

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.1
(420 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
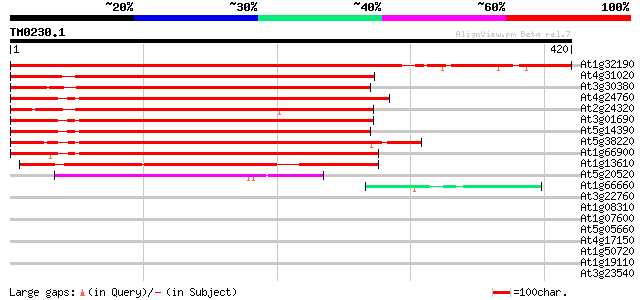
Score E
Sequences producing significant alignments: (bits) Value
At1g32190 unknown protein 576 e-165
At4g31020 unknown protein 348 3e-96
At3g30380 unknown protein 348 4e-96
At4g24760 unknown protein 343 8e-95
At2g24320 hypothetical protein 342 2e-94
At3g01690 unknown protein 336 2e-92
At5g14390 unknown protein 333 8e-92
At5g38220 unknown protein 332 2e-91
At1g66900 unknown protein 328 3e-90
At1g13610 hypothetical protein 274 6e-74
At5g20520 unknown protein 55 6e-08
At1g66660 hypothetical protein 42 6e-04
At3g22760 unknown protein 39 0.007
At1g08310 unknown protein 39 0.007
At1g07600 metallothionein-like protein 38 0.012
At5g05660 putative protein 37 0.027
At4g17150 unknown protein 37 0.027
At1g50720 hypothetical protein 36 0.035
At1g19110 unknown protein 36 0.035
At3g23540 unknown protein 35 0.059
>At1g32190 unknown protein
Length = 422
Score = 576 bits (1484), Expect = e-165
Identities = 286/431 (66%), Positives = 321/431 (74%), Gaps = 29/431 (6%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVP-IPHADDSSLDVLLVDT 59
MGCM S LAAKFAFFPPSPPTY L K DGKL+ VS+AS S P A D SLDV +V T
Sbjct: 1 MGCMFSHLAAKFAFFPPSPPTYHLTKTPDGKLSAVSSASSSSSTFPSAGDPSLDVKVVKT 60
Query: 60 KHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAST 119
+ GNK+ AFYL+NP ARLTLLYSHGNAADLGQL+DLFVQLKVNLRVNLMGYDYSGYGAST
Sbjct: 61 RRGNKVTAFYLRNPNARLTLLYSHGNAADLGQLFDLFVQLKVNLRVNLMGYDYSGYGAST 120
Query: 120 GKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSG 179
GKPSE TYADIEA YECL+++YGVGQED+ILYGQSVGSGPTLHLA+KLPRLRGVVLHSG
Sbjct: 121 GKPSEYDTYADIEAAYECLQTDYGVGQEDLILYGQSVGSGPTLHLASKLPRLRGVVLHSG 180
Query: 180 ILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESY 239
ILSGLRVLCHVKF FC DIY N+NKIKKVKCPVLVIHGTEDDVVNWLHGN LWKMA+E Y
Sbjct: 181 ILSGLRVLCHVKFKFCCDIYSNVNKIKKVKCPVLVIHGTEDDVVNWLHGNRLWKMAKEPY 240
Query: 240 DPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLKKLRQSQGPQSNSSSCFSCSC 299
+PLWIKGGGHCNLE+YPDYIRHL +FIQ+ME+ TT+ RLK + Q + S+ C
Sbjct: 241 EPLWIKGGGHCNLEIYPDYIRHLYRFIQDMENTTTKSRLKTIWQEIRRRDESTGC----- 295
Query: 300 CRMKCCSNNCTGCSNCSCINPEC----CNCSCINPECFNCSCINCSC-SLK-CTQCCCLP 353
CCS C +CSC P C C+C C C +C C CSC +LK C CC P
Sbjct: 296 ----CCSGLCR--PSCSCPKPRCPKPSCSCGC---GCGDCGCFKCSCPTLKGCFSCCKKP 346
Query: 354 SCIKCCHLPKF--STCFGSSCCTKCSRPSCCFKC--CCNLRCARPSCCCMSCFCWRCCVG 409
