

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0234.6
(1379 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
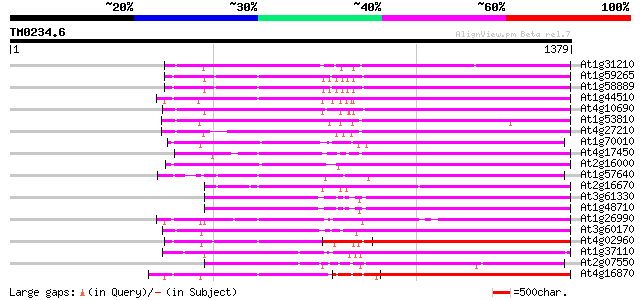
At1g31210 putative reverse transcriptase 772 0.0
At1g59265 polyprotein, putative 769 0.0
At1g58889 polyprotein, putative 768 0.0
At1g44510 polyprotein, putative 754 0.0
At4g10690 retrotransposon like protein 751 0.0
At1g53810 724 0.0
At4g27210 putative protein 689 0.0
At1g70010 hypothetical protein 614 e-175
At4g17450 retrotransposon like protein 596 e-170
At2g16000 putative retroelement pol polyprotein 595 e-170
At1g57640 590 e-168
At2g16670 putative retroelement pol polyprotein 586 e-167
At3g61330 copia-type polyprotein 582 e-166
At1g48710 hypothetical protein 578 e-165
At1g26990 polyprotein, putative 536 e-152
At3g60170 putative protein 534 e-151
At4g02960 putative polyprotein of LTR transposon 527 e-149
At1g37110 500 e-141
At2g07550 putative retroelement pol polyprotein 498 e-140
At4g16870 retrotransposon like protein 497 e-140
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 772 bits (1993), Expect = 0.0
Identities = 427/1026 (41%), Positives = 581/1026 (56%), Gaps = 45/1026 (4%)
Query: 381 VGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPY 440
VG G +PI +G TT+ +S PL+ VL P I K+L+SV +L D V FD
Sbjct: 353 VGDGTYLPITHTGSTTIKSSNGKIPLN--EVLVVPNIQKSLLSVSKLCDDYPCGVYFDAN 410
Query: 441 GFSVIDFQTGIPLMRCNSPGDLYPVTPSFPFAGL--------AQSLWHSRLGHPSSSALQ 492
+ID QT + LY V + F L + +WH RLGH +S ALQ
Sbjct: 411 KVCIIDLQTQKVVTTGPRRNGLY-VLENQEFVALYSNRQCAATEEVWHHRLGHANSKALQ 469
Query: 493 SLRSNKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSA 551
L+++K I +SPVCE C GK RLPF+ S++ + P D +H DLW SPV+S+
Sbjct: 470 HLQNSKAIQINKSRTSPVCEPCQMGKSSRLPFLISDSRVLHPLDRIHCDLWGPSPVVSNQ 529
Query: 552 GHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNGREFDNK 611
G ++Y +F+DD++ + W +PL NKS+ +FIS + + IK Q D G EF +
Sbjct: 530 GLKYYAIFVDDYSRYSWFYPLHNKSEFLSVFISFQKLVENQLNTKIKVFQSDGGGEFVSN 589
Query: 612 SFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMAT 671
+ + +G+ R SCP+T QNG AERK R + + ++L H+ P FW + A
Sbjct: 590 KLKTHLSEHGIHHRISCPYTPQQNGLAERKHRHLVELGLSMLFHSHTPQKFWVESFFTAN 649
Query: 672 YLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCVFLG 731
Y++N +P L NLSP + L+ P Y+ LRVFG CYP + NK PRS CVFLG
Sbjct: 650 YIINRLPSSVLKNLSPYEALFGEKPDYSSLRVFGSACYPCLRPLAQNKFDPRSLQCVFLG 709
Query: 732 YPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFANLTPTPSSTYEWLSDDIHPSVIHRWT 791
Y ++GY+CF KV ISR+VIF+E++ PF Y+ L ++ W
Sbjct: 710 YNSQYKGYRCFYPPTGKVYISRNVIFNESELPF-------KEKYQSLVPQYSTPLLQAWQ 762
Query: 792 TQTPSPDLQPTPVAPSATATPP------TSTASSSSPSDPSPSSSTPQSPPQPAPPVR-- 843
S P AP + P + + +DP P+S+ S + P
Sbjct: 763 HNKISEI--SVPAAPVQLFSKPIDLNTYAGSQVTEQLTDPEPTSNNEGSDEEVNPVAEEI 820
Query: 844 -----------TMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQ 892
M TRS GI KP + L I + PK A+ P W A+
Sbjct: 821 AANQEQVINSHAMTTRSKAGIQKPNTRYAL---ITSRMNTAEPKTLASAMKHPGWNEAVH 877
Query: 893 SEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDC 952
E + + +TW LVP D+NI+ W+F+ K +G ++ KARLV G Q GVD
Sbjct: 878 EEINRVHMLHTWSLVPPTDDMNILSSKWVFKTKLHPDGSIDKLKARLVAKGFDQEEGVDY 937
Query: 953 DETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNHPD 1012
ETFS VV+ ATIR VL ++ S+ WPI QLDV NAFLHG+L E V+M+QP GF DP P
Sbjct: 938 LETFSPVVRTATIRLVLDVSTSKGWPIKQLDVSNAFLHGELQEPVFMYQPSGFIDPQKPT 997
Query: 1013 YVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDI 1072
+VCRL K++YGLKQAPRAW+ F++++ GF S SD SLF+ + + YLLLYVDDI
Sbjct: 998 HVCRLTKAIYGLKQAPRAWFDTFSNFLLDYGFVCSKSDPSLFVCHQDGKILYLLLYVDDI 1057
Query: 1073 ILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAG 1132
+L S L + ++ L + F+MKDLGP YFLGI + +A GLFL Q+ YA DI+ +AG
Sbjct: 1058 LLTGSDQSLLEDLLQALKNRFSMKDLGPPRYFLGIQIEDYANGLFLHQTAYATDILQQAG 1117
Query: 1133 MASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQVCLHMH 1192
M+ C+P TP+ Q+L +PT +RSL G LQYLT TRPDI +AV +C MH
Sbjct: 1118 MSDCNPMPTPL--PQQLDNLNSELFAEPTYFRSLAGKLQYLTITRPDIQFAVNFICQRMH 1175
Query: 1193 APRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFL 1252
+P T LKRILRY++GT+ +GL + + L +Y+D+D GC +TRRST+G+C+ L
Sbjct: 1176 SPTTSDFGLLKRILRYIKGTIGMGLPIKRNSTLTLSAYSDSDHAGCKNTRRSTTGFCILL 1235
Query: 1253 GDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVS 1312
G NLISWS+KRQPT+S SS EAEYR + E W+ LL +L P T VYCDN+S
Sbjct: 1236 GSNLISWSAKRQPTVSNSSTEAEYRALTYAAREITWISFLLRDLGIPQYLPTQVYCDNLS 1295
Query: 1313 AIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGLPRVLFDDF 1372
A+YLS NP H R+KH + D H++RE+VA G H+ + Q+AD+FTK LPR F D
Sbjct: 1296 AVYLSANPALHNRSKHFDTDYHYIREQVALGLIETQHISATFQLADVFTKSLPRRAFVDL 1355
Query: 1373 RSSLSV 1378
RS L V
Sbjct: 1356 RSKLGV 1361
>At1g59265 polyprotein, putative
Length = 1466
Score = 769 bits (1986), Expect = 0.0
Identities = 464/1109 (41%), Positives = 605/1109 (53%), Gaps = 121/1109 (10%)
Query: 381 VGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPY 440
V G IPI +G T+L T K++PL+L ++L+ P I KNLISV +L N VSV F P
Sbjct: 362 VADGSTIPISHTGSTSLST--KSRPLNLHNILYVPNIHKNLISVYRLCNANGVSVEFFPA 419
Query: 441 GFSVIDFQTGIPLMRCNSPGDLY--PVTPSFPFAGLAQ-------SLWHSRLGHPSSSAL 491
F V D TG+PL++ + +LY P+ S P + A S WH+RLGHP+ S L
Sbjct: 420 SFQVKDLNTGVPLLQGKTKDELYEWPIASSQPVSLFASPSSKATHSSWHARLGHPAPSIL 479
Query: 492 QSLRSNKFISYEHLNSSPV---CESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVL 548
S+ SN +S LN S C C+ K ++PF S + P + ++SD+W+SP+L
Sbjct: 480 NSVISNYSLSV--LNPSHKFLSCSDCLINKSNKVPFSQSTINSTRPLEYIYSDVWSSPIL 537
Query: 549 SSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNGREF 608
S +R+YV+F+D FT + W +PL KSQV E FI+ N + F I DNG EF
Sbjct: 538 SHDNYRYYVIFVDHFTRYTWLYPLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEF 597
Query: 609 DNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQ 668
+ +Y + +G+ S PHT NG +ERK R I TLL+HAS+P ++W +A
Sbjct: 598 --VALWEYFSQHGISHLTSPPHTPEHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFA 655
Query: 669 MATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCV 728
+A YL+N +P L SP Q L+ P+Y LRVFGC CYP + +KL +S CV
Sbjct: 656 VAVYLINRLPTPLLQLESPFQKLFGTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCV 715
Query: 729 FLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPF----ANLTP--------------- 769
FLGY L Y C L ++ ISRHV FDE FPF A L+P
Sbjct: 716 FLGYSLTQSAYLCLHLQTSRLYISRHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPH 775
Query: 770 ------TPSSTYEWLSDDIH-------PSVIHRWTTQTPSPDLQ----------PTPVAP 806
TP SD H PS R +Q S +L P P AP
Sbjct: 776 TTLPTRTPVLPAPSCSDPHHAATPPSSPSAPFR-NSQVSSSNLDSSFSSSFPSSPEPTAP 834
Query: 807 SATATPPTS----------------------------TASSSSPSDPSP----------S 828
PT+ S S+P+ S S
Sbjct: 835 RQNGPQPTTQPTQTQTQTHSSQNTSQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASS 894
Query: 829 SSTPQSPP----QPAPPV--------------RTMATRSMRGIYKPRKLFNLSVTIDDPT 870
SST +PP P PP+ +M TR+ GI KP ++L+V++
Sbjct: 895 SSTSPTPPSILIHPPPPLAQIVNNNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSL---A 951
Query: 871 ISPLPKNPKLALSDPNWKSAMQSEFDALIRNNTWDLV-PRPCDVNIIRCMWIFRHKTKAN 929
P+ AL D W++AM SE +A I N+TWDLV P P V I+ C WIF K ++
Sbjct: 952 AESEPRTAIQALKDERWRNAMGSEINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSD 1011
Query: 930 GCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFL 989
G RYKARLV G +Q G+D ETFS V+K +IR VL +A+ RSWPI QLDV NAFL
Sbjct: 1012 GSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFL 1071
Query: 990 HGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTS 1049
G L + VYM QP GF D + P+YVC+LRK+LYGLKQAPRAWY +Y+ TIGF +S S
Sbjct: 1072 QGTLTDDVYMSQPPGFIDKDRPNYVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVS 1131
Query: 1050 DHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAV 1109
D SLF+ +RG + Y+L+YVDDI++ + L + + L+ F++KD L YFLGI
Sbjct: 1132 DTSLFVLQRGKSIVYMLVYVDDILITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEA 1191
Query: 1110 TRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGA 1169
R GL LSQ Y D++AR M + P TP+ KLS +GT DPT YR +VG+
Sbjct: 1192 KRVPTGLHLSQRRYILDLLARTNMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGS 1251
Query: 1170 LQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLIS 1229
LQYL FTRPDISYAV ++ MH P EH+ ALKRILRY+ GT G+ L L +
Sbjct: 1252 LQYLAFTRPDISYAVNRLSQFMHMPTEEHLQALKRILRYLAGTPNHGIFLKKGNTLSLHA 1311
Query: 1230 YTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWL 1289
Y+DADW G D ST+GY V+LG + ISWSSK+Q + RSS EAEYR VAN SE W+
Sbjct: 1312 YSDADWAGDKDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWI 1371
Query: 1290 RNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLH 1349
+LL EL L+ ++YCDNV A YL NPV H R KHI +D HF+R +V G RV+H
Sbjct: 1372 CSLLTELGIRLTRPPVIYCDNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVH 1431
Query: 1350 VPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
V + Q+AD TK L R F +F S + V
Sbjct: 1432 VSTHDQLADTLTKPLSRTAFQNFASKIGV 1460
>At1g58889 polyprotein, putative
Length = 1466
Score = 768 bits (1982), Expect = 0.0
Identities = 463/1109 (41%), Positives = 604/1109 (53%), Gaps = 121/1109 (10%)
Query: 381 VGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPY 440
V G IPI +G T+L T K++PL+L ++L+ P I KNLISV +L N VSV F P
Sbjct: 362 VADGSTIPISHTGSTSLST--KSRPLNLHNILYVPNIHKNLISVYRLCNANGVSVEFFPA 419
Query: 441 GFSVIDFQTGIPLMRCNSPGDLY--PVTPSFPFAGLAQ-------SLWHSRLGHPSSSAL 491
F V D TG+PL++ + +LY P+ S P + A S WH+RLGHP+ S L
Sbjct: 420 SFQVKDLNTGVPLLQGKTKDELYEWPIASSQPVSLFASPSSKATHSSWHARLGHPAPSIL 479
Query: 492 QSLRSNKFISYEHLNSSPV---CESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVL 548
S+ SN +S LN S C C+ K ++PF S + P + ++SD+W+SP+L
Sbjct: 480 NSVISNYSLSV--LNPSHKFLSCSDCLINKSNKVPFSQSTINSTRPLEYIYSDVWSSPIL 537
