

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.7
(399 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
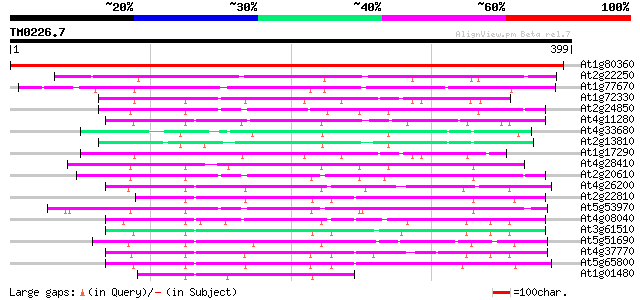
Score E
Sequences producing significant alignments: (bits) Value
At1g80360 putative aspartate aminotransferase 585 e-167
At2g22250 putative aspartate aminotransferase 139 3e-33
At1g77670 aminotransferase like protein 91 9e-19
At1g72330 alanine aminotransferase (ALAAT2) 76 3e-14
At2g24850 putative tyrosine aminotransferase 75 6e-14
At4g11280 ACC synthase (AtACS-6) 74 1e-13
At4g33680 unknown protein 74 1e-13
At2g13810 putative aspartate aminotransferase 72 4e-13
At1g17290 alanine aminotransferase (ALAAT1) 70 2e-12
At4g28410 unknown protein 67 1e-11
At2g20610 putative tyrosine aminotransferase 67 2e-11
At4g26200 1-aminocyclopropane-1-carboxylate synthase -like protein 66 3e-11
At2g22810 1-aminocyclopropane-1-carboxylate synthase (ACS4) 64 1e-10
At5g53970 tyrosine aminotransferase 64 2e-10
At4g08040 putative protein 63 2e-10
At3g61510 1-aminocyclopropane-1-carboxylate synthase-like protein 60 3e-09
At5g51690 ACS12 (MIO24.18) 59 4e-09
At4g37770 1-aminocyclopropane-1-carboxylate synthase - like pro... 59 5e-09
At5g65800 1-aminocyclopropane-1-carboxylate synthase ACS5 (pir||... 57 1e-08
At1g01480 1-aminocyclopropane-1-carboxylate synthase (ACC2) 56 3e-08
>At1g80360 putative aspartate aminotransferase
Length = 394
Score = 585 bits (1508), Expect = e-167
Identities = 274/394 (69%), Positives = 335/394 (84%)
Query: 1 MGSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPS 60
MGS+ L+RR L T+MP+M Q++ ++ N +SLAQGVV+WQPP+ AL+KVKELVW+P
Sbjct: 1 MGSFGMLSRRTLGTDMPVMAQIRSLMAELTNPMSLAQGVVHWQPPQKALEKVKELVWDPI 60
Query: 61 VSRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
+S YG DEGLPELR AL+KKLR+EN L S VMVTAGANQAFVNLV+TLCDAGDSVVMF
Sbjct: 61 ISSYGPDEGLPELRQALLKKLREENKLTNSQVMVTAGANQAFVNLVITLCDAGDSVVMFE 120
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
PYYFN+YM+FQMTG+TNI VGPG DTLYPD DWLER L+ESKP PK+VTVVNPGNP+GT
Sbjct: 121 PYYFNSYMAFQMTGVTNIIVGPGQSDTLYPDADWLERTLSESKPTPKVVTVVNPGNPSGT 180
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEGNHIVNIFSFSKAYGMMGW 240
Y+PE LL+RIA +CK+AG WL+VDNTYEYFM+DGLKH CVEG+HIVN+FSFSK YGMMGW
Sbjct: 181 YVPEPLLKRIAQICKDAGCWLIVDNTYEYFMYDGLKHCCVEGDHIVNVFSFSKTYGMMGW 240
Query: 241 RVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNRE 300
R+GYIAY ++G + +L+K+QDNIPICA+IISQ LA+++LE G W+ ERV++L KNR+
Sbjct: 241 RLGYIAYSERLDGFATELVKIQDNIPICAAIISQRLAVYALEEGSGWITERVKSLVKNRD 300
Query: 301 IVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN 360
IV EAL PLG+ +VKGGEGAIYL+AKLP+G DDF VVRWLA+RHG+ VIPG A G+ G
Sbjct: 301 IVKEALEPLGKENVKGGEGAIYLWAKLPEGHRDDFKVVRWLAHRHGVVVIPGCASGSPGY 360
Query: 361 LRISFGGLTESDCRAAAERLKKGLEELVANGLVQ 394
LR+SFGGL E + RAAA RL+KG+EEL+ +G+V+
Sbjct: 361 LRVSFGGLQEVEMRAAAARLRKGIEELLHHGMVE 394
>At2g22250 putative aspartate aminotransferase
Length = 475
Score = 139 bits (350), Expect = 3e-33
Identities = 108/379 (28%), Positives = 184/379 (48%), Gaps = 31/379 (8%)
Query: 33 VSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQENNLHKS-- 90
+ LA G + PK + + E +RY + G+ ELR A+ +KL++EN L +
Sbjct: 102 IRLAAGEPDFDTPKVVAEAGINAIRE-GFTRYTLNAGITELREAICRKLKEENGLSYAPD 160
Query: 91 SVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYP 150
++V+ GA Q+ + VL +C GD V++ APY+ + ++ T + + +
Sbjct: 161 QILVSNGAKQSLLQAVLAVCSPGDEVIIPAPYWVSYTEQARLADATPVVIPTKISNNFLL 220
Query: 151 DVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIAN-LCKNAGSWLLVDNTYEY 209
D LE LTE +L+ + +P NPTG+ P+SLL+ IA + K+ +L D YE+
