

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0228a.2
(510 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
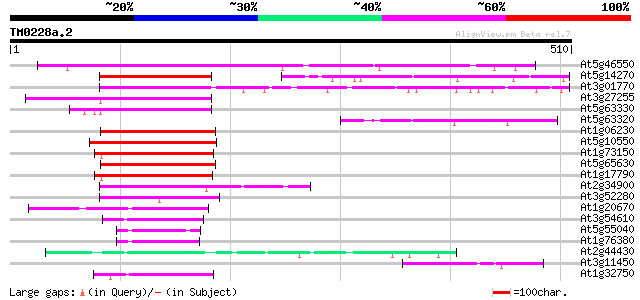
Score E
Sequences producing significant alignments: (bits) Value
At5g46550 unknown protein 241 9e-64
At5g14270 kinase - like protein 140 2e-33
At3g01770 unknown protein 135 5e-32
At3g27255 unknown protein 132 6e-31
At5g63330 unknown protein 120 1e-27
At5g63320 putative protein 120 2e-27
At1g06230 Ring3-like bromodomain protein 116 3e-26
At5g10550 bromodomain protein - like 103 2e-22
At1g73150 unknown protein 103 2e-22
At5g65630 unknown protein 102 5e-22
At1g17790 unknown protein (At1g17790) 98 9e-21
At2g34900 RING3 protein-like 92 5e-19
At3g52280 putative protein 75 1e-13
At1g20670 hypothetical protein 63 3e-10
At3g54610 histone acetyltransferase (HAT1) 62 8e-10
At5g55040 putative protein 56 4e-08
At1g76380 unknown protein 56 5e-08
At2g44430 unknown protein 51 2e-06
At3g11450 putative cell division related protein 47 2e-05
At1g32750 unknown protein 46 6e-05
>At5g46550 unknown protein
Length = 494
Score = 241 bits (614), Expect = 9e-64
Identities = 168/486 (34%), Positives = 256/486 (52%), Gaps = 42/486 (8%)
Query: 26 PGQKCEFGQRGLQTDENGRCNSAEKL----------FKPDSIKRGPPEGF-EGQKEKRQK 74
P K +FG +G +S++K+ S KRG P+ E Q +K+Q+
Sbjct: 5 PNIKIKFGPQGSVRTFQTLSDSSKKIEHVVTEDLSQSSEKSKKRGGPKELDEVQPKKKQR 64
Query: 75 IDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNV 134
+D S+QC A+L+ LM + W+F +PVDPV + IPDYF +I KPMDLGT+ SKL KNV
Sbjct: 65 LDCDWSSQCLALLRFLMEHRGGWLFKEPVDPVKMEIPDYFNVIQKPMDLGTVKSKLLKNV 124
Query: 135 YSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSV 194
YS +EFAADVRLTF+NAM YNP N+VH +AKE+N+IF+ +W+ + KK +
Sbjct: 125 YSNADEFAADVRLTFANAMHYNPLWNEVHTIAKEINEIFEVRWESLMKKKVLRLSWNEVR 184
Query: 195 TGTIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKISSLAARDAKVEVPK-------- 246
G ++ + C+ K + SL+++ KV+ K
Sbjct: 185 EGYKRQPVERDCSRRSSTGTSASSGVGLTKPAKENSEKGSLSSKPVKVQSKKNTPAVTPK 244
Query: 247 -LSQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSPLERKS-DPDSDGAVSSLD 304
L+ C I +K V Q+K+ S + DP S+G+ +S
Sbjct: 245 ALATCKCGRIICICLKSCSSFG----SDVCSLTDCQLKNISGAQASELDPQSNGSDTSKK 300
Query: 305 SEHVYPGSQHATPATDASSGEVWSTPDFPV--QLSPKKALRAAMLKSRFADTILKAQQKT 362
+ SQ P+ G T FP + P+KALRAA+LK+++A TI+KA+ +
Sbjct: 301 ERNGSLKSQLDKPSNSDLLGNELKTA-FPALPPVPPEKALRAAILKAQYAGTIIKAKHRI 359
Query: 363 LLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQREKEREA 422
+L +K D +++Q+EKE++ER QREE+ARIEA+++ A+ A R RA++EL+Q+RE +
Sbjct: 360 VLGQNNKADLIRIQIEKEQMERAQREEKARIEAEMRAAKVAERMRAQDELKQKRESQ--- 416