Query: 549 SSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNGREF 608
S +R+YV+F+D FT + W +PL KSQV E FI+ N + F I DNG EF
Sbjct: 538 SHDNYRYYVIFVDHFTRYTWLYPLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEF 597
Query: 609 DNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQ 668
+ +Y + +G+ S PHT NG +ERK R I TLL+HAS+P ++W +A
Sbjct: 598 --VALWEYFSQHGISHLTSPPHTPEHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFA 655
Query: 669 MATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCV 728
+A YL+N +P L SP Q L+ P+Y LRVFGC CYP + +KL +S CV
Sbjct: 656 VAVYLINRLPTPLLQLESPFQKLFGTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCV 715
Query: 729 FLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPF----ANLTP--------------- 769
FLGY L Y C L ++ ISRHV FDE FPF A L+P
Sbjct: 716 FLGYSLTQSAYLCLHLQTSRLYISRHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPH 775
Query: 770 ------TPSSTYEWLSDDIH-------PSVIHRWTTQTPSPDLQ----------PTPVAP 806
TP SD H PS R +Q S +L P P AP
Sbjct: 776 TTLPTRTPVLPAPSCSDPHHAATPPSSPSAPFR-NSQVSSSNLDSSFSSSFPSSPEPTAP 834
Query: 807 SATATPPTS----------------------------TASSSSPSDPSP----------S 828
PT+ S S+P+ S S
Sbjct: 835 RQNGPQPTTQPTQTQTQTHSSQNTSQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASS 894
Query: 829 SSTPQSPP----QPAPPV--------------RTMATRSMRGIYKPRKLFNLSVTIDDPT 870
SST +PP P PP+ +M TR+ GI KP ++L+V++
Sbjct: 895 SSTSPTPPSILIHPPPPLAQIVNNNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSL---A 951
Query: 871 ISPLPKNPKLALSDPNWKSAMQSEFDALIRNNTWDLV-PRPCDVNIIRCMWIFRHKTKAN 929
P+ AL D W++AM SE +A I N+TWDLV P P V I+ C WIF K ++
Sbjct: 952 AESEPRTAIQALKDERWRNAMGSEINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSD 1011
Query: 930 GCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFL 989
G RYKAR V G +Q G+D ETFS V+K +IR VL +A+ RSWPI QLDV NAFL
Sbjct: 1012 GSLNRYKARFVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFL 1071
Query: 990 HGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTS 1049
G L + VYM QP GF D + P+YVC+LRK+LYGLKQAPRAWY +Y+ TIGF +S S
Sbjct: 1072 QGTLTDDVYMSQPPGFIDKDRPNYVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVS 1131
Query: 1050 DHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAV 1109
D SLF+ +RG + Y+L+YVDDI++ + L + + L+ F++KD L YFLGI
Sbjct: 1132 DTSLFVLQRGKSIVYMLVYVDDILITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEA 1191
Query: 1110 TRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGA 1169
R GL LSQ Y D++AR M + P TP+ KLS +GT DPT YR +VG+
Sbjct: 1192 KRVPTGLHLSQRRYILDLLARTNMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGS 1251
Query: 1170 LQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLIS 1229
LQYL FTRPDISYAV ++ MH P EH+ ALKRILRY+ GT G+ L L +
Sbjct: 1252 LQYLAFTRPDISYAVNRLSQFMHMPTEEHLQALKRILRYLAGTPNHGIFLKKGNTLSLHA 1311
Query: 1230 YTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWL 1289
Y+DADW G D ST+GY V+LG + ISWSSK+Q + RSS EAEYR VAN SE W+
Sbjct: 1312 YSDADWAGDKDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWI 1371
Query: 1290 RNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLH 1349
+LL EL L+ ++YCDNV A YL NPV H R KHI +D HF+R +V G RV+H
Sbjct: 1372 CSLLTELGIRLTRPPVIYCDNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVH 1431
Query: 1350 VPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
V + Q+AD TK L R F +F S + V
Sbjct: 1432 VSTHDQLADTLTKPLSRTAFQNFASKIGV 1460
>At1g44510 polyprotein, putative
Length = 1459
Score = 754 bits (1946), Expect = 0.0
Identities = 449/1116 (40%), Positives = 608/1116 (54%), Gaps = 104/1116 (9%)
Query: 362 APGNLTSYSNLSHLNQK------LYVGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTP 415
A ++TS N L+Q + + G G+ I+ +G T L + + + L+L VL+ P
Sbjct: 343 ATHHITSDLNALSLHQPYNGGEYVMIADGTGLTIKQTGSTFLPS--QNRDLALHKVLYVP 400
Query: 416 QIVKNLISVRQLTTDNNVSVCFDPYGFSVIDFQTGIPLMRCNSPGDLY-------PVTPS 468
I KNLISV +L N VSV F P F V D TG L++ + DLY P T
Sbjct: 401 DIRKNLISVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDDLYEWPVTNPPATAL 460
Query: 469 F--PFAGLAQSLWHSRLGHPSSSALQSLRSNKFISYEHLNSSPV-CESCVFGKHVRLPFV 525
F P S WHSRLGHPS+S L +L S + +S+ C C+ K +LPF
Sbjct: 461 FTSPSPKTTLSSWHSRLGHPSASILNTLLSKFSLPVSVASSNKTSCSDCLINKSHKLPFA 520
Query: 526 SSNNVTVMPFDILHSDLWTSPVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISL 585
+S+ + P + + +D+WTSP++S +++Y++ +D +T + W +PL KSQV FI+
Sbjct: 521 TSSIHSSSPLEYIFTDVWTSPIISHDNYKYYLVLVDHYTRYTWLYPLQQKSQVKATFIAF 580
Query: 586 SNQIRTHFSQTIKCLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTI 645
+ F I+ L DNG EF + D+ +NG+ S PHT NG +ERK R I
Sbjct: 581 KALVENRFQAKIRTLYSDNGGEFI--ALRDFLVSNGISHLTSPPHTPEHNGLSERKHRHI 638
Query: 646 NNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFG 705
TLL ASVP +W +A A YL+N +P L SP Q L+ P+Y LRVFG
Sbjct: 639 VETGLTLLTQASVPREYWTYAFATAVYLINRMPTPVLCLQSPFQKLFGSSPNYQRLRVFG 698
Query: 706 CLCYPLVPSSTINKLQPRSTPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFA 765
CLC+P + T NKL+ RS CVFLGY L Y C D+ + ++ SRHV+FDE+ +PFA
Sbjct: 699 CLCFPWLRPYTRNKLEERSKRCVFLGYSLTQTAYLCLDVDNNRLYTSRHVMFDESTYPFA 758
Query: 766 N----------LTPTPSSTYEWLSDDIHPSVIHRWTT------QTPSPDLQP--TPVAPS 807
+TP SS+ ++ P + R + +TPSP Q +PV+P
Sbjct: 759 ASIREQSQSSLVTPPESSSSSSPANSGFPCSVLRLQSPPASSPETPSPPQQQNDSPVSPR 818
Query: 808 ATATP--------------PTSTASSSSPSDP---------------------------- 825
T +P P+ + S+S P+ P
Sbjct: 819 QTGSPTPSHHSQVRDSTLSPSPSVSNSEPTAPHENGPEPEAQSNPNSPFIGPLPNPNPET 878
Query: 826 SPSSSTPQSPPQPA-----PPVRT------------------MATRSMRGIYKPRKLFNL 862
+PSSS Q P + PP +T M TRS I KP+ +L
Sbjct: 879 NPSSSIEQRPVDKSTTTALPPNQTTIAATSNSRSQPPKNNHQMKTRSKNNITKPKTKTSL 938
Query: 863 SVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIF 922
+V + P +S P AL D W+ AM EFDA RN+TWDLVP +++ C W+F
Sbjct: 939 TVALTQPHLSE-PNTVTQALKDKKWRFAMSDEFDAQQRNHTWDLVPPNPTQHLVGCRWVF 997
Query: 923 RHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQL 982
+ K NG ++YKARLV G +Q GVD ETFS V+K TIR VL +A+ ++WP+ QL
Sbjct: 998 KLKYLPNGLIDKYKARLVAKGFNQQYGVDYAETFSPVIKATTIRVVLDVAVKKNWPLKQL 1057
Query: 983 DVQNAFLHGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTI 1042
DV NAFL G L E VYM QP GF D + P +VCRLRK++YGLKQAPRAWY ++ I
Sbjct: 1058 DVNNAFLQGTLTEEVYMAQPPGFVDKDRPSHVCRLRKAIYGLKQAPRAWYMELKQHLLNI 1117
Query: 1043 GFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLS 1102
GF +S +D SLFIY G+ + YLL+YVDDII+ S H ++++ LA F++KD L
Sbjct: 1118 GFVNSLADTSLFIYSHGTTLLYLLVYVDDIIVTGSDHKSVSAVLSSLAERFSIKDPTDLH 1177
Query: 1103 YFLGIAVTRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTL 1162
YFLGI TR GL L Q Y D++A+ M P ATP+ T KL+ GT +D +
Sbjct: 1178 YFLGIEATRTNTGLHLMQRKYMTDLLAKHNMLDAKPVATPLPTSPKLTLHGGTKLNDASE 1237
Query: 1163 YRSLVGALQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPS 1222
YRS+VG+LQYL FTRPDI++AV ++ MH P ++H A KR+LRY+ GT G+ L S
Sbjct: 1238 YRSVVGSLQYLAFTRPDIAFAVNRLSQFMHQPTSDHWQAAKRVLRYLAGTTTHGIFLNSS 1297
Query: 1223 PIEKLISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANV 1282
L +++DADW G ST+ Y ++LG N ISWSSK+Q +SRSS E+EYR VAN
Sbjct: 1298 SPIHLHAFSDADWAGDSADYVSTNAYVIYLGRNPISWSSKKQRGVSRSSTESEYRAVANA 1357
Query: 1283 VSESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVAR 1342
SE WL +LL ELH L ++CDN+ A Y+ NPV H R KHI +D HFVR +
Sbjct: 1358 ASEIRWLCSLLTELHIRLPHGPTIFCDNIGATYICANPVFHSRMKHIALDYHFVRGMIQS 1417
Query: 1343 GQARVLHVPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
RV HV + Q+AD TK L R F RS + V
Sbjct: 1418 RALRVSHVSTNDQLADALTKSLSRPHFLSARSKIGV 1453
>At4g10690 retrotransposon like protein
Length = 1515
Score = 751 bits (1938), Expect = 0.0
Identities = 435/1084 (40%), Positives = 593/1084 (54%), Gaps = 93/1084 (8%)
Query: 376 NQKLYVGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSV 435
+ + VG+G +PI G L S T PL VL P I K+L+SV +LT D S
Sbjct: 350 DDSVIVGNGDFLPITHIGTIPLNISQGTLPLE--DVLVCPGITKSLLSVSKLTDDYPCSF 407
Query: 436 CFDPYGFSVIDFQTGIPLMRCNSPGDLYPVTPSFPFAGLAQS--------LWHSRLGHPS 487
FD + D +T L + N LY V PF + +WH RLGHP+
Sbjct: 408 TFDSDSVVIKDKRTQQLLTQGNKHKGLY-VLKDVPFQTYYSTRQQSSDDEVWHQRLGHPN 466
Query: 488 SSALQSLRSNKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSP 546
LQ L K I SS +CE+C GK RLPFV+S V+ P + +H DLW +P
Sbjct: 467 KEVLQHLIKTKAIVVNK-TSSNMCEACQMGKVCRLPFVASEFVSSRPLERIHCDLWGPAP 525
Query: 547 VLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNGR 606
V S+ G ++YV+F+D+++ F W +PL KS F +F+ + + I QCD G
Sbjct: 526 VTSAQGFQYYVIFIDNYSRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHKIAMFQCDGGG 585
Query: 607 EFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHA 666
EF + F + A+ G+ SCPHT QNG AER+ R + + +L+ H+ VP W A
Sbjct: 586 EFVSYKFVAHLASCGIKQLISCPHTPQQNGIAERRHRYLTELGLSLMFHSKVPHKLWVEA 645
Query: 667 LQMATYLLNIIPRKNLS-NLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRST 725
+ +L N++P LS N SP ++L+ P YT LRVFG CYP + NK P+S
Sbjct: 646 FFTSNFLSNLLPSSTLSDNKSPYEMLHGTPPVYTALRVFGSACYPYLRPYAKNKFDPKSL 705
Query: 726 PCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFANL------------------ 767
CVFLGY ++GY+C KV I RHV+FDE +FP++++
Sbjct: 706 LCVFLGYNNKYKGYRCLHPPTGKVYICRHVLFDERKFPYSDIYSQFQTISGSPLFTAWQK 765
Query: 768 --------TPTPSSTYEWLSDDIHPSVIHRWTTQTP-SPDLQPTPVAPSATAT------- 811
TPS+ E D I PS + T +P++ T AP
Sbjct: 766 GFSSTALSRETPSTNVE---DIIFPSATVSSSVPTGCAPNIAETATAPDVDVAAAHDMVV 822
Query: 812 PPTSTASSSSPSDPSPSSS-----------------TPQS------PPQPAPPVRT---- 844
PP+ S+S P+ P S+S TPQS PP+++
Sbjct: 823 PPSPITSTSLPTQPEESTSDQNHYSTDSETAISSAMTPQSINVSLFEDSDFPPLQSVISS 882
Query: 845 ----------MATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSE 894
M TR+ GI KP + L + P PK+ K AL D W +AM E
Sbjct: 883 TTAAPETSHPMITRAKSGITKPNPKYALFSVKSN---YPEPKSVKEALKDEGWTNAMGEE 939
Query: 895 FDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDCDE 954