Sbjct: 221 DPKDLESKLTEKS---RLLILCSPSNPTGSVYPKSLLEEIARIIAKHPRLLVLSDEIYEH 277
Query: 210 FMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDN 264
++ H+ + + FSKA+ M GWR+GY+A P + A K+Q
Sbjct: 278 IIYAPATHTSFASLPDMYERTLTVNGFSKAFAMTGWRLGYLAGPKHI---VAACSKLQGQ 334
Query: 265 IPICASIISQHLALHSLEMGP---EWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAI 321
+ AS I+Q + +L +G E V E V+ + R+ + ++L + + +GA
Sbjct: 335 VSSGASSIAQKAGVAALGLGKAGGETVAEMVKAYRERRDFLVKSLGDIKGVKISEPQGAF 394
Query: 322 YLFAKL------PKGGY----DDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTES 371
YLF G+ D + + ++ +A++PG A G +RIS+ T
Sbjct: 395 YLFIDFSAYYGSEAEGFGLINDSSSLALYFLDKFQVAMVPGDAFGDDSCIRISYA--TSL 452
Query: 372 D-CRAAAERLKKGLEELVA 389
D +AA E+++K LE L A
Sbjct: 453 DVLQAAVEKIRKALEPLRA 471
>At1g77670 aminotransferase like protein
Length = 440
Score = 91.3 bits (225), Expect = 9e-19
Identities = 94/397 (23%), Positives = 165/397 (40%), Gaps = 28/397 (7%)
Query: 7 LARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEP---SVSR 63
+A+R + + I QM +L ++L QG + P D VKE + ++
Sbjct: 57 VAKRLEKFKTTIFTQMS-ILAVKHGAINLGQGFPNFDGP----DFVKEAAIQAIKDGKNQ 111
Query: 64 YGADEGLPELRAALVKKLRQENNL---HKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
Y G+P+L +A+ + R++ L + V VT+G +A +L L + GD V++FA
Sbjct: 112 YARGYGIPQLNSAIAARFREDTGLVVDPEKEVTVTSGCTEAIAAAMLGLINPGDEVILFA 171
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
P+Y + + M G + PD P LE + + + + P NPTG
Sbjct: 172 PFYDSYEATLSMAGAKVKGITLRPPDFSIP----LEELKAAVTNKTRAILMNTPHNPTGK 227
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMF--DGLKHSCVEG--NHIVNIFSFSKAYG 236
L+ IA+LC + D Y+ F D + + + G V + S K +
Sbjct: 228 MFTREELETIASLCIENDVLVFSDEVYDKLAFEMDHISIASLPGMYERTVTMNSLGKTFS 287
Query: 237 MMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLA 296
+ GW++G+ P L+ + + + S +Q A+ +L+ + +E +
Sbjct: 288 LTGWKIGWAIAPPH---LTWGVRQAHSYLTFATSTPAQWAAVAALKAPESYFKELKRDYN 344
Query: 297 KNREIVSEALSPLGEGSVKGGEGAIYLFA-KLPKGGYDDFDVVRWLANRHGIAVIPGSAC 355
+E + + L +G +V G ++ A P G +D +L G+ IP S
Sbjct: 345 VKKETLVKGLKEVG-FTVFPSSGTYFVVADHTPFGMENDVAFCEYLIEEVGVVAIPTSVF 403
Query: 356 ---GAAGNLRISFGGL-TESDCRAAAERLKKGLEELV 388
G + F E R A ER+K+ L+ V
Sbjct: 404 YLNPEEGKNLVRFAFCKDEETLRGAIERMKQKLKRKV 440
>At1g72330 alanine aminotransferase (ALAAT2)
Length = 540
Score = 76.3 bits (186), Expect = 3e-14
Identities = 82/337 (24%), Positives = 146/337 (42%), Gaps = 49/337 (14%)
Query: 64 YGADEGLPELRAALVKKLRQENNL--HKSSVMVTAGANQAF-VNLVLTLCDAGDSVVMFA 120
Y +G+ LR + + + + + +T GA+ A + + L L D ++
Sbjct: 170 YSHSQGIKGLRDVIAAGIEARDGFPADPNDIFLTDGASPAVHMMMQLLLSSEKDGILSPI 229
Query: 121 PYY--FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVP---KLVTVVNPG 175
P Y ++A ++ + ++ + L ++ L++ L E++ + + V+NPG
Sbjct: 230 PQYPLYSASIALHGGSLVPYYLDEATGWGL--EISDLKKQLEEARSKGISVRALVVINPG 287
Query: 176 NPTGTYIPESLLQRIANLCKNAGSWLLVDNTYE---------YFMFDGLKHSCVEGNHIV 226
NPTG + E + I N CK G LL D Y+ + F + S G +
Sbjct: 288 NPTGQVLAEENQRDIVNFCKQEGLVLLADEVYQENVYVPDKKFHSFKKVARSLGYGEKDI 347
Query: 227 NIFSFSKA----YGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLE 282
++ SF YG G R GY+ + Q+ K+ ++ +C++I Q LA SL
Sbjct: 348 SLVSFQSVSKGYYGECGKRGGYMEVTGFTSDVREQIYKMA-SVNLCSNISGQILA--SLV 404
Query: 283 MGP---------EWVRER---VQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAK--LP 328
M P ++ ER + ++AK + + +AL+ L + EGA+YLF + LP
Sbjct: 405 MSPPKPGDDSYDSYMAERDGILSSMAKRAKTLEDALNSLEGVTCNRAEGAMYLFPRINLP 464
Query: 329 KGGYD---------DFDVVRWLANRHGIAVIPGSACG 356
+ + D + L N G+ V+PGS G
Sbjct: 465 QKAIEAAEAEKTAPDAFYCKRLLNATGVVVVPGSGFG 501
