

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0239.11
(307 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
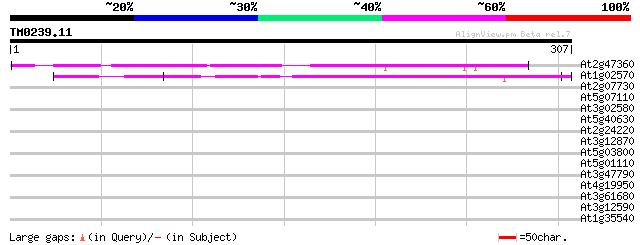
Score E
Sequences producing significant alignments: (bits) Value
At2g47360 unknown protein 213 1e-55
At1g02570 unknown protein 154 8e-38
At2g07730 putative non-LTR retroelement reverse transcriptase 34 0.087
At5g07110 unknown protein 30 1.6
At3g02580 sterol-C5-desaturase 30 1.6
At5g40630 unknown protein 30 2.1
At2g24220 hypothetical protein 30 2.1
At3g12870 hypothetical protein 28 4.8
At5g03800 putative protein 28 6.2
At5g01110 unknown protein (At5g01110) 28 6.2
At3g47790 ABC-type transport protein-like protein 28 6.2
At4g19950 putative protein 28 8.2
At3g61680 unknown protein 28 8.2
At3g12590 hypothetical protein 28 8.2
At1g35540 auxin response factor, putative 28 8.2
>At2g47360 unknown protein
Length = 303
Score = 213 bits (542), Expect = 1e-55
Identities = 124/298 (41%), Positives = 175/298 (58%), Gaps = 45/298 (15%)
Query: 2 GRWQQKAKEFFSNHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFL 61
G+W+++A++ F L+T TLLSLLLPLSF LL+ LS+A +F L
Sbjct: 6 GKWRKEAEQLVVK---------PFRLVTTTLLSLLLPLSFLLLSRLSSA-----SFLFSL 51
Query: 62 HSPQQQQQPFSFLFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS 121
Q Q + F+F+L L+ NP I+Y +VS +S+ TL+ GL KIT D S P
Sbjct: 52 TKSQPQTESSFFVFSLFLRANPAIVYAVVSSISVYTLVLGLTTKITA-TDPKHSIAFYPH 110
Query: 122 LYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGL 181
+ IAW+ L Q VG+G+E +I+ G+ +ER+ LS++ F GL
Sbjct: 111 VSIAWLTLFLVQISVGIGLETTISNGLI---------------IGSERNFLSRLVFFFGL 155
Query: 182 HETTQVWYRVVVRPVVDDTVFGG----AKKEKWIERVAVAASLGVLWWWRLREEVESLVV 237
HE +WYRV+VRPVVD+T+ GG ++E +ERVA+A S G LWWW+LR+EVE+LV
Sbjct: 156 HEVMLLWYRVIVRPVVDNTLLGGEDGQRREETVVERVALAVSCGTLWWWKLRDEVEALVG 215
Query: 238 MAEAKKSEQF-----GELELK------DFIGWWLYYVTVTIGMVRIVKGLMWMFMVAL 284
+AEAK++ G + + DF+ WWLYY+ VTIGMVRI+KG +W M+ L
Sbjct: 216 VAEAKRALLLLLPIDGNVNVSFDVGTVDFVNWWLYYMVVTIGMVRIIKGSLWFGMILL 273
>At1g02570 unknown protein
Length = 431
Score = 154 bits (388), Expect = 8e-38
Identities = 102/281 (36%), Positives = 150/281 (53%), Gaps = 51/281 (18%)
Query: 25 FHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPC 84
F +++I+LLSLL+PLSF L+ LS S S +F+L Q +
Sbjct: 20 FQMISISLLSLLVPLSFLFLSRLSV-------------SSSSAPVTVSGVFSLLHQADVG 66
Query: 85 ILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLYIAWILLCAFQFCVGLGIEASI 144
ILY ++S++ ++TLI+ L GK VL LYI WI+L Q CV GIE ++
Sbjct: 67 ILYTILSLIIVSTLIHILSGK-------PECSVLHSHLYICWIVLFIVQACVAFGIEGTM 119