Query: 423 ARAAIEEMKRTVEIEHN--LEIVKELETLSGCTLSYKA--------LGGRNDYKVALETL 472
R I +MK+ + E N ++ K+ + GC KA L +NDY LE +
Sbjct: 417 -RLEIAKMKKGFDFERNNHSKLKKKFVKVCGCFSLTKARLLLEELGLVLKNDYCPELEVI 475
Query: 473 DKLQFE 478
+F+
Sbjct: 476 GSEKFD 481
>At5g14270 kinase - like protein
Length = 688
Score = 140 bits (352), Expect = 2e-33
Identities = 65/102 (63%), Positives = 77/102 (74%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC A+LK LMS+ Y WVFN PVD V LNI DYF +I PMDLGT+ +KL YSC EF
Sbjct: 140 QCEALLKRLMSHQYGWVFNTPVDVVKLNILDYFNVIEHPMDLGTVKNKLTSGTYSCPSEF 199
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
AADVRLTFSNAMTYNPP NDV++MA L + F+ +WK ++KK
Sbjct: 200 AADVRLTFSNAMTYNPPGNDVYVMADTLRKFFEVRWKTLEKK 241
Score = 119 bits (297), Expect = 5e-27
Identities = 96/290 (33%), Positives = 155/290 (53%), Gaps = 35/290 (12%)
Query: 248 SQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSPLERKS------DPDSDGAVS 301
S + +IEKDL+ G+ + + G+V+ K S+ P K +P DGA S
Sbjct: 405 SSISPVTIEKDLVLGNSNGNSL--GSVSGDPKM---SSLPRASKGLGTIDLEPMLDGATS 459
Query: 302 SLDSEHVYPGS----QHATP----ATDASSGEVWSTPDFPVQLSPKKALRAAMLKSRFAD 353
+ + G + A+P + +A + ++ QL P+K+ RAA+LK+RFAD
Sbjct: 460 ASPTRGSSVGGLDQLESASPEKISSVEADCQQDGNSAQNEKQLPPEKSYRAAILKNRFAD 519
Query: 354 TILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAAR-------A 406
ILKA++K L ++ D DP K+Q E+E LE +++E+AR++A+ K AE A R A
Sbjct: 520 IILKAREKPLNQN-DTRDPEKLQREREELELQKKKEKARLQAEAKAAEEARRKAEAQAAA 578
Query: 407 RAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSY--KALGGRND 464
A E +++ E EREAAR A+ EM+++VE+ N + +++LE L + + +
Sbjct: 579 EAAAEAKRKLELEREAARQALMEMEQSVELNENAKFLEDLELLKTVDTDHLTNTIEEEDG 638
Query: 465 YKVALETLDKLQFENPLEQLGLFIKDELADEDEEVING-----TVEEGEI 509
V L + NPLEQLGLF+K + +E+ + + +EEGEI
Sbjct: 639 PDVGLRSF-SFGGSNPLEQLGLFMKQDEDEEEADPLTSPAPEIDIEEGEI 687
>At3g01770 unknown protein
Length = 620
Score = 135 bits (340), Expect = 5e-32
Identities = 143/505 (28%), Positives = 209/505 (41%), Gaps = 94/505 (18%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC ++LK LMS + W+FN PVD V LNIPDYF II PMDLGT+ SKL YS EF
Sbjct: 132 QCESLLKRLMSQQHCWLFNTPVDVVKLNIPDYFTIIKHPMDLGTVKSKLTSGTYSSPSEF 191
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKET 201
+ADVRLTF NAMTYNP N+V+ A L++ F+ +WK ++KK KS +
Sbjct: 192 SADVRLTFRNAMTYNPSDNNVYRFADTLSKFFEVRWKTIEKK----SSGTKSEPSNLATL 247
Query: 202 ARKSCNVMHP----RQKDTFPKKSQVSEHKRVL----------KISSLAARDAKVEVPKL 247
A K + P R+ + + S + KRV+ + SL + V++
Sbjct: 248 AHKDIAIPEPVAKKRKMNAVKRNSLLEPAKRVMTDEDRVKLGRDLGSLT--EFPVQIINF 305
