct: 63 GNATAANNVTNWSGTCYITPLIGAFIADAYLGRYWTIATFVFIYVSGMTLLTLSASVPGL 122
Query: 122 GP-TCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVS 180
P C C S+ Q A F++++Y IA+G+G +KP +S+FGADQFD+ EK K S
Sbjct: 123 KPGNCNADTCHPNSS-QTAVFFVALYMIALGTGGIKPCVSSFGADQFDENDENEKIKKSS 181
Query: 181 FFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRK 240
FFNW+ F+ G L + +V+I+ GW G+ + + ++A FF G+ YR R
Sbjct: 182 FFNWFYFSINVGALIAATVLVWIQMNVGWGWGFGVPTVAMVIAVCFFFFGSRFYR-LQRP 240
Query: 241 ARSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAI 300
S +V V AFR ++VP + S L E + G RKL HT +F DKAA+
Sbjct: 241 GGSPLTRIFQVIVAAFRKISVKVPEDKSLLFETADDESNIKGSRKLVHTDNLKFFDKAAV 300
Query: 301 TESNTDN-----SNP--PCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTT 353
ES +D+ NP C+VTQVE+ K +I + +W ++ + ++ T+FV QG T
Sbjct: 301 -ESQSDSIKDGEVNPWRLCSVTQVEELKSIITLLPVWATGIVFATVYSQMSTMFVLQGNT 359
Query: 354 MDRSLGPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIA 413
MD+ +G +F P+ASL F +++ P+YD F IP R+ T N RG LQR+GIG+
Sbjct: 360 MDQHMGKNFEIPSASLSLFDTVSVLFWTPVYDQFIIPLARKFTRNERGFTQLQRMGIGLV 419
Query: 414 IQIIGIAVTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEF 473
+ I + +E+ R+ ++ HN K+ + MSIFW +PQ ++ G A F G LEF
Sbjct: 420 VSIFAMITAGVLEVVRLDYVKTHNAYDQKQ-IHMSIFWQIPQYLLIGCAEVFTFIGQLEF 478
Query: 474 FYDQSPEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHY 533
FYDQ+P+ M+ L + +T+A GNY ++ LVTVV K+T+K WI DNLN HLD++
Sbjct: 479 FYDQAPDAMRSLCSALSLTTVALGNYLSTVLVTVVMKITKKNGKPGWIPDNLNRGHLDYF 538
Query: 534 YAFLFVIAIFNFGVFLWVSNGYIYKK 559
+ L ++ NF V+LW+S Y YKK
Sbjct: 539 FYLLATLSFLNFLVYLWISKRYKYKK 564
>At2g02040 histidine transport protein (PTR2-B)
Length = 585
Score = 417 bits (1071), Expect = e-116
Identities = 219/565 (38%), Positives = 330/565 (57%), Gaps = 9/565 (1%)
Query: 2 EAQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHK 61
E + + DG+VD G P + TG WKAC FIL + ER AYYGI +L+ Y+T LH+
Sbjct: 20 EVKLYAEDGSVDFNGNPPLKEKTGNWKACPFILGNECCERLAYYGIAGNLITYLTTKLHQ 79
Query: 62 NVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL 121
V++ T+V+ W G ++TP++GA +AD+Y GR+WTI IY +G+ L L++S+ L
Sbjct: 80 GNVSAATNVTTWQGTCYLTPLIGAVLADAYWGRYWTIACFSGIYFIGMSALTLSASVPAL 139
Query: 122 GPT-CENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVS 180
P C C A+ Q A F+ +Y IA+G+G +KP +S+FGADQFDD E+ K S
Sbjct: 140 KPAECIGDFCPSATPAQYAMFFGGLYLIALGTGGIKPCVSSFGADQFDDTDSRERVRKAS 199
Query: 181 FFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRK 240
FFNW+ F+ G L S+ +V+I+E GW LG+ I + +A +FF GTP+YR + +
Sbjct: 200 FFNWFYFSINIGALVSSSLLVWIQENRGWGLGFGIPTVFMGLAIASFFFGTPLYRFQ-KP 258
Query: 241 ARSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAI 300
S +V V +FR ++VP + + L+E + ++ +G RK+ HT ++LDKAA+
Sbjct: 259 GGSPITRISQVVVASFRKSSVKVPEDATLLYETQDKNSAIAGSRKIEHTDDCQYLDKAAV 318
Query: 301 TESNTD------NSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTM 354
NS CTVTQVE+ K++I MF IW +I S +A T+FV+QG M
Sbjct: 319 ISEEESKSGDYSNSWRLCTVTQVEELKILIRMFPIWASGIIFSAVYAQMSTMFVQQGRAM 378
Query: 355 DRSLGPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAI 414
+ +G F P A+L +F +++I +P+YD F +P R+ TG +G +QR+GIG+ +
Sbjct: 379 NCKIGS-FQLPPAALGTFDTASVIIWVPLYDRFIVPLARKFTGVDKGFTEIQRMGIGLFV 437
Query: 415 QIIGIAVTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFF 474
++ +A VEI R+H+ +V VP+S+ W +PQ I G A F G LEFF
Sbjct: 438 SVLCMAAAAIVEIIRLHMANDLGLVESGAPVPISVLWQIPQYFILGAAEVFYFIGQLEFF 497
Query: 475 YDQSPEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYY 534
YDQSP+ M+ L + T A GNY +S ++T+V T + WI DNLN HLD+++
Sbjct: 498 YDQSPDAMRSLCSALALLTNALGNYLSSLILTLVTYFTTRNGQEGWISDNLNSGHLDYFF 557
Query: 535 AFLFVIAIFNFGVFLWVSNGYIYKK 559
L +++ N V+ + + Y KK
Sbjct: 558 WLLAGLSLVNMAVYFFSAARYKQKK 582
>At1g62200 unknown protein (At1g62200)
Length = 590
Score = 407 bits (1045), Expect = e-113
Identities = 219/561 (39%), Positives = 329/561 (58%), Gaps = 20/561 (3%)
Query: 9 DGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSIT 68
DG++D+ G P TG WKAC FIL + ER AYYGI +L+ Y T LH++ V++ +
Sbjct: 38 DGSIDIYGNPPSKKKTGNWKACPFILGNECCERLAYYGIAKNLITYYTSELHESNVSAAS 97
Query: 69 SVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCENG 128
V W G +ITP++GA IADSY GR+WTI IY +G+ LL L++SL L P G