SC+ C P F S+CFG C KCS C+KC C + C R SCCC CF W CC G
Sbjct: 347 SCVSSCCCPTFKCSSCFGKPKCPKCS----CWKCLKCPDTECCRSSCCCSGCFSWLCCCG 402
Query: 410 KHSGRDEKQSG 420
++ ++ G
Sbjct: 403 GGRRKEGERRG 413
>At4g31020 unknown protein
Length = 294
Score = 348 bits (893), Expect = 3e-96
Identities = 166/274 (60%), Positives = 213/274 (77%), Gaps = 10/274 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD-GKLTVVSAASPSVPIPHADDSSLDVLLVDT 59
MG + S +AAKFAFFPP P TY + K D+ GKL ++ D +++V + T
Sbjct: 1 MGNVTSNVAAKFAFFPPEPATYGVTKDDETGKLVFAGVSA---------DKNVEVHQLTT 51
Query: 60 KHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAST 119
K GNK+VA + ++P+AR TLLYSHGNAADLGQ+ +LF++L+ +LRVN+M YDYSGYGAST
Sbjct: 52 KSGNKVVATFWRHPFARFTLLYSHGNAADLGQMVELFIELRAHLRVNIMSYDYSGYGAST 111
Query: 120 GKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSG 179
GKPSE +TY DIEA+Y CL S+YG+ QE+IILYGQSVGSGPTLH+A++L RLRGVVLHS
Sbjct: 112 GKPSEFNTYYDIEAVYSCLRSDYGIKQEEIILYGQSVGSGPTLHMASRLKRLRGVVLHSA 171
Query: 180 ILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESY 239
ILSG+RVL VK + FDI+KNI+KI+ V VLVIHGT D++V+ HG LW++A+E Y
Sbjct: 172 ILSGIRVLYPVKMTLWFDIFKNIDKIRHVNSQVLVIHGTNDEIVDLSHGKRLWELAKEKY 231
Query: 240 DPLWIKGGGHCNLELYPDYIRHLCKFIQEMESIT 273
DPLW+KGGGHCNLE YP+YI+HL KF+ ME ++
Sbjct: 232 DPLWVKGGGHCNLETYPEYIKHLKKFVNAMEKLS 265
>At3g30380 unknown protein
Length = 399
Score = 348 bits (892), Expect = 4e-96
Identities = 165/270 (61%), Positives = 212/270 (78%), Gaps = 9/270 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPP+PP+Y ++ + GKL ++ + +++VL + TK
Sbjct: 1 MGAVTSSMAAKFAFFPPNPPSYGVEVVE-GKLRLIGVENVK--------ENVEVLKLKTK 51
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
GN++VA Y+KNP A LTLLYSHGNAADLGQ+++LF +L ++LRVNL+GYDYSGYG S+G
Sbjct: 52 RGNQVVAAYIKNPTASLTLLYSHGNAADLGQMFELFSELSLHLRVNLIGYDYSGYGRSSG 111
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KPSE +TY+DIEA+Y CLE +YGV ++D+ILYGQSVGSGPTL LA++LP LR VVLHS I
Sbjct: 112 KPSEQNTYSDIEAVYRCLEEKYGVKEQDVILYGQSVGSGPTLELASRLPNLRAVVLHSAI 171
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
SGLRV+ VK ++ FDIYKN+ KI VKCPVLVIHGT DDVVNW HG L+++ +E Y+
Sbjct: 172 ASGLRVMYPVKRTYWFDIYKNVEKISFVKCPVLVIHGTSDDVVNWSHGKQLFELCKEKYE 231
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEME 270
PLWIKGG HC+LELYP YI+HL KF+ +E
Sbjct: 232 PLWIKGGNHCDLELYPQYIKHLRKFVSAIE 261
>At4g24760 unknown protein
Length = 365
Score = 343 bits (881), Expect = 8e-95