Query: 248 SQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSP--LERKSDPDSDGAVSSLDS 305
+ H E + ++ I ++ Q++ L DS+G L+
Sbjct: 306 LRDHSSKEE----RSGDDEIEIDINDLSHDALFQLRDLFDEFLRENQKKDSNGEPCVLEL 361
Query: 306 EHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLKSRFADTILKAQQK---- 361
H GS T G D + + +++ D++ Q
Sbjct: 362 LH---GSGPGNSLTQHCDGSELEDEDVDIGNYEHPISHISTVRTE-KDSVGGLNQMEDAS 417
Query: 362 ----TLLEHGD-----KGDPLKMQLEKERLERIQREERARIEAQIKTAEAAA---RARAE 409
+L+E D P + +L E+ R + + +K E R
Sbjct: 418 RGKLSLIEGADGHQDGNSAPKEKELPPEKRYRAALLKNRFADIILKAQEITLNQNEKRDP 477
Query: 410 EELRQQRE-----KEREAAR--------------AAIEEMKRTVEIE------------- 437
E L++++E K++E AR A +E KR +E+E
Sbjct: 478 ETLQREKEELELQKKKEKARLQAEAKEAEEARRKAEAQEAKRKLELEREAARQALLEMEK 537
Query: 438 -----HNLEIVKELETLSGCTLSYKALGGRNDYKVALETLDKLQF--ENPLEQLGLFIKD 490
N +K+LE L T++ L D + L F NPLEQLGLF+K
Sbjct: 538 SVEINENTRFLKDLELLK--TVNTDQLRNLRDVGSESDGLAVFGFGGSNPLEQLGLFMKH 595
Query: 491 ELADEDEEVI------NGTVEEGEI 509
E DEDE + VEEGEI
Sbjct: 596 E-EDEDESDMLAFPDPGNEVEEGEI 619
>At3g27255 unknown protein
Length = 503
Score = 132 bits (331), Expect = 6e-31
Identities = 70/174 (40%), Positives = 97/174 (55%), Gaps = 5/174 (2%)
Query: 15 IKFSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKLFKPDSIKRGPPEGFEGQKEKRQK 74
+ ++ R+ GQK + + S +K+ + RG G G+ E ++
Sbjct: 107 VSSTSDRVGFSTGQKISSRVSNSKKPSDFAVGSGKKVRHQNGTSRGWNRGTSGKFESSKE 166
Query: 75 IDRKGST-----QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSK 129
QC +L+ L S+P+SWVF PVD V LNIPDY I PMDLGT+
Sbjct: 167 TMTSTPNITLMKQCDTLLRKLWSHPHSWVFQAPVDVVKLNIPDYLTTIKHPMDLGTVKKN 226
Query: 130 LKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
L VYS EFAADVRLTF+NAMTYNPP +DVH+M L+++F+++WK + KK
Sbjct: 227 LASGVYSSPHEFAADVRLTFTNAMTYNPPGHDVHIMGDILSKLFEARWKTIKKK 280
>At5g63330 unknown protein
Length = 477
Score = 120 bits (302), Expect = 1e-27
Identities = 67/154 (43%), Positives = 88/154 (56%), Gaps = 25/154 (16%)
Query: 55 DSIKRGPPEGFE----GQKEKRQKI------DRKGST---------------QCAAILKC 89
D +R PPE F Q +KR + ++KG + +C +L
Sbjct: 112 DGPRRPPPENFATFVGSQGKKRPPVRSDKQRNKKGPSRLNVPTSYTVASVMKECETLLNR 171
Query: 90 LMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTF 149
L S+ W F PVDPV LNIPDYF +I PMDLGTI S+L K YS +FAADVRLTF
Sbjct: 172 LWSHKSGWPFRTPVDPVMLNIPDYFNVIKHPMDLGTIRSRLCKGEYSSPLDFAADVRLTF 231
Query: 150 SNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
SN++ YNPP N H MA+ +++ F+S WK ++KK
Sbjct: 232 SNSIAYNPPGNQFHTMAQGISKYFESGWKSIEKK 265
>At5g63320 putative protein
Length = 569
Score = 120 bits (301), Expect = 2e-27