Sbjct: 98 DVMIWQGTCYITPLIGAVIADSYWGRYWTIASFSAIYFIGMALLTLSASLPVLKPAACAG 157
Query: 129 V----CKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNW 184
V C A+ Q A F+ +Y IA+G+G +KP +S+FGADQFDD P E+ K SFFNW
Sbjct: 158 VAAALCSPATTVQYAVFFTGLYLIALGTGGIKPCVSSFGADQFDDTDPRERVRKASFFNW 217
Query: 185 WVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSS 244
+ F+ G+ S+ +V+++E GW LG+ I + V+ +FF+GTP+YR + + S
Sbjct: 218 FYFSINIGSFISSTLLVWVQENVGWGLGFLIPTVFMGVSIASFFIGTPLYRFQ-KPGGSP 276
Query: 245 AKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAITESN 304
+V V A+R KL +P + S L+E ++ + +G RK+ HT ++FLDKAA+
Sbjct: 277 ITRVCQVLVAAYRKLKLNLPEDISFLYETREKNSMIAGSRKIQHTDGYKFLDKAAVISEY 336
Query: 305 TDN----SNP--PCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSL 358
SNP CTVTQVE+ K +I MF IW ++ S ++ T+FV+QG +M+R +
Sbjct: 337 ESKSGAFSNPWKLCTVTQVEEVKTLIRMFPIWASGIVYSVLYSQISTLFVQQGRSMNRII 396
Query: 359 GPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIG 418
F P AS F ++IS+PIYD F +PF+RR TG +G+ LQR+GIG+ + ++
Sbjct: 397 -RSFEIPPASFGVFDTLIVLISIPIYDRFLVPFVRRFTGIPKGLTDLQRMGIGLFLSVLS 455
Query: 419 IAVTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQS 478
IA VE R+ + ++ V MSIFW +PQ ++ G+A F G +EFFYD+S
Sbjct: 456 IAAAAIVETVRLQL--------AQDFVAMSIFWQIPQYILMGIAEVFFFIGRVEFFYDES 507
Query: 479 PEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLF 538
P+ M+ + + A G+Y +S ++T+V T W+ D+LN+ HLD+++ L
Sbjct: 508 PDAMRSVCSALALLNTAVGSYLSSLILTLVAYFTALGGKDGWVPDDLNKGHLDYFFWLLV 567
Query: 539 VIAIFNFGVFLWVSNGYIYKK 559
+ + N V+ + + KK
Sbjct: 568 SLGLVNIPVYALICVKHTKKK 588
>At3g53960 transporter like protein
Length = 620
Score = 384 bits (985), Expect = e-106
Identities = 203/577 (35%), Positives = 319/577 (55%), Gaps = 15/577 (2%)
Query: 2 EAQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHK 61
+ Q LD + D G + + TG W+A FI+ + ER +Y+GI +LV Y+T LH+
Sbjct: 16 DQQKWVLDSSTDSRGEIPLRAQTGAWRAALFIIGIEFSERLSYFGISTNLVVYLTTILHQ 75
Query: 62 NVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL 121
++ ++ + + WSG++ + P+LG ++AD+YLGR+ T+ L+ IY MGL LL L+ + GL
Sbjct: 76 DLKMAVKNTNYWSGVTTLMPLLGGFVADAYLGRYGTVLLATTIYLMGLILLTLSWFIPGL 135
Query: 122 GPTCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSF 181
C +C E F+++IY I+IG+G KP++ +FGADQF+D P E++ K+S+
Sbjct: 136 -KACHEDMCVEPRKAHEIAFFIAIYLISIGTGGHKPSLESFGADQFEDGHPEERKMKMSY 194
Query: 182 FNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKA 241
FNWW G L + +VYIE++ GW + I I +F F +G P YR+++ +
Sbjct: 195 FNWWNAGLCAGILTAVTVIVYIEDRIGWGVASIILTIVMATSFFIFRIGKPFYRYRA-PS 253
Query: 242 RSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAIT 301
S ++V V A R L PS+ S LHE E Y R L + +FLDKAA+
Sbjct: 254 GSPLTPMLQVFVAAIAKRNLPCPSDSSLLHELTNEEYTKG--RLLSSSKNLKFLDKAAVI 311
Query: 302 ESNTDNSNPP-------CTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTM 354
E +N+ TVT+VE+ K++I M IW L T+F+KQ M
Sbjct: 312 EDRNENTKAEKQSPWRLATVTKVEEVKLLINMIPIWFFTLAFGVCATQSSTLFIKQAIIM 371
Query: 355 DRSL-GPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIA 413
DR + G F P ASL+S +++I++ IY+ +P +RR TGN RG+ +LQR+G+G+
Sbjct: 372 DRHITGTSFIVPPASLFSLIALSIIITVTIYEKLLVPLLRRATGNERGISILQRIGVGMV 431
Query: 414 IQIIGIAVTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEF 473
+ + + +E +R+ ++H++ + + +S WL PQ ++ GVA+ F GL E+
Sbjct: 432 FSLFAMIIAALIEKKRLDYAKEHHM---NKTMTLSAIWLAPQFLVLGVADAFTLVGLQEY 488
Query: 474 FYDQSPEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHY 533
FYDQ P+ M+ LG FY S + ++ N+ L+TV D + ++ W G +LN S LD +
Sbjct: 489 FYDQVPDSMRSLGIAFYLSVLGAASFVNNLLITVSDHLAEEISGKGWFGKDLNSSRLDRF 548
Query: 534 YAFLFVIAIFNFGVFLWVSNGYIYKKECTSTTEPNDG 570
Y L + N F+ V+ Y YK S DG
Sbjct: 549 YWMLAALTAANICCFVIVAMRYTYKTVQPSLAVVADG 585
>At2g37900 putative peptide/amino acid transporter
Length = 575
Score = 381 bits (978), Expect = e-106
Identities = 209/575 (36%), Positives = 320/575 (55%), Gaps = 24/575 (4%)
Query: 2 EAQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHK 61
+ Q LD ++D GR + + TG W+A FI+ + ER +Y+G+ +LV Y+T L++
Sbjct: 17 DQQKWVLDSSLDSRGRVPLRARTGAWRAALFIIAIEFSERLSYFGLATNLVVYLTTILNQ 76
Query: 62 NVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL 121
++ +I +V+ WSG++ + P+LG +IAD+YLGR+ T+ ++ IY MGL LL ++ + GL