+ +TWDLVP ++ C W+F+ K ++G +R KARLV G Q GVD E
Sbjct: 940 MGTMHETDTWDLVPPEMVDRLLGCKWVFKTKLNSDGSLDRLKARLVARGYEQEEGVDYVE 999
Query: 955 TFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNHPDYV 1014
T+S VV+ AT+R++L +A W + QLDV+NAFLH +L ETV+M QP GF DP+ PDYV
Sbjct: 1000 TYSPVVRSATVRSILHVATINKWSLKQLDVKNAFLHDELKETVFMTQPPGFEDPSRPDYV 1059
Query: 1015 CRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIIL 1074
C+L+K++Y LKQAPRAW+ F+ Y+ GF S SD SLF+Y +G D+ +LLLYVDD+IL
Sbjct: 1060 CKLKKAIYDLKQAPRAWFDKFSSYLLKYGFICSFSDPSLFVYLKGRDVMFLLLYVDDMIL 1119
Query: 1075 ISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAGMA 1134
++ L + ++ +L++EF MKD+G L YFLGI H GLFLSQ Y D++ AGM+
Sbjct: 1120 TGNNDVLLQQLLNILSTEFRMKDMGALHYFLGIQAHYHNDGLFLSQEKYTSDLLVNAGMS 1179
Query: 1135 SCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQVCLHMHAP 1194
C S+ P + L P +PT +R L G LQYLT TRPDI +AV VC MHAP
Sbjct: 1180 DC--SSMPTPLQLDLLQGNNKPFPEPTYFRRLAGKLQYLTLTRPDIQFAVNFVCQKMHAP 1237
Query: 1195 RTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFLGD 1254
LKRIL Y++GT+ +G++L + L Y+D+DW GC DTRRST G+C FLG
Sbjct: 1238 TMSDFHLLKRILHYLKGTMTMGINLSSNTDSVLRCYSDSDWAGCKDTRRSTGGFCTFLGY 1297
Query: 1255 NLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVSAI 1314
N+ISWS+KR PT+S+SS EAEYR ++ SE W+ LL E+ P +YCDN+SA+
Sbjct: 1298 NIISWSAKRHPTVSKSSTEAEYRTLSFAASEVSWIGFLLQEIGLPQQQIPEMYCDNLSAV 1357
Query: 1315 YLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGLPRVLFDDFRS 1374
YLS NP H R+KH ++D ++VRE+VA G V H+P+ Q+ADIFTK LP+ F D R
Sbjct: 1358 YLSANPALHSRSKHFQVDYYYVRERVALGALTVKHIPASQQLADIFTKSLPQAPFCDLRF 1417
Query: 1375 SLSV 1378
L V
Sbjct: 1418 KLGV 1421
>At1g53810
Length = 1522
Score = 724 bits (1870), Expect = 0.0
Identities = 418/1085 (38%), Positives = 575/1085 (52%), Gaps = 86/1085 (7%)
Query: 374 HLNQKLYVGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNV 433
H + + V G +PI +G ++ +S PL VL P IVK+L+SV +LT+D
Sbjct: 350 HGSDSIMVADGNFLPITHTGSGSIASSSGKIPLK--EVLVCPDIVKSLLSVSKLTSDYPC 407
Query: 434 SVCFDPYGFSVIDFQTGIPLMRCNSPGDLYP-------VTPSFPFAGLAQSLWHSRLGHP 486
SV FD + D T L+ + LY V S + +WH RLGH
Sbjct: 408 SVEFDADSVRINDKATKKLLVMGRNRDGLYSLEEPKLQVLYSTRQNSASSEVWHRRLGHA 467
Query: 487 SSSALQSLRSNKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TS 545
++ L L S+K I + VCE+C GK RLPF+ S P + +H DLW S
Sbjct: 468 NAEVLHQLASSKSIIIINKVVKTVCEACHLGKSTRLPFMLSTFNASRPLERIHCDLWGPS 527
Query: 546 PVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNG 605
P S G R+YV+F+D ++ F W +PL KS F F+ + IK QCD G
Sbjct: 528 PTSSVQGFRYYVVFIDHYSRFTWFYPLKLKSDFFSTFVMFQKLVENQLGHKIKIFQCDGG 587
Query: 606 REFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHH 665
EF + F + +G+ SCP+T QNG AERK R I + +++ + +P +W
Sbjct: 588 GEFISSQFLKHLQDHGIQQNMSCPYTPQQNGMAERKHRHIVELGLSMIFQSKLPLKYWLE 647
Query: 666 ALQMATYLLNIIPRKNL-SNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRS 724
+ A +++N++P +L +N SP Q LY + P Y+ LRVFGC CYP + K PRS
Sbjct: 648 SFFTANFVINLLPTSSLDNNESPYQKLYGKAPEYSALRVFGCACYPTLRDYASTKFDPRS 707
Query: 725 TPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPF----ANLTPTPSSTY--EWL 778
CVFLGY ++GY+C ++ ISRHV+FDE PF ++L P + W
Sbjct: 708 LKCVFLGYNEKYKGYRCLYPPTGRIYISRHVVFDENTHPFESIYSHLHPQDKTPLLEAWF 767
Query: 779 SDDIH-----PSVIHRWTTQTPSPDLQPTPVAPSATATP--------------------- 812
H P + P P+ AP++ A
Sbjct: 768 KSFHHVTPTQPDQSRYPVSSIPQPETTDLSAAPASVAAETAGPNASDDTSQDNETISVVS 827
Query: 813 --------------------PTSTASSSSPSDPSPSSSTPQSPPQPAP------PV---R 843
PT+ +S SP+ SP+SS SP Q AP PV
Sbjct: 828 GSPERTTGLDSASIGDSYHSPTADSSHPSPARSSPASSPQGSPIQMAPAQQVQAPVTNEH 887
Query: 844 TMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNNT 903
M TR GI KP K + V + P PK AL P W +AMQ E T
Sbjct: 888 AMVTRGKEGISKPNKRY---VLLTHKVSIPEPKTVTEALKHPGWNNAMQEEMGNCKETET 944
Query: 904 WDLVPRPCDVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPA 963
W LVP ++N++ MW+FR K A+G ++ KARLV G Q G+D ET+S VV+
Sbjct: 945 WTLVPYSPNMNVLGSMWVFRTKLHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRTP 1004
Query: 964 TIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYG 1023
T+R +L +A W + Q+DV+NAFLHGDL ETVYM QP GF D + PD+VC L KSLYG
Sbjct: 1005 TVRLILHVATVLKWELKQMDVKNAFLHGDLTETVYMRQPAGFVDKSKPDHVCLLHKSLYG 1064
Query: 1024 LKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRK 1083
LKQ+PRAW+ F++++ GF S D SLF+Y +D+ LLLYVDD+++ ++
Sbjct: 1065 LKQSPRAWFDRFSNFLLEFGFICSLFDPSLFVYSSNNDVILLLLYVDDMVITGNNSQSLT 1124
Query: 1084 SIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAGMASCHPSATPV 1143
++A L EF MKD+G + YFLGI + + GGLF+SQ YA D++ A MA+C P TP+
Sbjct: 1125 HLLAALNKEFRMKDMGQVHYFLGIQIQTYDGGLFMSQQKYAEDLLITASMANCSPMPTPL 1184
Query: 1144 DTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALK 1203
+ ++ DPT +RSL G LQYLT TRPDI +AV VC MH P LK
Sbjct: 1185 PLQLDRVSNQDEVFSDPTYFRSLAGKLQYLTLTRPDIQFAVNFVCQKMHQPSVSDFNLLK 1244
Query: 1204 RILRYVQGTLQLGLHLYPSPIEKLIS----------YTDADWGGCLDTRRSTSGYCVFLG 1253
RILRY++GT+ +G+ Y S ++S Y+D+D+ C +TRRS GYC F+G
Sbjct: 1245 RILRYIKGTVSMGIQ-YNSNSSSVVSAYESDYDLSAYSDSDYANCKETRRSVGGYCTFMG 1303
Query: 1254 DNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVSA 1313
N+ISWSSK+QPT+SRSS EAEYR ++ SE W+ ++L E+ L ++CDN+SA
Sbjct: 1304 QNIISWSSKKQPTVSRSSTEAEYRSLSETASEIKWMSSILREIGVSLPDTPELFCDNLSA 1363
Query: 1314 IYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGLPRVLFDDFR 1373
+YL+ NP H RTKH ++D H++RE+VA V H+P Q+ADIFTK LP F R
Sbjct: 1364 VYLTANPAFHARTKHFDVDHHYIRERVALKTLVVKHIPGHLQLADIFTKSLPFEAFTRLR 1423
Query: 1374 SSLSV 1378
L V
Sbjct: 1424 FKLGV 1428
>At4g27210 putative protein
Length = 1318
Score = 689 bits (1779), Expect = 0.0
Identities = 408/1063 (38%), Positives = 551/1063 (51%), Gaps = 104/1063 (9%)
Query: 374 HLNQKLYVGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNV 433
H + V G +PI +G T L +S T PL+ VL P I K+L+S+ +LT D
Sbjct: 201 HGTDAIMVDDGNYLPITHTGSTNLASSSGTVPLT--DVLVCPSITKSLLSMSKLTQDFPC 258
Query: 434 SVCFDPYGFSVIDFQTGIPLMRCNSPGDLYPVTPSFPFAGL--------AQSLWHSRLGH 485
+V F+ G V D T L+ ++ LY + F + +WH RLGH
Sbjct: 259 TVEFEYDGVRVNDKATKKLLLMGSNRDGLYCLKDDKQFQAFFSTRQRSASDEVWHRRLGH 318
Query: 486 PSSSALQSLRSNKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTS 545
P LQ P + +H DLW
Sbjct: 319 PHPQILQ-----------------------------------------PLERVHCDLWGP 337
Query: 546 PVLSSA-GHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDN 604
++S G R+Y +F+D ++ F W +PL KS + +F++ + SQ I QCD
Sbjct: 338 TTITSVQGFRYYAVFIDHYSRFSWIYPLKLKSDFYNIFLAFHKLVENQLSQKISVFQCDG 397
Query: 605 GREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWH 664
G EF + F + ++G+ + SCPHT QNG AERK R + + ++L + VP FW
Sbjct: 398 GGEFVSHKFLQHLQSHGIQQQLSCPHTPQQNGLAERKHRHLVELGLSMLFQSHVPHKFWV 457
Query: 665 HALQMATYLLNIIPRKNLS-NLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPR 723
A A +L+N++P L ++SP + LY + P YT LR FG C+P + NK P
Sbjct: 458 EAFFTANFLINLLPTSALKESISPYEKLYDKKPDYTSLRSFGSACFPTLRDYAENKFNPC 517
Query: 724 STPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFAN----LTPTPSSTY--EW 777
S CVFLGY ++GY+C ++ ISRHVIFDE+ +PF++ L P P + W
Sbjct: 518 SLKCVFLGYNEKYKGYRCLYPPTGRLYISRHVIFDESVYPFSHTYKHLHPQPRTPLLAAW 577
Query: 778 L--SDDIHPSVIHRWTTQTP---SPDLQP-----TPVAPSATATPPTSTAS--------- 818
L SD PS ++++P S D P TP+ P+ S AS
Sbjct: 578 LRSSDSPAPSTSTSPSSRSPLFTSADFPPLPQRKTPLLPTLVPISSVSHASNITTQQSPD 637
Query: 819 ----------SSSPSDPSPSSSTPQSPPQP-------------APPVRTMATRSMRGIYK 855
S+S D S SS + + V M TR+ GI K
Sbjct: 638 FDSERTTDFDSASIGDSSHSSQAGSDSEETIQQASVNVHQTHASTNVHPMVTRAKVGISK 697
Query: 856 PRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNNTWDLVPRPCDVNI 915
P + V + P PK AL P W AM E TW LVP D+++
Sbjct: 698 PNPRY---VFLSHKVSYPEPKTVTAALKHPGWTGAMTEEIGNCSETQTWSLVPYKSDMHV 754
Query: 916 IRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSR 975
+ W+FR K A+G + KAR+V G Q G+D ET+S VV+ T+R VL +A +
Sbjct: 755 LGSKWVFRTKLHADGTLNKLKARIVAKGFLQEEGIDYLETYSPVVRTPTVRLVLHLATAL 814
Query: 976 SWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCF 1035
+W I Q+DV+NAFLHGDL ETVYM QP GF DP+ PD+VC L KS+YGLKQ+PRAW+ F
Sbjct: 815 NWDIKQMDVKNAFLHGDLKETVYMTQPAGFVDPSKPDHVCLLHKSIYGLKQSPRAWFDKF 874
Query: 1036 ADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFAM 1095
+ ++ GF S SD SLFIY +++ LLLYVDD+++ +S S++A L EF M
Sbjct: 875 STFLLEFGFFCSKSDPSLFIYAHNNNLILLLLYVDDMVITGNSSQTLTSLLAALNKEFRM 934
Query: 1096 KDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTSAGT 1155
D+G L YFLGI V R GLF+SQ YA D++ A M C P TP+ +
Sbjct: 935 TDMGQLHYFLGIQVQRQQNGLFMSQQKYAEDLLIAASMEHCTPLPTPLPVQLDRVPHQEE 994
Query: 1156 PCDDPTLYRSLVGALQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQL 1215
DPT +RS+ G LQYLT TRPDI +AV VC MH P LKRILRY++GT+ +
Sbjct: 995 LFSDPTYFRSIAGKLQYLTLTRPDIQFAVNFVCQKMHQPTISDFHLLKRILRYIKGTITM 1054
Query: 1216 GLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAE 1275
G+ L +Y+D+DWG C TRRS G C F+G NL+SWSSK+ PT+SRSS EAE
Sbjct: 1055 GISYSRDSPTLLQAYSDSDWGNCKQTRRSVGGLCTFMGTNLVSWSSKKHPTVSRSSTEAE 1114
Query: 1276 YRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHF 1335
Y+ +++ SE WL LL EL PL ++CDN+SA+YL+ NP H RTKH ++D HF
Sbjct: 1115 YKSLSDAASEILWLSTLLRELRIPLPDTPELFCDNLSAVYLTANPAFHARTKHFDIDFHF 1174
Query: 1336 VREKVARGQARVLHVPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
VRE+VA V H+P QIADIFTK LP F R L V
Sbjct: 1175 VRERVALKALVVKHIPGSEQIADIFTKSLPYEAFIHLRGKLGV 1217
>At1g70010 hypothetical protein
Length = 1315
Score = 614 bits (1583), Expect = e-175
Identities = 367/1016 (36%), Positives = 533/1016 (52%), Gaps = 72/1016 (7%)
Query: 388 PIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVIDF 447
PI GS H + L L+ VL PQ NL+SV LT + FD + D
Sbjct: 312 PISGSVHLG-------RHLILNDVLFIPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDA 364
