

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.8
(518 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
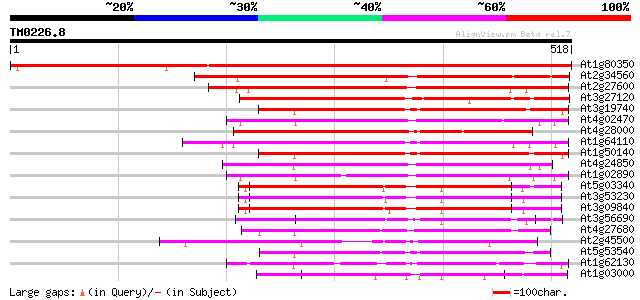
Score E
Sequences producing significant alignments: (bits) Value
At1g80350 CAD ATPase (AAA1) 865 0.0
At2g34560 katanin like protein 286 2e-77
At2g27600 putative AAA-type ATPase 263 1e-70
At3g27120 spastin protein, putative 259 3e-69
At3g19740 ATPase, putative 223 2e-58
At4g02470 221 6e-58
At4g28000 putative protein 218 6e-57
At1g64110 unknown protein 217 1e-56
At1g50140 hypothetical protein, 5' partial 216 3e-56
At4g24850 unknown protein (At4g24850) 204 1e-52
At1g02890 hypothetical protein 201 1e-51
At5g03340 transitional endoplasmic reticulum ATPase 196 3e-50
At3g53230 CDC48 - like protein 194 8e-50
At3g09840 putative transitional endoplasmic reticulum ATPase 193 2e-49
At3g56690 calmodulin-binding protein 179 3e-45
At4g27680 unknown protein 179 4e-45
At2g45500 hypothetical protein 176 4e-44
At5g53540 26S proteasome regulatory particle chain RPT6-like pro... 166 2e-41
At1g62130 hypothetical protein 166 3e-41
At1g03000 putative peroxisome assembly factor-2 164 1e-40
>At1g80350 CAD ATPase (AAA1)
Length = 523
Score = 865 bits (2234), Expect = 0.0
Identities = 437/524 (83%), Positives = 468/524 (88%), Gaps = 7/524 (1%)
Query: 1 MVGSS--LGGLQEHLKLAREYALEGQYDTSIIFFDGAVAQINKHLNSLDDPLLRSKWMNV 58
MVGSS L GLQ+HLKLAREYALEG YDTS+IFFDGA+AQINKHLN+LDDPL R+KWMNV
Sbjct: 1 MVGSSNSLAGLQDHLKLAREYALEGSYDTSVIFFDGAIAQINKHLNTLDDPLARTKWMNV 60
Query: 59 KKALSEETEVVKQLDAERSAFKNNPIGRRPPSPPISTKSSFVFQPLDEYPTSSSG-PMDD 117
KKA+ EETEVVKQLDAER AFK P GRR SPPI+TKSSFVFQPLDEYPTSS G PMDD
Sbjct: 61 KKAIMEETEVVKQLDAERRAFKEAPTGRRAASPPINTKSSFVFQPLDEYPTSSGGGPMDD 120
Query: 118 PDVWRPPSRD-TSRRPTRPGQMSTRKS--DGAWARGATPRGGATGRGAKAGATSKVNTGT 174
PDVWRPP+RD TSRRP R GQ TRKS DGAWARG T R G RG + GATSK G
Sbjct: 121 PDVWRPPTRDVTSRRPARAGQTGTRKSPQDGAWARGPTTRTGPASRGGRGGATSKSTAGA 180
Query: 175 RASTTGKKSGVSGKAGKTDSVISDAEESKSKKSQYEGPDPELAEMLERDVLETSPGVRWE 234
R+ST GKK G + K+ K +S+ DAE+ KSK+ YEGPD +LA MLERDVL+++PGVRW+
Sbjct: 181 RSSTAGKK-GAASKSNKAESMNGDAEDGKSKRGLYEGPDEDLAAMLERDVLDSTPGVRWD 239
Query: 235 DVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTT 294
DVAGL+EAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTT
Sbjct: 240 DVAGLSEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTT 299
Query: 295 FFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRV 354
FFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRG SGEHESSRRV
Sbjct: 300 FFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGGSGEHESSRRV 359
Query: 355 KSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRK 414
KSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLP+FESRK
Sbjct: 360 KSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPDFESRK 419
Query: 415 ELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNM 474
LI INL+TVEVA DVNI++VARRT+GYSGDDLTNVCRDAS+NGMRRKIAGKTRDEIKNM
Sbjct: 420 ALININLRTVEVASDVNIEDVARRTEGYSGDDLTNVCRDASMNGMRRKIAGKTRDEIKNM 479
Query: 475 SKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKWFHEFGSA 518
SKDDIS DPVAMCDFEEA+ KVQ SVS +DIE+HEKW EFGSA
Sbjct: 480 SKDDISNDPVAMCDFEEAIRKVQPSVSSSDIEKHEKWLSEFGSA 523
>At2g34560 katanin like protein
Length = 384
Score = 286 bits (731), Expect = 2e-77
Identities = 148/354 (41%), Positives = 226/354 (63%), Gaps = 16/354 (4%)
Query: 171 NTGTRASTTGKKSGVSGKAGKTDSVISDAEESKSKKSQY----EGPDPELAEMLERDVLE 226
+ G S V+ + G T + K KKS + LAE L RD++
Sbjct: 36 SNGNNGDVNNNSSPVTNQDGNTALANGNVIREKPKKSMFPPFESAETRTLAESLSRDIIR 95
Query: 227 TSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKA 286
+P ++WE + GL AK+LL+EAVV+P+ P YF G+ PWKG+L+FGPPGTGKT+LAKA
Sbjct: 96 GNPNIKWESIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKA 155
Query: 287 VATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASG 346
VATEC TTFFN+S++++ SKWRG+SE+++R LFDLAR +APSTIF+DEID++ + RG G
Sbjct: 156 VATECNTTFFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEG 215
Query: 347 --EHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIY 404
EHE+SRR+K+ELL+Q+DG+ T ++V VLAATN PW++D A+ RRLEKRI