Query: 145 AAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLHETTQVWYRVVVRPVVDDTVFGG 204
+ I S + SF +A ER VVV+PV+DDTVFG
Sbjct: 120 STTI------SIDTDKSFSLAAQER---------------------VVVKPVIDDTVFGV 152
Query: 205 -AKKEKWIERVAVAASLGVLWWWRLREEVESLVVMAEAKKSEQFGELELKDFIGWWLYYV 263
++E+W ER VA + G++WWWRLR+EVESLVV+AE K++ Q LE DF+ WW+YY+
Sbjct: 153 YVEEERWSERAVVAVTFGLMWWWRLRDEVESLVVVAEVKRNLQI-RLEGLDFVNWWMYYI 211
Query: 264 TVTIGM--VRIVKGLMWMFMVALCRRRVTEVPVSPVVESSL 302
V IG+ V ++ ++++ +V ++ P V+ S L
Sbjct: 212 CVGIGLADVGVLYTILFLIIVFTLIHSLSGKPECSVLHSHL 252
Score = 145 bits (366), Expect = 3e-35
Identities = 83/224 (37%), Positives = 129/224 (57%), Gaps = 15/224 (6%)
Query: 85 ILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLYIAWILLCAFQFCVGLGIEASI 144
+LY ++ ++ + TLI+ L GK VL LYI WI+L Q C GI+ ++
Sbjct: 222 VLYTILFLIIVFTLIHSLSGK-------PECSVLHSHLYICWIVLFIAQACA-FGIKRTM 273
Query: 145 AAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLHETTQVWYRVVVRPVVDDTVFGG 204
+ + S + + + ER +L +V F LGLHE +W+RVVV+PVVD+T++G
Sbjct: 274 STTM------SINPDKNLFLATHERWMLVRVLFFLGLHEVMLMWFRVVVKPVVDNTIYGV 327
Query: 205 AKKEKWIERVAVAASLGVLWWWRLREEVESLVVMAEAKKSEQFGELELKDFIGWWLYYVT 264
+E+W ER VA + G++WWWRLR+EVESLVV+ A + LE +F+ W +YY+
Sbjct: 328 YVEERWSERAVVAVTFGIMWWWRLRDEVESLVVVVTADRLNLPIRLEGLNFVNWCMYYIC 387
Query: 265 VTIGMVRIVKGLM-WMFMVALCRRRVTEVPVSPVVESSLNDDKV 307
V IG+++I KG + ++ + L +R + S V + NDD V
Sbjct: 388 VGIGLMKIFKGFLDFVNTLTLSIKRSRKGCESCVFDDMCNDDHV 431
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 34.3 bits (77), Expect = 0.087
Identities = 23/67 (34%), Positives = 37/67 (54%), Gaps = 6/67 (8%)
Query: 218 ASLGVLWWWRLREEVESLVVMAEAKKSEQFGELE----LKDFIGWWLYYVTVTIGMVRIV 273
A+L L WWRL + E+L +KK + GE++ LK W + +VT+G+ +V
Sbjct: 508 ATLDSLDWWRLLHDKETLWARVLSKK-YKVGEMQNSNWLKPKSSWSSTWRSVTVGLREVV 566
Query: 274 -KGLMWM 279
KG+ W+
Sbjct: 567 AKGVRWV 573
>At5g07110 unknown protein
Length = 216
Score = 30.0 bits (66), Expect = 1.6
Identities = 44/163 (26%), Positives = 80/163 (48%), Gaps = 46/163 (28%)
Query: 30 ITLLSLLL-------PLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQP-------FSFLF 75
ITL+++LL P + FLLASL+A+ LFL+ + QP FS L
Sbjct: 81 ITLVAILLAASLLTHPFALFLLASLAAS-------WLFLYFFRPADQPLVIGGRTFSDLE 133
Query: 76 TLA-LQINPCILYVLVSIVSI--ATLINGLMGKITLLNDSSTSPVLQPSLYIAWILLCAF 132
TL L ++ ++ + S+ S+ +TL G+MG ++ + +P L+
Sbjct: 134 TLGILCLSTVVVMFMTSVGSLLMSTLAVGIMG--VAIHGAFRAP---EDLF--------- 179
Query: 133 QFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKV 175
L + +I +G+F F N+++S++++ +A S +S+V