Sbjct: 77 DLKMAIRNVNYWSGVTTLMPLLGGFIADAYLGRYATVLVATTIYLMGLVLLTMSWFIPGL 136
Query: 122 GPTCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSF 181
P C VC E F+++IY I+IG+G KP++ +FGADQFDD E++ K+SF
Sbjct: 137 KP-CHQEVCVEPRKAHEVAFFIAIYLISIGTGGHKPSLESFGADQFDDDHVEERKMKMSF 195
Query: 182 FNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKA 241
FNWW + G L + V YIE++ GW + I + ++ I FF+G P YR+++ +
Sbjct: 196 FNWWNVSLCAGILTAVTAVAYIEDRVGWGVAGIILTVVMAISLIIFFIGKPFYRYRT-PS 254
Query: 242 RSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAIT 301
S ++V V A R L PS+PS LHE + S R L HT +FLDKAAI
Sbjct: 255 GSPLTPILQVFVAAIAKRNLPYPSDPSLLHEVSKTEFTSG--RLLCHTEHLKFLDKAAII 312
Query: 302 ESNT----DNSNP--PCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMD 355
E + +P T+T+VE+ K++I + IW L T F+KQ TMD
Sbjct: 313 EDKNPLALEKQSPWRLLTLTKVEETKLIINVIPIWFSTLAFGICATQASTFFIKQAITMD 372
Query: 356 RSLGPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQ 415
R +G F P AS+++ T++ISL +Y+ +P +R T N RG+ +LQR+G G+
Sbjct: 373 RHIG-GFTVPPASMFTLTALTLIISLTVYEKLLVPLLRSITRNQRGINILQRIGTGMIFS 431
Query: 416 IIGIAVTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFY 475
+I + + VE QR+ + PMS+ WL PQ ++ G A+ F GL E+FY
Sbjct: 432 LITMIIAALVEKQRLDRTNNNK--------PMSVIWLAPQFMVIGFADAFTLVGLQEYFY 483
Query: 476 DQSPEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYA 535
Q P+ M+ LG FY S + ++ N+ L+T VD + SW G +LN S LD +Y
Sbjct: 484 HQVPDSMRSLGIAFYLSVIGAASFLNNLLITAVDTLAENFSGKSWFGKDLNSSRLDRFYW 543
Query: 536 FLFVIAIFNFGVFLWVSNGYIYKKECTSTTEPNDG 570
FL + N VF+ V+ YK + +P+ G
Sbjct: 544 FLAGVIAANICVFVIVAKRCPYK-----SVQPSQG 573
>At1g22540 Similar to peptide transporter
Length = 557
Score = 374 bits (959), Expect = e-103
Identities = 203/573 (35%), Positives = 321/573 (55%), Gaps = 35/573 (6%)
Query: 2 EAQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHK 61
EA L TVD +P + S +G W++ FI+ +V ERFAYYGI ++L+ Y+T L +
Sbjct: 9 EAGTPLLAVTVDYRNKPAVKSSSGGWRSAGFIIGVEVAERFAYYGISSNLITYLTGPLGQ 68
Query: 62 NVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL 121
+ + +V+ WSG + + P+LGA++ADS+LGRF TI + +Y +GLG+L L++ +
Sbjct: 69 STAAAAANVNAWSGTASLLPLLGAFVADSFLGRFRTILAASALYIVGLGVLTLSAMIPS- 127
Query: 122 GPTCE--NGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKV 179
C+ N + + FQ+ F+ ++Y +A+ G KP + FGADQFD+ +P E + K
Sbjct: 128 --DCKVSNLLSSCSPRFQVITFFSALYLVALAQGGHKPCVQAFGADQFDEKEPEECKAKS 185
Query: 180 SFFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSR 239
SFFNWW F GTL + + YI++ WALG+ I I +VA + LGT YR R
Sbjct: 186 SFFNWWYFGMCFGTLTTLWVLNYIQDNLSWALGFGIPCIAMVVALVVLLLGTCTYRFSIR 245
Query: 240 KARSSAKGFIRVP---VVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPC----- 291
+ S F+R+ V A +N + V++ +L C
Sbjct: 246 REDQSP--FVRIGNVYVAAVKNWSVSALD-------------VAAAEERLGLVSCSSSQQ 290
Query: 292 FRFLDKAAITESNTDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQG 351
F FL+KA + + N C++ ++E+AK V+ + IWL L+ + +A T F KQG
Sbjct: 291 FSFLNKALVAK------NGSCSIDELEEAKSVLRLAPIWLTCLVYAVVFAQSPTFFTKQG 344
Query: 352 TTMDRSLGPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIG 411
TM+RS+ P + A+L SF ++VI +PIYD IP R T G+ +LQR+G G
Sbjct: 345 ATMERSITPGYKISPATLQSFISLSIVIFIPIYDRVLIPIARSFTHKPGGITMLQRIGTG 404
Query: 412 IAIQIIGIAVTYAVEIQRMHVIRKHNIV-GPKEVVPMSIFWLLPQNVIFGVANTFLASGL 470
I + + + V VE++R+ + +V P VPMS++WL+PQ V+FG+ + F GL
Sbjct: 405 IFLSFLAMVVAALVEMKRLKTAADYGLVDSPDATVPMSVWWLVPQYVLFGITDVFAMVGL 464
Query: 471 LEFFYDQSPEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHL 530
EFFYDQ P E++ +G Y S GN+ +SF++++++K T + SW +NLN++HL
Sbjct: 465 QEFFYDQVPNELRSVGLALYLSIFGIGNFLSSFMISIIEKATSQSGQASWFANNLNQAHL 524
Query: 531 DHYYAFLFVIAIFNFGVFLWVSNGYIYKKECTS 563
D++Y L ++ +L+V+ Y+ K+ TS
Sbjct: 525 DYFYWLLACLSFIGLASYLYVAKSYVSKRLDTS 557
>At1g72125 oligopeptide like transporter
Length = 557
Score = 369 bits (947), Expect = e-102
Identities = 194/560 (34%), Positives = 316/560 (55%), Gaps = 20/560 (3%)
Query: 4 QAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNV 63
Q + VD G S TG+W+A FI+ +V ERFAYYGIG++L++Y+T L ++
Sbjct: 10 QEEYVTDAVDHRGLAARRSNTGRWRAALFIIGVEVAERFAYYGIGSNLISYLTGPLGEST 69
Query: 64 VTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGP 123
+ +V+ WSG++ + P+LGA++AD++LGR+ TI +S LIY +GL L L++ L +