>At2g24850 putative tyrosine aminotransferase
Length = 445
Score = 75.1 bits (183), Expect = 6e-14
Identities = 85/338 (25%), Positives = 149/338 (43%), Gaps = 27/338 (7%)
Query: 64 YGADEGLPELRAALVKKLRQE--NNLHKSSVMVTAGANQAFVNLVLTLC-DAGDSVVMFA 120
Y G+ + R A+ + L E L V +T G NQA ++ +L + ++++
Sbjct: 87 YAPSPGVFKARRAVAEYLNGELPTKLKAEDVYITGGCNQAIEIVIDSLAGNPSANILLPR 146
Query: 121 PYY--FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPT 178
P Y ++A + I + P S + ++D LE E+ + ++NP NP
Sbjct: 147 PGYPHYDARAVYSGLEIRKYDLLPESDWEI--NLDGLEAAADENTVA---MVIINPNNPC 201
Query: 179 GTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEGNH-----IVNIFSFSK 233
G L ++A + + G ++ D Y++ ++ G K G ++ + S SK
Sbjct: 202 GNVYTYDHLNKVAEMARKLGIMIISDEVYDHVVY-GDKPFIPMGKFASIAPVITLGSISK 260
Query: 234 AYGMMGWRVGYIAY--PSEVEGLSAQLLKVQDNIPIC--ASIISQHLALHSLEMGP-EWV 288
+ GWRVG+IA P+ + + + ++D + + S I Q LE P E+
Sbjct: 261 GWVNPGWRVGWIAMNDPNGIFVSTGVVQAIEDFLDLTPQPSFILQEALPDILEKTPKEFF 320
Query: 289 RERVQTLAKNREIVSEALSPLG-EGSVKGGEGAIYLFAKLPKGGY----DDFDVVRWLAN 343
++++ + +N E+ E L + K E YL+ KL +DFD L +
Sbjct: 321 EKKIKAMRRNVELSCERLKDIPCLFCPKKPESCSYLWLKLDTSMLNNIKNDFDFCTKLVS 380
Query: 344 RHGIAVIPGSACGAAGNLRISFGGLTESDCRAAAERLK 381
+ +IPG A GA +RIS G ES + +RLK
Sbjct: 381 EESLILIPGVALGAENWVRISI-GTDESVVQEIFDRLK 417
>At4g11280 ACC synthase (AtACS-6)
Length = 495
Score = 74.3 bits (181), Expect = 1e-13
Identities = 88/344 (25%), Positives = 143/344 (40%), Gaps = 39/344 (11%)
Query: 69 GLPELRAALVKKLRQENN----LHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY- 123
GLPE R A+ K + + N ++++ GA A + L + GD ++ PYY
Sbjct: 98 GLPEFRQAVAKFMEKTRNNKVKFDPDRIVMSGGATGAHETVAFCLANPGDGFLVPTPYYP 157
Query: 124 -FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESK----PVPKLVTVVNPGNPT 178
F+ + ++ TG+ + V S + V+ LE ++ PV L+ V NP NP
Sbjct: 158 GFDRDLRWR-TGVNLVPVTCHSSNGFKITVEALEAAYENARKSNIPVKGLL-VTNPSNPL 215
Query: 179 GTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCV----------EGNHIVNI 228
GT + L+ + N + G L+ D Y F + V + I +
Sbjct: 216 GTTLDRECLKSLVNFTNDKGIHLIADEIYAATTFGQSEFISVAEVIEEIEDCNRDLIHIV 275
Query: 229 FSFSKAYGMMGWRVGYI-AYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSL---EMG 284
+S SK G+ G RVG + +Y V Q+ + + + +S +QHL L E
Sbjct: 276 YSLSKDMGLPGLRVGIVYSYNDRV----VQIARKMSSFGLVSS-QTQHLIAKMLSDEEFV 330
Query: 285 PEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLF---AKLPKGGYDDFDVVRWL 341
E++RE LA ++ L LG G +K G ++L+ L K D + W
Sbjct: 331 DEFIRESKLRLAARHAEITTGLDGLGIGWLKAKAG-LFLWMDLRNLLKTATFDSETELWR 389
Query: 342 ANRHGIA--VIPGSA--CGAAGNLRISFGGLTESDCRAAAERLK 381
H + V PG + C G R+ F + A ER++
Sbjct: 390 VIVHQVKLNVSPGGSFHCHEPGWFRVCFANMDHKTMETALERIR 433
>At4g33680 unknown protein
Length = 461
Score = 73.9 bits (180), Expect = 1e-13
Identities = 93/353 (26%), Positives = 137/353 (38%), Gaps = 52/353 (14%)
Query: 51 KVKELVWEPSVSRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLC 110
K EL S YGA++G LRAA+ K + V V+ GA C
Sbjct: 116 KAHELSTIEGYSGYGAEQGAKPLRAAIAKTFYGGLGIGDDDVFVSDGAK----------C 165
Query: 111 DAGDSVVMFA------------PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERI 158
D VMF P Y ++ + TG N V Y +++++ R
Sbjct: 166 DISRLQVMFGSNVTIAVQDPSYPAYVDSSVIMGQTGQFNTDVQK------YGNIEYM-RC 218
Query: 159 LTESKPVPKLVTV--------VNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYF 210
E+ P L TV +P NPTG L ++ K GS ++ D+ Y +
Sbjct: 219 TPENGFFPDLSTVGRTDIIFFCSPNNPTGAAATREQLTQLVEFAKKNGSIIVYDSAYAMY 278
Query: 211 MFDGLKHSCVE----GNHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIP 266
M D S E + SFSK G G R+G+ P ++ + N