Sbjct: 180 -----LEEQEAIGSGLFAFFNNNASNAAA---AAIATSAMSRV 214
>At3g02580 sterol-C5-desaturase
Length = 281
Score = 30.0 bits (66), Expect = 1.6
Identities = 19/68 (27%), Positives = 32/68 (46%), Gaps = 7/68 (10%)
Query: 213 RVAVAASLGVLWWWRLREEV-ESLVVMAEAKKSEQFGELELKDFIGWWLYYVTVTIGMVR 271
R+ + ++ + W+ L V ES++ K GE GW LY+V + I +V
Sbjct: 85 RLQMFVAMKAMPWYTLLPTVSESMIERGWTKCFASIGEF------GWILYFVYIAIYLVF 138
Query: 272 IVKGLMWM 279
+ G+ WM
Sbjct: 139 VEFGIYWM 146
>At5g40630 unknown protein
Length = 165
Score = 29.6 bits (65), Expect = 2.1
Identities = 18/42 (42%), Positives = 24/42 (56%), Gaps = 1/42 (2%)
Query: 129 LCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERS 170
L A +FC+G G I G + +SSSSSSSSF ++ S
Sbjct: 4 LMAKRFCIGFGC-GRIGTGNNKRSSSSSSSSSSFSSLSSSSS 44
>At2g24220 hypothetical protein
Length = 315
Score = 29.6 bits (65), Expect = 2.1
Identities = 32/112 (28%), Positives = 46/112 (40%), Gaps = 12/112 (10%)
Query: 21 PKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQ-------PFSF 73
P F + T L+ L LS+ +L LSAA + + + + + P FS
Sbjct: 43 PTYIFQKIKPTPLNTKLVLSYVVLGFLSAADNLMYAYA-YAYLPASTSSLLASSSLAFSA 101
Query: 74 LFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLYIA 125
LF + NP V+ SIV +I G M I L + S + S Y A
Sbjct: 102 LFGYLIVKNPLNASVINSIV----VITGAMAIIALDSSSDRYSYISNSQYFA 149
>At3g12870 hypothetical protein
Length = 206
Score = 28.5 bits (62), Expect = 4.8
Identities = 17/43 (39%), Positives = 26/43 (59%), Gaps = 2/43 (4%)
Query: 205 AKKEKWIERV-AVAASLGVLWWWRLREEVES-LVVMAEAKKSE 245
A K WI + A+ +SLG++W R + EVES L + E +K +
Sbjct: 94 ACKRSWIPSLCALLSSLGIIWAVRYKSEVESHLEKLLEREKED 136
>At5g03800 putative protein
Length = 1388
Score = 28.1 bits (61), Expect = 6.2
Identities = 18/38 (47%), Positives = 23/38 (60%), Gaps = 2/38 (5%)
Query: 146 AGIFEFDNSSSSSSSSFGESAAE--RSLLSKVFFLLGL 181
A I E +SSSSSSSSF + E S++ F+LL L
Sbjct: 52 ATIHECSSSSSSSSSSFDKEETEDIESVIDGFFYLLRL 89
>At5g01110 unknown protein (At5g01110)
Length = 729
Score = 28.1 bits (61), Expect = 6.2
Identities = 15/32 (46%), Positives = 21/32 (64%)
Query: 148 IFEFDNSSSSSSSSFGESAAERSLLSKVFFLL 179
+FE +SSSSSSSS S ++ L+ K+ F L
Sbjct: 31 VFEPSSSSSSSSSSASFSVSDSFLVEKICFSL 62
>At3g47790 ABC-type transport protein-like protein
Length = 727
Score = 28.1 bits (61), Expect = 6.2
Identities = 46/184 (25%), Positives = 65/184 (35%), Gaps = 11/184 (5%)
Query: 5 QQKAKEFFSNHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHY-LQTFTLFLHS 63
QQ+ + HG P L+S+L L F + SL + T F
Sbjct: 212 QQRLRIMMKMHGLGDVPYWIVSYTYFLLISILYMLCFAIFGSLIVLDAFTFSTGLNFFRL 271
Query: 64 PQQQQQPFSFLFTLALQINPCIL-YVLVSIVSIATLI-------NGLMGKITLLNDSSTS 115