Query: 448 QTGIPLMRCNSPGDLYPVT----------PSFPFAGL-AQSLWHSRLGHPSSSALQSLRS 496
+ + +LY V S A + + LWH RLGHPS LQ + S
Sbjct: 365 TRELMVGMGKQVANLYIVDLDSLSHPGTDSSITVASVTSHDLWHKRLGHPSVQKLQPMSS 424
Query: 497 NKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTS-PVLSSAGHRF 555
+ N+ C C K LPFVS NN + PFD++H D W V + G+R+
Sbjct: 425 LLSFPKQKNNTDFHCRVCHISKQKHLPFVSHNNKSSRPFDLIHIDTWGPFSVQTHDGYRY 484
Query: 556 YVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNGREFDNKSFHD 615
++ +DD++ W + L NKS V + + + F TIK ++ DN E + F+
Sbjct: 485 FLTIVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPELNFTQFYH 544
Query: 616 YCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLN 675
+ G++ SCP T QN ERK + I N+ R+L + +P S+W + A YL+N
Sbjct: 545 ---SKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLIN 601
Query: 676 IIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCVFLGYPLH 735
+P L + P ++L + P+Y H++VFGCLCY +K PR+ C F+GYP
Sbjct: 602 RLPAPILEDKCPFEVLTKTVPTYDHIKVFGCLCYASTSPKDRHKFSPRAKACAFIGYPSG 661
Query: 736 HRGYKCFDLSHRKVIISRHVIFDETQFPFANLTPTPSSTYEWLSDDIHPSVIHRWTTQTP 795
+GYK DL +I+SRHV+F E FPF L D+ Q
Sbjct: 662 FKGYKLLDLETHSIIVSRHVVFHEELFPF-------------LGSDLSQE------EQNF 702
Query: 796 SPDLQPTPVAPSATATPPTSTASSSSPSDPSPSSSTPQSPPQPAPPVRTMATRSMRGIY- 854
PDL PTP ++ + + SSS + PS++ + P+P+ V+T ++ + Y
Sbjct: 703 FPDLNPTPPM-QRQSSDHVNPSDSSSSVEILPSANPTNNVPEPS--VQTSHRKAKKPAYL 759
Query: 855 --------------KPRKLFNLSVTIDDPTISPL--------PKNPKLALSDPNWKSAMQ 892
+ RK + I+DP ++ L P N A W+ AM
Sbjct: 760 QDYYCHSVVSSTPHEIRKFLSYD-RINDPYLTFLACLDKTKEPSNYTEAEKLQVWRDAMG 818
Query: 893 SEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDC 952
+EFD L +TW++ P D I C WIF+ K ++G ERYKARLV G +Q G+D
Sbjct: 819 AEFDFLEGTHTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYTQKEGIDY 878
Query: 953 DETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFR----DP 1008
+ETFS V K +++ +L +A + QLD+ NAFL+GDL E +YM P G+ D
Sbjct: 879 NETFSPVAKLNSVKLLLGVAARFKLSLTQLDISNAFLNGDLDEEIYMRLPQGYASRQGDS 938
Query: 1009 NHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLY 1068
P+ VCRL+KSLYGLKQA R WY F+ + +GF S DH+ F+ +L+Y
Sbjct: 939 LPPNAVCRLKKSLYGLKQASRQWYLKFSSTLLGLGFIQSYCDHTCFLKISDGIFLCVLVY 998
Query: 1069 VDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDII 1128
+DDII+ S++ + + + S F ++DLG L YFLG+ + R G+ +SQ YA D++
Sbjct: 999 IDDIIIASNNDAAVDILKSQMKSFFKLRDLGELKYFLGLEIVRSDKGIHISQRKYALDLL 1058
Query: 1129 ARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQVC 1188
G C PS+ P+D + +G + YR L+G L YL TRPDI++AV ++
Sbjct: 1059 DETGQLGCKPSSIPMDPSMVFAHDSGGDFVEVGPYRRLIGRLMYLNITRPDITFAVNKLA 1118
Query: 1189 LHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGY 1248
APR H+ A+ +IL+Y++GT+ GL + +L Y +AD+ C D+RRSTSGY
Sbjct: 1119 QFSMAPRKAHLQAVYKILQYIKGTIGQGLFYSATSELQLKVYANADYNSCRDSRRSTSGY 1178
Query: 1249 CVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVYC 1308
C+FLGD+LI W S++Q +S+SSAEAEYR ++ E WL N L EL PLS TL++C
Sbjct: 1179 CMFLGDSLICWKSRKQDVVSKSSAEAEYRSLSVATDELVWLTNFLKELQVPLSKPTLLFC 1238
Query: 1309 DNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGL 1364
DN +AI+++ N V H+RTKHIE D H VRE++ +G + H+ + QIAD FTK L
Sbjct: 1239 DNEAAIHIANNHVFHERTKHIESDCHSVRERLLKGLFELYHINTELQIADPFTKPL 1294
>At4g17450 retrotransposon like protein
Length = 1433
Score = 596 bits (1536), Expect = e-170
Identities = 352/983 (35%), Positives = 498/983 (49%), Gaps = 50/983 (5%)
Query: 406 LSLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVIDFQTGIPLMRCNSPGDLYPV 465
+SL +VL+ P+ NLIS +LT + + V+DF + + P
Sbjct: 482 ISLHNVLYIPEFKFNLIS--ELTKELMIGRGSQVGNLYVLDFNENNHTVSLKGTTSMCPE 539
Query: 466 TPSFPFAGLAQSLWHSRLGHPSSSALQSLRSN-----KFISYEHLNSSPVCESCVFGKHV 520
+ WH RLGHP+ S + L K I+ EH VC C K
Sbjct: 540 FSVCSSVVVDSVTWHKRLGHPAYSKIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQK 599
Query: 521 RLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFE 580
L F S N+ FD++H D W F D W + L NKS V
Sbjct: 600 HLSFQSRQNMCSAAFDLVHIDTWGP-------------FSVPTNDATWIYLLKNKSDVLH 646
Query: 581 MFISLSNQIRTHFSQTIKCLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAER 640
+F + N + T + +K ++ DN E F D AA+G++ SCP T QN ER
Sbjct: 647 VFPAFINMVHTQYQTKLKSVRSDNAHEL---KFTDLFAAHGIVAYHSCPETPEQNSVVER 703
Query: 641 KIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSYTH 700
K + I N+ R LL +++P FW + A +L+N +P L+N SP + L P+Y
Sbjct: 704 KHQHILNVARALLFQSNIPLEFWGDCVLTAVFLINRLPTPVLNNKSPYEKLKNIPPAYES 763
Query: 701 LRVFGCLCYPLVPSSTINKLQPRSTPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDET 760
L+ FGCLCY +K +PR+ CVFLGYPL ++GYK D+ V ISRHVIF E
Sbjct: 764 LKTFGCLCYSSTSPKQRHKFEPRARACVFLGYPLGYKGYKLLDIETHAVSISRHVIFHED 823
Query: 761 QFPFANLTPTPSSTYEWLSDDIHPSV-IHRWTTQTPSPDLQPTPVAPSATATPPTSTASS 819
FPF + T + DDI + ++ +T L+ T + T P SS
Sbjct: 824 IFPFISST---------IKDDIKDFFPLLQFPARTDDLPLEQTSIID----THPHQDVSS 870
Query: 820 SSPSDPSPSSSTPQSPPQPAPPVRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPK 879
S P P S Q PP +P F I++ T + +P+
Sbjct: 871 SKALVPFD----PLSKRQKKPPKHLQDFHCYNNTTEPFHAF-----INNITNAVIPQRYS 921
Query: 880 LALSDPNWKSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCFERYKARL 939
A W AM+ E A++R NTW +V P + I C W+F K A+G ERYKARL
Sbjct: 922 EAKDFKAWCDAMKEEIGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKARL 981
Query: 940 VGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYM 999
V G +Q G+D +ETFS V K ++R +L +A W +HQLD+ NAFL+GDL E +YM
Sbjct: 982 VAKGYTQEEGLDYEETFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEIYM 1041
Query: 1000 HQPLGFRD----PNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFI 1055
P G+ D P +CRL KS+YGLKQA R WY ++ + +GFQ S +DH+LFI
Sbjct: 1042 KIPPGYADLVGEALPPHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTLFI 1101
Query: 1056 YRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGG 1115
+ +L+YVDDI+++S+S D A L S F ++DLG YFLGI + R G
Sbjct: 1102 KYANGVLMGVLVYVDDIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSEKG 1161
Query: 1116 LFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTF 1175
+ + Q Y ++++ G PS+ P+D KL+ G P D T YR LVG L YL
Sbjct: 1162 ISICQRKYILELLSTTGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYLQI 1221
Query: 1176 TRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLISYTDADW 1235
TRPDI+YAV +C HAP + H+ A+ ++LRY++GT+ GL L YTD+D+
Sbjct: 1222 TRPDIAYAVNTLCQFSHAPTSVHLSAVHKVLRYLKGTVGQGLFYSADDKFDLRGYTDSDF 1281
Query: 1236 GGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLE 1295
G C D+RR + YC+F+GD L+SW SK+Q T+S S+AEAE+R ++ E WL L +
Sbjct: 1282 GSCTDSRRCVAAYCMFIGDYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLFDD 1341
Query: 1296 LHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQ 1355
P +YCDN +A+++ N V H+RTK +E+D + RE V G + + V + Q
Sbjct: 1342 FKVPFIPPAYLYCDNTAALHIVNNSVFHERTKFVELDCYKTREAVESGFLKTMFVETGEQ 1401
Query: 1356 IADIFTKGLPRVLFDDFRSSLSV 1378
+AD TK + F + V
Sbjct: 1402 VADPLTKAIHPAQFHKLIGKMGV 1424
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 595 bits (1535), Expect = e-170
Identities = 356/1017 (35%), Positives = 524/1017 (51%), Gaps = 54/1017 (5%)
Query: 383 SGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGF 442
+G + I G G L + L +VL P+ NLIS+ LT D V FD
Sbjct: 463 TGPTVKISGVGTLKL-----NDDILLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSC 517
Query: 443 SVIDFQTGIPLMRCNSPGDLYPVTPSFPFAGLAQ----SLWHSRLGHPSSSALQSLRSNK 498
+ D G L + +LY + + S+WH RLGH S L ++ +
Sbjct: 518 EIQDLIKGRMLGQGRRVANLYLLDVGDQSISVNAVVDISMWHRRLGHASLQRLDAISDSL 577
Query: 499 FISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTS-PVLSSAGHRFYV 557
+ S C C K +L F +SN V FD+LH D+W V + G+++++
Sbjct: 578 GTTRHKNKGSDFCHVCHLAKQRKLSFPTSNKVCKEIFDLLHIDVWGPFSVETVEGYKYFL 637
Query: 558 LFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNGREFDNKSFHDYC 617
+DD + W + L KS+V +F + Q+ + +K ++ DN E SF+
Sbjct: 638 TIVDDHSRATWMYLLKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDNAPELKFTSFY--- 694
Query: 618 AANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNII 677
A G++ SCP T QN ERK + I N+ R L+ + VP S W + A +L+N
Sbjct: 695 AEKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRT 754
Query: 678 PRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCVFLGYPLHHR 737
P + L N +P ++L P Y LR FGCLCY +K QPRS C+FLGYP ++
Sbjct: 755 PSQLLMNKTPYEILTGTAPVYEQLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYK 814
Query: 738 GYKCFDLSHRKVIISRHVIFDETQFPFANLTPTPSSTYEWLSDDIHPSVIHRWTTQTPSP 797
GYK DL V ISR+V F E FP A + SS +
Sbjct: 815 GYKLMDLESNTVFISRNVQFHEEVFPLAKNPGSESSLKLF-------------------- 854
Query: 798 DLQPTPVAP------SATATPPTSTASSSSPSDPSPSSSTPQSPPQPAPPVR--TMATRS 849
TP+ P S T P+S S S P SS + PP TM +
Sbjct: 855 ----TPMVPVSSGIISDTTHSPSSLPSQISDLPPQISSQRVRKPPAHLNDYHCNTMQSDH 910
Query: 850 MRGIYKPRKLFNLSVT----IDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNNTWD 905
I +S + I++ T P+P N A W A+ +E A+ + NTW+
Sbjct: 911 KYPISSTISYSKISPSHMCYINNITKIPIPTNYAEAQDTKEWCEAVDAEIGAMEKTNTWE 970
Query: 906 LVPRPCDVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATI 965
+ P + C W+F K A+G ERYKARLV G +Q G+D +TFS V K TI
Sbjct: 971 ITTLPKGKKAVGCKWVFTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTI 1030
Query: 966 RTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNH----PDYVCRLRKSL 1021
+ +L ++ S+ W + QLDV NAFL+G+L E ++M P G+ + + V RL++S+
Sbjct: 1031 KLLLKVSASKKWFLKQLDVSNAFLNGELEEEIFMKIPEGYAERKGIVLPSNVVLRLKRSI 1090
Query: 1022 YGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSSHDL 1081
YGLKQA R W++ F+ + ++GF+ + DH+LF+ + +L+YVDDI++ S+S
Sbjct: 1091 YGLKQASRQWFKKFSSSLLSLGFKKTHGDHTLFLKMYDGEFVIVLVYVDDIVIASTSEAA 1150
Query: 1082 RKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAGMASCHPSAT 1141
+ L F ++DLG L YFLG+ V R G+ + Q YA +++ GM +C P +