Sbjct: 279 MSDDNPRSIFEIPGAEEVAMETASFSKYAGFTGVRLGWTVIPKKLLYSDGFPVAKDFNRI 338
Query: 267 IC-----ASIISQHLALHSL-EMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGA 320
IC AS ISQ AL L G E + + + +N I+ + + LG V GG+ A
Sbjct: 339 ICTCFNGASNISQAGALACLTPEGLEAMHKVIGFYKENTNIIIDTFTSLGY-DVYGGKNA 397
Query: 321 IYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRISFGGLTES 371
Y++ P +DV + + + PGS G G +R+S G E+
Sbjct: 398 PYVWVHFP--NQSSWDVFAEILEKTHVVTTPGSGFGPGGEGFVRVSAFGHREN 448
>At2g13810 putative aspartate aminotransferase
Length = 456
Score = 72.4 bits (176), Expect = 4e-13
Identities = 83/341 (24%), Positives = 133/341 (38%), Gaps = 54/341 (15%)
Query: 64 YGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA--- 120
YG ++G LR A+ + ++ ++ + V V+ GA L L L G +V +
Sbjct: 108 YGLEQGNKTLRKAIAETFYRDLHVKSNEVFVSDGAQSDISRLQLLL---GSNVTIAVQDP 164
Query: 121 --PYYFNAYMSFQMTGITN---------IHVGPGSPDTLYPDVDWLERILTESKPVPKLV 169
P Y ++ + TG + +++ G ++ +PD+ P ++
Sbjct: 165 TFPAYIDSSVIIGQTGHFHEKTKKYQNVVYMPCGPNNSFFPDL--------AMTPRTDVI 216
Query: 170 TVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVE----GNHI 225
+P NPTG L ++ + K GS ++ D+ Y F+ DG S E
Sbjct: 217 FFCSPNNPTGYVASRKQLHQLVDFAKTNGSIIIFDSAYAAFIEDGSPRSIYEIPGAREVA 276
Query: 226 VNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPIC-------------ASII 272
+ + SFSK G G R+G+ P E L + PI AS I
Sbjct: 277 IEVSSFSKFAGFTGVRLGWSIIPDE--------LLYSNGFPIINDFHRIVTTSFNGASNI 328
Query: 273 SQHLALHSLEMGP-EWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGG 331
+Q L L G + +R +NR+I+ + L LG V GG A YL+ G
Sbjct: 329 AQAGGLACLSSGGLKEIRSVNNYYKENRKILMDTLVSLGL-KVYGGVNAPYLWVHFK--G 385
Query: 332 YDDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTESD 372
+DV + I +PGS G G + G D
Sbjct: 386 SKSWDVFNEILENTHIITVPGSGFGPGGEEYLRISGFGRRD 426
>At1g17290 alanine aminotransferase (ALAAT1)
Length = 488
Score = 70.1 bits (170), Expect = 2e-12
Identities = 81/346 (23%), Positives = 142/346 (40%), Gaps = 47/346 (13%)
Query: 51 KVKELVWEPSVSRYGADEGLPELRAALVKKLRQENNL--HKSSVMVTAGANQAF-VNLVL 107
K+ + + + Y +G+ LR A+ + + + + +T GA+ + + L
Sbjct: 105 KILDQIPGRATGAYSHSQGIKGLRDAIADGIEARDGFPADPNDIFMTDGASPGVHMMMQL 164
Query: 108 TLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKP--- 164
+ D ++ P Y S + G T + ++ L++ L +++
Sbjct: 165 LITSEKDGILCPIPQYPLYSASIALHGGTLVPYYLDEASGWGLEISELKKQLEDARSKGI 224
Query: 165 VPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYE---------YFMFDGL 215
+ + V+NPGNPTG + E + + CK G LL D Y+ + F +
Sbjct: 225 TVRALAVINPGNPTGQVLSEENQRDVVKFCKQEGLVLLADEVYQENVYVPDKKFHSFKKV 284
Query: 216 KHSCVEGNHIVNIFSFSKA----YGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASI 271
S G + + SF YG G R GY+ + Q+ K+ ++ +C++I
Sbjct: 285 ARSMGYGEKDLALVSFQSVSKGYYGECGKRGGYMEVTGFTSDVREQIYKMA-SVNLCSNI 343
Query: 272 ISQHLALHSLEMGP---------EWVRER---VQTLAKNREIVSEALSPLGEGSVKGGEG 319
Q LA SL M P ++ E+ + +LA+ + + EAL+ L + EG
Sbjct: 344 SGQILA--SLIMSPPKPGDDSYESYIAEKDGILSSLARRAKTLEEALNKLEGVTCNRAEG 401
Query: 320 AIYLF------------AKLPKGGYDDFDVVRWLANRHGIAVIPGS 353
A+YLF A+ K D+F R L GI V+PGS
Sbjct: 402 AMYLFPCLHLPQKAIAAAEAEKTAPDNFYCKR-LLKATGIVVVPGS 446
>At4g28410 unknown protein
Length = 447
Score = 67.4 bits (163), Expect = 1e-11
Identities = 75/346 (21%), Positives = 141/346 (40%), Gaps = 29/346 (8%)
Query: 42 WQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQE--NNLHKSSVMVTAGAN 99
+Q A + V E + + + Y G+ R A+ L ++ + +H + +T G
Sbjct: 84 FQTSVDAEEAVVESLRSGAANSYAPGVGILPARRAVANYLNRDLPHKIHSDDIFMTVGCC 143
Query: 100 QAFVNLVLTLCDAGDSVVMFAPYY--FNAYMSFQMTGITNIHVGPGSPDTLYPDVDW--- 154
Q ++ L ++++ Y +N++ + I + L PD+DW