Sbjct: 216 RSEHEASRRLKTELLIQMDGLQKT-------NELVFVLAATNLPWELDAAMLRRLEKRIL 268
Query: 405 IPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIA 464
+PLP+ E+R+ + + + + + D + +++GYSG D+ +C++A++ +RR +A
Sbjct: 269 VPLPDPEARRGMFEMLIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLA 328
Query: 465 GKTRDEIKNMSKDDISK-DPVAMCDFEEALVKVQRSVSQADIERHEKWFHEFGS 517
D + +D++ K P+ D + AL + S + ++K+ ++GS
Sbjct: 329 -ILEDREDVVPEDELPKIGPILPEDIDRALSNTRPS-AHLHAHLYDKFNDDYGS 380
>At2g27600 putative AAA-type ATPase
Length = 435
Score = 263 bits (673), Expect = 1e-70
Identities = 141/364 (38%), Positives = 226/364 (61%), Gaps = 38/364 (10%)
Query: 184 GVSGKAGKTDSVISDAEESKSKKSQ---YEGPDPELAEM---LERDVLETSPGVRWEDVA 237
G SG D+ ++ ++K K + +G DPE +++ L ++ P ++W DVA
Sbjct: 76 GGSGPGSNGDAAVATRPKTKPKDGEGGGKDGEDPEQSKLRAGLNSAIVREKPNIKWSDVA 135
Query: 238 GLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFN 297
GL AK+ L+EAV+LP+ P++F G RRPW+ L++GPPGTGK+ LAKAVATE +TFF+
Sbjct: 136 GLESAKQALQEAVILPVKFPQFFTGKRRPWRAFLLYGPPGTGKSYLAKAVATEADSTFFS 195
Query: 298 VSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSE 357
VSS+ L SKW GESE++V LF++AR APS IF+DEIDSLC +RG E E+SRR+K+E
Sbjct: 196 VSSSDLVSKWMGESEKLVSNLFEMARESAPSIIFVDEIDSLCGTRGEGNESEASRRIKTE 255
Query: 358 LLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELI 417
LLVQ+ GV + + + V+VLAATN P+ +D+A+RRR +KRIYIPLP ++R+ +
Sbjct: 256 LLVQMQGVGH-------NDEKVLVLAATNTPYALDQAIRRRFDKRIYIPLPEAKARQHMF 308
Query: 418 RINL-KTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRR--------------- 461
+++L T + + + + ++T+G+SG D++ +D +R+
Sbjct: 309 KVHLGDTPHNLTEPDFEYLGQKTEGFSGSDVSVCVKDVLFEPVRKTQDAMFFFKSPDGTW 368
Query: 462 -----KIAGKTRDEIKNMS----KDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKWF 512
+ G + +++++ + I P+ DFE+ L + + +VS++D++ HE++
Sbjct: 369 MPCGPRHPGAIQTTMQDLATKGLAEKIIPPPITRTDFEKVLARQRPTVSKSDLDVHERFT 428
Query: 513 HEFG 516
EFG
Sbjct: 429 QEFG 432
>At3g27120 spastin protein, putative
Length = 327
Score = 259 bits (662), Expect = 3e-69
Identities = 133/307 (43%), Positives = 208/307 (67%), Gaps = 13/307 (4%)
Query: 213 DPELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLM 272
+P L E + ++++ P VRW+D+AGL AK+ + E V+ PL P+ F+G R P KG+L+
Sbjct: 29 EPRLIEHVSNEIMDRDPNVRWDDIAGLEHAKKCVTEMVIWPLLRPDIFKGCRSPGKGLLL 88
Query: 273 FGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFI 332
FGPPGTGKT++ KA+A E TFF +S+++L SKW GE E++VR LF +A P+ IF+
Sbjct: 89 FGPPGTGKTMIGKAIAGEAKATFFYISASSLTSKWIGEGEKLVRALFGVASCRQPAVIFV 148
Query: 333 DEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDID 392
DEIDSL + R + GEHESSRR+K++ L++++G + GS +I+++ ATN P ++D
Sbjct: 149 DEIDSLLSQRKSDGEHESSRRLKTQFLIEMEGF------DSGSEQILLI-GATNRPQELD 201
Query: 393 EALRRRLEKRIYIPLPNFESRKELIRINLKT--VEVAPDVNIDEVARRTDGYSGDDLTNV 450
EA RRRL KR+YIPLP+ E+R +I+ LK + D +++ + T+GYSG D+ N+
Sbjct: 202 EAARRRLTKRLYIPLPSSEARAWIIQNLLKKDGLFTLSDDDMNIICNLTEGYSGSDMKNL 261
Query: 451 CRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEK 510
+DA++ +R + K +I N++KDD+ V + DF++AL +V+ SVSQ ++ +E
Sbjct: 262 VKDATMGPLREAL--KRGIDITNLTKDDMRL--VTLQDFKDALQEVRPSVSQNELGIYEN 317
Query: 511 WFHEFGS 517
W ++FGS
Sbjct: 318 WNNQFGS 324
>At3g19740 ATPase, putative
Length = 439
Score = 223 bits (568), Expect = 2e-58
Identities = 115/291 (39%), Positives = 185/291 (63%), Gaps = 13/291 (4%)
Query: 230 GVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTLLAKAV 287
GV+++D+ L K+ L E V+LP+ PE F + RP KG+L+FGPPGTGKTLLAKA+
Sbjct: 146 GVKFDDIGALEHVKKTLNELVILPMRRPELFTRGNLLRPCKGILLFGPPGTGKTLLAKAL 205
Query: 288 ATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGE 347
ATE G F +++ +TL SKW G++E++ + LF A AP IF+DE+DSL +RG + E
Sbjct: 206 ATEAGANFISITGSTLTSKWFGDAEKLTKALFSFASKLAPVIIFVDEVDSLLGARGGAFE 265
Query: 348 HESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPL 407
HE++RR+++E + DG+ ++D R +++L ATN P+D+D+A+ RRL +RIY+ L
Sbjct: 266 HEATRRMRNEFMAAWDGL----RSKDSQR--ILILGATNRPFDLDDAVIRRLPRRIYVDL 319
Query: 408 PNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIAGKT 467
P+ E+R ++++I L + D++A+ T+GYSG DL N+C A+ ++ + +