Identities = 164/284 (57%), Positives = 219/284 (76%), Gaps = 8/284 (2%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAK AFFPP+PP+Y+L + + +L ++S P PH ++ +D+L + T+
Sbjct: 1 MGGVTSSMAAKLAFFPPNPPSYKLVRDETTELFLMS------PFPHREN--VDILRLPTR 52
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
G +IVA Y++ P A TLLYSHGNAAD+GQ+Y+LF++L ++LRVNLMGYDYSGYG S+G
Sbjct: 53 RGTEIVAMYIRYPMAVTTLLYSHGNAADIGQMYELFIELSIHLRVNLMGYDYSGYGQSSG 112
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KP+E +TYADIEA Y+CLE YG QE+IILYGQSVGSGPT+ LAA+LPRLR +LHS I
Sbjct: 113 KPTEQNTYADIEAAYKCLEENYGAKQENIILYGQSVGSGPTVDLAARLPRLRASILHSPI 172
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
LSGLRV+ VK ++ FDIYKNI+KI V+CPVLVIHGT DDVV++ HG LW++ +E Y+
Sbjct: 173 LSGLRVMYPVKRTYWFDIYKNIDKITLVRCPVLVIHGTADDVVDFSHGKQLWELCQEKYE 232
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLKKLRQS 284
PLW+KGG HC+LEL+P+YI HL KF+ +E +++ R+S
Sbjct: 233 PLWLKGGNHCDLELFPEYIGHLKKFVSAVEKSASKRNSSFSRRS 276
>At2g24320 hypothetical protein
Length = 316
Score = 342 bits (878), Expect = 2e-94
Identities = 166/295 (56%), Positives = 216/295 (72%), Gaps = 32/295 (10%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPP PPTY + K ++ + + +P + S+DV + TK
Sbjct: 1 MGNVTSNMAAKFAFFPP-PPTYDVGKDEETGKLMFTGITP--------EKSMDVHQLTTK 51
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
GNK++A + K+P++R TLLYSHGNAADLGQ+ DLF++L+ +LRVN+M YDYSGYGASTG
Sbjct: 52 SGNKVIATFWKHPFSRFTLLYSHGNAADLGQMVDLFIELRAHLRVNIMSYDYSGYGASTG 111
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KP+E +TY DIEA+Y CL +EYG+ QE++ILYGQSVGSGPTLHLA+++ RLRG+VLHS I
Sbjct: 112 KPTELNTYYDIEAVYNCLRTEYGIMQEEMILYGQSVGSGPTLHLASRVKRLRGIVLHSAI 171
Query: 181 LSGLRVLCHVKFSFCFDIYK-----------------------NINKIKKVKCPVLVIHG 217
LSGLRVL VK +F FD+YK NI+KI+ V CPVLVIHG
Sbjct: 172 LSGLRVLYPVKMTFWFDMYKVSLISLVSGYYYRVSLSNSGILQNIDKIRHVTCPVLVIHG 231
Query: 218 TEDDVVNWLHGNGLWKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESI 272
T+DD+VN HG LW++A++ YDPLW+KGGGHCNLE YP+YI+H+ KF+ ME +
Sbjct: 232 TKDDIVNMSHGKRLWELAKDKYDPLWVKGGGHCNLETYPEYIKHMRKFMNAMEKL 286
>At3g01690 unknown protein
Length = 361
Score = 336 bits (861), Expect = 2e-92
Identities = 160/272 (58%), Positives = 212/272 (77%), Gaps = 8/272 (2%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPPSPP+Y++ + L ++S P PH ++ ++++ + T+
Sbjct: 1 MGGVTSSVAAKFAFFPPSPPSYKVVTDELTGLLLLS------PFPHREN--VEIVKLRTR 52
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
G +IV Y+++P A TLLYSHGNAADLGQ+Y+LF++L ++L+VNLMGYDYSGYG STG