Sbjct: 144 QGIETMIHALAGPKANILLPTLIYPLYNSHAIHSLVEIRKYN--------LLPDLDWEID 195
Query: 155 LERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDG 214
L+ + + V ++NP NP G L+++A + + G ++ D Y ++
Sbjct: 196 LQGVEAMADENTIAVVIMNPHNPCGNVYTYEHLKKVAEVARKLGIMVISDEVYNQTIYGE 255
Query: 215 LKHSCV----EGNHIVNIFSFSKAYGMMGWRVGYIAY--PSEVEGLSAQLLKVQDNIPIC 268
K + +V + S SK + + GWR+G+IA P V + + +++++ I
Sbjct: 256 NKFVPMGIFSSITPVVTLGSISKGWLVPGWRIGWIAMNDPKNVFKTTRVVESIKEHLDIS 315
Query: 269 --ASIISQHLALHSLE-MGPEWVRERVQTLAKNREIVSEALSPLG-EGSVKGGEGAIYLF 324
S I Q + LE E+ + L++N + +AL + K E YL
Sbjct: 316 PDPSTILQFALPNILEKTKKEFFEKNNSILSQNVDFAFDALKDIPCLTCPKKPESCTYLV 375
Query: 325 AKLP----KGGYDDFDVVRWLANRHGIAVIPGSACGAAGNLRISFG 366
KL + +DFD LA + +PG G +R S G
Sbjct: 376 TKLDLSLLEDITNDFDFCMKLAQEENLVFLPGEVLGLKNWVRFSIG 421
>At2g20610 putative tyrosine aminotransferase
Length = 462
Score = 67.0 bits (162), Expect = 2e-11
Identities = 80/355 (22%), Positives = 147/355 (40%), Gaps = 30/355 (8%)
Query: 48 ALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQE--NNLHKSSVMVTAGANQAFVNL 105
A D V +++ + YG G+ R A+ + ++ + L + +TAG NQ +
Sbjct: 88 AEDAVVDVLRSGKGNSYGPGAGILPARRAVADYMNRDLPHKLTPEDIFLTAGCNQGIEIV 147
Query: 106 VLTLCDAGDSVVMFAPYY--FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESK 163
+L ++++ P + ++A ++ + + P + D++ +E I E+
Sbjct: 148 FESLARPNANILLPRPGFPHYDARAAYSGLEVRKFDLLPEKEWEI--DLEGIEAIADENT 205
Query: 164 PVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFD-------GLK 216
+ V+NP NP G L+++A + G ++ D Y+ +F G
Sbjct: 206 VA---MVVINPNNPCGNVYSHDHLKKVAETARKLGIMVISDEVYDRTIFGDNPFVSMGKF 262
Query: 217 HSCVEGNHIVNIFSFSKAYGMMGWRVGYIAY--PSEVEGLSAQLLKVQDNI---PICASI 271
S V ++ + SK + + GW++G+IA P V + L ++ N+ P A+I
Sbjct: 263 ASIVP---VLTLAGISKGWVVPGWKIGWIALNDPEGVFETTKVLQSIKQNLDVTPDPATI 319
Query: 272 ISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLG-EGSVKGGEGAIYLFAKLP-- 328
I L + + ++ + L N ++V + L + K E YL KL
Sbjct: 320 IQAALPAILEKADKNFFAKKNKILKHNVDLVCDRLKDIPCVVCPKKPESCTYLLTKLELS 379
Query: 329 --KGGYDDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTESDCRAAAERLK 381
DD D LA + +PG A G +RI+ G+ A ERLK
Sbjct: 380 LMDNIKDDIDFCVKLAREENLVFLPGDALGLKNWMRITI-GVEAHMLEDALERLK 433
>At4g26200 1-aminocyclopropane-1-carboxylate synthase -like protein
Length = 447
Score = 66.2 bits (160), Expect = 3e-11
Identities = 79/352 (22%), Positives = 145/352 (40%), Gaps = 46/352 (13%)
Query: 69 GLPELRAALVKKLRQ----ENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY- 123
GL R A+ + Q + +++TAGA A L L D D++++ PYY
Sbjct: 102 GLKTFRQAMASFMEQIRGGKARFDPDRIVLTAGATAANELLTFILADPNDALLVPTPYYP 161
Query: 124 -FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVP---KLVTVVNPGNPTG 179
F+ + ++ TG+ + + S + + LE ++ + V + NP NP G
Sbjct: 162 GFDRDLRWR-TGVKIVPIHCDSSNHFQITPEALESAYQTARDANIRVRGVLITNPSNPLG 220
Query: 180 TYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCV-------------EGNHIV 226
+ + +L+ + + C L+ D Y +F + + V E HIV
Sbjct: 221 ATVQKKVLEDLLDFCVRKNIHLVSDEIYSGSVFHASEFTSVAEIVENIDDVSVKERVHIV 280
Query: 227 NIFSFSKAYGMMGWRVGYI-AYPSEV----EGLSAQLLKVQDNIPICASIISQHLALHSL 281
+S SK G+ G+RVG I +Y V +S+ L + AS++S
Sbjct: 281 --YSLSKDLGLPGFRVGTIYSYNDNVVRTARRMSSFTLVSSQTQHMLASMLSDE------ 332
Query: 282 EMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAI------YLFAKLPKGGYDDF 335
E +++R + L + + + E L G +KG G +L K K G +
Sbjct: 333 EFTEKYIRINRERLRRRYDTIVEGLKKAGIECLKGNAGLFCWMNLGFLLEKKTKDG--EL 390
Query: 336 DVVRWLANRHGIAVIPGSA--CGAAGNLRISFGGLTESDCRAAAERLKKGLE 385
+ + + + PGS+ C G R+ F ++E+ A +R+ + ++
