

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0227.11
(425 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
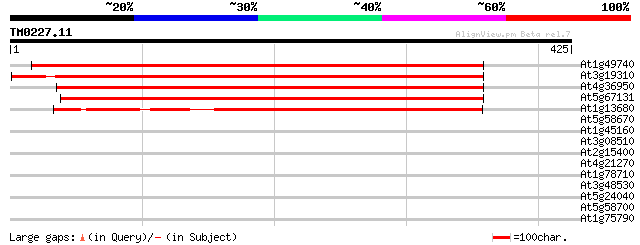
Score E
Sequences producing significant alignments: (bits) Value
At1g49740 unknown protein 539 e-153
At3g19310 unknown protein 516 e-147
At4g36950 MAP3K-like protein kinase 471 e-133
At5g67131 unknown protein 399 e-111
At1g13680 hypothetical protein 303 9e-83
At5g58670 phosphoinositide specific phospholipase C (AtPLC1) 39 0.005
At1g45160 similar to IRE homolog 1 dbj|BAA89784.1 33 0.23
At3g08510 phosphoinositide specific phospholipase C (AtPLC2) 32 0.51
At2g15400 DNA-directed RNA polymerase II, third largest subunit 32 0.51
At4g21270 kinesin-related protein katA 30 1.9
At1g78710 hypothetical protein 30 2.5
At3g48530 unknown protein 29 5.6
At5g24040 putative protein 28 7.4
At5g58700 phosphoinositide-specific phospholipase C4 (PLC4) 28 9.6
At1g75790 polymer protein, putative 28 9.6
>At1g49740 unknown protein
Length = 359
Score = 539 bits (1389), Expect = e-153
Identities = 246/343 (71%), Positives = 289/343 (83%)
Query: 17 ILVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKV 76
I +LL SS L+ +EG+ C+ N NC++GLHCETC+ N + RPRC+RTQPINP +K
Sbjct: 11 IALLLQSSFLLEISSALKEGKTCITNSNCDAGLHCETCIANTDFRPRCSRTQPINPITKA 70
Query: 77 KGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFEN 136
KGLPFN+YSWLTTHNSFA LG+ S TGS IL+PTNQQD+IT QLNNGVRG MLD+YDF+N
Sbjct: 71 KGLPFNKYSWLTTHNSFARLGEVSRTGSAILAPTNQQDSITSQLNNGVRGFMLDMYDFQN 130
Query: 137 DVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDA 196
D+WLCHSF G C+N+TAFQPAIN+L+E QVFLE N E+VTIIIEDYV SPKGLTKVFDA
Sbjct: 131 DIWLCHSFDGTCFNFTAFQPAINILREFQVFLEKNKEEVVTIIIEDYVKSPKGLTKVFDA 190
Query: 197 AGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQY 256
AGLRK+ FPVSRMPKNGGDWP++DDMV+KNQRL+VFTS + KEA+EGIAY+W+Y+VENQY
Sbjct: 191 AGLRKFMFPVSRMPKNGGDWPRLDDMVRKNQRLLVFTSDSHKEATEGIAYQWKYMVENQY 250
Query: 257 GNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAA 316
GNGG+K G CPNRA+S M+ S+SLVLVN F D DV +CK NSA LL + TC QAA
Sbjct: 251 GNGGLKVGVCPNRAQSAPMSDKSKSLVLVNHFPDAADVIVACKQNSASLLESIKTCYQAA 310
Query: 317 GKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCK 359
G+RWPNFIAVDFYKRSDGGGAP+AVDVANG+L+CGC N A CK
Sbjct: 311 GQRWPNFIAVDFYKRSDGGGAPQAVDVANGNLICGCDNFAACK 353
>At3g19310 unknown protein
Length = 413
Score = 516 bits (1329), Expect = e-147
Identities = 240/359 (66%), Positives = 289/359 (79%), Gaps = 7/359 (1%)
Query: 2 MMQKPTLAIATTLFT-ILVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTNGNV 60
M Q+ T + L IL++L S ALK EG+ C+ +KNC+ GLHCE+C+ + +