Sbjct: 1151 AAQLTEELDQRFKLRDLGDLKYFLGLEVARTTAGISICQRKYALELLQSTGMLACKPVSV 1210
Query: 1142 PVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQVCLHMHAPRTEHMLA 1201
P+ K+ G +D YR +VG L YLT TRPDI++AV ++C APRT H+ A
Sbjct: 1211 PMIPNLKMRKDDGDLIEDIEQYRRIVGKLMYLTITRPDITFAVNKLCQFSSAPRTTHLTA 1270
Query: 1202 LKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSS 1261
R+L+Y++GT+ GL S L + D+DW C D+RRST+ + +F+GD+LISW S
Sbjct: 1271 AYRVLQYIKGTVGQGLFYSASSDLTLKGFADSDWASCQDSRRSTTSFTMFVGDSLISWRS 1330
Query: 1262 KRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPV 1321
K+Q T+SRSSAEAEYR +A E WL LL+ L ++Y D+ +AIY++ NPV
Sbjct: 1331 KKQHTVSRSSAEAEYRALALATCEMVWLFTLLVSLQASPP-VPILYSDSTAAIYIATNPV 1389
Query: 1322 QHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
H+RTKHI++D H VRE++ G+ ++LHV + Q+ADI TK L F+ +S +S+
Sbjct: 1390 FHERTKHIKLDCHTVRERLDNGELKLLHVRTEDQVADILTKPLFPYQFEHLKSKMSI 1446
>At1g57640
Length = 1444
Score = 590 bits (1520), Expect = e-168
Identities = 373/1052 (35%), Positives = 539/1052 (50%), Gaps = 89/1052 (8%)
Query: 364 GNLTSYSNLSHLNQKLYVGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLIS 423
GNL S++ ++ L + + + S T L S+ L L V + ++ +LIS
Sbjct: 410 GNLELLSDMRSMSPVLIILADGNKRVAVSEGTVRLGSH----LILKSVFYVKELESDLIS 465
Query: 424 VRQLTTDNNVSVCFDPYGFSVIDFQTGIPLMRCNSPGDLYPVTPSFPFAGLAQSLWHSRL 483
V Q+ +N+ C + ++ + PF LWH RL
Sbjct: 466 VGQMMDENH-----------------------CVNAAAVHTSVKA-PF-----DLWHRRL 496
Query: 484 GHPSSSALQSLRSNKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW 543
GH S + L S + + + VC++C+ K R F S+N ++ F ++H D+W
Sbjct: 497 GHASDKIVNLLPRELLSSGKEILEN-VCDTCMRAKQTRDTFPLSDNRSMDSFQLIHCDVW 555
Query: 544 TS-PVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQC 602
S +G R+++ +DD++ +W + +++KS+ + + F IK ++
Sbjct: 556 GPYRAPSYSGARYFLTIVDDYSRGVWVYLMTDKSETQKHLKDFIALVERQFDTEIKIVRS 615
Query: 603 DNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSF 662
DNG EF +Y G+ SC T QNG+ ERK R I N+ R L + +P F
Sbjct: 616 DNGTEF--LCMREYFLHKGIAHETSCVGTPHQNGRVERKHRHILNIARALRFQSYLPIQF 673
Query: 663 WHHALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQP 722
W + A YL+N P L SP ++LY+ P Y+HLRVFG LCY + +K
Sbjct: 674 WGECILSAAYLINRTPSMLLQGKSPYEMLYKTAPKYSHLRVFGSLCYAHNQNHKGDKFAA 733
Query: 723 RSTPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFANLTPTPSSTY------- 775
RS CVF+GYP +G++ FDL +K +SR VIF ET+FP++ ++
Sbjct: 734 RSRRCVFVGYPHGQKGWRLFDLEEQKFFVSRDVIFQETEFPYSKMSCNEEDERVLVDCVG 793
Query: 776 -EWLSDDIHPSVI--HRWTTQTPSPDLQPTPVAPSAT--ATPPTSTASSSSPSDPSPSSS 830
++ + I P I T P++ P+ P ++ P+ S SS DP +SS
Sbjct: 794 PPFIEEAIGPRTIIGRNIGEATVGPNVATGPIIPEINQESSSPSEFVSLSS-LDPFLASS 852
Query: 831 TPQSPPQP---APPVRTMATRSMRGIYKPRKLFN-LSVTIDDPTISPLPKNPKL------ 880
T Q+ P P RS R KP KL N ++ T+ +ISP + L
Sbjct: 853 TVQTADLPLSSTTPAPIQLRRSSRQTQKPMKLKNFVTNTVSVESISPEASSSSLYPIEKY 912
Query: 881 ----------------------------ALSDPNWKSAMQSEFDALIRNNTWDLVPRPCD 912
A+ D W+ AM +E ++L N T+ +V P
Sbjct: 913 VDCHRFTSSHKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQTFSIVNLPPG 972
Query: 913 VNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIA 972
+ W+++ K +++G ERYKARLV G Q GVD DETF+ V K +T+R L +A
Sbjct: 973 KRALGNKWVYKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKMSTVRLFLGVA 1032
Query: 973 LSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWY 1032
+R W +HQ+DV NAFLHGDL E VYM P GF+ + P VCRL KSLYGLKQAPR W+
Sbjct: 1033 AARDWHVHQMDVHNAFLHGDLKEEVYMKLPQGFQCDD-PSKVCRLHKSLYGLKQAPRCWF 1091
Query: 1033 QCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASE 1092
+ + GF S SD+SLF Y ++L+YVDD+I+ S D + L S
Sbjct: 1092 SKLSSALKQYGFTQSLSDYSLFSYNNDGIFVHVLVYVDDLIISGSCPDAVAQFKSYLESC 1151
Query: 1093 FAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTS 1152
F MKDLG L YFLGI V+R+A G +LSQ Y DII+ G+ PSA P++ KLS S
Sbjct: 1152 FHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLGARPSAFPLEQNHKLSLS 1211
Query: 1153 AGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGT 1212
D + YR LVG L YL TRP++SY+V + M PR +H A R++RY++
Sbjct: 1212 TSPLLSDSSRYRRLVGRLIYLVVTRPELSYSVHTLAQFMQNPRQDHWNAAIRVVRYLKSN 1271
Query: 1213 LQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSA 1272
G+ L + ++ + D+D+ C TRRS +GY V LGD ISW +K+QPT+SRSSA
Sbjct: 1272 PGQGILLSSTSTLQINGWCDSDYAACPLTRRSLTGYFVQLGDTPISWKTKKQPTVSRSSA 1331
Query: 1273 EAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMD 1332
EAEYR +A + E WL+ +L +L A ++ D+ SAI LS NPVQH+RTKH+E+D
Sbjct: 1332 EAEYRAMAFLTQELMWLKRVLYDLGVSHVQAMRIFSDSKSAIALSVNPVQHERTKHVEVD 1391
Query: 1333 IHFVREKVARGQARVLHVPSRHQIADIFTKGL 1364
HF+R+ + G VPS Q+ADI TK L
Sbjct: 1392 CHFIRDAILDGIIATSFVPSHKQLADILTKAL 1423
>At2g16670 putative retroelement pol polyprotein
Length = 1333
Score = 586 bits (1511), Expect = e-167
Identities = 347/944 (36%), Positives = 510/944 (53%), Gaps = 51/944 (5%)
Query: 478 LWHSRLGHPSSSALQSL-RSNKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFD 536
LWHSR+GHP++ + + S+ +S HLN + C+ C K R F S N T+ F+
Sbjct: 391 LWHSRMGHPAARVVSLIPESSVSVSSTHLNKA--CDVCHRAKQTRNSFPLSINKTLRIFE 448
Query: 537 ILHSDLWTS-PVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQV---FEMFISLSNQIRTH 592
+++ DLW S G R+++ +DD++ +W + L++KS+ + F +++++
Sbjct: 449 LIYCDLWGPYRTPSHTGARYFLTIIDDYSRGVWLYLLNDKSEAPCHLKNFFAMTDR---Q 505
Query: 593 FSQTIKCLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTL 652
F+ IK ++ DNG EF + G+I SC T +N + ERK R + N+ R L
Sbjct: 506 FNVKIKTVRSDNGTEF--LCLTKFFQEQGVIHERSCVATPERNDRVERKHRHLLNVARAL 563
Query: 653 LAHASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLV 712
A++P FW + A YL+N P L++ +P + L+++ P + HLRVFG LCY
Sbjct: 564 RFQANLPIQFWGECVLTAAYLINRTPSSVLNDSTPYERLHKKQPRFDHLRVFGSLCYAHN 623
Query: 713 PSSTINKLQPRSTPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFAN------ 766
+ +K RS CVF+GYP +G++ FDL + +SR V+F E +FPF
Sbjct: 624 RNRGGDKFAERSRRCVFVGYPHGQKGWRLFDLEQNEFFVSRDVVFSELEFPFRISHEQNV 683
Query: 767 LTPTPSSTYEWLSDDIHPSVIHRWTTQTPSPDL-QPTPVAPSATA--------------- 810
+ + + + D + +H P+P + +P++PSAT+
Sbjct: 684 IEEEEEALWAPIVDGLIEEEVHLGQNAGPTPPICVSSPISPSATSSRSEHSTSSPLDTEV 743
Query: 811 --TPPTSTASSSSPSDPS-----------PSSSTPQSPPQPAPPVRTMATRSMRGIYKPR 857
TP TST S+SSPS P+ P+++ +PP PP R +
Sbjct: 744 VPTPATSTTSASSPSSPTNLQFLPLSRAKPTTAQAVAPPAVPPPRRQSTRNKAPPVTLKD 803
Query: 858 KLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEF---DALIRNNTWDLVPRPCDVN 914
+ N +V + P+ D SA + + DA N+TW + P
Sbjct: 804 FVVNTTVCQESPSKLNSILYQLQKRDDTRRFSASHTTYVAIDAQEENHTWTIEDLPPGKR 863
Query: 915 IIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALS 974
I W+++ K ++G ERYKARLV G Q G D ETF+ V K AT+R L +A+
Sbjct: 864 AIGSQWVYKVKHNSDGSVERYKARLVALGNKQKEGEDYGETFAPVAKMATVRLFLDVAVK 923
Query: 975 RSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQC 1034
R+W IHQ+DV NAFLHGDL E VYM P GF + +HP+ VCRLRK+LYGLKQAPR W++
Sbjct: 924 RNWEIHQMDVHNAFLHGDLREEVYMKLPPGF-EASHPNKVCRLRKALYGLKQAPRCWFEK 982
Query: 1035 FADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFA 1094
+ GFQ S +D+SLF +GS +L+YVDD+I+ +S + LAS F
Sbjct: 983 LTTALKRYGFQQSLADYSLFTLVKGSVRIKILIYVDDLIITGNSQRATQQFKEYLASCFH 1042
Query: 1095 MKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTSAG 1154
MKDLGPL YFLGI V R G+++ Q YA DII+ G+ P+ P++ KL S
Sbjct: 1043 MKDLGPLKYFLGIEVARSTTGIYICQRKYALDIISETGLLGVKPANFPLEQNHKLGLSTS 1102
Query: 1155 TPCDDPTLYRSLVGALQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQ 1214
DP YR LVG L YL TR D++++V + M PR +H A R++RY++
Sbjct: 1103 PLLTDPQRYRRLVGRLIYLAVTRLDLAFSVHILARFMQEPREDHWAAALRVVRYLKADPG 1162
Query: 1215 LGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEA 1274
G+ L S ++ + D+DW G +RRS +GY V GD+ ISW +K+Q T+S+SSAEA
Sbjct: 1163 QGVFLRRSGDFQITGWCDSDWAGDPMSRRSVTGYFVQFGDSPISWKTKKQDTVSKSSAEA 1222
Query: 1275 EYRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIH 1334
EYR ++ + SE WL+ LL L ++ CD+ SAIY++ NPV H+RTKHIE+D H
Sbjct: 1223 EYRAMSFLASELLWLKQLLFSLGVSHVQPMIMCCDSKSAIYIATNPVFHERTKHIEIDYH 1282
Query: 1335 FVREKVARGQARVLHVPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
FVR++ +G HV + Q+ADIFTK L R F FR L +
Sbjct: 1283 FVRDEFVKGVITPRHVGTTSQLADIFTKPLGRDCFSAFRIKLGI 1326
>At3g61330 copia-type polyprotein
Length = 1352
Score = 582 bits (1499), Expect = e-166
Identities = 329/911 (36%), Positives = 492/911 (53%), Gaps = 46/911 (5%)
Query: 478 LWHSRLGHPSSSALQSLRSNKFISYEHLNSSP--VCESCVFGKHVRLPFVS-SNNVTVMP 534
LWH R GH + L+ L + + + P VCE C+ GK ++ F S++ P
Sbjct: 467 LWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGKQFKMSFPKESSSRAQKP 526
Query: 535 FDILHSDLWTSPVLSSAGH-RFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHF 593
+++H+D+ S G +++LF+DDF+ W + L KS+VFE+F +
Sbjct: 527 LELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSEVFEIFKKFKAHVEKES 586
Query: 594 SQTIKCLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLL 653
IK ++ D G EF +K F YC NG+ + + P + QNG ERK RTI M R++L
Sbjct: 587 GLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGVVERKNRTILEMARSML 646
Query: 654 AHASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVP 713
+P W A+ A YLLN P K++S +P + R P +HLRVFG + + VP
Sbjct: 647 KSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKPGVSHLRVFGSIAHAHVP 706
Query: 714 SSTINKLQPRSTPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFANLTPTPSS 773
+KL +S +F+GY + +GYK ++ +K IISR+++FDE
Sbjct: 707 DEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVFDE------------EG 754
Query: 774 TYEWLSDDIHPSVIHRWTTQTPSPDLQPTPVAPSATATPPTSTASSSSPSDPSPSSSTPQ 833
++W S++ + + P P + P TPPTS SS + S S TP+
Sbjct: 755 EWDWNSNEEDYNFFPHFEEDEPEPTREEPP--SEEPTTPPTSPTSSQI--EESSSERTPR 810
Query: 834 SPPQPAPPVRTMATRSMRGIYKPRK------LFNLSVTIDDPTISPLPKNPKLALSDPNW 887
RS++ +Y+ + LF L + P + + A+ W
Sbjct: 811 F-------------RSIQELYEVTENQENLTLFCLFAECE-------PMDFQKAIEKKTW 850