Sbjct: 391 QLWDVILKELNLNISPGSSCHCSEVGWFRVCFANMSENTLEIALKRIHEFMD 442
>At2g22810 1-aminocyclopropane-1-carboxylate synthase (ACS4)
Length = 474
Score = 64.3 bits (155), Expect = 1e-10
Identities = 72/317 (22%), Positives = 136/317 (42%), Gaps = 27/317 (8%)
Query: 90 SSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY--FNAYMSFQMTGITNIHVGPGSPDT 147
+++++TAGA A L+ L D GD+ ++ PYY F+ + ++ TG+ + + S +
Sbjct: 112 NNLVLTAGATSANETLMFCLADPGDAFLLPTPYYPGFDRDLKWR-TGVEIVPIQSSSTNG 170
Query: 148 LYPDVDWLERILTESKPVP---KLVTVVNPGNPTGTYIPESLLQRIAN-LCKNAGSWLLV 203
LE ++K + K + + NP NP GT ++ L + + + KN L+
Sbjct: 171 FRITKLALEEAYEQAKKLDLNVKGILITNPSNPLGTTTTQTELNILFDFITKNKNIHLVS 230
Query: 204 DNTYEYFMFDG---------LKHSCVEGNHIVN----IFSFSKAYGMMGWRVGYIAYPSE 250
D Y +F+ LK++ +E ++N + S SK G+ G+RVG I Y ++
Sbjct: 231 DEIYSGTVFNSSEFISVMEILKNNQLENTDVLNRVHIVCSLSKDLGLPGFRVGAI-YSND 289
Query: 251 VEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLG 310
+ +SA + + L + ++RE + L + + L +G
Sbjct: 290 KDVISAATKMSSFGLVSSQTQYLLSSLLSDKKFTKNYLRENQKRLKNRQRKLVLGLEAIG 349
Query: 311 EGSVKGGEGAIYLFAKLP----KGGYDDFDVVRWLANRHGIAVIPGSACGA--AGNLRIS 364
+K G P K + D+ + + + + PGS+C G R+
Sbjct: 350 IKCLKSNAGLFCWVDMRPLLRSKTFEAEMDLWKKIVYEVKLNISPGSSCHCEEPGWFRVC 409
Query: 365 FGGLTESDCRAAAERLK 381
F + + + A +RLK
Sbjct: 410 FANMIDETLKLALKRLK 426
>At5g53970 tyrosine aminotransferase
Length = 414
Score = 63.5 bits (153), Expect = 2e-10
Identities = 83/380 (21%), Positives = 159/380 (41%), Gaps = 37/380 (9%)
Query: 28 GAKNCVSLAQG--VVY--WQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQ 83
G K +SL G +Y ++ + +L V + + Y GLP+ R A+ + L +
Sbjct: 31 GGKRVISLGMGDPTLYSCFRTTQVSLQAVSDSLLSNKFHGYSPTVGLPQARRAIAEYLSR 90
Query: 84 E--NNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY--FNAYMSFQMTGITNIH 139
+ L + V +T+G QA + L ++++ P + + F+ + +
Sbjct: 91 DLPYKLSQDDVFITSGCTQAIDVALSMLARPRANILLPRPGFPIYELCAKFRHLEVRYVD 150
Query: 140 VGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGS 199
+ P + + D+D +E + E+ + V+NPGNP G L +IA K G
Sbjct: 151 LLPENGWEI--DLDAVEALADENTVA---LVVINPGNPCGNVYSYQHLMKIAESAKKLGF 205
Query: 200 WLLVDNTYEYFMFD-------GLKHSCVEGNHIVNIFSFSKAYGMMGWRVGYIAY--PS- 249
++ D Y + F G+ S V ++ + S SK + + GWR+G+ PS
Sbjct: 206 LVIADEVYGHLAFGSKPFVPMGVFGSIVP---VLTLGSLSKRWIVPGWRLGWFVTTDPSG 262
Query: 250 --EVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALS 307
+ + + K D + A+ I + + + ++ + +L + +I + +
Sbjct: 263 SFKDPKIIERFKKYFDILGGPATFIQAAVPTILEQTDESFFKKTLNSLKNSSDICCDWIK 322
Query: 308 PLG-EGSVKGGEGAIYLFAKLP----KGGYDDFDVVRWLANRHGIAVIPGSACGAAGNLR 362
+ S EG++ + KL + DD D LA + ++PG+A G LR
Sbjct: 323 EIPCIDSSHRPEGSMAMMVKLNLSLLEDVSDDIDFCFKLAREESVILLPGTAVGLKNWLR 382
Query: 363 ISFGGLTESDCRAAAERLKK 382
I+F +D + E K+
Sbjct: 383 ITFA----ADATSIEEAFKR 398
>At4g08040 putative protein
Length = 460
Score = 63.2 bits (152), Expect = 2e-10
Identities = 87/344 (25%), Positives = 144/344 (41%), Gaps = 39/344 (11%)
Query: 69 GLPELRAALVK---KLRQENNLHKSSVMV-TAGANQAFVNLVLTLCDAGDSVVMFAPYY- 123
GLP + A+ K K+R+ ++ MV TAG+ A L+ L + GD+ ++ APYY
Sbjct: 85 GLPAFKDAMAKFMGKIRENKVKFDTNKMVLTAGSTSANETLMFCLANPGDAFLIPAPYYP 144
Query: 124 -FNAYMSFQMTG--ITNIHVGPGSPDTLYPDV--DWLERILTESKPVPKLVTVVNPGNPT 178
F+ + ++ TG I IH + + D D ER L + V K V + NP NP
Sbjct: 145 GFDRDLKWR-TGVEIVPIHCVSSNGYKITEDALEDAYERALKHNLNV-KGVLITNPSNPL 202
Query: 179 GTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCV-----------EGN-HIV 226
GT L + ++ D Y +FD + + V +G H+V
Sbjct: 203 GTSTTREELDLLLTFTSTKKIHMVSDEIYSGTVFDSPEFTSVLEVAKDKNMGLDGKIHVV 262
Query: 227 NIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSL---EM 283
