

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0234.12
(607 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
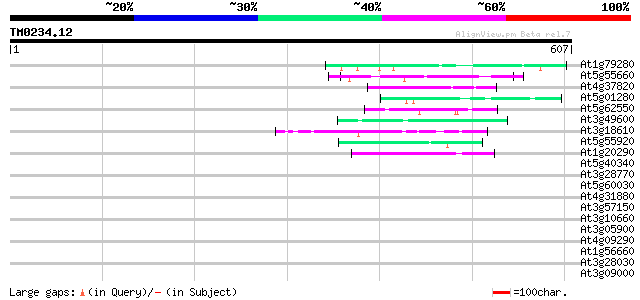
Score E
Sequences producing significant alignments: (bits) Value
At1g79280 hypothetical protein 50 4e-06
At5g55660 putative protein 47 2e-05
At4g37820 unknown protein 45 9e-05
At5g01280 putative protein 44 2e-04
At5g62550 unknown protein 44 3e-04
At3g49600 putative protein 44 3e-04
At3g18610 unknown protein 44 3e-04
At5g55920 nucleolar protein-like 43 4e-04
At1g20290 hypothetical protein 43 4e-04
At5g40340 unknown protein 42 0.001
At3g28770 hypothetical protein 40 0.003
At5g60030 KED - like protein 40 0.004
At4g31880 unknown protein 39 0.006
At3g57150 putative pseudouridine synthase (NAP57) 39 0.006
At3g10660 calmodulin-domain protein kinase CDPK isoform 2 39 0.006
At3g05900 unknown protein 39 0.008
At4g09290 putative protein 39 0.011
At1g56660 hypothetical protein 39 0.011
At3g28030 UV hypersensitive protein (UVH3) 38 0.014
At3g09000 unknown protein 38 0.014
>At1g79280 hypothetical protein
Length = 2111
Score = 50.1 bits (118), Expect = 4e-06
Identities = 70/306 (22%), Positives = 113/306 (36%), Gaps = 65/306 (21%)
Query: 342 DPITLTEFVNTLP--ESP-------PKLAT-RKSRRATSSKPSD----PKGKGILKEEPA 387
+P T+T ++ P +SP PK+A+ K +R S KPS P G+ I++ +
Sbjct: 1668 EPSTMTRVPSSTPLIKSPVATTQQLPKVASDNKEKRLISQKPSTEFRRPSGRRIVRPQLV 1727
Query: 388 KKKAASKTVII-----------REPSPERPLKRTSAQ--------QQQMSDSESSEDTWK 428
K + + K + ++P+ P + + +++ +DS SE
Sbjct: 1728 KPEESPKVDVDMPEAEGTGDEGKQPAAHEPESQVTTSVRPVQTLVRKRQADSLVSEPQQD 1787
Query: 429 DSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVV 488
+ ET E P A ++ E D + + + I E DA +TD D
Sbjct: 1788 SLTQGETSSEIAPPASKKAKGSESHPDTSEGENLAKE--PAIDELMDATTTTDGD----- 1840
Query: 489 KRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASL-TRTSPIRSLSKVI 547
+ET AE A E T E ++ +P + T T P S+
Sbjct: 1841 ------------NEETEAENAEEKTEEYVEAQQDNEADEPVEESPTETETIPTEEESRDQ 1888
Query: 548 TESINSDSALVVIPEQPMPISTSLPT-----------STQTEPIQPETQPQTQAEPQAPK 596
TE N + L + L T S P + E QP+T A P
Sbjct: 1889 TEEENQEP-LTDMESDKEEGELDLDTLEDLEEGTDVASMMRSPEKEEVQPETLATPTQSP 1947
Query: 597 SPIQTA 602
S ++TA
Sbjct: 1948 SRMETA 1953
>At5g55660 putative protein
Length = 759
Score = 47.4 bits (111), Expect = 2e-05
Identities = 43/191 (22%), Positives = 82/191 (42%), Gaps = 13/191 (6%)
Query: 359 KLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMS 418
K +KS A S S K K E A + +K + K ++++
Sbjct: 467 KRTPKKSSPAAGSSSSKRSAKSQKKTEEATR--TNKKSVAHSDDESEEEKEDDEEEEKEQ 524
Query: 419 DSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEE 478
+ E E+ ++ +++EDE L++ + + EEE E++ + +G R +SD +E
Sbjct: 525 EVEEEEEENENGIPDKSEDEAPQLSESEENVESEEESEEE----TKKKKRGSRTSSDKKE 580
Query: 479 STDSDAI--PVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKR--TSDPARAASL 534
S V + P + Q++ + S+ S+ + SKR T PA+ +
Sbjct: 581 SAGKSRSKKTAVPTKSSPPKKATQKRSAGKRKKSDDDSDTSPKASSKRKKTEKPAKEQA- 639
Query: 535 TRTSPIRSLSK 545
+P++S+SK
Sbjct: 640 --AAPLKSVSK 648
Score = 42.7 bits (99), Expect = 6e-04
Identities = 49/229 (21%), Positives = 98/229 (42%), Gaps = 42/229 (18%)
Query: 346 LTEFVNTLPESPPKLATRKSR------------RATSSKPSDPKGKGILKEEPAKKKAAS 393
L EF + S K T+K AT+ + K KG+ ++ KK +
Sbjct: 417 LLEFCDLFDISVAKATTKKEDIVTKLVEFLEKPHATTDVLVNEKEKGVKRKRTPKKSS-- 474
Query: 394 KTVIIREPSPERPLKRTSAQQQQMSDSESSED----TWKDSSSEETEDEDMPLAKRRKII 449
P+ + SA+ Q+ ++ + + D SEE +++D K +++
Sbjct: 475 -------PAAGSSSSKRSAKSQKKTEEATRTNKKSVAHSDDESEEEKEDDEEEEKEQEV- 526
Query: 450 LEEEEDEQDDHEIIQNVIQGIREASDAEESTDS--DAIPVVKRRKLPLRGPLQQKETAAE 507
EEE+E++++ I + S++EE+ +S ++ K++K R +KE+A +
Sbjct: 527 --EEEEEENENGIPDKSEDEAPQLSESEENVESEEESEEETKKKKRGSRTSSDKKESAGK 584
Query: 508 QASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSA 556
RSK+T+ P +++ + + RS K +SD++
Sbjct: 585 S------------RSKKTAVPTKSSPPKKATQKRSAGKRKKSDDDSDTS 621
Score = 30.0 bits (66), Expect = 3.9
Identities = 31/180 (17%), Positives = 69/180 (38%), Gaps = 18/180 (10%)
Query: 340 QGDPITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIR 399
+GD + E V + E + A ++ A ++ + KG +E K+ +
Sbjct: 173 KGDDVDEAEKVENVDEDDKEEALKEKNEAELAEEEETN-KGEEVKEANKEDDVEADTKVA 231
Query: 400 EPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDD 459