Query: 888 KSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQI 947
++AM E ++ +N+TW+L P I W+++ K + G ERYKARLV G SQ
Sbjct: 851 RNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQR 910
Query: 948 AGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRD 1007
G+D DE F+ V + T+R ++++A W IHQ+DV++AFL+GDL E VY+ QP G+
Sbjct: 911 VGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIV 970
Query: 1008 PNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLL 1067
D V RL+K LYGLKQAPRAW Y F +H+L+I + D+ L
Sbjct: 971 KGEEDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACL 1030
Query: 1068 YVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDI 1127
YVDD+I ++ + + + EF M D+G +SY+LGI V + G+F++Q YA+++
Sbjct: 1031 YVDDLIFTGNNPSIFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEV 1090
Query: 1128 IARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQV 1187
+ + + +P TP++ KLS DPT ++SLVG+L+YLT TRPDI YAV V
Sbjct: 1091 LKKFKIDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVV 1150
Query: 1188 CLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSG 1247
+M P T H A KRILRY++GT+ GLH + KL+ Y+D+DWGG +D R+STSG
Sbjct: 1151 SRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSG 1210
Query: 1248 YCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVY 1307
+ ++GD +W SK+QP ++ S+ EAEY + V + WLRNLL EL P T ++
Sbjct: 1211 FVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIF 1270
Query: 1308 CDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGLPRV 1367
DN SAI L+ NPV H R+KHI+ H++RE V++ ++ +V + Q+AD FTK L R
Sbjct: 1271 VDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADFFTKPLKRE 1330
Query: 1368 LFDDFRSSLSV 1378
F RS L V
Sbjct: 1331 NFIKMRSLLGV 1341
>At1g48710 hypothetical protein
Length = 1352
Score = 578 bits (1491), Expect = e-165
Identities = 330/911 (36%), Positives = 492/911 (53%), Gaps = 46/911 (5%)
Query: 478 LWHSRLGHPSSSALQSLRSNKFISYEHLNSSP--VCESCVFGKHVRLPFVS-SNNVTVMP 534
LWH R GH + L+ L + + + P VCE C+ GK ++ F S++
Sbjct: 467 LWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGKQFKMSFPKESSSRAQKS 526
Query: 535 FDILHSDLWTSPVLSSAGH-RFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHF 593
+++H+D+ S G +++LF+DDF+ W + L KS+VFE+F +
Sbjct: 527 LELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSEVFEIFKKFKAHVEKES 586
Query: 594 SQTIKCLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLL 653
IK ++ D G EF +K F YC NG+ + + P + QNG AERK RTI M R++L
Sbjct: 587 GLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGVAERKNRTILEMARSML 646
Query: 654 AHASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVP 713
+P W A+ A YLLN P K++S +P + R +HLRVFG + + VP
Sbjct: 647 KSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKSGVSHLRVFGSIAHAHVP 706
Query: 714 SSTINKLQPRSTPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFANLTPTPSS 773
+KL +S +F+GY + +GYK ++ +K IISR+++FDE
Sbjct: 707 DEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVFDE------------EG 754
Query: 774 TYEWLSDDIHPSVIHRWTTQTPSPDLQPTPVAPSATATPPTSTASSSSPSDPSPSSSTPQ 833
++W S++ + + P P + P TPPTS SS + S S TP+
Sbjct: 755 EWDWNSNEEDYNFFPHFEEDEPEPTREEPP--SEEPTTPPTSPTSSQI--EESSSERTPR 810
Query: 834 SPPQPAPPVRTMATRSMRGIYKPRK------LFNLSVTIDDPTISPLPKNPKLALSDPNW 887
RS++ +Y+ + LF L + P + + A+ W
Sbjct: 811 F-------------RSIQELYEVTENQENLTLFCLFAECE-------PMDFQEAIEKKTW 850
Query: 888 KSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQI 947
++AM E ++ +N+TW+L P I W+++ K + G ERYKARLV G Q
Sbjct: 851 RNAMDEEIKSIQKNDTWELTSLPNGHKTIGVKWVYKAKKNSKGEVERYKARLVAKGYIQR 910
Query: 948 AGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRD 1007
AG+D DE F+ V + T+R ++++A W IHQ+DV++AFL+GDL E VY+ QP G+
Sbjct: 911 AGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIV 970
Query: 1008 PNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLL 1067
D V RL+K+LYGLKQAPRAW Y F +H+L+I + D+ L
Sbjct: 971 KGEEDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACL 1030
Query: 1068 YVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDI 1127
YVDD+I ++ + + + EF M D+G +SY+LGI V + G+F++Q YA+++
Sbjct: 1031 YVDDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEV 1090
Query: 1128 IARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQV 1187
+ + M +P TP++ KLS DPT ++SLVG+L+YLT TRPDI YAV V
Sbjct: 1091 LKKFKMDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVV 1150
Query: 1188 CLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSG 1247
+M P T H A KRILRY++GT+ GLH + KL+ Y+D+DWGG +D R+STSG
Sbjct: 1151 SRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSG 1210
Query: 1248 YCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVY 1307
+ ++GD +W SK+QP + S+ EAEY + V + WLRNLL EL P T ++
Sbjct: 1211 FVFYIGDTAFTWMSKKQPIVVLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIF 1270
Query: 1308 CDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGLPRV 1367
DN SAI L+ NPV H R+KHI+ H++RE V++ ++ +V + Q+ADIFTK L R
Sbjct: 1271 VDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADIFTKPLKRE 1330
Query: 1368 LFDDFRSSLSV 1378
F RS L V
Sbjct: 1331 DFIKMRSLLGV 1341
>At1g26990 polyprotein, putative
Length = 1436
Score = 536 bits (1381), Expect = e-152
Identities = 341/1052 (32%), Positives = 520/1052 (49%), Gaps = 85/1052 (8%)
Query: 362 APGNLTSYSNLSHLNQKL-----YVGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQ 416
A ++T NL H + L + +G + I+G+G L T LSL +VL P+
Sbjct: 427 ASHHVTHERNLYHTYKALDRTFVRLPNGHTVKIEGTGFIQL-----TDALSLHNVLFIPE 481
Query: 417 IVKNLISVRQLTTDNNVSVCFDPYGFSVIDFQTGIPLMRCNSPGDLYPVT--------PS 468
NL+SV LT V F + + L + + G+LY + S
Sbjct: 482 FKFNLLSVSVLTKTLQSKVSFTSDECMIQALTKELMLGKGSQVGNLYILNLDKSLVDVSS 541
Query: 469 FPFAGLAQS------LWHSRLGHPSSSALQSLRSNKFISYEHLNS-SPVCESCVFGKHVR 521
FP + S +WH RLGHPS + + +L + + +N S C C K
Sbjct: 542 FPGKSVCSSVKNESEMWHKRLGHPSFAKIDTLSDVLMLPKQKINKDSSHCHVCHLSKQKH 601
Query: 522 LPFVSSNNVTVMPFDILHSDLW---TSPVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQV 578
LPF S N++ F+++H D W + P + S +R+++ +DDF+ W + L KS V
Sbjct: 602 LPFKSVNHIREKAFELVHIDTWGPFSVPTVDS--YRYFLTIVDDFSRATWIYLLKQKSDV 659
Query: 579 FEMFISLSNQIRTHFSQTIKCLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKA 638
+F S + T + + ++ DN E F++ A G+ CP T QN
Sbjct: 660 LTVFPSFLKMVETQYHTKVCSVRSDNAHEL---KFNELFAKEGIKADHPCPETPEQNFVV 716
Query: 639 ERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSY 698
ERK + + N+ R L+ + +P +W + A +L+N + ++N +P + L + P Y
Sbjct: 717 ERKHQHLLNVARALMFQSGIPLEYWGDCVLTAVFLINRLLSPVINNETPYERLTKGKPDY 776
Query: 699 THLRVFGCLCYPLVPSSTINKLQPRSTPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFD 758
+ L+ FGCLCY + K PR+ C+FLGYP+ ++GYK D+ V ISRHVIF
Sbjct: 777 SSLKAFGCLCYCSTSPKSRTKFDPRAKACIFLGYPMGYKGYKLLDIETYSVSISRHVIFY 836
Query: 759 ETQFPFANLTPTPSSTYEWLSDDIHPSVIHRWTTQTPSPDL-QPTPVAPSATATPPTSTA 817
E FPFA+ T ++ D P + P+P+ + P+ S++ P
Sbjct: 837 EDIFPFASSNITDAAK------DFFPHIY------LPAPNNDEHLPLVQSSSDAPHNHDE 884
Query: 818 SSSSPSDPSPSSSTPQ----SPPQPAPPVRTMATRSMRGIYKPRKLFNLSVT-------I 866
SSS PS ST Q S Q T + Y + S I
Sbjct: 885 SSSMIFVPSEPKSTRQRKLPSHLQDFHCYNNTPTTTKTSPYPLTNYISYSYLSEPFGAFI 944
Query: 867 DDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKT 926
+ T + LP+ A D W AM E A +R TW + P + C WI K
Sbjct: 945 NIITATKLPQKYSEARLDKVWNDAMGKEISAFVRTGTWSICDLPAGKVAVGCKWIITIKF 1004
Query: 927 KANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQN 986
A+G ER+KARLV G +Q G+D TFS V K T++ +L++A W +HQLD+ N
Sbjct: 1005 LADGSIERHKARLVAKGYTQQEGIDFFNTFSPVAKMVTVKVLLSLAPKMKWYLHQLDISN 1064
Query: 987 AFLHGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQH 1046
A L+GDL E +YM P G+ + + G + +P A +C D
Sbjct: 1065 ALLNGDLEEEIYMKLPPGYSE-------------IQGQEVSPNA--KCHGD--------- 1100
Query: 1047 STSDHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLG 1106
H+LF+ + +L+YVDDI++ S++ + + L+S F ++DLG +FLG
Sbjct: 1101 ----HTLFVKAQDGFFLVVLVYVDDILIASTTEAASAELTSQLSSFFQLRDLGEPKFFLG 1156
Query: 1107 IAVTRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSL 1166
I + R+A G+ L Q Y D++A + + C PS+ P++ QKLS GT +D YR +
Sbjct: 1157 IEIARNADGISLCQRKYVLDLLASSDFSDCKPSSIPMEPNQKLSKDTGTLLEDGKQYRRI 1216
Query: 1167 VGALQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEK 1226
+G LQYL TRPDI++AV ++ + AP H+ AL +ILRY++GT+ GL
Sbjct: 1217 LGKLQYLCLTRPDINFAVSKLAQYSSAPTDIHLQALHKILRYLKGTIGQGLFYGADTNFD 1276
Query: 1227 LISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSES 1286
L ++D+DW C DTRR +G+ +F+G++L+SW SK+Q +S SSAEAEYR ++ E
Sbjct: 1277 LRGFSDSDWQTCPDTRRCVTGFAIFVGNSLVSWRSKKQDVVSMSSAEAEYRAMSVATKEL 1336
Query: 1287 CWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQAR 1346
WL +L P + +YCDN +A++++ N V H+RTKHIE D H VRE + G +
Sbjct: 1337 IWLGYILTAFKIPFTHPAYLYCDNEAALHIANNSVFHERTKHIENDCHKVRECIEAGILK 1396
Query: 1347 VLHVPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
+ V + +Q+AD TK L F + S L +
Sbjct: 1397 TIFVRTDNQLADTLTKPLYPKPFRENNSKLGL 1428
>At3g60170 putative protein
Length = 1339
Score = 534 bits (1375), Expect = e-151
Identities = 336/1034 (32%), Positives = 526/1034 (50%), Gaps = 57/1034 (5%)
Query: 376 NQKLYVGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSV 435
N+ + +G+ + + G G + + T+ + V + P++ NL+S+ QL +
Sbjct: 325 NRTVKLGNDTRMSVVGKGSVKVKVNGVTQVIP--EVYYVPELRNNLLSLGQLQERGLAIL 382
Query: 436 CFDPYGFSVIDFQTGIPLMRCNSPGD-LYPVTPSFP-----------FAGLAQSLWHSRL 483
D G + + +M N G+ ++ + S P LWH R
Sbjct: 383 IRD--GTCKVYHPSKGAIMETNMSGNRMFFLLASKPQKNSLCLQTEEVMDKENHLWHCRF 440
Query: 484 GHPSSSALQSLRSNKFISYEHL--NSSPVCESCVFGKHVRLPFVSSNN-VTVMPFDILHS 540
GH + L+ L K + + + +C C+ GK R + + ++HS
Sbjct: 441 GHLNQEGLKLLAHKKMVIGLPILKATKEICAICLTGKQHRESMSKKTSWKSSTQLQLVHS 500
Query: 541 DLWTSPV--LSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIK 598