Identities = 78/210 (37%), Positives = 125/210 (59%), Gaps = 25/210 (11%)
Query: 301 SSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLKSRFADTILKAQQ 360
+++D+ + P + A P S PD SP K RAA LK+RFADTI+KA++
Sbjct: 38 TTMDAVVLVPDEETAPPERQIS-------PD-----SPDKRYRAAFLKNRFADTIMKARE 85
Query: 361 KTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAA---ARARAEEELRQQRE 417
K + G+KGDP K+++E+E E+ REE+ R++A+ K AE A A+A A E+ R++RE
Sbjct: 86 KAFTK-GEKGDPEKLRIEREEFEKRLREEKERLQAEAKAAEEARRKAKAEAAEKARRERE 144
Query: 418 KEREAARAAIEEMKRTVEIEHNLEIVKELETLSG-------CTLSYKALGGR-NDYKVAL 469
+EREAAR A+++M++TVEI + +++L+ L S + + + ++ + L
Sbjct: 145 QEREAARQALQKMEKTVEINEGIRFMEDLQMLRATGTEGDQLPTSMEVMSPKFSEDMLGL 204
Query: 470 ETLDKLQFENPLEQLGLFIK-DELADEDEE 498
+ NPLE LGL++K DE DE+E+
Sbjct: 205 GSFKMESNSNPLEHLGLYMKMDEDEDEEED 234
>At1g06230 Ring3-like bromodomain protein
Length = 766
Score = 116 bits (291), Expect = 3e-26
Identities = 56/105 (53%), Positives = 70/105 (66%)
Query: 83 CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFA 142
C+A+L+ LM + + WVFN PVD L + DY+ II PMDLGTI S L KN+Y EFA
Sbjct: 425 CSALLERLMKHKHGWVFNAPVDVKGLGLLDYYTIIEHPMDLGTIKSALMKNLYKSPREFA 484
Query: 143 ADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCE 187
DVRLTF NAMTYNP DVH+MA L QIF+ +W ++ + E
Sbjct: 485 EDVRLTFHNAMTYNPEGQDVHLMAVTLLQIFEERWAVIEADYNRE 529
>At5g10550 bromodomain protein - like
Length = 678
Score = 103 bits (258), Expect = 2e-22
Identities = 51/116 (43%), Positives = 75/116 (63%)
Query: 73 QKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKK 132
+K+ + T C IL LM + +SWVF PVD V L + DY I+ KPMDLGT+ L+K
Sbjct: 243 EKVLKSMMTTCGQILVKLMKHKWSWVFLNPVDVVGLGLHDYHRIVDKPMDLGTVKMNLEK 302
Query: 133 NVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCED 188
+Y +FA+DVRLTF+NAM+YNP DV++MA++L FD + K+++ ++
Sbjct: 303 GLYRSPIDFASDVRLTFTNAMSYNPKGQDVYLMAEKLLSQFDVWFNPTLKRFEAQE 358
>At1g73150 unknown protein
Length = 461
Score = 103 bits (258), Expect = 2e-22
Identities = 51/112 (45%), Positives = 73/112 (64%), Gaps = 4/112 (3%)
Query: 78 KGSTQ----CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKN 133
KG+ Q C +L LM + W+FN PVD V L + DY II +PMDLGT+ ++L K+
Sbjct: 114 KGTVQILKSCNNLLTKLMKHKSGWIFNTPVDVVTLGLHDYHNIIKEPMDLGTVKTRLSKS 173
Query: 134 VYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWK 185
+Y EFA DVRLTF+NAM YNP +DV+ MA+ L +F+ KW ++ +++
Sbjct: 174 LYKSPLEFAEDVRLTFNNAMLYNPVGHDVYHMAEILLNLFEEKWVPLETQYE 225
>At5g65630 unknown protein
Length = 590
Score = 102 bits (254), Expect = 5e-22
Identities = 50/105 (47%), Positives = 65/105 (61%)