EP E K+T ++ + E ED +D E +D+ ED+++D
Sbjct: 232 EPEVED--KKTESKDENEDKEEEKEDEKEDEKEESNDDK---------------EDKKED 274
Query: 460 HEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRE 519
+ +G E + + +D + + + P++++++ + + RE
Sbjct: 275 IKKSNKRGKGKTEKTRGKTKSDEEKKDIEPKTPFFSDRPVRERKSVERLVAVVDKDSSRE 334
>At4g37820 unknown protein
Length = 532
Score = 45.4 bits (106), Expect = 9e-05
Identities = 34/139 (24%), Positives = 66/139 (47%), Gaps = 2/139 (1%)
Query: 388 KKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRK 447
+K A+S +E PER K S+ Q + + E + +DSSS+E E+ P K ++
Sbjct: 319 EKDASSSQDESKEEKPERKKKEESSSQGEGKEEEPEKREKEDSSSQEESKEEEPENKEKE 378
Query: 448 IILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAE 507
+EE+E + EI + +E ++ +E T+ + ++ ++Q E + +
Sbjct: 379 ASSSQEENEIKETEIKEKEESSSQEGNENKE-TEKKSSESQRKENTNSEKKIEQVE-STD 436
Query: 508 QASEPTSEVQRERRSKRTS 526
++ + Q+ SKR S
Sbjct: 437 SSNTQKGDEQKTDESKRES 455
Score = 38.5 bits (88), Expect = 0.011
Identities = 44/174 (25%), Positives = 80/174 (45%), Gaps = 14/174 (8%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
ES K + + + A+SS+ + K E KK+ +S +E PE+ K S+ Q
Sbjct: 309 ESKVKESGKNEKDASSSQDESKEEK----PERKKKEESSSQGEGKEEEPEKREKEDSSSQ 364
Query: 415 QQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREAS 474
++ S E E+ K++SS + E+E K +I +EE Q+ +E + S
Sbjct: 365 EE-SKEEEPENKEKEASSSQEENE----IKETEIKEKEESSSQEGNE--NKETEKKSSES 417
Query: 475 DAEESTDSD-AIPVVKR--RKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRT 525
+E+T+S+ I V+ +G Q+ + + ++ TS + E S +T
Sbjct: 418 QRKENTNSEKKIEQVESTDSSNTQKGDEQKTDESKRESGNDTSNKETEDDSSKT 471
Score = 31.2 bits (69), Expect = 1.7
Identities = 25/118 (21%), Positives = 50/118 (42%), Gaps = 5/118 (4%)
Query: 364 KSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESS 423
K + +SS+ + + K +++K + + E T +Q +D ES
Sbjct: 394 KEKEESSSQEGNENKETEKKSSESQRKENTNSEKKIEQVESTDSSNTQKGDEQKTD-ESK 452
Query: 424 EDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTD 481
++ D+S++ETED+ +K EE + + E QN + + A + + D
Sbjct: 453 RESGNDTSNKETEDDSSKTESEKK----EENNRNGETEETQNEQEQTKSALEISHTQD 506
>At5g01280 putative protein
Length = 460
Score = 44.3 bits (103), Expect = 2e-04
Identities = 51/208 (24%), Positives = 82/208 (38%), Gaps = 32/208 (15%)
Query: 402 SPERPLKRTSAQQQQMSDSESSEDTW----KDSSSEET-------EDEDMPLAKRRKIIL 450
S + P++RT+ + SD E S+ W S S +D M L R +
Sbjct: 3 SQQYPVRRTAGENLLYSDGEKSDYEWLVTPPGSPSRNVTNHLNAPDDNLMTLISRLENYS 62
Query: 451 EEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQAS 510
+EE + Q + + GIR S + S + P P ++ +T A++ S
Sbjct: 63 KEESEHQTTSLHSSSSVSGIRRPSSSSSSRSTSRPPT----------PTRKSKTPAKRPS 112
Query: 511 EPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTS 570
PTS R+ T+ A S + TS RS S+ + S S + + + +T
Sbjct: 113 TPTS------RATSTTTRATLTSSSTTSSTRSWSRPSSSSGTGTSRVTLTAAR----ATR 162
Query: 571 LPTSTQTEPIQPETQPQTQAEP-QAPKS 597
TST + T ++ P AP S
Sbjct: 163 PTTSTDQQTTGSATSTRSNNRPMSAPNS 190
>At5g62550 unknown protein
Length = 487
Score = 43.9 bits (102), Expect = 3e-04
Identities = 38/156 (24%), Positives = 72/156 (45%), Gaps = 20/156 (12%)
Query: 385 EPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPL-- 442
E K+K A + + E S E + Q ++++ ED +DSS ET E+ L
Sbjct: 285 ETTKEKDALQDSSVTETSKEEG----ALQDSSVTETTKEEDALQDSSVTETTKEEQALET 340
Query: 443 ---AKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEEST----DSD----AIPVVKRR 491
+ RK + ++++D E++Q +G+R ++D + SD V +
Sbjct: 341 VTQGRTRKSLEVINVNQENDSEVVQESEEGLRPSADGVQIVTVVKPSDKKRARKETVPKN 400
Query: 492 KLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSD 527
LP+R +K A A+ T +V ++ +S++ S+
Sbjct: 401 NLPVR---TKKSLATNSANSKTVQVNKDDKSQKKSE 433
Score = 35.0 bits (79), Expect = 0.12
Identities = 41/185 (22%), Positives = 77/185 (41%), Gaps = 17/185 (9%)
Query: 408 KRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVI 467
++ + Q ++++ +D +DSS ET E+ L +EED D +V
Sbjct: 274 EKDALQDSSVTETTKEKDALQDSSVTETSKEEGALQDSSVTETTKEEDALQD----SSVT 329
Query: 468 QGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASE----PTSEVQRERRSK 523
+ +E E T R+ L + Q+ ++ Q SE P+++ +
Sbjct: 330 ETTKEEQALETVTQGRT-----RKSLEVINVNQENDSEVVQESEEGLRPSADGVQIVTVV 384
Query: 524 RTSDPARAASLT---RTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPI 580
+ SD RA T P+R+ + T S NS + V ++ S + T +T+ +