+S SK G+ G+RVG I Y + + +SA + S +QHL + L
Sbjct: 263 --YSLSKDLGLPGFRVGLI-YSNNEKVVSAATKMSSFGL---ISSQTQHLLANLLSDERF 316
Query: 284 GPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYL--FAKLPKGGYDDFDVVRW- 340
++ E + L + ++ + L G +K G L K + + W
Sbjct: 317 TTNYLEENKKRLRERKDRLVSGLKEAGISCLKSNAGLFCWVDLRHLLKSNTFEAEHSLWT 376
Query: 341 -LANRHGIAVIPGSA--CGAAGNLRISFGGLTESDCRAAAERLK 381
+ G+ + PGS+ C G R+ F +++ A +R+K
Sbjct: 377 KIVCEVGLNISPGSSCHCDEPGWFRVCFANMSDQTMEVAMDRVK 420
>At3g61510 1-aminocyclopropane-1-carboxylate synthase-like protein
Length = 488
Score = 59.7 bits (143), Expect = 3e-09
Identities = 74/337 (21%), Positives = 138/337 (39%), Gaps = 28/337 (8%)
Query: 69 GLPELRAALVKKLRQENN----LHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY- 123
GL + R A+ + + V+++ GA A ++ L D GD+ ++ PYY
Sbjct: 94 GLKQFRQAIATFMERARGGRVRFEAERVVMSGGATGANETIMFCLADPGDAFLVPTPYYA 153
Query: 124 -FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYI 182
F+ + ++ TG+ I V S + LE +++ + + NP GT +
Sbjct: 154 AFDRDLRWR-TGVRIIPVECSSSNNFQITKQALESAYLKAQETGIKIKGLIISNPLGTSL 212
Query: 183 PESLLQRIANLCKNAGSWLLVDNTYEYFMFDG----LKHSCVEGNHIVN------IFSFS 232
L+ + + + L+ D Y +F ++ + VN ++S S
Sbjct: 213 DRETLESLVSFINDKQIHLVCDEIYAATVFAEPGFISVAEIIQEMYYVNRDLIHIVYSLS 272
Query: 233 KAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLA--LHSLEMGPEWVRE 290
K G+ G+RVG + ++V A+ + + + +S LA L ++ E
Sbjct: 273 KDMGLPGFRVGVVYSYNDVVVSCARRM---SSFGLVSSQTQSFLAAMLSDQSFVDNFLVE 329
Query: 291 RVQTLAKNREIVSEALSPLGEGSVKGGEGAIYL--FAKLPKGGYDDFDVVRW--LANRHG 346
+ +AK + +E L +G ++ G L + K D ++ W + N+
Sbjct: 330 VSKRVAKRHHMFTEGLEEMGISCLRSNAGLFVLMDLRHMLKDQTFDSEMALWRVIINKVK 389
Query: 347 IAVIPGSA--CGAAGNLRISFGGLTESDCRAAAERLK 381
I V PGS+ C G R+ F + E + A ER+K
Sbjct: 390 INVSPGSSFHCSEPGWFRVCFANMDEDTLQIALERIK 426
>At5g51690 ACS12 (MIO24.18)
Length = 495
Score = 59.3 bits (142), Expect = 4e-09
Identities = 79/352 (22%), Positives = 143/352 (40%), Gaps = 36/352 (10%)
Query: 60 SVSRYGADEGLPELRAALVKKLRQ----ENNLHKSSVMVTAGANQAFVNLVLTLCDAGDS 115
S++ Y EGL ELR A + + + S++++TAG A L L D G++
Sbjct: 144 SIAMYKPFEGLLELRVAFADFMSRIMGGNVSFDPSNMVITAGGTPAIEVLAFCLADHGNA 203
Query: 116 VVMFAPYY--FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVV- 172
++ PYY F+ + F+ TG+ I V S D V LE+ L +++ V+ +
Sbjct: 204 FLIPTPYYPGFDRDIKFR-TGVELIPVHCRSSDNFTVTVSALEQALNQARKRGSKVSGIL 262
Query: 173 --NPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCV---------E 221
NP NP G + L I + ++ D + ++ + + +
Sbjct: 263 FSNPSNPVGNILSRETLCDILRFAQEKNIHVISDEIFAGSVYGDKEFVSMAEIAGSGEFD 322
Query: 222 GNHIVNIFSFSKAYGMMGWRVGYI-AYPSEVEGLSAQLLKVQDNIPICASIISQHLALHS 280
+ I+ SK + G+R G I ++ +V + +L++ ++P+ I L L
Sbjct: 323 KTRVHIIYGLSKDLSIPGFRAGVIYSFHEDVVNAAKKLMRF-SSVPVLVQRILISL-LSD 380
Query: 281 LEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKL--------PKGGY 332
+ ++ Q + E L LG + G G +Y + + KG
Sbjct: 381 VRFIEGYMAAHRQRIRDKHIRFVEGLKQLGIPCAESG-GGLYCWVDMSSLLTSYSEKGEL 439
Query: 333 DDFDVVRWLANRHGIAVIPGSACGA--AGNLRISFGGLTESDCRAAAERLKK 382
+ F+ + +A I PG+AC G R F L + D ER+++
Sbjct: 440 ELFEKLLTVAK---INATPGTACYCIEPGWFRCCFTALADEDIPVIMERIRQ 488
>At4g37770 1-aminocyclopropane-1-carboxylate synthase - like
protein
Length = 469
Score = 58.9 bits (141), Expect = 5e-09
Identities = 78/349 (22%), Positives = 146/349 (41%), Gaps = 44/349 (12%)
Query: 69 GLPELRAALVKKLRQEN----NLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY- 123
GLP + A+ + + + + + +++TAGA A L+ L D GD+ ++ PYY
Sbjct: 87 GLPSFKNAMADFMSENRGNRVSFNPNKLVLTAGATPANETLMFCLADPGDAFLLPTPYYP 146
Query: 124 -FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPV---PKLVTVVNPGNPTG 179