Sbjct: 70 AVAAANVNAWSGIATLLPVLGAFVADAFLGRYRTIIISSLIYVLGLAFLTLSAFL--IPN 127
Query: 124 TCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFN 183
T E V S+F F+ S+Y +AIG KP + FGADQFD+ EK + SFFN
Sbjct: 128 TTE--VTSSTSSFLNVLFFFSLYLVAIGQSGHKPCVQAFGADQFDEKDSQEKSDRSSFFN 185
Query: 184 WWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARS 243
WW + + G + L VVYI+E+F WA G+ I + +++ + F G +YR+ R+
Sbjct: 186 WWYLSLSAGICFAILVVVYIQEEFSWAFGFGIPCVFMVISLVLFVSGRRIYRYSKRRHEE 245
Query: 244 SAKGFIRVP---VVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAI 300
F R+ VA +N++L + S+L + E+E S ++ F +KA +
Sbjct: 246 EINPFTRIGRVFFVALKNQRL----SSSDLCKVELEANTSPEKQS--------FFNKALL 293
Query: 301 TESNTDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGP 360
+++ + VE A +I + +W L + +A +T F KQG TMDR++ P
Sbjct: 294 VPNDSSQGENASKSSDVEDATALIRLIPVWFTTLAYAIPYAQYMTFFTKQGVTMDRTILP 353
Query: 361 HFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIA 420
P ASL F ++V+ +PIYD F+P R T G+ L+R+G GI + I +
Sbjct: 354 GVKIPPASLQVFIGISIVLFVPIYDRVFVPIARLITKEPCGITTLKRIGTGIVLSTITMV 413
Query: 421 VTYAVEIQRMHVIRKHNIVG-PKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSP 479
+ VE +R+ ++H ++ P+ +PMSI+WL+PQ ++ G+A+ + G+ EFFY Q P
Sbjct: 414 IAALVEFKRLETAKEHGLIDQPEATLPMSIWWLIPQYLLLGLADVYTLVGMQEFFYSQVP 473
Query: 480 EEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFV 539
E++ +G Y S + G+ +S L++++D T +SW NLN +HLD++Y L +
Sbjct: 474 TELRSIGLALYLSALGVGSLLSSLLISLIDLATGGDAGNSWFNSNLNRAHLDYFYWLLAI 533
Query: 540 IAIFNFGVFLWVSNGYIYKK 559
++ F FL++S YIY++
Sbjct: 534 VSAVGFFTFLFISKSYIYRR 553
>At1g22570 Similar to LeOPT1 [Lycopersicon esculentum
Length = 565
Score = 368 bits (944), Expect = e-102
Identities = 199/556 (35%), Positives = 317/556 (56%), Gaps = 19/556 (3%)
Query: 11 TVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITSV 70
+VD G P S TG W++ FI+ +V ERFAY+GI +L+ Y+T L ++ + +V
Sbjct: 18 SVDHRGFPAGKSSTGGWRSAWFIIGVEVAERFAYFGIACNLITYLTGPLGQSTAKAAVNV 77
Query: 71 SNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCE-NGV 129
+ WSG + I PILGA++AD+YLGR+ TI ++ LIY +GLGLL L++SL +G + + N
Sbjct: 78 NTWSGTASILPILGAFVADAYLGRYRTIVVASLIYILGLGLLTLSASLIIMGLSKQRNDA 137
Query: 130 CKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWVFTT 189
+ S + F+ S+Y +AIG G KP + FGADQFD P E + SFFNWW +
Sbjct: 138 SAKPSIWVNTLFFCSLYLVAIGQGGHKPCVQAFGADQFDAEDPKEVIARGSFFNWWFLSL 197
Query: 190 ACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYR-----HKSRKARSS 244
+ G S + V Y++E WA G+ I + ++A F LG +YR H+ + ++
Sbjct: 198 SAGISISIIVVAYVQENVNWAFGFGIPCLFMVMALAIFLLGRKIYRYPKGHHEEVNSSNT 257
Query: 245 AKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAITESN 304
RV V+AF+NRKL++ + EL + +E S R+ FL KA I+
Sbjct: 258 FARIGRVFVIAFKNRKLRLEHSSLELDQGLLEDGQSEKRKDR-----LNFLAKAMISREG 312
Query: 305 TDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHFHF 364
+ PC+ V+ AK ++ + IW+ ++ + +A +T F KQG T+DR + P
Sbjct: 313 VE----PCSGRDVDDAKALVRLIPIWITYVVSTIPYAQYITFFTKQGVTVDRRILPGVEI 368
Query: 365 PAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYA 424
PAASL SF +++IS+P+Y+ F+P R+ T G+ +LQR+G G+ + + + +
Sbjct: 369 PAASLLSFVGVSILISVPLYERVFLPIARKITKKPFGITMLQRIGAGMVLSVFNMMLAAL 428
Query: 425 VEIQRMHVIRKHNIVG-PKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMK 483
VE +R+ + R+H +V P VPMSI+W +PQ ++ G+ + F G EFFYDQ P E++
Sbjct: 429 VESKRLKIAREHGLVDKPDVTVPMSIWWFVPQYLLLGMIDLFSMVGTQEFFYDQVPTELR 488
Query: 484 VLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVIAIF 543
+G + S M ++ + FL++++D T K W NLN +H+D++Y L
Sbjct: 489 SIGLSLSLSAMGLSSFLSGFLISLIDWATGK---DGWFNSNLNRAHVDYFYWLLAAFTAI 545
Query: 544 NFGVFLWVSNGYIYKK 559
F FL++S Y+Y++
Sbjct: 546 AFFAFLFISKMYVYRR 561
>At1g72140 peptide transporter like protein (PTR2-B)
Length = 555
Score = 362 bits (929), Expect = e-100
Identities = 200/557 (35%), Positives = 308/557 (54%), Gaps = 37/557 (6%)
Query: 11 TVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITSV 70
+VD G P I S +G WK+ F + +V E+FAY+GI ++L+ Y T L ++ + ++V
Sbjct: 22 SVDFRGNPSIRSSSGAWKSSGFTMCAEVAEKFAYFGIASNLITYFTEALGESTAVAASNV 81
Query: 71 SNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCENGVC 130
+ W G + P++ IADS+LGRF TI L+ Y MGLGLL ++++ L C +