Sbjct: 1 MFQRFTFFLTALLIPCILIILSPSYALK------EGETCIVSKNCDRGLHCESCLASDSF 54
Query: 61 RPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQL 120
RPRC+R QPINPT+KVKGLP+N+YSWLTTHNSFA +G KS TGS+IL+P+NQQD+IT QL
Sbjct: 55 RPRCSRMQPINPTTKVKGLPYNKYSWLTTHNSFARMGAKSGTGSMILAPSNQQDSITSQL 114
Query: 121 NNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIII 180
NGVRG MLDLYDF+ND+WLCHS+GG C+NYTAFQPA+N+LKE QVFL+ N +VT+I+
Sbjct: 115 LNGVRGFMLDLYDFQNDIWLCHSYGGNCFNYTAFQPAVNILKEFQVFLDKNKDVVVTLIL 174
Query: 181 EDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEA 240
EDYV SP GLT+VFDA+GLR + FPVSRMPKNG DWP +DDM+ +NQRL+VFTS KEA
Sbjct: 175 EDYVKSPNGLTRVFDASGLRNFMFPVSRMPKNGEDWPTLDDMICQNQRLLVFTSNPQKEA 234
Query: 241 SEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKD 300
SEGIA+ WRY++ENQYG+GGMKAG C NR ES +M SRSL+LVN+F D DV SCK
Sbjct: 235 SEGIAFMWRYMIENQYGDGGMKAGVCTNRPESVAMGDRSRSLILVNYFPDTADVIGSCKQ 294
Query: 301 NSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCK 359
NSAPLL V C +A+GKRWPNFIAVDFYKRSDGGGAP+AVDVANGH VCGC +IA CK
Sbjct: 295 NSAPLLDTVKNCQEASGKRWPNFIAVDFYKRSDGGGAPKAVDVANGHAVCGCEDIAACK 353
>At4g36950 MAP3K-like protein kinase
Length = 799
Score = 471 bits (1211), Expect = e-133
Identities = 214/324 (66%), Positives = 261/324 (80%)
Query: 36 GQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFAL 95
G+ C + C++GL C++C NGN CTR QP+NPTSKV GLPFN+YSWLTTHNS+A+
Sbjct: 340 GETCSSTSECDAGLSCQSCPANGNTGSTCTRIQPLNPTSKVNGLPFNKYSWLTTHNSYAI 399
Query: 96 LGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQ 155
G S TGS ++SP NQ+D+IT+QL NGVRG+MLD YDF+ND+WLCHS GG C+N+TAFQ
Sbjct: 400 TGANSATGSFLVSPKNQEDSITNQLKNGVRGIMLDTYDFQNDIWLCHSTGGTCFNFTAFQ 459
Query: 156 PAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGD 215
PAIN LKEI FLE+N SEIVTII+EDYV S GLT VF+A+GL K+ P+SRMPK+G D
Sbjct: 460 PAINALKEINDFLESNLSEIVTIILEDYVKSQMGLTNVFNASGLSKFLLPISRMPKDGTD 519
Query: 216 WPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSM 275
WP VDDMV++NQRLVVFTSK KEASEG+AY+W Y+VENQYGN GMK GSC +R+ES S+
Sbjct: 520 WPTVDDMVKQNQRLVVFTSKKDKEASEGLAYQWNYMVENQYGNDGMKDGSCSSRSESSSL 579
Query: 276 NTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGG 335
+T SRSLV N+F P+ Q+C DNS+PL+ M+ TC++AAGKRWPNFIAVDFY+RSD G
Sbjct: 580 DTMSRSLVFQNYFETSPNSTQACADNSSPLIEMMRTCHEAAGKRWPNFIAVDFYQRSDSG 639
Query: 336 GAPEAVDVANGHLVCGCGNIATCK 359
GA EAVD ANG L CGC ++ CK
Sbjct: 640 GAAEAVDEANGRLTCGCDSLVYCK 663
>At5g67131 unknown protein
Length = 426
Score = 399 bits (1025), Expect = e-111
Identities = 183/321 (57%), Positives = 237/321 (73%)
Query: 39 CVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFALLGQ 98