Sbjct: 385 KPSDKKRARKETVPKNNLPVRTKKSLATNSANSKTVQVNKDDKSQKKSERI-TKPRTKRV 443
Query: 581 QPETQ 585
Q E++
Sbjct: 444 QEESK 448
>At3g49600 putative protein
Length = 1672
Score = 43.9 bits (102), Expect = 3e-04
Identities = 36/184 (19%), Positives = 74/184 (39%), Gaps = 8/184 (4%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
+ P K ++SRR S + + ++ K + + K + S + L+ +Q
Sbjct: 275 KKPTKTTKKRSRRKRSVSSESEE---VESDDSKKLRKSHKKSLPSNRSGSKELRDKHDEQ 331
Query: 415 QQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREAS 474
+ D S E+ED PL K+ + ++ ++DD ++ + ++
Sbjct: 332 SRAGRKRHDSDV----SEPESEDNKQPLRKKEEAYRGGQKQKRDDEDVEADHLKDRYTRD 387
Query: 475 DAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASL 534
D + + DSD + + K LR ++ Q + +V + + K SD +R +
Sbjct: 388 DKKAARDSDDSEIEYQNKKQLRSKVEVYSAGMSQKRKEEEDVTKHGKDKYRSD-SRGKEV 446
Query: 535 TRTS 538
R S
Sbjct: 447 ARDS 450
Score = 33.9 bits (76), Expect = 0.27
Identities = 48/202 (23%), Positives = 75/202 (36%), Gaps = 35/202 (17%)
Query: 364 KSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSES- 422
K RR S SD G+ + +KKKA + K+ S SDSES
Sbjct: 219 KKRRHDDSSESDEHGRD--RRRRSKKKAKGR-------------KQESESDSSSSDSESD 263
Query: 423 --SEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQ--------GIRE 472
S+D K + T+ ++R + E EE E DD + ++ + G +E
Sbjct: 264 SDSDDGKKRGRKKPTKTTKKRSRRKRSVSSESEEVESDDSKKLRKSHKKSLPSNRSGSKE 323
Query: 473 ASDAEEST--------DSD-AIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSK 523
D + DSD + P + K PLR + +Q + + +
Sbjct: 324 LRDKHDEQSRAGRKRHDSDVSEPESEDNKQPLRKKEEAYRGGQKQKRDDEDVEADHLKDR 383
Query: 524 RTSDPARAASLTRTSPIRSLSK 545
T D +AA + S I +K
Sbjct: 384 YTRDDKKAARDSDDSEIEYQNK 405
>At3g18610 unknown protein
Length = 636
Score = 43.5 bits (101), Expect = 3e-04
Identities = 53/235 (22%), Positives = 101/235 (42%), Gaps = 27/235 (11%)
Query: 288 KKMQIIEKIQVAPKEDTSDEVRQRDYPIDDFPIWSKKDNPACILEYVRMLRRQGDPITLT 347
K+ ++EK ++ + +SD+ D D+ + KK +LE + D + +
Sbjct: 168 KQPAVLEKAKI--ESSSSDD----DSSSDEETVPMKKQT--AVLEKAKAESSSSDDGSSS 219
Query: 348 EFVNTLPESPPKLATRKSRRATSSKPSDP----KGKGILKEEPAKKKAASK-TVIIREPS 402
+ T + P + + S +SS P K ++K+ A+ ++ + + EP+
Sbjct: 220 DEEPTPAKKEPIVVKKDSSDESSSDEETPVVKKKPTTVVKDAKAESSSSEEESSSDDEPT 279
Query: 403 PERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEI 462
P + + DS SSE+ DS EE++DE P K+ K+ + + E E
Sbjct: 280 PAKKPTVVKNAKPAAKDSSSSEE---DSDEEESDDEKPP-TKKAKVSSKTSKQESSSDE- 334
Query: 463 IQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQ 517
+ SD EES D P K++ + ++++ A+Q PT++ Q
Sbjct: 335 -------SSDESDKEESKDEKVTP--KKKDSDVEMVDAEQKSNAKQPKTPTNQTQ 380
Score = 40.4 bits (93), Expect = 0.003
Identities = 57/267 (21%), Positives = 109/267 (40%), Gaps = 47/267 (17%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEE-----PAKKKAASKTVIIREPSPERPLKR 409
+S K T A+ +KP KGK +E+ K+K V+ +E + + K+
Sbjct: 3 KSSKKSVTEVETPASMTKPLK-KGKRDAEEDLDMQVTKKQKKELIDVVQKEKAEKTVPKK 61
Query: 410 TSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEI--IQNVI 467
+ SDS+ E T + S + E EEE+D D EI +
Sbjct: 62 VESSSSDASDSDEEEKTKETPSKLKDESSS-----------EEEDDSSSDEEIAPAKKRP 110
Query: 468 QGIREA------SDAEESTDSDAIPVVKR----RKLPLRGPLQQKETAAEQASEPTSE-- 515
+ I++A SD + ++D + PV K+ K + ++++++ + P +
Sbjct: 111 EPIKKAKVESSSSDDDSTSDEETAPVKKQPAVLEKAKVESSSSDDDSSSDEETVPVKKQP 170
Query: 516 -VQRERRSKRTSDPARAASLTRTSPIRS----LSKVITESINSDSALVVIPEQPMPISTS 570
V + + + +S ++S T P++ L K ES +SD S
Sbjct: 171 AVLEKAKIESSSSDDDSSSDEETVPMKKQTAVLEKAKAESSSSDDG---------SSSDE 221
Query: 571 LPTSTQTEPI--QPETQPQTQAEPQAP 595
PT + EPI + ++ ++ ++ + P
Sbjct: 222 EPTPAKKEPIVVKKDSSDESSSDEETP 248
>At5g55920 nucleolar protein-like
Length = 682
Score = 43.1 bits (100), Expect = 4e-04
Identities = 35/161 (21%), Positives = 65/161 (39%), Gaps = 6/161 (3%)
Query: 356 SPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQ 415
+PP KS+ + KG + K K V+ EP + + S +
Sbjct: 17 TPPTKQLTKSKTPPMKPQTSMLKKGAKSQNKPPLKKQKKEVVEEEPLEDYEVTDDSDEDD 76
Query: 416 QMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIRE--- 472
++SD +D SEE ++ D ++ ++EE E D+ + + ++G +
Sbjct: 77 EVSDGSDEDDISPAVESEEIDESDDGENGSNQLFSDDEE-ENDEETLGDDFLEGSGDEDE 135
Query: 473 --ASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASE 511
+ DA+ DSD +V + R QK+ AA + +