Sbjct: 82 NLWLGTAAFLPLIWGSIADSFLGRFRTILLTSSFYIMGLGLLTFSATIPSL---CNDQET 138
Query: 131 KEA--SNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWVFT 188
+E+ S ++ F+ ++Y IA+G G K + FGADQFD+ P E + K S+FNW F
Sbjct: 139 RESCVSQVKVIIFFCALYLIALGEGGFKVCLRAFGADQFDEQDPNESKAKSSYFNWLYFA 198
Query: 189 TACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKS----RKARSS 244
+ G L + L Y++E WALGYAI + ++A F LG YR + R+ +
Sbjct: 199 ISIGILTTRLVTNYVQENLSWALGYAIPCLSMMLALFLFLLGIKTYRFSTGGEGRQGKKH 258
Query: 245 AKGFIRVP---VVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAIT 301
F+R+ V A RNR+ Q PS+ L E T FRFLD+A I+
Sbjct: 259 DNPFVRIGRVFVAAARNRR-QTPSDTCLLLPNES-------------TKKFRFLDRAVIS 304
Query: 302 ESNTDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPH 361
C +VE+AK V+ + IWL L+ +A T F KQG+TMDRS+
Sbjct: 305 ----------CDSYEVEEAKAVLSLIPIWLCSLVFGIVFAQSPTFFTKQGSTMDRSISST 354
Query: 362 FHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAV 421
PAA+L F +++ +PIYD F+P R T G+ LQR+ GI + II + +
Sbjct: 355 LQVPAATLQCFISLAILVFIPIYDRLFVPIARSITRKPAGITTLQRISTGIFLSIISMVI 414
Query: 422 TYAVEIQRMHVIRKHNIV-GPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPE 480
VE++R+ R H +V PK VPMS+ WL+PQ ++FGV++ F GL EFFY + P
Sbjct: 415 AALVEMKRLKTARDHGLVDSPKATVPMSVCWLIPQYILFGVSDVFTMVGLQEFFYGEVPP 474
Query: 481 EMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVI 540
+++ +G Y S + GN+ +SF+V+V+++ T + SW +NLN++HLD++Y L +
Sbjct: 475 QLRSMGLALYLSIIGIGNFLSSFMVSVIEEATSQSGQVSWFSNNLNQAHLDYFYWLLACL 534
Query: 541 AIFNFGVFLWVSNGYIY 557
+ F ++ + Y+Y
Sbjct: 535 SSLAFIFTVYFAKSYLY 551
>At1g72115 oligopeptide like transporter
Length = 561
Score = 362 bits (929), Expect = e-100
Identities = 194/552 (35%), Positives = 304/552 (54%), Gaps = 16/552 (2%)
Query: 12 VDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITSVS 71
VD G S+TG+W+A FI+ +V ERFA YGIG++L++Y+T L ++ + +V+
Sbjct: 18 VDHRGFSARRSITGRWRAAWFIIGVEVAERFANYGIGSNLISYLTGPLGQSTAVAAANVN 77
Query: 72 NWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCENGVCK 131
WSG+S I P+LGA++AD++LGR+ TI ++ IY +GL L L++ L V
Sbjct: 78 AWSGISTILPLLGAFVADAFLGRYITIIIASFIYVLGLAFLTLSAFLI----PNNTEVTS 133
Query: 132 EASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWVFTTAC 191
S+F A F+ S+Y +AIG KP + FGADQFD+ P E + SFFNWW +
Sbjct: 134 SPSSFLNALFFFSLYLVAIGQSGHKPCVQAFGADQFDEKNPQENSDRSSFFNWWYLSMCA 193
Query: 192 GTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSSAKGFIRV 251
G + L VVYI+E WALG+ I + +++ + F LG YR + F R+
Sbjct: 194 GIGLAILVVVYIQENVSWALGFGIPCVFMVISLVLFVLGRKSYRFSKTRQEEETNPFTRI 253
Query: 252 P---VVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAITESNTDNS 308
VAF+N++L N S+L + E+ R FL+KA + +++D
Sbjct: 254 GRVFFVAFKNQRL----NSSDLCKVEL----IEANRSQESPEELSFLNKALLVPNDSDEG 305
Query: 309 NPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHFHFPAAS 368
C VE A ++ + +WL L + +A +T F KQG TM+R++ P P AS
Sbjct: 306 EVACKSRDVEDATALVRLIPVWLTTLAYAIPFAQYMTFFTKQGVTMERTIFPGVEIPPAS 365
Query: 369 LWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYAVEIQ 428
L ++V+ +PIYD +P R T + G+ L+R+G G+ + + + V VE +
Sbjct: 366 LQVLISISIVLFVPIYDRVLVPIGRSITKDPCGITTLKRIGTGMVLATLTMVVAALVESK 425
Query: 429 RMHVIRKHNIVG-PKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMKVLGT 487
R+ +++ ++ PK +PMSI+WL PQ ++ G+A+ G+ EFFY Q P E++ LG
Sbjct: 426 RLETAKEYGLIDQPKTTLPMSIWWLFPQYMLLGLADVHTLVGMQEFFYSQVPTELRSLGL 485
Query: 488 TFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVIAIFNFGV 547
Y S M G+ +S L+ ++D T +SW NLN +HLD++Y L V++ F
Sbjct: 486 AIYLSAMGVGSLLSSLLIYLIDLATGGDAGNSWFNSNLNRAHLDYFYWLLAVVSAVGFFT 545
Query: 548 FLWVSNGYIYKK 559
FL++S YIY++
Sbjct: 546 FLFISKSYIYRR 557
>At1g72130 peptide transporter PTR2-B -like protein
Length = 538
Score = 360 bits (924), Expect = 1e-99
Identities = 194/558 (34%), Positives = 306/558 (54%), Gaps = 37/558 (6%)
Query: 10 GTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITS 69
G G + + +G WK+ I+V Q+ ERFAY+GI ++L+ Y+T L ++ + +
Sbjct: 12 GNTTEGFLEIRENTSGGWKSARLIIVVQMAERFAYFGIASNLIMYLTGPLGESTAAAAAN 71
Query: 70 VSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCENGV 129
V+ W+G P+LG ++ADSYLGRF TI +S +Y +GLGLL ++ + P+ ++
Sbjct: 72 VNAWTGTVAFLPLLGGFLADSYLGRFRTIIISSSLYILGLGLLSFSTMI----PSHQS-- 125