C + +C SGL+C C G +P CTR Q +PTS + GLPFN+Y+WL THN+F+
Sbjct: 40 CSSATDCVSGLYCGDCPAVGRSKPVCTRGQATSPTSIINGLPFNKYTWLMTHNAFSNANA 99
Query: 99 KSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAI 158
+ G ++ NQ+DTIT+QL NGVRGLMLD+YDF ND+WLCHS GQC+N+TAFQPAI
Sbjct: 100 PLLPGVERITFYNQEDTITNQLQNGVRGLMLDMYDFNNDIWLCHSLRGQCFNFTAFQPAI 159
Query: 159 NVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGDWPK 218
N+L+E++ FL NP+EIVTIIIEDYV PKGL+ +F AGL KYWFPVS+MP+ G DWP
Sbjct: 160 NILREVEAFLSQNPTEIVTIIIEDYVHRPKGLSTLFANAGLDKYWFPVSKMPRKGEDWPT 219
Query: 219 VDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSMNTT 278
V DMVQ+N RL+VFTS A+KE EG+AY+WRY+VEN+ G+ G+K GSCPNR ES +N+
Sbjct: 220 VTDMVQENHRLLVFTSVAAKEDEEGVAYQWRYMVENESGDPGVKRGSCPNRKESQPLNSK 279
Query: 279 SRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGGGAP 338
S SL L+N+F P +CK++SAPL MV TC ++ G R PNF+AV+FY RSDGGG
Sbjct: 280 SSSLFLMNYFPTYPVEKDACKEHSAPLAEMVGTCLKSGGNRMPNFLAVNFYMRSDGGGVF 339
Query: 339 EAVDVANGHLVCGCGNIATCK 359
E +D NG ++CGC ++ C+
Sbjct: 340 EILDRMNGPVLCGCETLSACQ 360
>At1g13680 hypothetical protein
Length = 300
Score = 303 bits (777), Expect = 9e-83
Identities = 158/326 (48%), Positives = 206/326 (62%), Gaps = 28/326 (8%)
Query: 34 QEGQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKG-LPFNRYSWLTTHNS 92
Q G C ++++CN GL C C G RC R+ + S V +PFN+Y++LTTHNS
Sbjct: 2 QLGDQCSSDEDCNVGLGCFKC---GIDVARCVRSNITDQFSIVNNSMPFNKYAFLTTHNS 58
Query: 93 FALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYT 152
+A+ G+ L Q+DTI QLN+GVR LMLD YD+E D
Sbjct: 59 YAIEGKA-------LHVATQEDTIVQQLNSGVRALMLDTYDYEGD--------------- 96
Query: 153 AFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKN 212
AI+ KEI FL ANPSEIVT+I+EDYV S GLTKVF +GL+K+WFPV MP
Sbjct: 97 --NRAIDTFKEIFAFLTANPSEIVTLILEDYVKSQNGLTKVFTDSGLKKFWFPVQNMPIG 154
Query: 213 GGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAES 272
G DWP V DMV N RL+VFTS SK+ +EGIAY+W Y+VENQYG+ G+K C NRA+S
Sbjct: 155 GQDWPLVKDMVANNHRLIVFTSAKSKQETEGIAYQWNYMVENQYGDDGVKPDECSNRADS 214
Query: 273 PSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRS 332
+ +++LV VN F+ +P +C++NS LL M+ TC AAG RW NF+AV+FYKRS
Sbjct: 215 ALLTDKTKALVSVNHFKTVPVKILTCEENSEQLLDMIKTCYVAAGNRWANFVAVNFYKRS 274
Query: 333 DGGGAPEAVDVANGHLVCGCGNIATC 358
+GGG +A+D NG L+CG ++ C
Sbjct: 275 NGGGTFQAIDKLNGELLCGRDDVHAC 300
>At5g58670 phosphoinositide specific phospholipase C (AtPLC1)
Length = 561
Score = 38.9 bits (89), Expect = 0.005
Identities = 56/227 (24%), Positives = 87/227 (37%), Gaps = 41/227 (18%)
Query: 80 PFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVW 139