Sbjct: 320 PDAENRLKILKIFLTPENLETGFEFDKLAKETEGYSGSDLKNLCIAAAYRPVQELLQEEN 379
Query: 468 RDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHE--KWFHEFG 516
+D + N S D P+++ DF ++ KV SV+ +E KW ++G
Sbjct: 380 KDSVTNASPD---LRPLSLDDFIQSKAKVSPSVAYDATTMNELRKWNEQYG 427
>At4g02470
Length = 371
Score = 221 bits (564), Expect = 6e-58
Identities = 131/332 (39%), Positives = 197/332 (58%), Gaps = 23/332 (6%)
Query: 201 ESKSKKSQYEG--PDPELAEMLERDVLETSP-GVRWEDVAGLTEAKRLLEEAVVLPLWMP 257
E+KS K + + E + L DV+ S GV ++D+ L K L+E V+LPL P
Sbjct: 33 ENKSLKKSLKDVVTENEFEKKLLSDVIPPSDIGVSFDDIGALENVKETLKELVMLPLQRP 92
Query: 258 EYFQG--IRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMV 315
E F + +P KG+L+FGPPGTGKT+LAKAVATE G F N+S +++ SKW GE E+ V
Sbjct: 93 ELFDKGQLTKPTKGILLFGPPGTGKTMLAKAVATEAGANFINISMSSITSKWFGEGEKYV 152
Query: 316 RCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGS 375
+ +F LA APS IF+DE+DS+ R GEHE+ R++K+E +V DG+
Sbjct: 153 KAVFSLASKIAPSVIFVDEVDSMLGRRENPGEHEAMRKMKNEFMVNWDGLRTK------D 206
Query: 376 RKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEV 435
R+ V+VLAATN P+D+DEA+ RRL +R+ + LP+ +R +++ + L E+APDV+++ +
Sbjct: 207 RERVLVLAATNRPFDLDEAVIRRLPRRLMVNLPDATNRSKILSVILAKEEIAPDVDLEAI 266
Query: 436 ARRTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMC-------- 487
A TDGYSG DL N+C A+ + R+I K + E ++ P+ C
Sbjct: 267 ANMTDGYSGSDLKNLCVTAA-HFPIREILEKEKKEKTAAQAENRPTPPLYSCTDVRSLTM 325
Query: 488 -DFEEALVKVQRSVS--QADIERHEKWFHEFG 516
DF+ A +V SVS +++ ++W +G
Sbjct: 326 NDFKAAHDQVCASVSSDSSNMNELQQWNELYG 357
>At4g28000 putative protein
Length = 726
Score = 218 bits (555), Expect = 6e-57
Identities = 115/278 (41%), Positives = 178/278 (63%), Gaps = 9/278 (3%)
Query: 207 SQYEGPDPELAEMLERDVLETSP-GVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQG-IR 264
S+ PD E + + +V+ + GV + D+ L E K L+E V+LPL P+ F+G +
Sbjct: 386 SKEVAPDNEFEKRIRPEVIPANEIGVTFADIGSLDETKESLQELVMLPLRRPDLFKGGLL 445
Query: 265 RPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARA 324
+P +G+L+FGPPGTGKT++AKA+A E G +F NVS +T+ SKW GE E+ VR LF LA
Sbjct: 446 KPCRGILLFGPPGTGKTMMAKAIANEAGASFINVSMSTITSKWFGEDEKNVRALFTLAAK 505
Query: 325 YAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAA 384
+P+ IF+DE+DS+ R GEHE+ R++K+E + DG+ + A G R ++VLAA
Sbjct: 506 VSPTIIFVDEVDSMLGQRTRVGEHEAMRKIKNEFMTHWDGLMSNA----GDR--ILVLAA 559
Query: 385 TNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSG 444
TN P+D+DEA+ RR E+RI + LP+ ESR++++R L + E +++ E+A+ TDGYSG
Sbjct: 560 TNRPFDLDEAIIRRFERRIMVGLPSVESREKILR-TLLSKEKTENLDFQELAQMTDGYSG 618
Query: 445 DDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKD 482
DL N C A+ +R I + + + +++ K+
Sbjct: 619 SDLKNFCTTAAYRPVRELIKQECLKDQERRKREEAEKN 656
>At1g64110 unknown protein
Length = 824
Score = 217 bits (553), Expect = 1e-56
Identities = 140/387 (36%), Positives = 217/387 (55%), Gaps = 36/387 (9%)
Query: 160 RGAKAGATSKVNTGTRASTTGKKSGVSGKAG-KTDSV--ISDAEESKSK--------KSQ 208
R KAG K+ T+ ++ + S K KT+SV +S EE + + K+
Sbjct: 430 REGKAGGREKLKQKTKEESSKEVKAESIKPETKTESVTTVSSKEEPEKEAKAEKVTPKAP 489
Query: 209 YEGPDPELAEMLERDVLETSP-GVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQG-IRRP 266
PD E + + +V+ V ++D+ L E K L+E V+LPL P+ F G + +P
Sbjct: 490 EVAPDNEFEKRIRPEVIPAEEINVTFKDIGALDEIKESLQELVMLPLRRPDLFTGGLLKP 549
Query: 267 WKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYA 326
+G+L+FGPPGTGKT+LAKA+A E G +F NVS +T+ SKW GE E+ VR LF LA +
Sbjct: 550 CRGILLFGPPGTGKTMLAKAIAKEAGASFINVSMSTITSKWFGEDEKNVRALFTLASKVS 609
Query: 327 PSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATN 386
P+ IF+DE+DS+ R GEHE+ R++K+E + DG+ + G R ++VLAATN
Sbjct: 610 PTIIFVDEVDSMLGQRTRVGEHEAMRKIKNEFMSHWDGL----MTKPGER--ILVLAATN 663
Query: 387 FPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDD 446
P+D+DEA+ RR E+RI + LP E+R++++R L +V +++ E+A T+GY+G D
Sbjct: 664 RPFDLDEAIIRRFERRIMVGLPAVENREKILRTLLAKEKVDENLDYKELAMMTEGYTGSD 723
Query: 447 LTNVCRDASLNGMRRKI---------AGKTRDEIKNMSKDD------ISKDPVAMCDFEE 491
L N+C A+ +R I K R+ K +D+ I+ P+ DF+E
Sbjct: 724 LKNLCTTAAYRPVRELIQQERIKDTEKKKQREPTKAGEEDEGKEERVITLRPLNRQDFKE 783
Query: 492 ALVKVQRSVSQ--ADIERHEKWFHEFG 516