Sbjct: 53 RGTEIVGMYVRHPMATSTLLYSHGNAADLGQMYELFIELSIHLKVNLMGYDYSGYGQSTG 112
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KPSE +TYADIEA+Y+CLE +G QE +ILYGQSVGSGPTL LA++LP+LR VVLHS I
Sbjct: 113 KPSEHNTYADIEAVYKCLEETFGSKQEGVILYGQSVGSGPTLDLASRLPQLRAVVLHSPI 172
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
LSGLRV+ VK ++ FDIYKNI+KI V CPVL+IHGT D+VV+ HG LW++ ++ Y+
Sbjct: 173 LSGLRVMYSVKKTYWFDIYKNIDKIPYVDCPVLIIHGTSDEVVDCSHGKQLWELCKDKYE 232
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEMESI 272
PLW+KGG HC+LE YP+YIRHL KFI +E +
Sbjct: 233 PLWVKGGNHCDLEHYPEYIRHLKKFIATVERL 264
>At5g14390 unknown protein
Length = 369
Score = 333 bits (855), Expect = 8e-92
Identities = 163/270 (60%), Positives = 210/270 (77%), Gaps = 8/270 (2%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPPSP +Y+L + L +++ P PH ++ +++L + T+
Sbjct: 1 MGGVTSSVAAKFAFFPPSPSSYKLVYDELTGLLLMN------PFPHREN--VEILKLPTR 52
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
G +IVA Y+++P A TLLYSHGNAADLGQ+Y+LF++L ++L+VNLMGYDYSGYG STG
Sbjct: 53 RGTEIVAMYVRHPMATSTLLYSHGNAADLGQMYELFIELSIHLKVNLMGYDYSGYGQSTG 112
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KPSE TYADIEA Y+CLE YG QEDIILYGQSVGSGPTL LAA+LP+LR VLHS I
Sbjct: 113 KPSEHHTYADIEAAYKCLEETYGAKQEDIILYGQSVGSGPTLDLAARLPQLRAAVLHSPI 172
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
LSGLRV+ VK ++ FDI+KNI+KI V CPVLVIHGT D+VV+ HG LW++++E Y+
Sbjct: 173 LSGLRVMYPVKKTYWFDIFKNIDKIPLVNCPVLVIHGTCDEVVDCSHGKQLWELSKEKYE 232
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEME 270
PLW++GG HC+LE YP+YI+HL KFI +E
Sbjct: 233 PLWLEGGNHCDLEHYPEYIKHLKKFITTVE 262
>At5g38220 unknown protein
Length = 336
Score = 332 bits (852), Expect = 2e-91
Identities = 169/317 (53%), Positives = 222/317 (69%), Gaps = 22/317 (6%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPPSPP+Y D +L + +P DD +DVL + T+
Sbjct: 1 MGGVTSSIAAKFAFFPPSPPSYGFVS-DVDRLYITE-------VPRRDD--VDVLKLKTR 50
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
GN+IVA Y+K+P A TLLYSHGNAADLGQ+++LF++L LR+NLMGYDYSGYG STG
Sbjct: 51 RGNEIVAIYIKHPKANGTLLYSHGNAADLGQMFELFIELSNRLRLNLMGYDYSGYGQSTG 110
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
K SE +TYADI+A Y CL+ YGV + +ILYGQSVGSGPT+ LA++ P LRGVVLHS I
Sbjct: 111 KASECNTYADIDAAYTCLKEHYGVKDDQLILYGQSVGSGPTIDLASRTPNLRGVVLHSPI 170
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
LSG+RVL VK ++ FDIYKNI+KI V CPVLVIHGT D+VV+ HG LW++++E Y+
Sbjct: 171 LSGMRVLYPVKRTYWFDIYKNIDKIGAVTCPVLVIHGTADEVVDCSHGKQLWELSKEKYE 230