Q F + LQI+ L + S V IAT+I GL+G I L
Sbjct: 272 NDYSIQLVFFFICINLQISVAFLASAMFSDVKIATVIAYIYVFGTGLLG-IFLFQFFLED 330
Query: 116 PVLQPSLYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKV 175
P+ IA L F GL E S +A ++ FG E + + +
Sbjct: 331 PLFPRGWIIAMELYPGFSLYRGL-YELSQSAFAGDYRGIDGMKWRDFGNGMKEVTCIMLI 389
Query: 176 FFLL 179
+LL
Sbjct: 390 EWLL 393
>At4g19950 putative protein
Length = 306
Score = 27.7 bits (60), Expect = 8.2
Identities = 25/98 (25%), Positives = 40/98 (40%), Gaps = 4/98 (4%)
Query: 85 ILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLYIAWILLCAFQFCVGLGIEA-- 142
I L+ +S A L + L + L + Q L W +L FQFC + + A
Sbjct: 34 ITLTLIFPLSFAILAHSLFTQPILAQIDTYPQADQSQLQHEWTVLLVFQFCYIIFLFAFS 93
Query: 143 --SIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFL 178
S AA +F + + SF + + L+ K F+
Sbjct: 94 LLSTAAVVFTVASLYTGKPVSFSSTMSAIPLVLKRLFI 131
>At3g61680 unknown protein
Length = 658
Score = 27.7 bits (60), Expect = 8.2
Identities = 25/96 (26%), Positives = 43/96 (44%), Gaps = 18/96 (18%)
Query: 139 GIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLHETTQVWYRVVVRPVVD 198
G++ S + G+F F SSS S ++LL + E+ ++ + P +D
Sbjct: 69 GMKTSRSVGVFSFQISSSIIPSPI------KTLLFETDTSQDEQESDEI--EIETEPNLD 120
Query: 199 DTVFGGAKKEKWIERVAVAASLGVLWWWRLREEVES 234
GAKK W+ER+ L + W+ ++ ES
Sbjct: 121 -----GAKKANWVERL-----LEIRRQWKREQKTES 146
>At3g12590 hypothetical protein
Length = 1211
Score = 27.7 bits (60), Expect = 8.2
Identities = 14/37 (37%), Positives = 24/37 (64%), Gaps = 1/37 (2%)
Query: 37 LPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSF 73
LP+S F A+L + HY+++ + LH P++ QP +F
Sbjct: 162 LPVSSFASAALVKSLHYVRSL-VALHIPRRSFQPAAF 197
>At1g35540 auxin response factor, putative
Length = 661
Score = 27.7 bits (60), Expect = 8.2
Identities = 19/54 (35%), Positives = 26/54 (47%)
Query: 23 ATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFT 76
++ L + +LLSL L LS L SLS + F+LF SP S L +
Sbjct: 58 SSLSLYSFSLLSLSLSLSLSLSLSLSLSLSPFSLFSLFSLSPLSLSLSLSLLLS 111
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,496,842
Number of Sequences: 26719
Number of extensions: 248856
Number of successful extensions: 1165
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 1145
Number of HSP's gapped (non-prelim): 18
length of query: 307
length of database: 11,318,596
effective HSP length: 99
effective length of query: 208
effective length of database: 8,673,415
effective search space: 1804070320
effective search space used: 1804070320
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0239.11