P + Y T HNS+ Q + S+ + I L NGVR + LDL+ +
Sbjct: 110 PLSHYFLYTGHNSYLTGNQLNSNSSI--------EPIVKALRNGVRVIELDLWPNSSGKE 161
Query: 140 LCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTS------PKGLTKV 193
GG + Q +NV+KE + A P V + +ED++T K ++K
Sbjct: 162 AEVRHGGTLTSREDLQKCLNVVKENAFQVSAYP---VVLTLEDHLTPILQKKVAKMVSKT 218
Query: 194 FDAAGLR-----KYWFPVSRMPKN----GGDWPK-----------VDDMVQKNQRLVVFT 233
F + + FP KN PK D + +++
Sbjct: 219 FGGSLFQCTDETTECFPSPESLKNKILISTKPPKEYLQTQISKGSTTDESTRAKKISDAE 278
Query: 234 SKASKEASEGIAYEWRYLVENQYGN--GGMK--AGSCPNRAESPSMN 276
+ +E E +A E+R L+ GN GG+K PNR SM+
Sbjct: 279 EQVQEEDEESVAIEYRDLISIHAGNRKGGLKNCLNGDPNRVIRLSMS 325
>At1g45160 similar to IRE homolog 1 dbj|BAA89784.1
Length = 1067
Score = 33.5 bits (75), Expect = 0.23
Identities = 25/93 (26%), Positives = 40/93 (42%), Gaps = 11/93 (11%)
Query: 191 TKVFDAAGLRKYWFPVSRMPKNGGDW---PKVDDM------VQKNQRLVVFTSKASKEAS 241
T+ D + RK+ + R+P DW P+VDD Q+N+ F + +
Sbjct: 272 TEPIDESSFRKFKECLERIPALETDWGSTPRVDDSGSGYPEYQRNEAGQKFKRRDKESLE 331
Query: 242 EGIAYEWRYLVENQYGNGGMKAGSCPNRAESPS 274
A + Y+V N +GN + G + E PS
Sbjct: 332 SETALD--YVVPNDHGNNAAREGYAAAKQEFPS 362
>At3g08510 phosphoinositide specific phospholipase C (AtPLC2)
Length = 581
Score = 32.3 bits (72), Expect = 0.51
Identities = 42/169 (24%), Positives = 70/169 (40%), Gaps = 25/169 (14%)
Query: 80 PFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFEN--D 137
P + Y T HNS+ +TG+ + S ++ I D L GVR + LD++ N D
Sbjct: 108 PISHYFIFTGHNSY-------LTGNQLSSDCSEVPII-DALKKGVRVIELDIWPNSNKDD 159
Query: 138 VWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVT---SPKGLTKVF 194
+ + H T I LK I+ V + +ED++T K V
Sbjct: 160 IDVLHGM-----TLTTPVGLIKCLKAIRAHAFDVSDYPVVVTLEDHLTPDLQSKVAEMVT 214
Query: 195 DAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEG 243
+ G + PV K ++P + + +R ++ ++K KE EG
Sbjct: 215 EIFGEILFTPPVGESLK---EFPSPNSL----KRRIIISTKPPKEYKEG 256
>At2g15400 DNA-directed RNA polymerase II, third largest subunit
Length = 319
Score = 32.3 bits (72), Expect = 0.51
Identities = 24/83 (28%), Positives = 37/83 (43%), Gaps = 8/83 (9%)
Query: 1 MMMQKPTLAIATTLFTI--LVLLDSSLA--LKFKPLQQEG----QICVANKNCNSGLHCE 52
M+ + PT+AI + VL D +A L PL E + C ++CN HCE
Sbjct: 41 MISEVPTMAIHLVKIEVNSSVLNDEFIAQRLSLIPLTSERAMSMRFCQDCEDCNGDEHCE 100
Query: 53 TCVTNGNVRPRCTRTQPINPTSK 75
C + +C Q ++ TS+
Sbjct: 101 FCSVEFPLSAKCVTDQTLDVTSR 123
>At4g21270 kinesin-related protein katA
Length = 793
Score = 30.4 bits (67), Expect = 1.9
Identities = 20/61 (32%), Positives = 31/61 (50%), Gaps = 1/61 (1%)
Query: 222 MVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSMNTTSRS 281