Sbjct: 136 EGSLDADSDADSDDDDIVAKSDAIDRDLAMQKKDAAAELED 176
Score = 35.0 bits (79), Expect = 0.12
Identities = 19/84 (22%), Positives = 46/84 (54%), Gaps = 3/84 (3%)
Query: 442 LAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQ 501
+AK +K+ ++ E+ D + ++ V Q + D +E+ +AI ++ +P+R P ++
Sbjct: 558 VAKLKKMSNVKQSSEEGDDDAVETVEQAEVSSDDDDEA---EAIEETEKPSVPVRQPKER 614
Query: 502 KETAAEQASEPTSEVQRERRSKRT 525
KE ++ + E +R ++ K++
Sbjct: 615 KEKKNKEKLAKSKEDKRGKKDKKS 638
>At1g20290 hypothetical protein
Length = 497
Score = 43.1 bits (100), Expect = 4e-04
Identities = 29/155 (18%), Positives = 64/155 (40%), Gaps = 5/155 (3%)
Query: 370 SSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKD 429
S+K +P K P+ T++ E E + ++++ + E E+ ++
Sbjct: 334 STKWVNPSDSDFYKFTPSMVHVGEGTIVQAEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 393
Query: 430 SSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVK 489
EE E+E+ + + EEEE+E+++ E + + E + EE + + +
Sbjct: 394 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE-----E 448
Query: 490 RRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKR 524
+ +++E E+ E E + E K+
Sbjct: 449 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKK 483
Score = 39.7 bits (91), Expect = 0.005
Identities = 22/110 (20%), Positives = 51/110 (46%)
Query: 383 KEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPL 442
+EE +++ + E E + ++++ + E E+ ++ EE E+E+
Sbjct: 382 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 441
Query: 443 AKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRK 492
+ + EEEE+E+++ E + + E + EE + +VK++K
Sbjct: 442 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKKYMIVKKKK 491
Score = 37.7 bits (86), Expect = 0.019
Identities = 22/110 (20%), Positives = 51/110 (46%)
Query: 383 KEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPL 442
+EE +++ + E E + ++++ + E E+ ++ EE E+E+
Sbjct: 383 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 442
Query: 443 AKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRK 492
+ + EEEE+E+++ E + + E + EE + + V K++K
Sbjct: 443 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKKYMIVKKKKK 492
Score = 32.3 bits (72), Expect = 0.78
Identities = 18/98 (18%), Positives = 44/98 (44%)
Query: 383 KEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPL 442
+EE +++ + E E + ++++ + E E+ ++ EE E+E+
Sbjct: 400 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 459
Query: 443 AKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEEST 480
+ + EEEE+E+++ E + +++ ST
Sbjct: 460 EEEEEEEEEEEEEEEEEEEEEEKKYMIVKKKKKKRRST 497
>At5g40340 unknown protein
Length = 1008
Score = 42.0 bits (97), Expect = 0.001
Identities = 34/147 (23%), Positives = 64/147 (43%), Gaps = 9/147 (6%)
Query: 369 TSSKPSDPKGKGILKEEPAKKKAASK--------TVIIREPSPERPLKRTSAQQQQMSDS 420
T + ++ G +K+E +KK+ SK T S ++ KR ++ ++ SD
Sbjct: 715 TGKEENETDKHGKMKKERKRKKSESKKEGGEGEETQKEANESTKKERKRKKSESKKQSDG 774
Query: 421 ESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEEST 480
E S+++ P +K++ +EEEE ++ E + + + D EE
Sbjct: 775 EEETQKEPSESTKKERKRKNPESKKKAEAVEEEETRKESVESTKKERKRKKPKHDEEEVP 834
Query: 481 DSDAIPVVKRRKLPLRGPLQQKETAAE 507
+ P K++K G ++KET E
Sbjct: 835 NETEKP-EKKKKKKREGKSKKKETETE 860
Score = 31.2 bits (69), Expect = 1.7
Identities = 44/230 (19%), Positives = 76/230 (32%), Gaps = 53/230 (23%)
Query: 369 TSSKPSDPKGKGILKEEPAKKKAASKTVIIR---EPSPERPLKRTSAQQQQMSDSESSED 425
T KP +P L +E ++ ++ +R P + L R +A Q SS D
Sbjct: 590 TLDKPCEPSSDNGLGQEELSRELSNAVDFLRLGATPKEMQDLIRVAALGTQYPKDSSSRD 649
Query: 426 TWK--------------------------DSSSEETEDEDMPLAKRRKI----------- 448
+ D EE + P+ K ++
Sbjct: 650 MVREFMTIYRSFTYHDGANHKFLGSYDSSDKEKEELSEMGKPVTKGKEKKDKKGKAKQKA 709
Query: 449 ----ILEEEEDEQDDH-----EIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPL 499
+ +EE+E D H E + + +E + EE+ K RK
Sbjct: 710 EEIEVTGKEENETDKHGKMKKERKRKKSESKKEGGEGEETQKEANESTKKERKRKKSESK 769
Query: 500 QQKETAAEQASEPTSEVQRERRSKRTSDPARAASL----TRTSPIRSLSK 545
+Q + E EP+ ++ER+ K +A ++ TR + S K
Sbjct: 770 KQSDGEEETQKEPSESTKKERKRKNPESKKKAEAVEEEETRKESVESTKK 819
Score = 29.3 bits (64), Expect = 6.6
Identities = 27/129 (20%), Positives = 56/129 (42%), Gaps = 15/129 (11%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
E K A +++ K S+ K + +EE K+ + S + +PE K+ A +