F+ + ++ TG + + S + LE +++ + K V + NP NP G
Sbjct: 147 GFDRDLKWR-TGAEIVPIQCKSANGFRITKVALEEAYEQAQKLNLKVKGVLITNPSNPLG 205
Query: 180 TYIPESLLQRIANLCKNAGSWLLVDNTYEYFMF---------DGLKHSCVEGNHIVN--- 227
T + L + + L+ D Y +F + LK +E + +
Sbjct: 206 TTTTRTELNHLLDFISRKKIHLISDEIYSGTVFTNPGFISVMEVLKDRKLENTDVFDRVH 265
Query: 228 -IFSFSKAYGMMGWRVGYIAYPSEVEGLSAQ-------LLKVQDNIPICASIISQHLALH 279
++S SK G+ G+RVG I Y ++ +SA L+ Q + A + + +
Sbjct: 266 IVYSLSKDLGLPGFRVGVI-YSNDDFVVSAATKMSSFGLISSQTQYLLSALLSDKTFTKN 324
Query: 280 SLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYL--FAKLPKGGYDDFDV 337
LE +++ +++++VS L G +K G L K + ++
Sbjct: 325 YLE------ENQIRLKNRHKKLVS-GLEAAGIECLKSNAGLFCWVDMRHLLKSNTFEAEI 377
Query: 338 VRW--LANRHGIAVIPGSA--CGAAGNLRISFGGLTESDCRAAAERLKK 382
W + + + PGS+ C G R+ F L+E + A +RLK+
Sbjct: 378 ELWKKIVYEVKLNISPGSSCHCNEPGWFRVCFANLSEETLKVALDRLKR 426
>At5g65800 1-aminocyclopropane-1-carboxylate synthase ACS5
(pir||S71174)
Length = 470
Score = 57.4 bits (137), Expect = 1e-08
Identities = 71/345 (20%), Positives = 140/345 (40%), Gaps = 30/345 (8%)
Query: 69 GLPELRAALVKKLRQENN----LHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY- 123
G+PE + A+ + + + +++ AG+ A L+ L + GD+ ++ PYY
Sbjct: 87 GMPEFKKAMAEFMEEIRGNRVTFDPKKIVLAAGSTSANETLMFCLAEPGDAFLLPTPYYP 146
Query: 124 -FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVP---KLVTVVNPGNPTG 179
F+ + ++ TG + + S + L++ +++ + K V V NP NP G
Sbjct: 147 GFDRDLKWR-TGAEIVPIHCSSSNGFQITESALQQAYQQAQKLDLKVKGVLVTNPSNPLG 205
Query: 180 TYIPESLLQRIANLCKNAGSWLLVDNTYEYFMF---------DGLKHSCVEGNHIVN--- 227
T + L + + + L+ D Y MF D LK +E +
Sbjct: 206 TALTRRELNLLVDFITSKNIHLISDEIYSGTMFGFEQFISVMDVLKDKKLEDTEVSKRVH 265
Query: 228 -IFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPE 286
++S SK G+ G+RVG I Y ++ +SA + + L + +
Sbjct: 266 VVYSLSKDLGLPGFRVGAI-YSNDEMIVSAATKMSSFGLVSSQTQYLLSALLSDKKFTSQ 324
Query: 287 WVRERVQTLAKNREIVSEALSPLGEGSVKGGEGA---IYLFAKLPKGGYD-DFDVVRWLA 342
++ E + L + + L G ++ G + + L ++ + D+ + +
Sbjct: 325 YLEENQKRLKSRQRRLVSGLESAGITCLRSNAGLFCWVDMRHLLDTNTFEAELDLWKKIV 384
Query: 343 NRHGIAVIPGSACGAA--GNLRISFGGLTESDCRAAAERLKKGLE 385
+ + PGS+C G R+ F ++E A +RLK +E
Sbjct: 385 YNVKLNISPGSSCHCTEPGWFRVCFANMSEDTLDLALKRLKTFVE 429
>At1g01480 1-aminocyclopropane-1-carboxylate synthase (ACC2)
Length = 496
Score = 56.2 bits (134), Expect = 3e-08
Identities = 48/171 (28%), Positives = 76/171 (44%), Gaps = 18/171 (10%)
Query: 92 VMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY--FNAYMSFQMTGITNIHVGPGSPDTLY 149
V+++ GA A ++ L D GD ++ +PYY F+ + ++ TG+ I V S D
Sbjct: 122 VVMSGGATGANETIMFCLADPGDVFLIPSPYYAAFDRDLRWR-TGVEIIPVPCSSSDNFK 180
Query: 150 PDVD---WLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNT 206
VD W + ES K + + NP NP GT + + L + L+VD
Sbjct: 181 LTVDAAEWAYKKAQESNKKVKGLILTNPSNPLGTMLDKDTLTNLVRFVTRKNIHLVVDEI 240
Query: 207 YEYFMFDG------------LKHSCVEGNHIVNIFSFSKAYGMMGWRVGYI 245
Y +F G + S V + I ++S SK G+ G+RVG +
Sbjct: 241 YAATVFAGGDFVSVAEVVNDVDISEVNVDLIHIVYSLSKDMGLPGFRVGIV 291
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,207,706
Number of Sequences: 26719
Number of extensions: 399304
Number of successful extensions: 980
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 910
Number of HSP's gapped (non-prelim): 44
length of query: 399
length of database: 11,318,596
effective HSP length: 101
effective length of query: 298
effective length of database: 8,619,977
effective search space: 2568753146
effective search space used: 2568753146
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0226.7