A +V S + A + ++W +G
Sbjct: 784 AKNQVAASFAAEGAGMGELKQWNELYG 810
>At1g50140 hypothetical protein, 5' partial
Length = 601
Score = 216 bits (549), Expect = 3e-56
Identities = 115/291 (39%), Positives = 183/291 (62%), Gaps = 13/291 (4%)
Query: 230 GVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTLLAKAV 287
GV++ED+ L + K+ L E V+LP+ PE F + RP KG+L+FGPPGTGKTLLAKA+
Sbjct: 308 GVKFEDIGALEDVKKALNELVILPMRRPELFARGNLLRPCKGILLFGPPGTGKTLLAKAL 367
Query: 288 ATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGE 347
ATE G F +++ +TL SKW G++E++ + LF A AP IF+DEIDSL +RG S E
Sbjct: 368 ATEAGANFISITGSTLTSKWFGDAEKLTKALFSFATKLAPVIIFVDEIDSLLGARGGSSE 427
Query: 348 HESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPL 407
HE++RR+++E + DG+ ++D R +++L ATN P+D+D+A+ RRL +RIY+ L
Sbjct: 428 HEATRRMRNEFMAAWDGL----RSKDSQR--ILILGATNRPFDLDDAVIRRLPRRIYVDL 481
Query: 408 PNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIAGKT 467
P+ E+R ++++I L + D +++A+ T+GYSG DL N+C A+ ++ + +
Sbjct: 482 PDAENRLKILKIFLTPENLESDFQFEKLAKETEGYSGSDLKNLCIAAAYRPVQELLQEEQ 541
Query: 468 RDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHE--KWFHEFG 516
+ S S +++ DF ++ KV SV+ +E KW ++G
Sbjct: 542 KGARAEASPGLRS---LSLDDFIQSKAKVSPSVAYDATTMNELRKWNEQYG 589
>At4g24850 unknown protein (At4g24850)
Length = 442
Score = 204 bits (519), Expect = 1e-52
Identities = 117/315 (37%), Positives = 188/315 (59%), Gaps = 16/315 (5%)
Query: 197 SDAEESKSKKSQYEGPDPELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWM 256
+D + S S K + +L +L + V ++D+ L + K +L+E V+LPL
Sbjct: 103 NDLKGSTSSKKDIVVENVFEKRLLSDVILPSDIDVTFDDIGALEKVKDILKELVMLPLQR 162
Query: 257 PEYF--QGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERM 314
PE F + +P KG+L+FGPPGTGKT+LAKAVA E F N+S +++ SKW GE E+
Sbjct: 163 PELFCKGELTKPCKGILLFGPPGTGKTMLAKAVAKEADANFINISMSSITSKWFGEGEKY 222
Query: 315 VRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDG 374
V+ +F LA +PS IF+DE+DS+ R EHE+SR++K+E ++ DG++
Sbjct: 223 VKAVFSLASKMSPSVIFVDEVDSMLGRREHPREHEASRKIKNEFMMHWDGLTTQ------ 276
Query: 375 SRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDE 434
R+ V+VLAATN P+D+DEA+ RRL +R+ + LP+ +R ++++ L +++PD++I E
Sbjct: 277 ERERVLVLAATNRPFDLDEAVIRRLPRRLMVGLPDTSNRAFILKVILAKEDLSPDLDIGE 336
Query: 435 VARRTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDI------SKDPVAMC- 487
+A T+GYSG DL N+C A+ ++ + + R+ +++ + S D A+
Sbjct: 337 IASMTNGYSGSDLKNLCVTAAHRPIKEILEKEKRERDAALAQGKVPPPLSGSSDLRALNV 396
Query: 488 -DFEEALVKVQRSVS 501
DF +A V SVS
Sbjct: 397 EDFRDAHKWVSASVS 411
>At1g02890 hypothetical protein
Length = 1240
Score = 201 bits (510), Expect = 1e-51
Identities = 126/331 (38%), Positives = 194/331 (58%), Gaps = 25/331 (7%)
Query: 201 ESKSKKSQYEG--PDPELAEMLERDVLETSP-GVRWEDVAGLTEAKRLLEEAVVLPLWMP 257
E+KS K + + E + L DV+ S GV + D+ L K L+E V+LPL P
Sbjct: 906 ENKSTKKSLKDVVTENEFEKKLLSDVIPPSDIGVSFSDIGALENVKDTLKELVMLPLQRP 965
Query: 258 EYFQG--IRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMV 315
E F + +P KG+L+FGPPGTGKT+LAKAVATE G F N+S +++ SK E+ V
Sbjct: 966 ELFGKGQLTKPTKGILLFGPPGTGKTMLAKAVATEAGANFINISMSSITSK----GEKYV 1021
Query: 316 RCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGS 375
+ +F LA APS IF+DE+DS+ R GEHE+ R++K+E ++ DG+
Sbjct: 1022 KAVFSLASKIAPSVIFVDEVDSMLGRRENPGEHEAMRKMKNEFMINWDGLRTK------D 1075
Query: 376 RKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEV 435
++ V+VLAATN P+D+DEA+ RRL +R+ + LP+ +R +++ + L E+A DV+++ +
Sbjct: 1076 KERVLVLAATNRPFDLDEAVIRRLPRRLMVNLPDSANRSKILSVILAKEEMAEDVDLEAI 1135
Query: 436 ARRTDGYSGDDLTNVCRDASLNGMR------RKIAGKTRDEIKNMSKDDISKD--PVAMC 487
A TDGYSG DL N+C A+ +R +K + E + M + S D P+ M
Sbjct: 1136 ANMTDGYSGSDLKNLCVTAAHLPIREILEKEKKERSVAQAENRAMPQLYSSTDVRPLNMN 1195
Query: 488 DFEEALVKVQRSVS--QADIERHEKWFHEFG 516
DF+ A +V SV+ +++ ++W +G
Sbjct: 1196 DFKTAHDQVCASVASDSSNMNELQQWNELYG 1226
>At5g03340 transitional endoplasmic reticulum ATPase
Length = 810
Score = 196 bits (498), Expect = 3e-50
Identities = 118/303 (38%), Positives = 173/303 (56%), Gaps = 25/303 (8%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGI-RRPWKGVLMFGPPGTGK 280