D+ P+ +S +G R+ + F+DDFT W + L KS+ F F + +
Sbjct: 501 DI-CGPITPISHSGKRYILSFIDDFTRKTWVYFLHEKSEAFATFKIFKASVEKEIGAFLT 559
Query: 599 CLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASV 658
CL+ D G EF + F ++C ++G+ + + T QNG AERK RTI N +R++L+ V
Sbjct: 560 CLRTDRGGEFTSNEFGEFCRSHGISRQLTAAFTPQQNGVAERKNRTIMNAVRSMLSERQV 619
Query: 659 PPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTIN 718
P FW A + + ++ N P + ++P + R P + RVFGC+ Y +P +
Sbjct: 620 PKMFWSEATKWSVHIQNRSPTAAVEGMTPEEAWSGRKPVVEYFRVFGCIGYVHIPDQKRS 679
Query: 719 KLQPRSTPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFANLTPTPSSTYEWL 778
KL +S CVFLG + ++ +D +K++IS+ V+FDE + +++W
Sbjct: 680 KLDDKSKKCVFLGVSEESKAWRLYDPVMKKIVISKDVVFDEDK------------SWDWD 727
Query: 779 SDDIHPSVIHRWTTQTPSPDLQPTP--VAPSATATPPTSTASSSSPSDPSPSSSTPQSPP 836
D+ + T + D + V P A A+P + SD + SSS +P
Sbjct: 728 QADVEAKEV---TLECGDEDDEKNSEVVEPIAVASP------NHVGSDNNVSSSPILAPS 778
Query: 837 QPAP-PVRTMATRSMR-----GIYKPRK----LFNLSVTIDDPTISPLPKNPKLALSDPN 886
PAP PV TR R Y+ + NLSV + P A+ D
Sbjct: 779 SPAPSPVAAKVTRERRPPGWMADYETGEGEEIEENLSVMLLMMMTEADPIQFDDAVKDKI 838
Query: 887 WKSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQ 946
W+ AM+ E +++++NNTW+L P I W+++ K +G ++YKARLV G +Q
Sbjct: 839 WREAMEHEIESIVKNNTWELTTLPKGFTPIGVKWVYKTKLNEDGEVDKYKARLVAKGYAQ 898
Query: 947 IAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFR 1006
G+D E F+ V + T+RT+L I+ +W I QLDV++AFLHG+L E VY+ QP GF
Sbjct: 899 CYGIDYTEVFAPVARLDTVRTILAISSQFNWEIFQLDVKSAFLHGELKEEVYVRQPEGFI 958
Query: 1007 DPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLL 1066
+ V +LRK+LYGLKQAPRAWY Y F+ S+H+LF R ++ +
Sbjct: 959 REGEEEKVYKLRKALYGLKQAPRAWYSRIEAYFLKEEFERCPSEHTLFTKTRVGNILIVS 1018
Query: 1067 LYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARD 1126
LYVDD+I S + + EF M DLG + +FLGI V + GG+F+ Q YAR+
Sbjct: 1019 LYVDDLIFTGSDKAMCDEFKKSMMLEFEMSDLGKMKHFLGIEVKQSDGGIFICQRRYARE 1078
Query: 1127 IIARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQ 1186
++AR GM + P+ KL+ D T+++ LVG+L YLT TRPD+ Y V
Sbjct: 1079 VLARFGMDESNAVKNPIVPGTKLTKDENGEKVDETMFKQLVGSLMYLTVTRPDLMYGVCL 1138
Query: 1187 VCLHMHAPRTEHMLALKRILRYVQGTLQLGL--HLYPSPIEKLISYTDADWGGCLDTRRS 1244
+ M PR H LA KRILRY++GT++LG+ + KL+++TD+D+ G L+ RRS
Sbjct: 1139 ISRFMSNPRMSHWLAAKRILRYLKGTVELGIFYRRRKNRSLKLMAFTDSDYAGDLNDRRS 1198
Query: 1245 TSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELHFPLS*AT 1304
TSG+ + I W+SK+QP ++ S+ EAEY A + WLR +L +L AT
Sbjct: 1199 TSGFVFLMASGAICWASKKQPVVALSTTEAEYIAAAFCACQCVWLRKVLEKLGAEEKSAT 1258
Query: 1305 LVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGL 1364
++ CDN S I LS +PV H ++KHIE+ H++R+ V ++ + P+ Q+ADIFTK L
Sbjct: 1259 VINCDNSSTIQLSKHPVLHGKSKHIEVRFHYLRDLVNGDVVKLEYCPTEDQVADIFTKPL 1318
Query: 1365 PRVLFDDFRSSLSV 1378
F+ R+ L +
Sbjct: 1319 KLEQFEKLRALLGM 1332
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 527 bits (1358), Expect = e-149
Identities = 289/629 (45%), Positives = 385/629 (60%), Gaps = 22/629 (3%)
Query: 769 PTPSST-YEWLSDDIHPSVIHRWTTQTPSPDL----QPTPVAPSATATPPTSTASSSSPS 823
P P++ ++ + + + +++ +PSP+ P P +P ++ PT + S S P+
Sbjct: 818 PQPTAQPHQTQNSNSNSPILNNPNPNSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPN 877
Query: 824 DPSPSS-STPQSPPQ-PAPPV-----------RTMATRSMRGIYKPRKLFNLSVTIDDPT 870
PS SS STP PP PAPP+ +MATR+ GI KP + ++ + ++
Sbjct: 878 SPSSSSTSTPPLPPVLPAPPIIQVNAQAPVNTHSMATRAKDGIRKPNQKYSYATSL---A 934
Query: 871 ISPLPKNPKLALSDPNWKSAMQSEFDALIRNNTWDLVPRPC-DVNIIRCMWIFRHKTKAN 929
+ P+ A+ D W+ AM SE +A I N+TWDLVP P V I+ C WIF K ++
Sbjct: 935 ANSEPRTAIQAMKDDRWRQAMGSEINAQIGNHTWDLVPPPPPSVTIVGCRWIFTKKFNSD 994
Query: 930 GCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFL 989
G RYKARLV G +Q G+D ETFS V+K +IR VL +A+ RSWPI QLDV NAFL
Sbjct: 995 GSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFL 1054
Query: 990 HGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTS 1049
G L + VYM QP GF D + PDYVCRLRK++YGLKQAPRAWY Y+ T+GF +S S
Sbjct: 1055 QGTLTDEVYMSQPPGFVDKDRPDYVCRLRKAIYGLKQAPRAWYVELRTYLLTVGFVNSIS 1114
Query: 1050 DHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAV 1109
D SLF+ +RG + Y+L+YVDDI++ + L K + L+ F++K+ L YFLGI
Sbjct: 1115 DTSLFVLQRGRSIIYMLVYVDDILITGNDTVLLKHTLDALSQRFSVKEHEDLHYFLGIEA 1174
Query: 1110 TRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGA 1169
R GL LSQ Y D++AR M + P ATP+ T KL+ +GT DPT YR +VG+
Sbjct: 1175 KRVPQGLHLSQRRYTLDLLARTNMLTAKPVATPMATSPKLTLHSGTKLPDPTEYRGIVGS 1234
Query: 1170 LQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLIS 1229
LQYL FTRPD+SYAV ++ +MH P +H ALKR+LRY+ GT G+ L L +
Sbjct: 1235 LQYLAFTRPDLSYAVNRLSQYMHMPTDDHWNALKRVLRYLAGTPDHGIFLKKGNTLSLHA 1294
Query: 1230 YTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWL 1289
Y+DADW G D ST+GY V+LG + ISWSSK+Q + RSS EAEYR VAN SE W+
Sbjct: 1295 YSDADWAGDTDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSELQWI 1354
Query: 1290 RNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLH 1349
+LL EL LS ++YCDNV A YL NPV H R KHI +D HF+R +V G RV+H
Sbjct: 1355 CSLLTELGIQLSHPPVIYCDNVGATYLCANPVFHSRMKHIALDYHFIRNQVQSGALRVVH 1414
Query: 1350 VPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
V + Q+AD TK L RV F +F + V
Sbjct: 1415 VSTHDQLADTLTKPLSRVAFQNFSRKIGV 1443
Score = 303 bits (777), Expect = 3e-82
Identities = 203/547 (37%), Positives = 276/547 (50%), Gaps = 43/547 (7%)
Query: 381 VGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPY 440
+ G IPI +G +L TS ++ L L+ VL+ P I KNLISV +L N VSV F P
Sbjct: 341 IADGSTIPITHTGSASLPTS--SRSLDLNKVLYVPNIHKNLISVYRLCNTNRVSVEFFPA 398
Query: 441 GFSVIDFQTGIPLMRCNSPGDLY--PVTPS-------FPFAGLAQSLWHSRLGHPSSSAL 491
F V D TG+PL++ + +LY P+ S P + S WHSRLGHPS + L
Sbjct: 399 SFQVKDLNTGVPLLQGKTKDELYEWPIASSQAVSMFASPCSKATHSSWHSRLGHPSLAIL 458
Query: 492 QSLRSNKFISYEHLNSSPV---CESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVL 548
S+ SN S LN S C C K ++PF +S + P + ++SD+W+SP+L
Sbjct: 459 NSVISNH--SLPVLNPSHKLLSCSDCFINKSHKVPFSNSTITSSKPLEYIYSDVWSSPIL 516
Query: 549 SSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNGREF 608
S +R+YV+F+D FT + W +PL KSQV + FI + + F I L DNG EF
Sbjct: 517 SIDNYRYYVIFVDHFTRYTWLYPLKQKSQVKDTFIIFKSLVENRFQTRIGTLYSDNGGEF 576
Query: 609 DNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQ 668
DY + +G+ S PHT NG +ERK R I M TLL+HASVP ++W +A
Sbjct: 577 --VVLRDYLSQHGISHFTSPPHTPEHNGLSERKHRHIVEMGLTLLSHASVPKTYWPYAFS 634
Query: 669 MATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCV 728
+A YL+N +P L SP Q L+ + P+Y L+VFGC CYP + +KL+ +S C
Sbjct: 635 VAVYLINRLPTPLLQLQSPFQKLFGQPPNYEKLKVFGCACYPWLRPYNRHKLEDKSKQCA 694
Query: 729 FLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFANLTPTPSSTYEWLSDDIHPSVIH 788
F+GY L Y C + ++ SRHV FDE FPF+ S++ E SD H
Sbjct: 695 FMGYSLTQSAYLCLHIPTGRLYTSRHVQFDERCFPFSTTNFGVSTSQEQRSDSAPNWPSH 754
Query: 789 RWTTQTP---------SPDLQPTPVAPSATATPPTSTASSS---SPSDPSPSSSTPQSP- 835
TP P L +P PS+ + T+ SSS S S SPSSS P +P
Sbjct: 755 TTLPTTPLVLPAPPCLGPHLDTSPRPPSSPSPLCTTQVSSSNLPSSSISSPSSSEPTAPS 814
Query: 836 ---PQP-APPVRTMATRSMRGIY--------KPRKLFNLSVTIDDPTISPLPKNPKLALS 883
PQP A P +T + S I P S P SP P ++S
Sbjct: 815 HNGPQPTAQPHQTQNSNSNSPILNNPNPNSPSPNSPNQNSPLPQSPISSPHIPTPSTSIS 874
Query: 884 DPNWKSA 890
+PN S+
Sbjct: 875 EPNSPSS 881
>At1g37110
Length = 1356
Score = 500 bits (1287), Expect = e-141
Identities = 333/1041 (31%), Positives = 517/1041 (48%), Gaps = 62/1041 (5%)
Query: 376 NQKLYVGSGQGIPIQGSGHTTLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSV 435
N + +G + QG G + T T + L +V + P + +NLIS L + +
Sbjct: 333 NTTILLGDDHSVESQGQGTIRIDTHGGTIKI-LENVKYVPHLRRNLISTGTL---DKLGY 388
Query: 436 CFDPYGFSVIDFQTGIPLMRCNSPGDLYPVTPSFPFAGLAQS--------LWHSRLGHPS 487
+ V F+ +R + LY + S + L + LWHSRLGH S
Sbjct: 389 RHEGGEGKVRYFKNNKTALRGSLSNGLYVLDGSTVMSELCNAETDKVKTALWHSRLGHMS 448
Query: 488 SSALQSLRSNKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPV 547
+ L+ L I + +N CE CV GK ++ F + + +H+DLW SP
Sbjct: 449 MNNLKVLAGKGLIDRKEINELEFCEHCVMGKSKKVSFNVGKHTSEDALSYVHADLWGSPN 508
Query: 548 L--SSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNG 605
+ S +G ++++ +DD T +W + L +K + F+ F + + ++ +KCL+ DNG
Sbjct: 509 VTPSISGKQYFLSIIDDKTRKVWLYFLKSKDETFDKFCEWKSLVENQVNKKVKCLRTDNG 568
Query: 606 REFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHH 665
EF N F YC +G+ +C +T QNG AER RTI +R LL + V FW
Sbjct: 569 LEFCNSRFDSYCKEHGIERHRTCTYTPQQNGVAERMNRTIMEKVRCLLNKSGVEEVFWAE 628
Query: 666 ALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRST 725
A A YL+N P +++ P ++ R P Y HLR FG + Y KL+PR+
Sbjct: 629 AAATAAYLINRSPASAINHNVPEEMWLNRKPGYKHLRKFGSIAY---VHQDQGKLKPRAL 685
Query: 726 PCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFANLTPTPSSTYEWLSDDIHPS 785
FLGYP +GYK + L K +ISR+V+F E+ + +L T D+++
Sbjct: 686 KGFFLGYPAGTKGYKVWLLEEEKCVISRNVVFQES-VVYRDLKVKEDDT-----DNLNQK 739
Query: 786 VIHRWTTQTPSPDLQPTPVAPSATATPPTSTASSSSPSDPSPSSSTPQ-----SPPQPAP 840
+T S +++ A ++ + S S P SS + S
Sbjct: 740 -------ETTSSEVEQNKFAEASGSGGVIQLQSDSEPITEGEQSSDSEEEVEYSEKTQET 792
Query: 841 PVRTMAT-------RSMRGIYKPRKL-----FNLSVTIDDPTISPLPKNPKLALSDPN-- 886
P RT T R R I P + ++ + + I P++ + A+ +
Sbjct: 793 PKRTGLTTYKLARDRVRRNINPPTRFTEESSVTFALVVVENCIVQEPQSYQEAMESQDCE 852
Query: 887 -WKSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCF-ERYKARLVGDGR 944
W A E D+L++N TWDLV +P D II C W+F+ K+ G R+KARLV G
Sbjct: 853 KWDMATHDEMDSLMKNGTWDLVDKPKDRKIIGCRWLFKLKSGIPGVEPTRFKARLVAKGY 912
Query: 945 SQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLG 1004
+Q GVD E F+ VVK +IR ++++ + + + Q+DV+ FLHGDL E +YM QP G
Sbjct: 913 TQREGVDYQEIFAPVVKHVSIRILMSLVVDKDLELEQMDVKTTFLHGDLEEELYMEQPEG 972
Query: 1005 FRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFI-YRRGSDMA 1063
F + + VCRL+KSLYGLKQ+PR W + F ++S+ F S D +++ + D