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEM---------ESITTEKRLKKLRQSQGPQSNS 291
PLW+ GGGHCNLELYP++I+HL K++ + ++ TT+ K QS+ ++
Sbjct: 231 PLWVSGGGHCNLELYPEFIKHLKKYVISISKGPRTGSNKTATTDAAKK---QSKPAENGR 287
Query: 292 SSCFSCSCCRMKCCSNN 308
+ F CC + N+
Sbjct: 288 ADTFQLGCCLPEVSRNS 304
>At1g66900 unknown protein
Length = 272
Score = 328 bits (842), Expect = 3e-90
Identities = 163/278 (58%), Positives = 210/278 (74%), Gaps = 11/278 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD--GKLTVVSAASPSVPIPHADDSSLDVLLVD 58
MG + S +AAKFAFFPP+PP+Y++ D G+L + IP DD +D+L +
Sbjct: 1 MGGVTSSIAAKFAFFPPTPPSYEVIADDSCGGRLYIPE-------IPRRDD--VDILKLR 51
Query: 59 TKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAS 118
T+ GN+IVA Y+K+ A TLLYSHGNAADLGQ+++LFV+L LRVNLMGYDYSGYG S
Sbjct: 52 TRCGNEIVAVYVKHSKANGTLLYSHGNAADLGQMFELFVELSNRLRVNLMGYDYSGYGQS 111
Query: 119 TGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHS 178
TG+ SE +TYADIEA Y+CL+ +YGV + +I+YGQSVGSGPT+ LA++ P LRGVVL
Sbjct: 112 TGQASECNTYADIEASYKCLKEKYGVKDDQLIVYGQSVGSGPTVDLASRTPNLRGVVLQC 171
Query: 179 GILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARES 238
ILSG+RVL VK ++ FDIYKNI+KI V CPVLVIHGT D+VV+W HG LW++++E
Sbjct: 172 PILSGMRVLYPVKCTYWFDIYKNIDKIGSVTCPVLVIHGTADEVVDWSHGKRLWELSKEK 231
Query: 239 YDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEK 276
Y+PLWI GGGHC+LELYPD+IRHL KF+ + + E+
Sbjct: 232 YEPLWISGGGHCDLELYPDFIRHLKKFVVSLGNKQAEQ 269
>At1g13610 hypothetical protein
Length = 335
Score = 274 bits (701), Expect = 6e-74
Identities = 142/271 (52%), Positives = 184/271 (67%), Gaps = 25/271 (9%)
Query: 8 LAAKFAFFPPSPPTYQLKKRDD-GKLTVVSAASPSVPIPH-ADDSSLDVLLVDTKHGNKI 65
+AAK AFFPP+PP+Y + + GK+ + S +PH D +++V+ + TK GN+I
Sbjct: 1 MAAKLAFFPPNPPSYTVVTEESTGKMMI------STNLPHYLRDENIEVVKIRTKRGNEI 54
Query: 66 VAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTGKPSES 125
VA Y+KNP A+LT+L+SHGNA+DL Q++ + +L + L VNLMGYDYSGYG S+GKPSE
Sbjct: 55 VAMYVKNPTAKLTVLFSHGNASDLAQIFYILAEL-IQLNVNLMGYDYSGYGQSSGKPSEQ 113
Query: 126 STYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGILSGLR 185
TYADIEA Y L YG E IILYGQSVGSGP+L LA++LPRLR +VLHS LSGLR
Sbjct: 114 DTYADIEAAYNWLRQTYGTKDERIILYGQSVGSGPSLELASRLPRLRALVLHSPFLSGLR 173
Query: 186 VLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYDPLWIK 245
V+ VK SF FDIYK GT+DDVVN HG LW + +E Y+PLW+K
Sbjct: 174 VMYPVKHSFPFDIYK----------------GTDDDVVNISHGKHLWGLCKEKYEPLWLK 217
Query: 246 GGGHCNLELYPDYIRHLCKFIQEMESITTEK 276