R+ + P V WED+ GL KR L+E V P+ PE F+ P KGVL +GPPG GK
Sbjct: 468 RETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGK 527
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKA+A EC F +V L + W GESE VR +FD AR AP +F DE+DS+
Sbjct: 528 TLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIAT 587
Query: 341 SRG--ASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR- 397
RG A ++ RV ++LL ++DG++ ++K V ++ ATN P ID AL R
Sbjct: 588 QRGNSAGDAGGAADRVLNQLLTEMDGMN--------AKKTVFIIGATNRPDIIDSALLRP 639
Query: 398 -RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASL 456
RL++ IYIPLP+ +SR + + L+ VA DV++ +A+ T G+SG D+T +C+ A
Sbjct: 640 GRLDQLIYIPLPDEDSRLNIFKACLRKSPVAKDVDVTALAKYTQGFSGADITEICQRACK 699
Query: 457 NGMRRKIAGKTRDEIK----------NMSKDDISKDPVAMCDFEEALVKVQRSVSQADIE 506
+R I +E + +M D++S+ + FEE++ +RSVS ADI
Sbjct: 700 YAIRENIEKDIENERRRSQNPEAMEEDMVDDEVSE--IRAAHFEESMKYARRSVSDADIR 757
Query: 507 RHE 509
+++
Sbjct: 758 KYQ 760
Score = 168 bits (426), Expect = 6e-42
Identities = 104/258 (40%), Positives = 156/258 (60%), Gaps = 15/258 (5%)
Query: 212 PDPEL---AEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPW 267
PD E+ E ++R+ E V ++DV G+ + + E V LPL P+ F+ I +P
Sbjct: 182 PDTEIFCEGEPVKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPP 241
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
KG+L++GPPG+GKTL+A+AVA E G FF ++ + SK GESE +R F+ A AP
Sbjct: 242 KGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAP 301
Query: 328 STIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNF 387
S IFIDEIDS+ R + E RR+ S+LL +DG+ SR V+V+ ATN
Sbjct: 302 SIIFIDEIDSIAPKREKT-NGEVERRIVSQLLTLMDGLK--------SRAHVIVMGATNR 352
Query: 388 PWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGD 445
P ID ALRR R ++ I I +P+ R E++RI+ K +++A DV+++ +++ T GY G
Sbjct: 353 PNSIDPALRRFGRFDREIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERISKDTHGYVGA 412
Query: 446 DLTNVCRDASLNGMRRKI 463
DL +C +A+L +R K+
Sbjct: 413 DLAALCTEAALQCIREKM 430
>At3g53230 CDC48 - like protein
Length = 815
Score = 194 bits (494), Expect = 8e-50
Identities = 117/299 (39%), Positives = 170/299 (56%), Gaps = 19/299 (6%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGI-RRPWKGVLMFGPPGTGK 280
R+ + P V WED+ GL KR L+E V P+ PE F+ P KGVL +GPPG GK
Sbjct: 469 RETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGK 528
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKA+A EC F ++ L + W GESE VR +FD AR AP +F DE+DS+
Sbjct: 529 TLLAKAIANECQANFISIKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIAT 588
Query: 341 SRGAS--GEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR- 397
RG S ++ RV ++LL ++DG++ ++K V ++ ATN P ID AL R
Sbjct: 589 QRGNSVGDAGGAADRVLNQLLTEMDGMN--------AKKTVFIIGATNRPDIIDPALLRP 640
Query: 398 -RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASL 456
RL++ IYIPLP+ ESR ++ + L+ VA DV++ +A+ T G+SG D+T +C+ +
Sbjct: 641 GRLDQLIYIPLPDEESRYQIFKSCLRKSPVAKDVDLRALAKYTQGFSGADITEICQRSCK 700
Query: 457 NGMRRKIAGKTRDEIKN------MSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHE 509
+R I E K M +D+ + FEE++ +RSVS ADI +++
Sbjct: 701 YAIRENIEKDIEKERKRAESPEAMEEDEEEIAEIKAGHFEESMKYARRSVSDADIRKYQ 759
Score = 169 bits (428), Expect = 3e-42
Identities = 105/258 (40%), Positives = 156/258 (59%), Gaps = 15/258 (5%)
Query: 212 PDPEL---AEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPW 267
PD E+ E ++R+ E V ++DV G+ + + E V LPL P+ F+ I +P
Sbjct: 183 PDTEIFCEGEPIKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPP 242
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
KG+L++GPPG+GKTL+A+AVA E G FF ++ + SK GESE +R F+ A AP
Sbjct: 243 KGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAP 302
Query: 328 STIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNF 387
S IFIDEIDS+ R + E RR+ S+LL +DG+ SR V+V+ ATN
Sbjct: 303 SIIFIDEIDSIAPKREKT-HGEVERRIVSQLLTLMDGLK--------SRAHVIVMGATNR 353
Query: 388 PWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGD 445
P ID ALRR R ++ I I +P+ R E++RI+ K +++A DV+++ V++ T GY G
Sbjct: 354 PNSIDPALRRFGRFDREIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERVSKDTHGYVGA 413
Query: 446 DLTNVCRDASLNGMRRKI 463