Sbjct: 973 FVSDSSENKVCRLKKSLYGLKQSPRQWNKRFDRFMSSQQFIRSEHDACVYVKHVSEHDFI 1032
Query: 1064 YLLLYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAV--TRHAGGLFLSQS 1121
YLLLYVDD+++ +S + L++EF MKD+G S LGI + R G L LSQ
Sbjct: 1033 YLLLYVDDMLIAGASKAEINRVKEQLSTEFEMKDMGGASRILGIDIYRDRKGGVLKLSQE 1092
Query: 1122 TYARDIIARAGMASCHPSATPVDTKQKL-STSAGTPCDDPTL--YRSLVGALQY-LTFTR 1177
Y R ++ R M+ + PV KL + C D + Y S VG++ Y + TR
Sbjct: 1093 IYIRKVLDRFNMSGAKMTNAPVGAHFKLAAVREEDECVDTDVVPYSSAVGSIMYAMLGTR 1152
Query: 1178 PDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGG 1237
PD++YA+ + +M P + H A+K ++RY++G L L + Y D+++
Sbjct: 1153 PDLAYAICLISRYMSKPGSMHWEAVKWVMRYLKGAQDLNLVFTKEKDFTVTGYCDSNYAA 1212
Query: 1238 CLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELH 1297
LD RRS SGY +G N +SW + QP ++ S+ EAEY +A E+ W++ LL ++
Sbjct: 1213 DLDRRRSISGYVFTIGGNTVSWKASLQPVVAMSTTEAEYIALAEAAKEAMWIKGLLQDMG 1272
Query: 1298 FPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIA 1357
++CD+ SAI LS N V H+RTKHI++ +++R+ V G VL + +
Sbjct: 1273 MQQD-KVKIWCDSQSAICLSKNSVYHERTKHIDVRFNYIRDVVESGDVDVLKIHTSRNPV 1331
Query: 1358 DIFTKGLPRVLFDDFRSSLSV 1378
D TK +P + F+S+L V
Sbjct: 1332 DALTKCIP---VNKFKSALGV 1349
>At2g07550 putative retroelement pol polyprotein
Length = 1356
Score = 498 bits (1281), Expect = e-140
Identities = 322/921 (34%), Positives = 474/921 (50%), Gaps = 55/921 (5%)
Query: 478 LWHSRLGHPSSSALQSLRSNKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFDI 537
LWH RL H S ++ L F+ + ++S VCE C++GK R F +++ T +
Sbjct: 440 LWHQRLCHMSQKNMEILVRKGFLDKKKVSSLDVCEDCIYGKAKRKSFSLAHHDTKEKLEY 499
Query: 538 LHSDLWTSPV--LSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQ 595
+HSDLW +P LS ++++ +DDFT +W + + K + FE F+ N + +
Sbjct: 500 IHSDLWGAPFVPLSLGKCQYFMSIIDDFTRKVWVYFMKTKDEAFEKFVEWVNLVENQTDR 559
Query: 596 TIKCLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAH 655
+K L+ DNG EF NK F +C + G+ +C +T QNG AER RTI +R++L+
Sbjct: 560 RVKTLRTDNGLEFCNKLFDGFCESIGIHRHRTCAYTPQQNGVAERMNRTIMEKVRSMLSD 619
Query: 656 ASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSS 715
+ +P FW A L+N P L+ P + P Y++LR +GC+ + +
Sbjct: 620 SGLPKRFWAEATHTTVLLINKTPSSALNFEIPDKKWSGNPPVYSYLRRYGCVAF---VHT 676
Query: 716 TINKLQPRSTPCVFLGYPLHHRGYKCFDLSHRKVIISRHVIFDETQFPFANL-------- 767
KL+PR+ V +GYP+ +GYK + L RK ++SR++IF E + +L
Sbjct: 677 DDGKLEPRAKKGVLIGYPVGVKGYKVWILDERKCVVSRNIIFQENAV-YKDLMQRQENVS 735
Query: 768 TPTPSSTYEWLSDDIHPSVIHRWTTQTPSPDLQPTPVAPSATATPPTSTASSSSPSDPSP 827
T T +L D+ R ++ T AP +P ST ++ +D
Sbjct: 736 TEEDDQTGSYLEFDLE---AERDVISGGDQEMVNTIPAPE---SPVVSTPTTQDTND-DE 788
Query: 828 SSSTPQSPPQPAPPVRTMATRSMRGIYKPRKLFN-------LSVTIDDPTISPLPKNPKL 880
S QSP + R R I PR+ + L T D + P+N +
Sbjct: 789 DSDVNQSPLS----YHLVRDRDKREIRAPRRFDDEDYYAEALYTTEDGEAVE--PENYRK 842
Query: 881 ALSDPN---WKSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCFE-RYK 936
A D N WK AM E D+ +NNTW +V RP + II C WIF++K G E R+K
Sbjct: 843 AKLDANFDKWKLAMDEEIDSQEKNNTWTIVTRPENQRIIGCRWIFKYKLGILGVEEPRFK 902
Query: 937 ARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHET 996
ARLV G +Q G+D E F+ VVK +IR +L+I + QLDV+ AFLHG+L E
Sbjct: 903 ARLVAKGYAQKEGIDYHEIFAPVVKHVSIRVLLSIVAQEDLELEQLDVKTAFLHGELKEK 962
Query: 997 VYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLF-- 1054
+YM P G+ + VC L K+LYGLKQAP+ W + F +++ I F S D +
Sbjct: 963 IYMSPPEGYESMFKANEVCLLNKALYGLKQAPKQWNEKFDNFMKEICFVKSAYDSCAYTK 1022
Query: 1055 IYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRH-- 1112
+ GS M YLL+YVDDI++ S + + ++ A L F MKDLG LG+ + R
Sbjct: 1023 VLPDGSVM-YLLIYVDDILVASKNKEAITALKANLGMRFEMKDLGAAKKILGMEIIRDRT 1081
Query: 1113 AGGLFLSQSTYARDIIARAGMASCHPSATPVD--------TKQKLSTSAGTPCDDPTLYR 1164
G L+LSQ Y I+ MA P+ TP+ T+QKL P Y
Sbjct: 1082 LGVLWLSQEGYLNKILETYNMAEAKPAMTPLGAHFKFQAATEQKLIRDEDFMKSVP--YS 1139
Query: 1165 SLVGALQY-LTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSP 1223
S VG++ Y + TRPD++Y V + M P EH L +K +LRY++GTL+ L S
Sbjct: 1140 SAVGSIMYAMLGTRPDLAYPVGIISRFMSQPIKEHWLGVKWVLRYIKGTLKTRLCYKKSS 1199
Query: 1224 IEKLISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVV 1283
++ Y DAD+ LD RRS +G LG N ISW S Q +++S+ E+EY + V
Sbjct: 1200 SFSIVGYCDADYAADLDKRRSITGLVFTLGGNTISWKSGLQRVVAQSTTESEYMSLTEAV 1259
Query: 1284 SESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARG 1343
E+ WL+ LL + + + ++CD+ SAI LS N V H+RTKHI++ HF+RE ++ G
Sbjct: 1260 KEAIWLKGLLKDFGYEQK-SVEIFCDSQSAIALSKNNVHHERTKHIDVKYHFIREIISDG 1318
Query: 1344 QARVLHVPSRHQIADIFTKGL 1364
VL + + ADIFTK L
Sbjct: 1319 TVEVLKISTEKNPADIFTKVL 1339
>At4g16870 retrotransposon like protein
Length = 1474
Score = 497 bits (1279), Expect = e-140
Identities = 271/587 (46%), Positives = 355/587 (60%), Gaps = 8/587 (1%)
Query: 793 QTPSPDLQPTPVAPSATATPPTSTASSSSPSDPSPSSSTPQSPPQPAPPVRTMATRSMRG 852
+ P+ +++PTP AP PT+T ++ + + S +S P +P Q M TR+
Sbjct: 889 RNPTNEIEPTP-APHPKPVKPTTTTTTPNRTTVSDASHQPTAPQQNQ---HNMKTRAKNN 944
Query: 853 IYKPRKLFNLSVTIDDPTISPL-PKNPKLALSDPNWKSAMQSEFDALIRNNTWDLVPRPC 911
I KP F+L+ T+ P SP P N AL D W+ AM EFDA RN+TWDLVP
Sbjct: 945 IKKPNTKFSLTATL--PNRSPSEPTNVTQALKDKKWRFAMSDEFDAQQRNHTWDLVPHES 1002
Query: 912 DVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTI 971
+ ++ C W+F+ K NG ++YKARLV G +Q GVD ETFS V+K TIR VL +
Sbjct: 1003 QL-LVGCKWVFKLKYLPNGAIDKYKARLVAKGFNQQYGVDYAETFSPVIKSTTIRLVLDV 1061
Query: 972 ALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAW 1031
A+ + W I QLDV NAFL G L E VYM QP GF D + P +VCRLRK++YGLKQAPRAW
Sbjct: 1062 AVKKDWEIKQLDVNNAFLQGTLTEEVYMAQPPGFIDKDRPTHVCRLRKAIYGLKQAPRAW 1121
Query: 1032 YQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLAS 1091
Y ++ IGF +S SD SLFIY G+ Y+L+YVDDII+ S +++ LA
Sbjct: 1122 YMELKQHLFNIGFVNSLSDASLFIYCHGTTFVYVLVYVDDIIVTGSDKSSIDAVLTSLAE 1181
Query: 1092 EFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLST 1151
F++KD L YFLGI TR GL L Q Y +D++A+ MA P TP+ T KL+
Sbjct: 1182 RFSIKDPTDLHYFLGIEATRTKQGLHLMQRKYIKDLLAKHNMADAKPVLTPLPTSPKLTL 1241
Query: 1152 SAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQG 1211
GT +D + YRS+VG+LQYL FTRPDI+YAV ++ M P +H A KR+LRY+ G
Sbjct: 1242 HGGTKLNDASEYRSVVGSLQYLAFTRPDIAYAVNRLSQLMPQPTEDHWQAAKRVLRYLAG 1301
Query: 1212 TLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSS 1271
T G+ L + L +++DADW G D ST+ Y ++LG N ISWSSK+Q ++RSS
Sbjct: 1302 TSTHGIFLDTTSPLNLHAFSDADWAGDSDDYVSTNAYVIYLGKNPISWSSKKQRGVARSS 1361
Query: 1272 AEAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEM 1331
E+EYR VAN SE WL +LL +LH L ++CDN+ A YL NPV H R KHI +
Sbjct: 1362 TESEYRAVANAASEVKWLCSLLSKLHIRLPIRPSIFCDNIGATYLCANPVFHSRMKHIAI 1421
Query: 1332 DIHFVREKVARGQARVLHVPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
D HFVR + G RV HV +R Q+AD TK L R F R + V
Sbjct: 1422 DYHFVRNMIQSGALRVSHVSTRDQLADALTKPLSRAHFQSARFKIGV 1468
Score = 300 bits (768), Expect = 4e-81
Identities = 207/598 (34%), Positives = 291/598 (48%), Gaps = 45/598 (7%)
Query: 342 PPCPLILPLPCTP*RLLLQI-APGNLTSYSNLSHLNQK-----LYVGSGQGIPIQGSGHT 395
P L + P T LL A ++TS N L+Q + + G + I +G T
Sbjct: 318 PRANLAMGAPYTANNWLLDSGATHHITSDLNALALHQPYNGDDVMIADGTSLKITKTGST 377
Query: 396 TLLTSYKTKPLSLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVIDFQTGIPLMR 455
L ++ + L+L+ VL+ P I KNL+SV +L N VSV F P F V D TG L++
Sbjct: 378 FLPSN--ARDLTLNKVLYVPDIQKNLVSVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQ 435
Query: 456 CNSPGDLY--PVTP-------SFPFAGLAQSLWHSRLGHPSSSALQSLRSNKFISYEHLN 506
+ +LY PVT + P S WHSRLGHPSSS L +L S +
Sbjct: 436 GRTKDELYEWPVTNPKATALFTTPSPKTTLSSWHSRLGHPSSSILNTLISKFSLPVSVSA 495
Query: 507 SSPV-CESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRFYVLFLDDFTD 565
S+ + C C K +LPF S+ + P + + SD+W SP+LS +++Y++ +D T
Sbjct: 496 SNKLACSDCFINKSHKLPFSISSIKSTSPLEYIFSDVWMSPILSPDNYKYYLVLVDHHTR 555
Query: 566 FLWTFPLSNKSQVFEMFISLSNQIRTHFSQTIKCLQCDNGREFDNKSFHDYCAANGLIFR 625
+ W +PL KSQV FI+ + F I+ L DNG EF + ++ +NG+
Sbjct: 556 YTWLYPLQQKSQVKSTFIAFKALVENRFQAKIRTLYSDNGGEFI--ALREFLVSNGISHL 613
Query: 626 FSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKNLSNL 685
S PHT NG +ERK R I TLL ASVP +W +A A YL+N +P LS
Sbjct: 614 TSPPHTPEHNGLSERKHRHIVETGLTLLTQASVPREYWPYAFAAAVYLINRMPTPVLSME 673
Query: 686 SPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCVFLGYPLHHRGYKCFDLS 745
SP Q L+ P+Y LRVFGCLC+P + T NKL+ RS CVFLGY L Y CFD+
Sbjct: 674 SPFQKLFGSKPNYERLRVFGCLCFPWLRPYTHNKLEERSRRCVFLGYSLTQTAYLCFDVE 733
Query: 746 HRKVIISRHVIFDETQFPFANLT---PTPSSTYEWLSDDIHPSVIHRWTTQTPSPDLQPT 802
H+++ SRHV+FDE FPF+NLT P+ T+E S + +P L +
Sbjct: 734 HKRLYTSRHVVFDEASFPFSNLTSQNSLPTVTFEQSSSPL------------VTPILSSS 781
Query: 803 PVAPSATATPPTSTASSSSP--SDPSPSSSTPQSPPQPAPPVR-TMATRSMRGIYKPRKL 859
V PS ++P T P + SP SS P + P P P R T + + L
Sbjct: 782 SVLPSCLSSPCTVLHQQQPPVTTPNSPHSSQPTTSPAPLSPHRSTTMDFQVPQVRSSSPL 841
Query: 860 FNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALI-------RNNTWDLVPRP 910
+ S +++ +P P+ P +A I RN T ++ P P
Sbjct: 842 LSSSSSLNSEPTAPNENGPEPEAQSPPIGPLSNPTHEAFIGPLPNPNRNPTNEIEPTP 899
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.329 0.141 0.453
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,769,455
Number of Sequences: 26719
Number of extensions: 1434163
Number of successful extensions: 13492
Number of sequences better than 10.0: 381
Number of HSP's better than 10.0 without gapping: 235
Number of HSP's successfully gapped in prelim test: 152
Number of HSP's that attempted gapping in prelim test: 7319
Number of HSP's gapped (non-prelim): 2378
length of query: 1379
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1267
effective length of database: 8,326,068
effective search space: 10549128156
effective search space used: 10549128156
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0234.6


