

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0247.8
(170 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
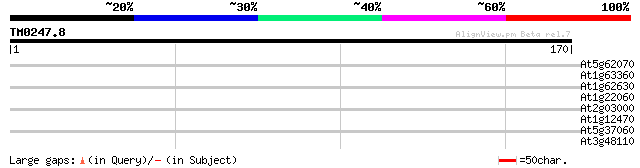
Score E
Sequences producing significant alignments: (bits) Value
At5g62070 unknown protein 30 0.82
At1g63360 disease resistance protein, putative 28 1.8
At1g62630 putative RPS-2 disease resistance protein (AA002294 28 3.1
At1g22060 hypothetical protein 28 3.1
At2g03000 hypothetical protein 27 4.1
At1g12470 hypothetical protein 27 5.3
At5g37060 putative transporter protein 26 9.1
At3g48110 glycine--tRNA ligase precursor, chloroplast (edd1) 26 9.1
>At5g62070 unknown protein
Length = 403
Score = 29.6 bits (65), Expect = 0.82
Identities = 30/107 (28%), Positives = 50/107 (46%), Gaps = 16/107 (14%)
Query: 27 LMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFHA 86
L +LKA K+++++R + + R Q M LR + TL + + + A SH+S FH+
Sbjct: 136 LRALKALVKLQALVRGHIV---RKQTADM-LRRMQTLVRLQSQARARASRSSHSSASFHS 191
Query: 87 FHKV---SRLSGVSSIH-------KVSRLSE--ASRLIPWMANKLLN 121
+ S S S+H +VS L S+ + W A + N
Sbjct: 192 STALLFPSSSSSPRSLHTRCVSNAEVSSLDHRGGSKRLDWQAEESEN 238
>At1g63360 disease resistance protein, putative
Length = 884
Score = 28.5 bits (62), Expect = 1.8
Identities = 16/70 (22%), Positives = 33/70 (46%), Gaps = 12/70 (17%)
Query: 79 HASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWMANKLLNERRSTNILRTYQTPQHL 138
H ++E+ K+ ++G+SS+H + L +PW N ++ +T +HL
Sbjct: 619 HLNLEYT--RKLESITGISSLHNLKVLKLFRSRLPW----------DLNTVKELETLEHL 666
Query: 139 DPTTLLVFPK 148
+ T + P+
Sbjct: 667 EILTTTIDPR 676
>At1g62630 putative RPS-2 disease resistance protein (AA002294
Length = 893
Score = 27.7 bits (60), Expect = 3.1
Identities = 14/60 (23%), Positives = 27/60 (44%), Gaps = 10/60 (16%)
Query: 89 KVSRLSGVSSIHKVSRLSEASRLIPWMANKLLNERRSTNILRTYQTPQHLDPTTLLVFPK 148
K+ + G+SS+H + L +PW N ++ +T +HL+ T + P+
Sbjct: 629 KLESIDGISSLHNLKVLKLYGSRLPW----------DLNTVKELETLEHLEILTTTIDPR 678
>At1g22060 hypothetical protein
Length = 1999
Score = 27.7 bits (60), Expect = 3.1
Identities = 25/99 (25%), Positives = 43/99 (43%), Gaps = 2/99 (2%)
Query: 9 SQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLTKSHA 68
S+ ++ + ++LA L SLK+ K++ LR N + +RV+ LT L + L ++
Sbjct: 1251 SRNVGAQHMNANIKLLADLDSLKSELKIERNLRNN--LDRRVEELTSELDEKHLLLENFD 1308
Query: 69 SVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSE 107
K E + RL V + H+ S E
Sbjct: 1309 LQKSQVELLEKMVAELESEKSFQRLEYVRNAHRESSFIE 1347
>At2g03000 hypothetical protein
Length = 535
Score = 27.3 bits (59), Expect = 4.1
Identities = 19/62 (30%), Positives = 29/62 (46%), Gaps = 3/62 (4%)
Query: 8 ISQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILR---QNTIIIKRVQPLTMTLRVINTLT 64
IS T P+S SPRVL MS + + S R Q ++ + P + + R + T T
Sbjct: 86 ISTTNVPSSTSNSPRVLQTSMSRRGTSPMSSSTRRSVQASMSARETVPSSTSTRSMQTST 145
Query: 65 KS 66
+
Sbjct: 146 ST 147
>At1g12470 hypothetical protein
Length = 950
Score = 26.9 bits (58), Expect = 5.3
Identities = 15/38 (39%), Positives = 17/38 (44%)
Query: 114 WMANKLLNERRSTNILRTYQTPQHLDPTTLLVFPKLWF 151
WMANK LN RR + Y + H T V L F
Sbjct: 515 WMANKNLNPRRLITAMMRYSSGPHAKNETHEVIKYLEF 552
>At5g37060 putative transporter protein
Length = 859
Score = 26.2 bits (56), Expect = 9.1
Identities = 11/60 (18%), Positives = 28/60 (46%)
Query: 28 MSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFHAF 87
+S ++P + ++LR ++ V M++ ++ + + V F A+T + + F
Sbjct: 202 LSFTSFPVIYTVLRDMNLLNSEVGKFAMSVALLGDMAGVYVIVIFEAMTHADVGGAYSVF 261
>At3g48110 glycine--tRNA ligase precursor, chloroplast (edd1)
Length = 1067
Score = 26.2 bits (56), Expect = 9.1
Identities = 16/46 (34%), Positives = 24/46 (51%)
Query: 53 LTMTLRVINTLTKSHASVKFHALTRSHASVEFHAFHKVSRLSGVSS 98
L +L +I + + HAS +F L RS + F + + R S VSS
Sbjct: 4 LHFSLPLIVSFLRPHASPRFFLLPRSLSQSPFLSRRRFHRTSAVSS 49
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,165,106
Number of Sequences: 26719
Number of extensions: 103868
Number of successful extensions: 249
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 246
Number of HSP's gapped (non-prelim): 8
length of query: 170
length of database: 11,318,596
effective HSP length: 92
effective length of query: 78
effective length of database: 8,860,448
effective search space: 691114944
effective search space used: 691114944
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0247.8