Sbjct: 747 EETQKEANESTKKERKRKKSESKKQSDGEEETQKEPSESTKKERKRKNPESK-KKAEAVE 805
Query: 415 QQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREAS 474
++ + ES E T K+ ++RK +EE+ ++ E + + RE
Sbjct: 806 EEETRKESVESTKKE--------------RKRKKPKHDEEEVPNETEKPEKKKKKKREGK 851
Query: 475 DAEESTDSD 483
++ T+++
Sbjct: 852 SKKKETETE 860
>At3g28770 hypothetical protein
Length = 2081
Score = 40.4 bits (93), Expect = 0.003
Identities = 35/172 (20%), Positives = 73/172 (42%), Gaps = 7/172 (4%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
E L +K T K K K+E K+ +K++ E E+ S +
Sbjct: 1053 EESRDLKAKKKEEETKEKKESENHKS-KKKEDKKEHEDNKSMKKEEDKKEKKKHEESKSR 1111
Query: 415 QQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREAS 474
++ D + E +S+++ ED++ + ++++E D+++ E + + +
Sbjct: 1112 KKEEDKKDMEKLEDQNSNKKKEDKNEKKKSQHVKLVKKESDKKEKKENEE------KSET 1165
Query: 475 DAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTS 526
ES+ S V K+ K + ++KE +++ E + E R K+TS
Sbjct: 1166 KEIESSKSQKNEVDKKEKKSSKDQQKKKEKEMKESEEKKLKKNEEDRKKQTS 1217
Score = 35.8 bits (81), Expect = 0.071
Identities = 46/275 (16%), Positives = 114/275 (40%), Gaps = 21/275 (7%)
Query: 332 EYVRMLRRQGDPITLTEFVNTLPESPPKLATRKSRR-------ATSSKPSDPKGKGILKE 384
++V++++++ D E N ++ + KS++ SSK K + +KE
Sbjct: 1142 QHVKLVKKESDKKEKKE--NEEKSETKEIESSKSQKNEVDKKEKKSSKDQQKKKEKEMKE 1199
Query: 385 EPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDS----SSEETEDEDM 440
KK ++ ++ S E K+ ++++ + ++T K S S E+E ++
Sbjct: 1200 SEEKKLKKNEEDRKKQTSVEENKKQKETKKEKNKPKDDKKNTTKQSGGKKESMESESKEA 1259
Query: 441 PLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQ 500
++ + + + DE +EI+ SD++ +D ++ +
Sbjct: 1260 ENQQKSQATTQADSDE-SKNEILMQADSQADSHSDSQADSDESKNEILMQADSQATTQRN 1318
Query: 501 QKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVI 560
+E +Q S ++ Q+E + ++ + T+ S + ES+ S+S
Sbjct: 1319 NEEDRKKQTSVAENKKQKETKEEKNKPKDDKKNTTKQSGGKK------ESMESESKEAEN 1372
Query: 561 PEQPMPISTSLPTSTQTEPI-QPETQPQTQAEPQA 594
++ + + ++ E + Q ++Q + ++ QA
Sbjct: 1373 QQKSQATTQADSDESKNEILMQADSQADSHSDSQA 1407
Score = 33.1 bits (74), Expect = 0.46
Identities = 39/248 (15%), Positives = 108/248 (42%), Gaps = 14/248 (5%)
Query: 288 KKMQIIEKIQVAPKEDTSDEVRQRDYPIDDFPIWSKKDNPACILEYVRMLRRQGDPITLT 347
K+ + +K + + D S RD+ ++ I +K + + +Y + +++G+
Sbjct: 873 KEESMKKKREEVQRNDKSSTKEVRDFA-NNMDIDVQKGSGESV-KYKKDEKKEGNKEENK 930
Query: 348 EFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPL 407
+ +NT ++++ + K + K + K+E KK+ + + +E + +
Sbjct: 931 DTINT--------SSKQKGKDKKKKKKESKNSNMKKKEEDKKEYVNNELKKQEDNKKETT 982
Query: 408 KRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAK-RRKIILEEEEDEQDDHEIIQNV 466
K +++ ++ + + +DS+S+ E ++ K + K ++E+ + D + +
Sbjct: 983 KSENSKLKEENKDNKEKKESEDSASKNREKKEYEEKKSKTKEEAKKEKKKSQDKKREEKD 1042
Query: 467 IQGIREASDAEESTDSDA---IPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSK 523
+ + + EES D A K +K ++KE E + + + +++ K
Sbjct: 1043 SEERKSKKEKEESRDLKAKKKEEETKEKKESENHKSKKKEDKKEHEDNKSMKKEEDKKEK 1102
Query: 524 RTSDPARA 531
+ + +++
Sbjct: 1103 KKHEESKS 1110
Score = 29.6 bits (65), Expect = 5.1
Identities = 48/256 (18%), Positives = 99/256 (37%), Gaps = 38/256 (14%)
Query: 354 PESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQ 413
P+ K T++S S S+ K E ++K+ + T + S L + +Q
Sbjct: 1345 PKDDKKNTTKQSGGKKESMESESK------EAENQQKSQATTQADSDESKNEILMQADSQ 1398
Query: 414 QQQMSDSESSEDTWKDS-------------SSEETEDEDMPLA--KRRKIILEEEEDEQD 458
SDS++ D K+ ++EE + +A K++K EE+ +D
Sbjct: 1399 ADSHSDSQADSDESKNEILMQADSQATTQRNNEEDRKKQTSVAENKKQKETKEEKNKPKD 1458
Query: 459 DHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQR 518
D + ++ +ES +S++ ++K + E+ E + S+
Sbjct: 1459 DKK------NTTEQSGGKKESMESESKEAENQQKSQATTQGESDESKNEILMQADSQADT 1512
Query: 519 ERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTE 578
S+ SD ++ L + S + T+S S + ++ + +QT+
Sbjct: 1513 HANSQGDSDESKNEILMQAD---SQADSQTDSDESKNEIL--------MQADSQADSQTD 1561
Query: 579 PIQPETQPQTQAEPQA 594
+ + + QA+ QA
Sbjct: 1562 SDESKNEILMQADSQA 1577
>At5g60030 KED - like protein
Length = 292
Score = 40.0 bits (92), Expect = 0.004
Identities = 37/197 (18%), Positives = 82/197 (40%), Gaps = 14/197 (7%)
Query: 363 RKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPE----------RPLKRTSA 412
RK R S S G ++E KK+ V+ + + + R K+
Sbjct: 95 RKIRYRNSEAVSVESVYGRERDEKKMKKSKDADVVDEKVNEKLEAEQRSEERRERKKEKK 154