DL +C +A+L +R K+
Sbjct: 414 DLAALCTEAALQCIREKM 431
>At3g09840 putative transitional endoplasmic reticulum ATPase
Length = 809
Score = 193 bits (491), Expect = 2e-49
Identities = 115/301 (38%), Positives = 169/301 (55%), Gaps = 21/301 (6%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGI-RRPWKGVLMFGPPGTGK 280
R+ + P V W D+ GL KR L+E V P+ PE F+ P KGVL +GPPG GK
Sbjct: 468 RETVVEVPNVSWNDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGK 527
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKA+A EC F +V L + W GESE VR +FD AR AP +F DE+DS+
Sbjct: 528 TLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIAT 587
Query: 341 SRGASGEHE---SSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR 397
RG + ++ RV ++LL ++DG++ ++K V ++ ATN P ID AL R
Sbjct: 588 QRGGGSGGDGGGAADRVLNQLLTEMDGMN--------AKKTVFIIGATNRPDIIDSALLR 639
Query: 398 --RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDAS 455
RL++ IYIPLP+ +SR + + L+ +A DV+I +A+ T G+SG D+T +C+ A
Sbjct: 640 PGRLDQLIYIPLPDEDSRLNIFKAALRKSPIAKDVDIGALAKYTQGFSGADITEICQRAC 699
Query: 456 LNGMRRKIAGKTRDEIKN------MSKDDISK-DPVAMCDFEEALVKVQRSVSQADIERH 508
+R I E + M +D + + + FEE++ +RSVS ADI ++
Sbjct: 700 KYAIRENIEKDIEKEKRRSENPEAMEEDGVDEVSEIKAAHFEESMKYARRSVSDADIRKY 759
Query: 509 E 509
+
Sbjct: 760 Q 760
Score = 169 bits (427), Expect = 4e-42
Identities = 104/258 (40%), Positives = 156/258 (60%), Gaps = 15/258 (5%)
Query: 212 PDPEL---AEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPW 267
PD E+ E ++R+ E V ++DV G+ + + E V LPL P+ F+ I +P
Sbjct: 182 PDTEIFCEGEPVKREDEERLDDVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPP 241
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
KG+L++GPPG+GKTL+A+AVA E G FF ++ + SK GESE +R F+ A AP
Sbjct: 242 KGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAP 301
Query: 328 STIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNF 387
S IFIDEIDS+ R + E RR+ S+LL +DG+ SR V+V+ ATN
Sbjct: 302 SIIFIDEIDSIAPKREKT-NGEVERRIVSQLLTLMDGLK--------SRAHVIVMGATNR 352
Query: 388 PWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGD 445
P ID ALRR R ++ I I +P+ R E++RI+ K +++A DV+++ +++ T GY G
Sbjct: 353 PNSIDPALRRFGRFDREIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERISKDTHGYVGA 412
Query: 446 DLTNVCRDASLNGMRRKI 463
DL +C +A+L +R K+
Sbjct: 413 DLAALCTEAALQCIREKM 430
>At3g56690 calmodulin-binding protein
Length = 1022
Score = 179 bits (455), Expect = 3e-45
Identities = 121/311 (38%), Positives = 174/311 (55%), Gaps = 24/311 (7%)
Query: 209 YEGPDPELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGI-RRPW 267
+E ++ R+V+ P V WEDV G E K L EAV P + F+ I RP
Sbjct: 699 FENAKTKIRPSAMREVILEVPKVNWEDVGGQNEVKNQLMEAVEWPQKHQDAFKRIGTRPP 758
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
G+LMFGPPG KTL+A+AVA+E F V L SKW GESE+ VR LF ARA AP
Sbjct: 759 SGILMFGPPGCSKTLMARAVASEAKLNFLAVKGPELFSKWVGESEKAVRSLFAKARANAP 818
Query: 328 STIFIDEIDSLCNSRGASGEHES-SRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATN 386
S IF DEIDSL + RG + S S RV S+LLV++DG+ R V V+AATN
Sbjct: 819 SIIFFDEIDSLASIRGKENDGVSVSDRVMSQLLVELDGLH--------QRVGVTVIAATN 870
Query: 387 FPWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSG 444
P ID AL R R ++ +Y+ PN R+ +++I+L+ + + D+ + E+A T GY+G
Sbjct: 871 RPDKIDSALLRPGRFDRLLYVGPPNETDREAILKIHLRKIPCSSDICLKELASITKGYTG 930
Query: 445 DDLTNVCRDASLNGMRRKIAGKTRDEIKNMS----KDDISK-DPVAMCDFEEALVKVQRS 499
D++ +CR+A++ + + E++ +S K IS+ +P + ++ K QR
Sbjct: 931 ADISLICREAAIAALEESL------EMEEISMRHLKAAISQIEPTEILSYKALSEKFQRL 984
Query: 500 VSQADIERHEK 510
V D +R E+
Sbjct: 985 V-HTDPQREEE 994
Score = 128 bits (321), Expect = 9e-30
Identities = 81/224 (36%), Positives = 134/224 (59%), Gaps = 17/224 (7%)
Query: 265 RPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARA 324
RP KGVL+ GPPGTGKT LA+ A G FF+V+ + S++ GESE+ + +F A
Sbjct: 416 RPTKGVLIHGPPGTGKTSLARTFARHSGVNFFSVNGPEIISQYLGESEKALDEVFRSASN 475
Query: 325 YAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAA 384
P+ +FID++D++ +R GE E S+R+ + LL +DG+S T DG V+V+AA
Sbjct: 476 ATPAVVFIDDLDAIAPARKEGGE-ELSQRMVATLLNLMDGISRT----DG----VVVIAA 526