Query: 83 CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFA 142
C+ IL LM + ++WVFN PVD V L + DY ++ KPMDLGT+ L K Y +FA
Sbjct: 173 CSQILVKLMKHKWAWVFNTPVDVVGLGLHDYHQVVKKPMDLGTVKLNLDKGFYVSPIDFA 232
Query: 143 ADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCE 187
DVRLTF NAMTYNP DV+ MA +L FD + KK++ +
Sbjct: 233 TDVRLTFDNAMTYNPKGQDVYFMADKLLDHFDGMFNPAFKKFEAQ 277
>At1g17790 unknown protein (At1g17790)
Length = 487
Score = 98.2 bits (243), Expect = 9e-21
Identities = 48/111 (43%), Positives = 72/111 (64%), Gaps = 4/111 (3%)
Query: 78 KGSTQ----CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKN 133
KG+ Q C ++L LM + +WVFN PVD L + DY I+ +PMDLGT+ +KL K+
Sbjct: 127 KGTVQIFKNCNSLLTKLMKHKSAWVFNVPVDAKGLGLHDYHNIVKEPMDLGTVKTKLGKS 186
Query: 134 VYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKW 184
+Y +FA DVRLTF+NA+ YNP +DV+ A+ L +F+ KW ++ ++
Sbjct: 187 LYKSPLDFAEDVRLTFNNAILYNPIGHDVYRFAELLLNMFEDKWVSIEMQY 237
>At2g34900 RING3 protein-like
Length = 386
Score = 92.4 bits (228), Expect = 5e-19
Identities = 63/204 (30%), Positives = 100/204 (48%), Gaps = 17/204 (8%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
Q A + + + + ++W F +PVD L + DY+ +I KPMDLGTI K++ + YS V E
Sbjct: 113 QFATMFRQIAQHKWAWPFLEPVDVKGLGLHDYYKVIEKPMDLGTIKKKMESSEYSNVREI 172
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKW------------KDMDKKWKCEDE 189
ADVRL F NAM YN DV++MA+ L + F+ KW K +D++ +
Sbjct: 173 YADVRLVFKNAMRYNEEKEDVYVMAESLLEKFEEKWLLIMPKLVEEEKKQVDEEAEKHAN 232
Query: 190 LGKSVTGTIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKISSLAARDAKVEVPKLSQ 249
++ E AR N ++ + D +K + S +R K+S+ + + +LS
Sbjct: 233 KQLTMEAAQAEMARDLSNELY--EIDLQLEKLRESVVQRCRKLSTQEKKGLSAALGRLSP 290
Query: 250 VHCKSIEKDLIKGSEERDRIAPGA 273
+ + K L SE GA
Sbjct: 291 ---EDLSKALKMVSESNPSFPAGA 311
>At3g52280 putative protein
Length = 369
Score = 74.7 bits (182), Expect = 1e-13
Identities = 40/112 (35%), Positives = 62/112 (54%), Gaps = 3/112 (2%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNV---YSCV 138
Q I + + + +W F PV+ L + DYF +I KPMD TI ++++ Y V
Sbjct: 97 QFGTIFRQITQHKCAWPFMHPVNVEGLGLHDYFEVIDKPMDFSTIKNQMEAKDGTGYKHV 156
Query: 139 EEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDEL 190
+ AD+RL F NAM YN +DV+ MAK+L + F+ KW K + E+++
Sbjct: 157 MQIYADMRLVFENAMNYNEETSDVYSMAKKLLEKFEEKWAHFLPKVQEEEKI 208
>At1g20670 hypothetical protein
Length = 652
Score = 63.2 bits (152), Expect = 3e-10
Identities = 47/164 (28%), Positives = 71/164 (42%), Gaps = 9/164 (5%)
Query: 18 STKRIEVDPGQKCEFGQRGLQTDENGR-CNSAEKLFKPDSIKRGPPEGFEGQKEKRQKID 76
S + +D + F +R L +G ++ EK K I +G P E