Query: 130 CKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWVFTT 189
K+++ Q F+ S+Y +AIG G P + FGADQFD E K SFFNW +F
Sbjct: 126 -KDSNQLQETIFFFSLYLVAIGQGGYNPCIKVFGADQFDGNDHKEARDKSSFFNWLMFGN 184
Query: 190 ACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKA--RSSAKG 247
L + L YI+E W+LG+ I ++ L++ F LGT YR + + ++
Sbjct: 185 CISILTTRLVSTYIQENLSWSLGFGIPSVSMLLSLFLFLLGTTSYRFSTERVGKKNPFAR 244
Query: 248 FIRVPVVAFRNRK---LQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAITESN 304
RV + A +NR+ L + + + + H S FRFLD+AAI+
Sbjct: 245 ISRVFMEALKNRRQPDLDIANANANETLLLLAHQSSKQ---------FRFLDRAAIS--- 292
Query: 305 TDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHFHF 364
C + ++E+AK V+ + IW+ ++ + A T F KQG TMDRS+ P
Sbjct: 293 -------CELAEIEEAKAVLRLIPIWITSVVYTIVHAQSPTFFTKQGATMDRSISPGLLV 345
Query: 365 PAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYA 424
PAA+L SF ++V+ +PIYD +PF R T N G+ LQR+G GI + I+ + +
Sbjct: 346 PAATLQSFINLSVVVFIPIYDRLLVPFARSFTQNSSGITTLQRIGTGIFLSILAMVLAAL 405
Query: 425 VEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMKV 484
VE +R+ R + +PMS++WL+PQ VIFGV++ F GL EFFY Q P E++
Sbjct: 406 VETKRLQAARD------ELSIPMSVWWLIPQYVIFGVSDMFTMVGLQEFFYGQVPSELRS 459
Query: 485 LGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVIAIFN 544
+G S GNY +SF+++V+DK+T + SW ++L+++HLD++Y L +
Sbjct: 460 VGMALNLSIYGAGNYLSSFMISVIDKITNQYGQRSWFDNDLDQAHLDYFYWLLACLGFIG 519
Query: 545 FGVFLWVSNGYIYKKECT 562
F +LW + Y+Y + T
Sbjct: 520 FAFYLWFAKSYVYSRSNT 537
>At1g22550 Similar to LeOPT1 [Lycopersicon esculentum
Length = 564
Score = 353 bits (906), Expect = 1e-97
Identities = 198/561 (35%), Positives = 315/561 (55%), Gaps = 22/561 (3%)
Query: 7 TLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTS 66
++ +VD G P S TG W++ +I+ +V ERFAY+GIG++L+ Y+T L ++ T+
Sbjct: 14 SVSDSVDHRGLPAGKSSTGGWRSAWYIIGVEVGERFAYFGIGSNLITYLTGPLGQSTATA 73
Query: 67 ITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCE 126
+V+ WSG + I P+LGA+IAD+YLGR+ TI ++ LIY +GLGLL L+S L +G + +
Sbjct: 74 AVNVNTWSGTASILPVLGAFIADAYLGRYRTIVVASLIYILGLGLLTLSSILILMGLSEQ 133
Query: 127 NGVCKEASN----FQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFF 182
+ AS + F+ S+Y +AIG G KP + FGADQFD P E+ + SFF
Sbjct: 134 RQHNRNASAKPFFWVNILFFCSLYLVAIGQGGHKPCVQAFGADQFDVGDPKERISRGSFF 193
Query: 183 NWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHK--SRK 240
NWW + + G S + VVY+++ WALG+ I + ++A F G YR+ R+
Sbjct: 194 NWWFLSLSAGITLSIIVVVYVQDNVNWALGFGIPCLFMVMALALFLFGRKTYRYPRGDRE 253
Query: 241 ARSSAKGFI-RVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAA 299
+++A I RV +VAF+NRKL++ H ++E V S ++ FL KA
Sbjct: 254 GKNNAFARIGRVFLVAFKNRKLKL------THSGQLE--VGSYKKCKGQ---LEFLAKAL 302
Query: 300 ITESNTDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLG 359
+ + PC+ VE A ++ + IW+ ++ + +A T F KQG T+DR +
Sbjct: 303 LP---GEGGVEPCSSRDVEDAMALVRLIPIWITSVVSTIPYAQYATFFTKQGVTVDRKIL 359
Query: 360 PHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGI 419
P F P AS + ++ IS+P Y+ F+P R T G+ +LQR+G G+ + + +
Sbjct: 360 PGFEIPPASFQALIGLSIFISVPTYERVFLPLARLITKKPSGITMLQRIGAGMVLSSLNM 419
Query: 420 AVTYAVEIQRMHVIRKHNIVG-PKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQS 478
V VE++R+ ++H +V P +PMSI+W +PQ ++ G+ + F G EFFYDQ
Sbjct: 420 VVAALVEMKRLETAKEHGLVDRPDATIPMSIWWFVPQYLLLGMIDVFSLVGTQEFFYDQV 479
Query: 479 PEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLF 538
P E++ +G S M ++ + FL+TV++ T K SW NLN +H+D++Y L
Sbjct: 480 PTELRSIGLALSLSAMGLASFLSGFLITVINWATGKNGGDSWFNTNLNRAHVDYFYWLLA 539
Query: 539 VIAIFNFGVFLWVSNGYIYKK 559
F FL +S Y+Y++
Sbjct: 540 AFTAIGFLAFLLLSRLYVYRR 560
>At1g68570 peptide transporter like
Length = 596
Score = 340 bits (873), Expect = 1e-93
Identities = 203/558 (36%), Positives = 308/558 (54%), Gaps = 18/558 (3%)
Query: 14 LGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITSVSNW 73
L GRP + G FI ++ E+ A G A++++Y+T LH + + +++N+
Sbjct: 16 LHGRP--NRPKGGLITMPFIFANEICEKLAVVGFHANMISYLTTQLHLPLTKAANTLTNF 73
Query: 74 SGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCENG--VCK 131
+G S +TP+LGA+IADS+ GRFWTIT + +IY +G+ LL +++ + L P G VC
Sbjct: 74 AGTSSLTPLLGAFIADSFAGRFWTITFASIIYQIGMTLLTISAIIPTLRPPPCKGEEVCV 133