Query: 385 TNFPWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVA-PDVNIDEVARRTDG 441
TN P I+ ALRR RL++ I I +P+ R +++ I L+ + + ++ ++++A T G
Sbjct: 527 TNRPDSIEPALRRPGRLDREIEIGVPSSTQRSDILHIILRGMRHSLSNIQVEQLAMATHG 586
Query: 442 YSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVA 485
+ G DL+ +C +A+ +RR + D+ + S + + P+A
Sbjct: 587 FVGADLSALCCEAAFVCLRRHL-----DQSSSSSNLPLEEAPIA 625
>At4g27680 unknown protein
Length = 398
Score = 179 bits (453), Expect = 4e-45
Identities = 105/302 (34%), Positives = 166/302 (54%), Gaps = 30/302 (9%)
Query: 215 ELAEMLERDVLETSP---------------GVRWEDVAGLTEAKRLLEEAVVLPLWMPEY 259
E+++ L R +++T+P V + + GL K+ L E V+LPL PE
Sbjct: 50 EISKRLGRPLVQTNPYEDVIACDVINPDHIDVEFGSIGGLETIKQALYELVILPLKRPEL 109
Query: 260 FQ--GIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRC 317
F + P KGVL++GPPGTGKT+LAKA+A E G F NV + L SKW G+++++V
Sbjct: 110 FAYGKLLGPQKGVLLYGPPGTGKTMLAKAIAKESGAVFINVRVSNLMSKWFGDAQKLVSA 169
Query: 318 LFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRK 377
+F LA P+ IFIDE++S R S +HE+ +K+E + DG S
Sbjct: 170 VFSLAYKLQPAIIFIDEVESFLGQR-RSTDHEAMANMKTEFMALWDGFST------DPHA 222
Query: 378 IVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVAR 437
VMVLAATN P ++DEA+ RRL + I +P+ R E++++ LK V PD++ D +AR
Sbjct: 223 RVMVLAATNRPSELDEAILRRLPQAFEIGIPDRRERAEILKVTLKGERVEPDIDFDHIAR 282
Query: 438 RTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQ 497
+GY+G D+ +C+ A+ +R + + + K + P++ D E+ L +
Sbjct: 283 LCEGYTGSDIFELCKKAAYFPIREIL------DAERKGKPCLDPRPLSQLDLEKVLATSK 336
Query: 498 RS 499
++
Sbjct: 337 KT 338
>At2g45500 hypothetical protein
Length = 481
Score = 176 bits (445), Expect = 4e-44
Identities = 131/385 (34%), Positives = 193/385 (50%), Gaps = 70/385 (18%)
Query: 139 STRKSDGAWARGATPRGGATGRG--------AKAGATSKVNTGTR-ASTTGKKSGVSGKA 189
S R+ W + R A G G A + S +T +R T +K+ V+
Sbjct: 111 SYREKISNWQNQVSERLQALGVGMSENKRTVAYPSSASVSSTASRYRKTLSQKTPVARGG 170
Query: 190 GKTDSVISDAEES-KSKKSQYEGPDPELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEE 248
T DA S K K D +L EM+ +++ SP V+W+DVAGL AK+ L E
Sbjct: 171 VATPRNPKDAAASPKPVKESGNVYDDKLVEMINTTIVDRSPSVKWDDVAGLNGAKQALLE 230
Query: 249 AVVLPLWMPEYFQGIRRPWK-----GVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATL 303
V+LP + F G+RRP + G+L+FGPPG GKT+LAKAVA+E TFFNVS+++L
Sbjct: 231 MVILPAKRRDLFTGLRRPARVTSLLGLLLFGPPGNGKTMLAKAVASESQATFFNVSASSL 290
Query: 304 ASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVD 363
SKW IDS+ ++R S E+E+SRR+KSE L+Q D
Sbjct: 291 TSKW---------------------------IDSIMSTRSTS-ENEASRRLKSEFLIQFD 322
Query: 364 GVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLK- 422
GV+ +N D +V+++ ATN P ++D+A+ RRL KRIY+PLP+ RK L + LK
Sbjct: 323 GVT---SNPD---DLVIIIGATNKPQELDDAVLRRLVKRIYVPLPDSNVRKLLFKTKLKC 376
Query: 423 TVEVAPDVNIDEVARRTDG--------------------YSGDDLTNVCRDASLNGMRRK 462
D +ID++ + T+G YSG DL +C +A++ +R
Sbjct: 377 QPHSLSDGDIDKIVKETEGKLYRLCIKKHRFISQVTDKRYSGSDLQALCEEAAMMPIREL 436
Query: 463 IAGKTRDEIKNMSKDDISKDPVAMC 487
A + + +S+ V +C
Sbjct: 437 GANILTIQANKVLNFSVSQINVEVC 461
>At5g53540 26S proteasome regulatory particle chain RPT6-like
protein
Length = 403
Score = 166 bits (421), Expect = 2e-41
Identities = 96/271 (35%), Positives = 154/271 (56%), Gaps = 15/271 (5%)
Query: 231 VRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTLLAKAVA 288
V + + GL K+ L E V+LPL PE F + P KGVL++GPPGTGKT+LAKA+A
Sbjct: 84 VEFGSIGGLESIKQALYELVILPLKRPELFAYGKLLGPQKGVLLYGPPGTGKTMLAKAIA 143
Query: 289 TECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEH 348
E F NV + L SKW G+++++V +F LA P+ IFIDE+DS R S ++
Sbjct: 144 RESEAVFINVKVSNLMSKWFGDAQKLVSAVFSLAYKLQPAIIFIDEVDSFLGQR-RSTDN 202
Query: 349 ESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLP 408
E+ +K+E + DG T + +R VMVLAATN P ++DEA+ RR + I +P
Sbjct: 203 EAMSNMKTEFMALWDGF----TTDQNAR--VMVLAATNRPSELDEAILRRFPQSFEIGMP 256
Query: 409 NFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIAGKTR 468
+ + R +++++ LK V D+N D +AR + Y+G D+ +C+ A+ +R + +
Sbjct: 257 DCQERAQILKVVLKGESVESDINYDRIARLCEDYTGSDIFELCKKAAYFPIREILEAEKE 316
Query: 469 DEIKNMSKDDISKDPVAMCDFEEALVKVQRS 499
+ ++ + P+ D E+ L +++