Sbjct: 120 SQSDLNLDQTPEPSFNRRNLSAAASGSDYHTGEKASKATDILQGSPV------ESGPTTP 173
Query: 77 RKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYS 136
IL L V++ PVDP L PDYF II PMD T+ +KL YS
Sbjct: 174 LPDKKLLLFILDRLQKKDTYGVYSDPVDPEEL--PDYFEIIKNPMDFSTLRNKLDSGAYS 231
Query: 137 CVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDM 180
+E+F DV L +NAM YN + A+ + ++ ++++
Sbjct: 232 TLEQFERDVFLICTNAMEYNSADTVYYRQARAIQELAKKDFENL 275
>At3g54610 histone acetyltransferase (HAT1)
Length = 568
Score = 62.0 bits (149), Expect = 8e-10
Identities = 33/93 (35%), Positives = 52/93 (55%), Gaps = 3/93 (3%)
Query: 85 AILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLK-KNVYSCVEEFAA 143
A+LK + + +W F +PVD + ++PDY+ II P+DL I+ +++ + Y ++ F A
Sbjct: 466 ALLKTMQDHADAWPFKEPVD--SRDVPDYYDIIKDPIDLKVIAKRVESEQYYVTLDMFVA 523
Query: 144 DVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSK 176
D R F+N TYN P + A L F SK
Sbjct: 524 DARRMFNNCRTYNSPDTIYYKCATRLETHFHSK 556
>At5g55040 putative protein
Length = 145
Score = 56.2 bits (134), Expect = 4e-08
Identities = 32/76 (42%), Positives = 42/76 (55%), Gaps = 5/76 (6%)
Query: 98 VFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNP 157
V+ +PVDP L PDY +I PMD T+ KL YS +EE +DV L SNAM YN
Sbjct: 66 VYAEPVDPEEL--PDYHDMIEHPMDFSTVRKKLANGSYSTLEELESDVLLICSNAMQYNS 123
Query: 158 PHNDVHMMAKELNQIF 173
+ K++N +F
Sbjct: 124 SDT---VYYKQVNHVF 136
>At1g76380 unknown protein
Length = 579
Score = 55.8 bits (133), Expect = 5e-08
Identities = 28/75 (37%), Positives = 42/75 (55%), Gaps = 2/75 (2%)
Query: 98 VFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNP 157
V++ P DP L PDY+ II PMD T+ KL+ Y+ +E+F DV L +NAM YN
Sbjct: 164 VYSDPADPEEL--PDYYEIIKNPMDFTTLRKKLESGAYTTLEQFEQDVFLICTNAMEYNS 221
Query: 158 PHNDVHMMAKELNQI 172
+ A+ + ++
Sbjct: 222 ADTVYYRQARAMLEL 236
>At2g44430 unknown protein
Length = 646
Score = 50.8 bits (120), Expect = 2e-06
Identities = 85/386 (22%), Positives = 151/386 (39%), Gaps = 45/386 (11%)
Query: 33 GQRGLQTDENGRCNSAEKLFKPDSIKRGPPEGFEGQKEKRQKIDRKGSTQCAAILKCLMS 92
G +DE G ++E +K K+G + K Q + ++L + S
Sbjct: 270 GSEASHSDELGESGTSESKWKRKRRKQGGAGEIRSAESKSQPL--------ISLLDLIRS 321
Query: 93 YPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVY-SCVEEFAADVRLTFSN 151
+P +F + + + DY +++ + +D+ TI KLK+ Y S F D++L F+N
Sbjct: 322 HPRGSLFERRLR--SQEAKDYKSMVKQHLDIETIQRKLKQGSYDSSSLIFYRDLQLLFTN 379
Query: 152 AMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHP 211
A+ + P + M A EL + + + E GK+ IK+ A + M
Sbjct: 380 AIVFFPLSSSESMAAHELRAVVSQEMR---------KETGKAGPRLIKQEA----SGMRS 426