Query: 413 QQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIRE 472
+++ D + ++ K+ +E + D K++K +ED D+ E +++ Q E
Sbjct: 155 KKKNNKDEDVVDEKVKEKLEDEQKSADRKERKKKKSKKNNDEDVVDEKEKLEDE-QKSAE 213
Query: 473 ASDAEESTDSDAIPVVKRRKL---PLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPA 529
+ +++ D D + ++ KL G ++++ ++ E +R+ + KR SD
Sbjct: 214 IKEKKKNKDEDVVDEKEKEKLEDEQRSGERKKEKKKKRKSDEEIVSEERKSKKKRKSDEE 273
Query: 530 RAASLTRTSPIRSLSKV 546
+ ++ R L ++
Sbjct: 274 MGSEERKSKKKRKLKEI 290
Score = 33.1 bits (74), Expect = 0.46
Identities = 35/193 (18%), Positives = 83/193 (42%), Gaps = 18/193 (9%)
Query: 335 RMLRRQGDPITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASK 394
+ +++ D + E VN E+ + R+ R+ K + K + ++ +E K+K +
Sbjct: 118 KKMKKSKDADVVDEKVNEKLEAEQRSEERRERKKEKKKKKNNKDEDVV-DEKVKEKLEDE 176
Query: 395 TVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEE 454
+ S +R ++++ S + ED + E E + + +++K E+
Sbjct: 177 -----QKSADR-----KERKKKKSKKNNDEDVVDEKEKLEDEQKSAEIKEKKKNKDEDVV 226
Query: 455 DEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTS 514
DE++ ++ G R+ + ++ SD V + RK ++K + E+
Sbjct: 227 DEKEKEKLEDEQRSGERK-KEKKKKRKSDEEIVSEERK------SKKKRKSDEEMGSEER 279
Query: 515 EVQRERRSKRTSD 527
+ +++R+ K D
Sbjct: 280 KSKKKRKLKEIDD 292
>At4g31880 unknown protein
Length = 873
Score = 39.3 bits (90), Expect = 0.006
Identities = 45/230 (19%), Positives = 89/230 (38%), Gaps = 30/230 (13%)
Query: 379 KGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ--QQMSDSESSEDTWKDSSSEETE 436
+G +K E +KAA + R +P+ ++ Q +DS D+ K +
Sbjct: 272 QGHIKRETEVEKAAEISTPERTDAPKDESGKSGVSNGVAQQNDSSVDTDSMKKQDDTGAK 331
Query: 437 DEDMPLAKRRKIIL----EEEEDEQDDHEIIQNVIQGIREA---SDAEESTDSDAIPVVK 489
DE L R L EE+ D + E +N +++A D++ +++ ++
Sbjct: 332 DEPQQLDNPRNTDLNNTTEEKPDVEHQIEEKENESSSVKQADLSKDSDIKEETEPAELLD 391
Query: 490 RRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITE 549
+ + P+ TAA + ++ + SK + D T
Sbjct: 392 SKDVLTSPPVDSSVTAATSSENEKNKSVQILPSKTSGDE-------------------TA 432
Query: 550 SINSDSALVVIPEQPMPISTS--LPTSTQTEPIQPETQPQTQAEPQAPKS 597
+++S S +PEQ +P T+ + TE ++P T+ + P +
Sbjct: 433 NVSSPSMAEELPEQSVPKKTANQKKKESSTEEVKPSASIATEEVSEEPNT 482
Score = 32.7 bits (73), Expect = 0.60
Identities = 26/93 (27%), Positives = 40/93 (42%), Gaps = 8/93 (8%)
Query: 355 ESPPKLATR------KSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLK 408
E PPK + K ++ SK K KEEP+K +SK+ P P K
Sbjct: 756 EEPPKTVGKSGSSRSKKDISSVSKSGKSKASSKKKEEPSKATTSSKSK--SGPVKSVPAK 813
Query: 409 RTSAQQQQMSDSESSEDTWKDSSSEETEDEDMP 441
+ + + S S S+ + S+ E+E E+ P
Sbjct: 814 SKTGKGKAKSGSASTPASKAKESASESESEETP 846
>At3g57150 putative pseudouridine synthase (NAP57)
Length = 565
Score = 39.3 bits (90), Expect = 0.006
Identities = 26/107 (24%), Positives = 55/107 (51%), Gaps = 9/107 (8%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
E+ K+ + K ++ + + G K+E KKK + VI SP+ K+ +
Sbjct: 468 EAEEKVKSSKKKKKKDKEEEKEEEAGSEKKEKKKKKDKKEEVIEEVASPKSEKKK----K 523
Query: 415 QQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHE 461
++ D+E++ D +S++E++E + K++K ++ +D +DD E
Sbjct: 524 KKSKDTEAAVDAEDESAAEKSEKK-----KKKKDKKKKNKDSEDDEE 565
Score = 30.4 bits (67), Expect = 3.0
Identities = 24/116 (20%), Positives = 53/116 (45%), Gaps = 9/116 (7%)
Query: 411 SAQQQQMSDSESSE-DTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQG 469
+A ++ +D+E+ E + +++ D P+ ++ E E +E ++
Sbjct: 422 AAPEEIKADAENGEAGEARKRKHDDSSDSPAPVTTKKSKTKEVEGEEAEEKVKSSKK--- 478
Query: 470 IREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRT 525
++ D EE + +A K +K +KE E+ + P SE +++++SK T
Sbjct: 479 -KKKKDKEEEKEEEAGSEKKEKKKKK----DKKEEVIEEVASPKSEKKKKKKSKDT 529
Score = 28.9 bits (63), Expect = 8.7
Identities = 29/114 (25%), Positives = 44/114 (38%), Gaps = 29/114 (25%)
Query: 355 ESPPKLATRKSR-----------RATSSKPSDPKGKGILKEEPA-------KKKAASKTV 396
+SP + T+KS+ + SSK K K KEE A KKK K
Sbjct: 449 DSPAPVTTKKSKTKEVEGEEAEEKVKSSKKKKKKDKEEEKEEEAGSEKKEKKKKKDKKEE 508
Query: 397 IIRE---PSPERPLKRTS--------AQQQQMSDSESSEDTWKDSSSEETEDED 439
+I E P E+ K+ S A+ + ++ + KD + + ED
Sbjct: 509 VIEEVASPKSEKKKKKKSKDTEAAVDAEDESAAEKSEKKKKKKDKKKKNKDSED 562
>At3g10660 calmodulin-domain protein kinase CDPK isoform 2
Length = 646
Score = 39.3 bits (90), Expect = 0.006