G GH ++E+ P+Y+ HL KFI +E + K
Sbjct: 218 GRGHSDIEMSPEYLPHLRKFISAIEKLPVPK 248
>At5g20520 unknown protein
Length = 308
Score = 55.5 bits (132), Expect = 6e-08
Identities = 55/225 (24%), Positives = 102/225 (44%), Gaps = 24/225 (10%)
Query: 34 VVSAASPSVPIPHADDSSL-DVLLVDTKHGNKIVAFYLKN-PYAR-LTLLYSHGNAADLG 90
V+ S S PI A + + + + + + G ++ A+++K P R T+L+ NA ++
Sbjct: 35 VLPGLSKSYPITPARLNLIYEDIWLQSSDGVRLHAWFIKMFPECRGPTILFFQENAGNIA 94
Query: 91 QLYDLFVQLKVNLRVNLMGYDYSGYGASTGKPSESSTYADIEAIYECLESEYGVGQEDII 150
++ + L+ N+ Y GYGAS G PS+ D +A + L + I+
Sbjct: 95 HRLEMVRIMIQKLKCNVFMLSYRGYGASEGYPSQQGIIKDAQAALDHLSGRTDIDTSRIV 154
Query: 151 LYGQSVGSGPTLHLAAKLP-RLRGVVLH---------SGIL----------SGLRVLCHV 190
++G+S+G L P ++ ++L +G+L SG + L +
Sbjct: 155 VFGRSLGGAVGAVLTKNNPDKVSALILENTFTSILDMAGVLLPFLKWFIGGSGTKSLKLL 214
Query: 191 KFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMA 235
F +K I+ I ++K PVL + G +D++V H L+ A
Sbjct: 215 NF-VVRSPWKTIDAIAEIKQPVLFLSGLQDEMVPPFHMKMLYAKA 258
>At1g66660 hypothetical protein
Length = 412
Score = 42.0 bits (97), Expect = 6e-04
Identities = 34/138 (24%), Positives = 56/138 (39%), Gaps = 20/138 (14%)
Query: 267 QEMESITTEKRLKKLRQSQGPQSNSSSCFSC-SCCR-----MKCCSNNCTGCSNCSCINP 320
+E E++TT+++ + SQ + SS C +CC + CSN CS+C
Sbjct: 140 EEEENVTTDEQSGSPKSSQPVKLQSSDVLDCPTCCEPLKRPIYQCSNGHLACSSC----- 194
Query: 321 ECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPS 380
C +N +C C C++ +C + I+ +P + G T S
Sbjct: 195 ----CQKLNKKCSF-----CRCNIGDIRCRAMEKVIEASIVPCPNAKHGCKETTTYCNQS 245
Query: 381 CCFKCCCNLRCARPSCCC 398
K C +RC+ P C
Sbjct: 246 SHEKVCKFVRCSCPVSNC 263
>At3g22760 unknown protein
Length = 609
Score = 38.5 bits (88), Expect = 0.007
Identities = 23/57 (40%), Positives = 28/57 (48%), Gaps = 7/57 (12%)
Query: 279 KKLRQSQGPQSNSSSCFSCSCCRMKCCSNNC-TGCSNCSCINPECCNCSCINPECFN 334
KK R+S+ SSC C+C + KC C + CI P CSCIN CFN
Sbjct: 312 KKRRKSEQSGEGDSSCKRCNCKKSKCLKLYCECFAAGFYCIEP----CSCIN--CFN 362
>At1g08310 unknown protein
Length = 315
Score = 38.5 bits (88), Expect = 0.007
Identities = 33/108 (30%), Positives = 57/108 (52%), Gaps = 6/108 (5%)
Query: 76 RLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTGKPSESSTYADIEAIY 135
++ L++ G++ D+ + +L L V L+ YD SGYGAS ++ S +++E I
Sbjct: 34 KIILVHGFGSSKDMN--FSASKELIEELEVYLLFYDRSGYGASDSN-TKRSLESEVEDIA 90
Query: 136 ECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLP-RLRGVVLHSGILS 182
E L + +G + L G S+GS PT +P RL GV + +++
Sbjct: 91 E-LADQLELGPK-FYLIGISMGSYPTWGCLRHIPHRLSGVAFVAPVVN 136