Query: 132 EASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWVFTTAC 191
A QL+ Y+++ A+GSG ++P + FGADQFD+ P + ++FNW+ F
Sbjct: 134 VADTAQLSILYVALLLGALGSGGIRPCVVAFGADQFDESDPNQTTKTWNYFNWYYFCMGA 193
Query: 192 GTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSSAKGFIRV 251
L + +V+I++ GW LG I + ++ IAF G +YRH A S I+V
Sbjct: 194 AVLLAVTVLVWIQDNVGWGLGLGIPTVAMFLSVIAFVGGFQLYRHLV-PAGSPFTRLIQV 252
Query: 252 PVVAFRNRKLQVPSNPSELH-EFEMEHYVSSGRRKLHHTPCFRFLDKAAITESNTDNSNP 310
V AFR RKL++ S+PS L+ E++ +S G KL HT FLDKAAI + DN P
Sbjct: 253 GVAAFRKRKLRMVSDPSLLYFNDEIDAPISLG-GKLTHTKHMSFLDKAAIV-TEEDNLKP 310
Query: 311 P--------CTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHF 362
TV +VE+ K VI M I ++ +A + T ++Q TM+R L F
Sbjct: 311 GQIPNHWRLSTVHRVEELKSVIRMGPIGASGILLITAYAQQGTFSLQQAKTMNRHLTNSF 370
Query: 363 HFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVT 422
PA S+ F M+ ++ YD F+ R+ TG RG+ L R+GIG I II V
Sbjct: 371 QIPAGSMSVFTTVAMLTTIIFYDRVFVKVARKFTGLERGITFLHRMGIGFVISIIATLVA 430
Query: 423 YAVEIQRMHVIRKHNIVG-PKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEE 481
VE++R V +H ++ P +VP+S WL+PQ + GVA F++ G LEFFYDQ+PE
Sbjct: 431 GFVEVKRKSVAIEHGLLDKPHTIVPISFLWLIPQYGLHGVAEAFMSIGHLEFFYDQAPES 490
Query: 482 MKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGD-NLNESHLDHYYAFLFVI 540
M+ T + ++ GNY ++ LVT+V K + K +W+ D NLN L+++Y + V+
Sbjct: 491 MRSTATALFWMAISIGNYVSTLLVTLVHKFSAKPDGSNWLPDNNLNRGRLEYFYWLITVL 550
Query: 541 AIFNFGVFLWVSNGYIYK 558
N +LW + Y YK
Sbjct: 551 QAVNLVYYLWCAKIYTYK 568
>At1g12110 nitrate/chlorate transporter CHL1
Length = 590
Score = 334 bits (857), Expect = 7e-92
Identities = 201/565 (35%), Positives = 311/565 (54%), Gaps = 27/565 (4%)
Query: 13 DLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITSVSN 72
D GRP S TG W + IL + ER GIG +LV Y+T +H T+ +V+N
Sbjct: 17 DFQGRPADRSKTGGWASAAMILCIEAVERLTTLGIGVNLVTYLTGTMHLGNATAANTVTN 76
Query: 73 WSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCENGV--- 129
+ G S++ +LG +IAD++LGR+ TI + I A G+ +L L++ + GL P N
Sbjct: 77 FLGTSFMLCLLGGFIADTFLGRYLTIAIFAAIQATGVSILTLSTIIPGLRPPRCNPTTSS 136
Query: 130 -CKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWVFT 188
C++AS QL YL++Y A+G+G VK ++S FG+DQFD+ +P E+ FFN + F
Sbjct: 137 HCEQASGIQLTVLYLALYLTALGTGGVKASVSGFGSDQFDETEPKERSKMTYFFNRFFFC 196
Query: 189 TACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSSAKGF 248
G+L + +VY+++ G GY I A ++A F GT YR K + S
Sbjct: 197 INVGSLLAVTVLVYVQDDVGRKWGYGICAFAIVLALSVFLAGTNRYRFK-KLIGSPMTQV 255
Query: 249 IRVPVVAFRNRKLQVPSNPSELHEFE---MEHYVSSGRRKLHHTPCFRFLDKAAITE--- 302
V V A+RNRKL++P++PS L++ + G++KL HT FR LDKAAI +
Sbjct: 256 AAVIVAAWRNRKLELPADPSYLYDVDDIIAAEGSMKGKQKLPHTEQFRSLDKAAIRDQEA 315
Query: 303 ---SNTDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVE---VTIFVKQGTTMDR 356
SN N T+T VE+ K ++ M IW ++ FW V T+ V Q T+DR
Sbjct: 316 GVTSNVFNKWTLSTLTDVEEVKQIVRMLPIWATCIL---FWTVHAQLTTLSVAQSETLDR 372
Query: 357 SLGPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQI 416
S+G F P AS+ F V ++++ +YD I ++ G++ LQR+G+G+
Sbjct: 373 SIG-SFEIPPASMAVFYVGGLLLTTAVYDRVAIRLCKKLFNYPHGLRPLQRIGLGLFFGS 431
Query: 417 IGIAVTYAVEIQRMHVIRKHNIVGPK-EVVPMSIFWLLPQNVIFGVANTFLASGLLEFFY 475
+ +AV VE++R+ H GP + +P+ + L+PQ +I G+ + +G L+FF
Sbjct: 432 MAMAVAALVELKRLRTAHAH---GPTVKTLPLGFYLLIPQYLIVGIGEALIYTGQLDFFL 488
Query: 476 DQSPEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYA 535
+ P+ MK + T ST+A G +F+S LVT+V+K T K H WI D+LN+ L ++Y
Sbjct: 489 RECPKGMKGMSTGLLLSTLALGFFFSSVLVTIVEKFTGKA--HPWIADDLNKGRLYNFYW 546
Query: 536 FLFVIAIFNFGVFLWVSNGYIYKKE 560
+ V+ NF +FL S Y+YK++
Sbjct: 547 LVAVLVALNFLIFLVFSKWYVYKEK 571
>At2g26690 putative nitrate transporter
Length = 586
Score = 331 bits (849), Expect = 6e-91
Identities = 195/555 (35%), Positives = 306/555 (55%), Gaps = 25/555 (4%)
Query: 7 TLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTS 66
T+ VD GRP S TG W IL +V ER + GI +LV Y+ +H TS
Sbjct: 17 TVADAVDYKGRPADKSKTGGWITAALILGIEVVERLSTMGIAVNLVTYLMETMHLPSSTS 76
Query: 67 ITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGP-TC 125
V+++ G S++ +LG ++ADS+LGRF TI + I A+G G L + + L L P TC
Sbjct: 77 ANIVTDFMGTSFLLCLLGGFLADSFLGRFKTIGIFSTIQALGTGALAVATKLPELRPPTC 136
Query: 126 ENG-VCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNW 184