Identities = 31/100 (31%), Positives = 46/100 (46%), Gaps = 8/100 (8%)
Query: 499 LQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALV 558
+ +T E +P E+Q E + ++ A T+ + SK S +V
Sbjct: 66 MPSSKTNPETKLKPDLEIQPEEKKEKV-----LAEETKQKVVPEESKQEVPPEESKREVV 120
Query: 559 VIPEQPMPI--STSLPTSTQTEPIQPETQPQTQAEPQAPK 596
V PE P S S P +T+ E ET+P+T+AEPQ PK
Sbjct: 121 VQPESAKPETKSESKPETTKPETTS-ETKPETKAEPQKPK 159
>At3g05900 unknown protein
Length = 660
Score = 38.9 bits (89), Expect = 0.008
Identities = 66/348 (18%), Positives = 124/348 (34%), Gaps = 63/348 (18%)
Query: 277 TIGDALNGIKLKKMQIIEKIQVAPKEDTSDEVRQRDYPIDDFPIWSKKDNPACILEYVRM 336
++ D I +K++ +E ++V PK +TS++V + LE R
Sbjct: 207 SVRDVSAEIAEEKVKDVEALEVEPKPETSEKVETQ-------------------LEKARE 247
Query: 337 LRRQGDPITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTV 396
L + + + E ++ +L + K S+ K E + + +
Sbjct: 248 LETEVEVVKAEETAEATEQAKVELEGKLEDVIVEEKDSEINSKDEKTSESGSALCSEEIL 307
Query: 397 IIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDE 456
+ S P+K T D+ D + + +EE D P EE+
Sbjct: 308 STIQESNTDPIKETE------GDASYPIDVIEKAITEEKHVVDEP--------ANEEKPS 353
Query: 457 QDDHEIIQNVIQGIREASDA--EESTDSDAI-------PVVKRRKLPLRGPLQQKETAAE 507
+ + + I + SD +E T+ DA + K + P + ET +E
Sbjct: 354 ESSAALSPEKVVPINQDSDTKPKEETEGDAAAPADVIEKAITEEKYVVDEP-SKDETTSE 412
Query: 508 QASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESIN-----------SDSA 556
S E ++P + +SP + K ITE + S+S
Sbjct: 413 SGSALCPEKAVPTNQDLDTEPKKETEEDVSSPADIIEKAITEEKHVVEEPSKDEKTSESG 472
Query: 557 LVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQTATT 604
+ PE+ +P + + TEP + +T+ + +P I+ A T
Sbjct: 473 SALSPEKVVPTN----QDSDTEP-----KKETEGDVPSPADVIEKAIT 511
>At4g09290 putative protein
Length = 376
Score = 38.5 bits (88), Expect = 0.011
Identities = 27/110 (24%), Positives = 55/110 (49%), Gaps = 12/110 (10%)
Query: 348 EFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPL 407
+F+ P+S + +RR + +K S PK K+EPAK+ + T + + + ++P
Sbjct: 241 DFIEKSPDSRRLRSMTAARRISKAKTS-PK-----KKEPAKRGRKAATKVTPKVTIKKPK 294
Query: 408 KRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQ 457
A+++ D+ S T + + E+ + ED +++ E +EDE+
Sbjct: 295 SPKEAKEKADGDTSSVPKTKPEEAKEKADAED------NRLLPESDEDEE 338
>At1g56660 hypothetical protein
Length = 522
Score = 38.5 bits (88), Expect = 0.011
Identities = 35/168 (20%), Positives = 68/168 (39%), Gaps = 16/168 (9%)
Query: 364 KSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESS 423
K T K PK + KEE +K K +E + L++ ++++ D
Sbjct: 182 KDESGTEEKKKKPKKEKKQKEE-SKSNEDKKVKGKKEKGEKGDLEKEDEEKKKEHDETDQ 240
Query: 424 EDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDH-EIIQNVIQGIREASDAEESTDS 482
E KDS + +++D A+ +K ++E+ E+D+ E ++G + + E D
Sbjct: 241 EMKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKGKKGKGEKPEKED- 299
Query: 483 DAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPAR 530
G ++ A EQ + + +E + K+ D A+
Sbjct: 300 -------------EGKKTKEHDATEQEMDDEAADHKEGKKKKNKDKAK 334
Score = 34.3 bits (77), Expect = 0.21
Identities = 32/148 (21%), Positives = 67/148 (44%), Gaps = 15/148 (10%)
Query: 377 KGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETE 436
KGK EE ++K K +E P ++ ++ + S++ EE E
Sbjct: 117 KGKEKKHEELEEEKEGKKKKNKKEKDESGPEEKNKKADKEKKHEDVSQE------KEELE 170
Query: 437 DEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLR 496
+ED K+ K ++E+DE E + + +E EES ++ V +++ +
Sbjct: 171 EED---GKKNK---KKEKDESGTEEKKK---KPKKEKKQKEESKSNEDKKVKGKKEKGEK 221
Query: 497 GPLQQKETAAEQASEPTSEVQRERRSKR 524
G L++++ ++ + T + +E+ SK+
Sbjct: 222 GDLEKEDEEKKKEHDETDQEMKEKDSKK 249
Score = 33.1 bits (74), Expect = 0.46
Identities = 27/138 (19%), Positives = 53/138 (37%), Gaps = 10/138 (7%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
E K K + S K + KE P + + S+ + + EP E+ + + ++
Sbjct: 355 EGETKQKKNKKKEKKSEKGEKDVKEDKKKENPLETEVMSRDIKLEEPEAEKKEEDDTEEK 414
Query: 415 QQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREAS 474
++ + K ++ + K K+ +EEE + D ++ I +
Sbjct: 415 KKSKVEGGESEEGKKKKKKDKKKNKKKDTKEPKMTEDEEEKKDDSKDV------KIEGSK 468
Query: 475 DAEESTDSDAIPVVKRRK 492
EE D D VK++K
Sbjct: 469 AKEEKKDKD----VKKKK 482
Score = 32.7 bits (73), Expect = 0.60
Identities = 33/164 (20%), Positives = 66/164 (40%), Gaps = 14/164 (8%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