>At1g07600 metallothionein-like protein
Length = 45
Score = 37.7 bits (86), Expect = 0.012
Identities = 20/40 (50%), Positives = 22/40 (55%), Gaps = 6/40 (15%)
Query: 313 SNCSCINPECC--NCSC---INPECFNCSC-INCSCSLKC 346
SNC C + C +CSC N EC NCSC NCSC C
Sbjct: 4 SNCGCGSSCKCGDSCSCEKNYNKECDNCSCGSNCSCGSNC 43
Score = 30.8 bits (68), Expect = 1.5
Identities = 19/50 (38%), Positives = 22/50 (44%), Gaps = 9/50 (18%)
Query: 289 SNSSSCFSCSCCRMKCCSNNCTG-CSNCSCINPECCNCSCINPECFNCSC 337
SN SC C C N C NCSC + NCSC + NC+C
Sbjct: 4 SNCGCGSSCKCGDSCSCEKNYNKECDNCSCGS----NCSCGS----NCNC 45
>At5g05660 putative protein
Length = 820
Score = 36.6 bits (83), Expect = 0.027
Identities = 33/141 (23%), Positives = 48/141 (33%), Gaps = 22/141 (15%)
Query: 296 SCSCCRMKCCSNNCTGCSNCSCINPECCNC-SCINPECF--------NCSCINCSCSLKC 346
SC + SNNC C C C +C + +CF C SC C
Sbjct: 185 SCGEVCERPLSNNCGHCCLLLCHPGPCASCPKLVKAKCFCGGVEDVRRCGHKQFSCGDVC 244
Query: 347 TQC--CCLPSCIKCCHLPKFSTC-----FGSSCCTKCSRPSCC---FKC---CCNLRCAR 393
+ C + +C + CH + C + SC CC F+C C N+
Sbjct: 245 ERVLDCNIHNCREICHDGECPPCRERAVYKCSCGKVKEEKDCCERVFRCEASCENMLNCG 304
Query: 394 PSCCCMSCFCWRCCVGKHSGR 414
C C C + + G+
Sbjct: 305 KHVCERGCHAGECGLCPYQGK 325
Score = 29.3 bits (64), Expect = 4.2
Identities = 29/113 (25%), Positives = 36/113 (31%), Gaps = 16/113 (14%)
Query: 287 PQSNSSSCFSCSCCRMK----CCSN--NC-TGCSNCSCINPECCNCSCINPECFNC---S 336
P + + CSC ++K CC C C N C C EC C
Sbjct: 265 PPCRERAVYKCSCGKVKEEKDCCERVFRCEASCENMLNCGKHVCERGCHAGECGLCPYQG 324
Query: 337 CINCSCSLKCTQ---C-CCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKC 385
+C C + Q C P C C K C C +C R C C
Sbjct: 325 KRSCPCGKRFYQGLSCDVVAPLCGGTC--DKVLGCGYHRCPERCHRGPCLETC 375
>At4g17150 unknown protein
Length = 502
Score = 36.6 bits (83), Expect = 0.027
Identities = 37/142 (26%), Positives = 65/142 (45%), Gaps = 23/142 (16%)
Query: 79 LLYSHGNA---ADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTGKPSESSTYA--DIEA 133
++Y HGN+ AD + + V L N+ V + D+SG G S G + D++
Sbjct: 78 VIYCHGNSGCRADANEA--VMVLLPSNITVFTL--DFSGSGLSEGDYVSLGWHEKDDLKT 133
Query: 134 IYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGILSGLRVLCHVKFS 193
+ L + V + I L+G+S+G+ +L A+ P + G+VL S FS
Sbjct: 134 VVSYLRNSNQVSR--IGLWGRSMGAVTSLLYGAEDPSIAGMVLDSA------------FS 179
Query: 194 FCFDIYKNINKIKKVKCPVLVI 215
FD+ + + K++ P +
Sbjct: 180 NLFDLMMELVDVYKIRLPKFTV 201
>At1g50720 hypothetical protein
Length = 154
Score = 36.2 bits (82), Expect = 0.035
Identities = 18/77 (23%), Positives = 32/77 (41%)
Query: 279 KKLRQSQGPQSNSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCI 338
+++ +SQG +++ +C S C + ++NC C N CC C+ C
Sbjct: 75 EEICKSQGMYNSTMACCSNKCVDLAYDNDNCGACKNQCKFTQTCCRGECVYLAYDKRHCG 134
Query: 339 NCSCSLKCTQCCCLPSC 355
C+ S + C C
Sbjct: 135 ECNHSCLVGEFCVYGLC 151
Score = 31.6 bits (70), Expect = 0.86
Identities = 12/35 (34%), Positives = 17/35 (48%)
Query: 322 CCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCI 356
CC+ C++ N +C C K TQ CC C+
Sbjct: 90 CCSNKCVDLAYDNDNCGACKNQCKFTQTCCRGECV 124
>At1g19110 unknown protein
Length = 754
Score = 36.2 bits (82), Expect = 0.035
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 10/64 (15%)
Query: 331 ECFNCSCINCSCSLKCTQCCCLPSCIKCCH------LPKFSTCFGSSCCTKCSRPSCCFK 384
E F + +C SL C +CCC+ C++CC + F+ F + C C CC
Sbjct: 691 EKFVKAASSCCVSL-CNKCCCM-CCVQCCSKLNDQCVLVFTQLFTAIACIACF--ECCST 746
Query: 385 CCCN 388
CC+
Sbjct: 747 VCCS 750
Score = 32.3 bits (72), Expect = 0.50
Identities = 22/71 (30%), Positives = 29/71 (39%), Gaps = 20/71 (28%)
Query: 304 CCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPK 363
CC + C C CC C C+ +C CS +N C L TQ +CI C
Sbjct: 700 CCVSLCNKC---------CCMC-CV--QC--CSKLNDQCVLVFTQLFTAIACIACFE--- 742
Query: 364 FSTCFGSSCCT 374
C + CC+
Sbjct: 743 ---CCSTVCCS 750
>At3g23540 unknown protein
Length = 568
Score = 35.4 bits (80), Expect = 0.059
Identities = 30/102 (29%), Positives = 52/102 (50%), Gaps = 8/102 (7%)
Query: 79 LLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTGKPSES--STYADIEAIYE 136
++Y HGN AD + V L N+ V + D+SG G S G+ + D++A+ E
Sbjct: 68 VIYCHGNRADGSEA--AIVLLPSNITVFTL--DFSGSGLSGGEHVTLGWNEKDDLKAVVE 123
Query: 137 CLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHS 178
L + + I L+G+S+G+ +L + P + G++L S
Sbjct: 124 FLRQDGNISL--IGLWGRSMGAVTSLMYGVEDPSIAGMILDS 163
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.139 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,702,429
Number of Sequences: 26719
Number of extensions: 509098
Number of successful extensions: 2197
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 61
Number of HSP's that attempted gapping in prelim test: 1987
Number of HSP's gapped (non-prelim): 173
length of query: 420
length of database: 11,318,596
effective HSP length: 102
effective length of query: 318
effective length of database: 8,593,258
effective search space: 2732656044
effective search space used: 2732656044
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0230.1