M +NQ + A KE GI+++ R VE Q G G +A S N+ + +MN+ S
Sbjct: 1 MASRNQNRPPRSPNAKKEGLGGISFDKRRKVETQGGTGRRQAFSAVNK-QDVTMNSDVGS 59
Query: 282 L 282
+
Sbjct: 60 I 60
>At1g78710 hypothetical protein
Length = 359
Score = 30.0 bits (66), Expect = 2.5
Identities = 26/101 (25%), Positives = 44/101 (42%), Gaps = 11/101 (10%)
Query: 118 DQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVT 177
D ++ G + L D+ F W H+ + ++Y FQ ++KE+ N E
Sbjct: 178 DSISRGNQWLGSDVAIFNTFHWWSHTGRAKTWDY--FQTGDKIVKEM------NRMEAFK 229
Query: 178 IIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGDWPK 218
I + T K + D + R ++ VS + NGG+W K
Sbjct: 230 IAL---TTWSKWIDHNIDPSKTRVFYQGVSPVHLNGGEWGK 267
>At3g48530 unknown protein
Length = 424
Score = 28.9 bits (63), Expect = 5.6
Identities = 13/43 (30%), Positives = 22/43 (50%)
Query: 272 SPSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQ 314
+P + RS+ NF + + + C D SAP++S V C +
Sbjct: 317 APEIYHDYRSITTKNFLVSVREHLEKCGDTSAPIMSGVIACTK 359
>At5g24040 putative protein
Length = 373
Score = 28.5 bits (62), Expect = 7.4
Identities = 9/23 (39%), Positives = 15/23 (65%)
Query: 200 RKYWFPVSRMPKNGGDWPKVDDM 222
+K WF +S + + DW +V+DM
Sbjct: 272 KKIWFEISELNEERNDWDQVEDM 294
>At5g58700 phosphoinositide-specific phospholipase C4 (PLC4)
Length = 597
Score = 28.1 bits (61), Expect = 9.6
Identities = 32/114 (28%), Positives = 48/114 (42%), Gaps = 13/114 (11%)
Query: 80 PFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVW 139
P + Y T HNS+ +TG+ + S ++ I D L GVR + LDL+ D
Sbjct: 119 PLSHYFIFTGHNSY-------LTGNQLSSNCSELP-IADALRRGVRVVELDLWPRGTDD- 169
Query: 140 LCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKV 193
+C G + +K + P V I +ED++T PK KV
Sbjct: 170 VCVKHGRTLTKEVKLGKCLESIKANAFAISKYP---VIITLEDHLT-PKLQFKV 219
>At1g75790 polymer protein, putative
Length = 545
Score = 28.1 bits (61), Expect = 9.6
Identities = 14/50 (28%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Query: 245 AYEWRYLVENQYGNGGMKAGSCPNRA-ESPSMNTTSRSLVLVNFFRDLPD 293
+Y+W ++ GG K N P +N T+ +++VN F +LP+
Sbjct: 28 SYQWVVSYSQRFILGGNKQVIVINDMFPGPILNATANDIIVVNIFNNLPE 77
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,726,270
Number of Sequences: 26719
Number of extensions: 417964
Number of successful extensions: 928
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 915
Number of HSP's gapped (non-prelim): 16
length of query: 425
length of database: 11,318,596
effective HSP length: 102
effective length of query: 323
effective length of database: 8,593,258
effective search space: 2775622334
effective search space used: 2775622334
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0227.11