E K K ++ S D K KG KK+ K + +E ++ + Q+
Sbjct: 189 EKKKKPKKEKKQKEESKSNEDKKVKG-------KKEKGEKGDLEKEDEEKKKEHDETDQE 241
Query: 415 QQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDE-------QDDHEIIQNVI 467
+ DS+ ++ KD S E + + K+ K E+ED+ + + ++
Sbjct: 242 MKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKGKKGKGEKPEKEDEG 301
Query: 468 QGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASE 511
+ +E E+ D +A + +K + ++KET ++ E
Sbjct: 302 KKTKEHDATEQEMDDEAADHKEGKKKKNKDKAKKKETVIDEVCE 345
Score = 30.8 bits (68), Expect = 2.3
Identities = 35/184 (19%), Positives = 70/184 (38%), Gaps = 11/184 (5%)
Query: 354 PESPPKLATRKSRRATSSKPSDP-----KGKGILKEEPAKKKAASKTVIIREPSPERPLK 408
PE + K AT + D +GK ++ AKKK + + + ++
Sbjct: 295 PEKEDEGKKTKEHDATEQEMDDEAADHKEGKKKKNKDKAKKKETVIDEVCEKETKDKDDD 354
Query: 409 RTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKR---RKIILEEEEDEQDDHEIIQN 465
+Q++ E + + E+ + E+ PL R I LEE E E+ + + +
Sbjct: 355 EGETKQKKNKKKEKKSEKGEKDVKEDKKKEN-PLETEVMSRDIKLEEPEAEKKEEDDTEE 413
Query: 466 VIQGIREASDAEESTDSDAIPVVKRRKLPLRGP--LQQKETAAEQASEPTSEVQRERRSK 523
+ E ++EE K +K + P + +E + + + E + + K
Sbjct: 414 KKKSKVEGGESEEGKKKKKKDKKKNKKKDTKEPKMTEDEEEKKDDSKDVKIEGSKAKEEK 473
Query: 524 RTSD 527
+ D
Sbjct: 474 KDKD 477
Score = 30.8 bits (68), Expect = 2.3
Identities = 26/145 (17%), Positives = 55/145 (37%), Gaps = 3/145 (2%)
Query: 384 EEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLA 443
+E AK++ + +E P+ + + + + S KD S +D
Sbjct: 5 QENAKEEKLHVKIKTQELDPKEKGENVEVEMEVKAKSIEKVKAKKDEESSGKSKKDKEKK 64
Query: 444 KRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKE 503
K + + E +ED+ DD + ++ E + + V + K +G ++ E
Sbjct: 65 KGKNVDSEVKEDKDDDKKKDGKMVSKKHEEGHGDLEVKESDVKVEEHEKEHKKGKEKKHE 124
Query: 504 TAAEQASEPTSEVQRERRSKRTSDP 528
E E + ++ ++ K S P
Sbjct: 125 ---ELEEEKEGKKKKNKKEKDESGP 146
>At3g28030 UV hypersensitive protein (UVH3)
Length = 1479
Score = 38.1 bits (87), Expect = 0.014
Identities = 40/185 (21%), Positives = 77/185 (41%), Gaps = 13/185 (7%)
Query: 371 SKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDS 430
S DPKGKG+L + + + + E+ + ++ S ++E+ +
Sbjct: 192 SVKDDPKGKGVLLDGDDLDNLVQDSSVQGKDYQEK-------LDEMLAASLAAEEERNFT 244
Query: 431 SSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKR 490
S T +P + +EEED D EI+ V+ G + + S + ++ +
Sbjct: 245 SKASTSAAAIPSEE------DEEEDSDGDEEILLPVMDGNIDPAVLASLPPSMQLDLLAQ 298
Query: 491 RKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITES 550
+ L +QK ++A E SE+Q E K + + R++ R++ V T
Sbjct: 299 MREKLMAENRQKYQKVKKAPEKFSELQIEAYLKTVAFRREINEVQRSAGGRAVGGVQTSR 358
Query: 551 INSDS 555
I S++
Sbjct: 359 IASEA 363
>At3g09000 unknown protein
Length = 541
Score = 38.1 bits (87), Expect = 0.014
Identities = 49/233 (21%), Positives = 89/233 (38%), Gaps = 29/233 (12%)
Query: 391 AASKTVIIREPSPER-PLKRTSAQQQQMSDSESSEDTW-----------KDSSSEETEDE 438
AA+ + + S +R PL+RT+A+ S++E S+ W K+S
Sbjct: 46 AAALSGVSETASSQRYPLRRTAAENFLYSENEKSDYDWLLTPPGTPQFEKESHRSVMNQH 105
Query: 439 DMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKR-RKLPLRG 497
D P R +L+ + N + I ++ + T S ++ ++R
Sbjct: 106 DAP--NSRPTVLKSR---------LGNCREDIVSGNNNKPQTSSSSVAGLRRPSSSGSSR 154
Query: 498 PLQQKETAAEQASEPTSEVQR----ERRSKRTSDPARAASLTRTSPIRSLSKVITESINS 553
+ T +++ PT+ R + R+S P A+LT S + T + +S
Sbjct: 155 STSRPATPTRRSTTPTTSTSRPVTTRASNSRSSTPTSRATLTAARATTSTAAPRTTTTSS 214
Query: 554 DSALVVIPEQPMPISTSLPTSTQ-TEPIQPETQPQTQAEPQAPKSPIQTATTA 605
SA P + P +S + + P P +P T P S + T+
Sbjct: 215 GSARSATPTRSNPRPSSASSKKPVSRPATPTRRPSTPTGPSIVSSKAPSRGTS 267
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.129 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,449,129
Number of Sequences: 26719
Number of extensions: 606331
Number of successful extensions: 4150
Number of sequences better than 10.0: 335
Number of HSP's better than 10.0 without gapping: 60
Number of HSP's successfully gapped in prelim test: 290
Number of HSP's that attempted gapping in prelim test: 3234
Number of HSP's gapped (non-prelim): 752
length of query: 607
length of database: 11,318,596
effective HSP length: 105
effective length of query: 502
effective length of database: 8,513,101
effective search space: 4273576702
effective search space used: 4273576702
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0234.12