+G C A+ FQ+ Y+S+Y IA+G+G +K ++S FG+DQFDD P EK H FFN
Sbjct: 137 HHGEACIPATAFQMTILYVSLYLIALGTGGLKSSISGFGSDQFDDKDPKEKAHMAFFFNR 196
Query: 185 WVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSS 244
+ F + GTL + +VY++++ G + Y I + +A + F GT YR+K + S
Sbjct: 197 FFFFISMGTLLAVTVLVYMQDEVGRSWAYGICTVSMAIAIVIFLCGTKRYRYKKSQG-SP 255
Query: 245 AKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAIT--- 301
+V AFR RK+++P + L+E E ++ HT F LDKAAI
Sbjct: 256 VVQIFQVIAAAFRKRKMELPQSIVYLYEDNPEGI------RIEHTDQFHLLDKAAIVAEG 309
Query: 302 --ESNTDNSNPP-----CTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTM 354
E D P +VT+VE+ K+++ + IW +I +A +T V+Q +TM
Sbjct: 310 DFEQTLDGVAIPNPWKLSSVTKVEEVKMMVRLLPIWATTIIFWTTYAQMITFSVEQASTM 369
Query: 355 DRSLGPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAI 414
R++G F PA SL F V ++I+L +YD +PF ++ G G LQR+ IG+ +
Sbjct: 370 RRNIG-SFKIPAGSLTVFFVAAILITLAVYDRAIMPFWKKWKGK-PGFSSLQRIAIGLVL 427
Query: 415 QIIGIAVTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFF 474
G+A VE +R+ V + + ++ +P+S+F L+PQ + G F+ +G L+FF
Sbjct: 428 STAGMAAAALVEQKRLSVAKSSS----QKTLPISVFLLVPQFFLVGAGEAFIYTGQLDFF 483
Query: 475 YDQSPEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYY 534
QSP+ MK + T + +T++ G + +SFLV++V +VT D W+ DN+N LD++Y
Sbjct: 484 ITQSPKGMKTMSTGLFLTTLSLGFFVSSFLVSIVKRVTSTSTDVGWLADNINHGRLDYFY 543
Query: 535 AFLFVIAIFNFGVFL 549
L +++ NF V++
Sbjct: 544 WLLVILSGINFVVYI 558
>At1g69870 unknown protein
Length = 620
Score = 324 bits (830), Expect = 9e-89
Identities = 191/545 (35%), Positives = 282/545 (51%), Gaps = 14/545 (2%)
Query: 25 GKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITSVSNWSGLSWITPILG 84
G W+A +FIL + ER G+ A+ + Y+T H V + ++ WSG + +TP++G
Sbjct: 53 GGWRAVSFILGNETLERLGSIGLLANFMVYLTKVFHLEQVDAANVINIWSGFTNLTPLVG 112
Query: 85 AYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCENGV----CKEASNFQLAF 140
AYI+D+Y+GRF TI + +GL + LT+S L P N C + Q+
Sbjct: 113 AYISDTYVGRFKTIAFASFATLLGLITITLTASFPQLHPASCNSQDPLSCGGPNKLQIGV 172
Query: 141 FYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWVFTTACGTLASTLFV 200
L + +++GSG ++P FG DQFD + SFFNW+ T + + V
Sbjct: 173 LLLGLCFLSVGSGGIRPCSIPFGVDQFDQRTEEGVKGVASFFNWYYMTFTVVLIITQTVV 232
Query: 201 VYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSSAKGFIRVPVVAFRNRK 260
VYI+++ W +G++I +A + FF G Y + + S G +V V A + RK
Sbjct: 233 VYIQDQVSWIIGFSIPTGLMALAVVMFFAGMKRYVYVKPEG-SIFSGIAQVIVAARKKRK 291
Query: 261 LQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAIT-ESNTDNSNPP------CT 313
L++P+ + SS KLH + FR LDKAA+ E + PP C+
Sbjct: 292 LKLPAEDDGTVTYYDPAIKSSVLSKLHRSNQFRCLDKAAVVIEGDLTPEGPPADKWRLCS 351
Query: 314 VTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHFHFPAASLWSFA 373
V +VE+ K +I + IW +I + T V Q MDR+LGP F PA SL +
Sbjct: 352 VQEVEEVKCLIRIVPIWSAGIISLAAMTTQGTFTVSQALKMDRNLGPKFEIPAGSLSVIS 411
Query: 374 VFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYAVEIQRMHVI 433
+ T+ I LP YD F+PFMRR TG+ G+ LLQR+G GI I + V VE RM I
Sbjct: 412 LLTIGIFLPFYDRVFVPFMRRITGHKSGITLLQRIGTGIVFAIFSMIVAGIVE--RMRRI 469
Query: 434 RKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMKVLGTTFYTST 493
R N P + PMS+FWL PQ ++ G+ F G +EFF Q PE M+ + + ++ +
Sbjct: 470 RSINAGDPTGMTPMSVFWLSPQLILMGLCEAFNIIGQIEFFNSQFPEHMRSIANSLFSLS 529
Query: 494 MAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVIAIFNFGVFLWVSN 553
AG +Y +SFLVTVV K + W+ NLN LD++Y + V+ + N F + +
Sbjct: 530 FAGSSYLSSFLVTVVHKFSGGHDRPDWLNKNLNAGKLDYFYYLIAVLGVVNLVYFWYCAR 589
Query: 554 GYIYK 558
GY YK
Sbjct: 590 GYRYK 594
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.140 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,089,467
Number of Sequences: 26719
Number of extensions: 555425
Number of successful extensions: 1725
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 54
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 1402
Number of HSP's gapped (non-prelim): 62
length of query: 573
length of database: 11,318,596
effective HSP length: 105
effective length of query: 468
effective length of database: 8,513,101
effective search space: 3984131268
effective search space used: 3984131268
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0248.6