Sbjct: 317 GKRVSVPR------PLTQLDLEKVLATSKKT 341
>At1g62130 hypothetical protein
Length = 372
Score = 166 bits (420), Expect = 3e-41
Identities = 117/341 (34%), Positives = 177/341 (51%), Gaps = 57/341 (16%)
Query: 201 ESKSKKSQYEGPDPELAEMLERDVLETSP-GVRWEDVAGLTEAKRLLEEAVVLPLWMPEY 259
E +SKKS + E+ D++ S GV ++D+ L K L+E V+LP PE
Sbjct: 50 EIESKKSLKDIVTENTFEI--SDIIPPSEIGVTFDDIGALENVKDTLKELVMLPFQWPEL 107
Query: 260 F----------------------QGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFN 297
F +P G+L+FGP GTGKT+LAKAVATE G N
Sbjct: 108 FCKGQLTKMLTLWIGGFLISLLLYFSTQPCNGILLFGPSGTGKTMLAKAVATEAGANLIN 167
Query: 298 VSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSE 357
+S S+W E E+ V+ +F LA +PS IF+DE++S+ H + K+E
Sbjct: 168 MSM----SRWFSEGEKYVKAVFSLASKISPSIIFLDEVESML--------HRYRLKTKNE 215
Query: 358 LLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELI 417
++ DG+ TNE ++ V+VLAATN P+D+DEA+ RRL R+ + LP+ SR +++
Sbjct: 216 FIINWDGLR---TNE---KERVLVLAATNRPFDLDEAVIRRLPHRLMVGLPDARSRSKIL 269
Query: 418 RINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKD 477
++ L +++PD +IDEVA T+GYSG+DL RDA++ R AG +++ +
Sbjct: 270 KVILSKEDLSPDFDIDEVASMTNGYSGNDLKE--RDAAVAEGRVPPAGSGGSDLRVLK-- 325
Query: 478 DISKDPVAMCDFEEALVKVQRSVSQADIERH--EKWFHEFG 516
M DF AL V S+S + +W ++G
Sbjct: 326 --------MEDFRNALELVSMSISSKSVNMTALRQWNEDYG 358
>At1g03000 putative peroxisome assembly factor-2
Length = 941
Score = 164 bits (415), Expect = 1e-40
Identities = 100/292 (34%), Positives = 169/292 (57%), Gaps = 13/292 (4%)
Query: 229 PGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVA 288
P V+W+DV GL + K + + V LPL + F R GVL++GPPGTGKTLLAKAVA
Sbjct: 653 PNVKWDDVGGLEDVKTSILDTVQLPLLHKDLFSSGLRKRSGVLLYGPPGTGKTLLAKAVA 712
Query: 289 TECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEH 348
TEC F +V L + + GESE+ VR +F+ AR+ P IF DE+DSL +RGASG+
Sbjct: 713 TECSLNFLSVKGPELINMYIGESEKNVRDIFEKARSARPCVIFFDELDSLAPARGASGDS 772
Query: 349 ES-SRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR--RLEKRIYI 405
RV S++L ++DG+S+++ + + ++ A+N P ID AL R R +K +Y+
Sbjct: 773 GGVMDRVVSQMLAEIDGLSDSSQD-------LFIIGASNRPDLIDPALLRPGRFDKLLYV 825
Query: 406 PL-PNFESRKELIRINLKTVEVAPDVNIDEVARRTDG-YSGDDLTNVCRDASLNGMRRKI 463
+ + R+ +++ + +++ DV++ VA++ ++G D+ +C DA +RK+
Sbjct: 826 GVNADASYRERVLKALTRKFKLSEDVSLYSVAKKCPSTFTGADMYALCADAWFQAAKRKV 885
Query: 464 AGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKWFHEF 515
+ ++ +DD V DF +A+ ++ S+S +++++E +F
Sbjct: 886 SKSDSGDMPT-EEDDPDSVVVEYVDFIKAMDQLSPSLSITELKKYEMLRDQF 936
Score = 51.6 bits (122), Expect = 1e-06
Identities = 54/210 (25%), Positives = 89/210 (41%), Gaps = 22/210 (10%)
Query: 270 VLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPST 329
VL+ G PG GK + K VA G S +L + ++ + F++AR Y+P+
Sbjct: 380 VLLHGIPGCGKRTVVKYVARRLGLHVVEFSCHSLLASSERKTSTALAQTFNMARRYSPTI 439
Query: 330 IFIDEID----------SLCNSRGASGEHESSRRVKSELLVQVDGV------SNTATNED 373
+ + D SL + G S E S R +E + D SN + NE
Sbjct: 440 LLLRHFDVFKNLGSQDGSLGDRVGVSFEIASVIRELTEPVSNGDSSMEEKSNSNFSENEV 499
Query: 374 GSRK--IVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLKTVEVAPDVN 431
G + V+++A+ I +RR I + N E R E++ +L+ V +++
Sbjct: 500 GKFRGHQVLLIASAESTEGISPTIRRCFSHEIRMGSLNDEQRSEMLSQSLQGVSQFLNIS 559
Query: 432 IDEVAR----RTDGYSGDDLTNVCRDASLN 457
DE + +T G+ DL + DA N
Sbjct: 560 SDEFMKGLVGQTSGFLPRDLQALVADAGAN 589
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.131 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,697,928
Number of Sequences: 26719
Number of extensions: 513145
Number of successful extensions: 2140
Number of sequences better than 10.0: 189
Number of HSP's better than 10.0 without gapping: 126
Number of HSP's successfully gapped in prelim test: 64
Number of HSP's that attempted gapping in prelim test: 1776
Number of HSP's gapped (non-prelim): 265
length of query: 518
length of database: 11,318,596
effective HSP length: 104
effective length of query: 414
effective length of database: 8,539,820
effective search space: 3535485480
effective search space used: 3535485480
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0226.8