Query: 212 RQKDTFPKKSQVSEHKRVLKISSLAARDAKVEVPKLSQVHCKSIEKDLIKG---SEERDR 268
+ D S +S K + + + + K S +KD K SEE+D
Sbjct: 427 GKADAETSDSSLSRQKSSGPL--VVCKKRRSVSAKASPSSSSFSQKDDTKEETLSEEKDN 484
Query: 269 IAPGAVAFKQKRQIKSTSPLERKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWS 328
IA G + R+ + + + G +E S+ + ++S +
Sbjct: 485 IATGV---RSSRRANKVAAVVANNTKTGKGRNKQKQTE-----SKTNSSNDNSSKQDTGK 536
Query: 329 TPDFPVQLSPKKALRAAM--LKSRFADTILKAQQKT--LLEHGDKGDPLKMQLEKERLER 384
T V KK++ + LK K Q K+ ++ K P +++ ++
Sbjct: 537 TEKKTVSADKKKSVADFLKRLKKNSPQKEAKDQNKSGGNVKKDSKTKPRELRSSSVGKKK 596
Query: 385 IQRE----ERARIEAQIKTAEAAARA 406
+ E +RA Q KTAEA A A
Sbjct: 597 AEVENTPVKRAPGRPQKKTAEATASA 622
>At3g11450 putative cell division related protein
Length = 663
Score = 47.4 bits (111), Expect = 2e-05
Identities = 39/132 (29%), Positives = 74/132 (55%), Gaps = 7/132 (5%)
Query: 358 AQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQRE 417
A+ +TL+++ + DP ++ ++E Q+++ A+I+A+ K E AA A EE+ R++ E
Sbjct: 298 ARIRTLVDNAYRKDPRIVKRKEEEKAEKQQKKDAKIQAKKKQEEDAAIAAEEEKRRKEEE 357
Query: 418 KEREAARAAIEEMKRTVEIEHNLEIVKE---LETLSGCTLSYKALG-GRNDYKVALETLD 473
++R A A ++ K+T E E L + KE L TLS ++ + L D + +L+
Sbjct: 358 EKRAAESA--QQQKKTKEREKKL-LRKERNRLRTLSAPLVAQRLLDISEEDIENLCMSLN 414
Query: 474 KLQFENPLEQLG 485
Q +N +++G
Sbjct: 415 TEQLQNLCDKMG 426
>At1g32750 unknown protein
Length = 1919
Score = 45.8 bits (107), Expect = 6e-05
Identities = 38/115 (33%), Positives = 53/115 (46%), Gaps = 8/115 (6%)
Query: 77 RKGSTQCAAILKCL-----MSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLK 131
+KG A IL+ + + S +F KPV PDY I+ PMDL TI K++
Sbjct: 1797 KKGEVGLANILERIVDTLRLKEEVSRLFLKPVSKK--EAPDYLDIVENPMDLSTIRDKVR 1854
Query: 132 KNVYSCVEEFAADVRLTFSNAMTYNPPHN-DVHMMAKELNQIFDSKWKDMDKKWK 185
K Y E+F DV NA YN N + +A +L +I D D + + K
Sbjct: 1855 KIEYRNREQFRHDVWQIKYNAHLYNDGRNPGIPPLADQLLEICDYLLDDYEDQLK 1909
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.130 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,526,371
Number of Sequences: 26719
Number of extensions: 503852
Number of successful extensions: 2644
Number of sequences better than 10.0: 232
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 165
Number of HSP's that attempted gapping in prelim test: 1904
Number of HSP's gapped (non-prelim): 548
length of query: 510
length of database: 11,318,596
effective HSP length: 104
effective length of query: 406
effective length of database: 8,539,820
effective search space: 3467166920
effective search space used: 3467166920
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0228a.2


