

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.6
(263 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
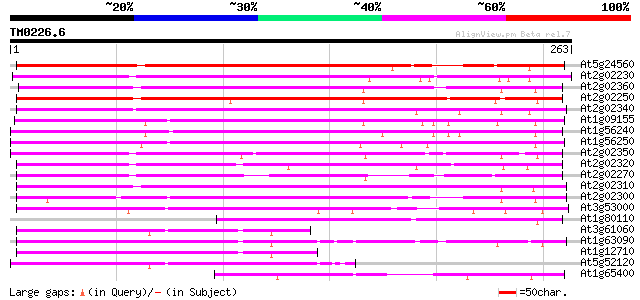
At5g24560 phloem-specific lectin-like protein 194 4e-50
At2g02230 putative phloem-specific lectin 193 7e-50
At2g02360 putative phloem-specific lectin 184 4e-47
At2g02250 lectin-like protein 184 4e-47
At2g02340 putative phloem-specific lectin 174 6e-44
At1g09155 unknown protein 173 1e-43
At1g56240 hypothetical protein 167 7e-42
At1g56250 hypothetical protein 161 4e-40
At2g02350 SKP1 interacting partner 3 (SKIP3) 155 2e-38
At2g02320 unknown protein 146 1e-35
At2g02270 putative phloem-specific lectin 144 4e-35
At2g02310 putative phloem-specific lectin 141 4e-34
At2g02300 putative phloem-specific lectin 135 2e-32
At3g53000 unknown protein 93 2e-19
At1g80110 unknown protein 92 3e-19
At3g61060 unknown protein 81 7e-16
At1g63090 unknown protein 78 4e-15
At1g12710 unknown protein 75 3e-14
At5g52120 unknown protein 73 1e-13
At1g65400 hypothetical protein 59 3e-09
>At5g24560 phloem-specific lectin-like protein
Length = 251
Score = 194 bits (493), Expect = 4e-50
Identities = 110/262 (41%), Positives = 161/262 (60%), Gaps = 24/262 (9%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CIA ++S T+P DA R+S VSK+ RSAADS+ W+RFLPSD + SLSR
Sbjct: 6 LPEECIATMISFTSPFDACRISAVSKLLRSAADSNTTWERFLPSDYRMYIDNSLSRF--- 62
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
S K+ +L + P++I++G+ SF ++K+SGKKC+ML AR L I W + + W ++P+S
Sbjct: 63 SNKQLFLRFCESPLLIEDGRTSFWMEKRSGKKCWMLSARKLDIVWVDSPEFWIWVSIPDS 122
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVN----N 179
RF +V L + W EI G I+T LS T Y A+LVFK + +F + L V+
Sbjct: 123 RFEEVAGLLMVCWFEIRGKISTSLLSKATNYSAYLVFKEQEMGSFGFESLPLEVSFRSTR 182
Query: 180 IGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSV 239
+N ++ +L + E R DGWLEIE+GE+++ G +DEE+EMSV
Sbjct: 183 TEVYNNRRV-FLKSGTQES--------------REDGWLEIELGEYYV-GFDDEEIEMSV 226
Query: 240 LQI-DGCFNVGLIIEGIEVRPK 260
L+ +G + G+I++GIE+RPK
Sbjct: 227 LETREGGWKGGIIVQGIEIRPK 248
>At2g02230 putative phloem-specific lectin
Length = 317
Score = 193 bits (491), Expect = 7e-50
Identities = 118/289 (40%), Positives = 168/289 (57%), Gaps = 31/289 (10%)
Query: 2 ELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRAN 61
+ LPE CI+ ++S T+P DA ++ VSK +SAA SD VW+ FLPS+ S+V QS AN
Sbjct: 33 DALPEDCISKVISHTSPRDACVVASVSKSVKSAAQSDLVWEMFLPSEYSSLVLQS---AN 89
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
SKK +L L+D+ V+++NGKKSF ++K SGKKC+ML A L I W + Y W T+P
Sbjct: 90 HLSKKEIFLSLADNSVLVENGKKSFWVEKASGKKCYMLSAMELTIIWGDSPAYWKWITVP 149
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPN--FHEVLVVLSVNN 179
ES+F V LR++ W E+ G I+ LS T Y ++VFK + + F V V V
Sbjct: 150 ESKFEKVAELRNVCWFEVRGKISCGMLSKGTHYSVYVVFKTANGRSYGFDLVPVEAGVGF 209
Query: 180 IGGHNITKIAYL---NPDS----------------LEGEVHDRWDRVQGPSVRSDGWLEI 220
+G K Y N DS +EGE +R V GP R DGW E+
Sbjct: 210 VGKVATKKSVYFESGNADSRSATSHYSGISEEEEEVEGE-RERGMNVVGPKERVDGWSEV 268
Query: 221 EMGEFFIS--GLED---EEVEMSVLQI-DGCFNVGLIIEGIEVRPKNDN 263
E+G+F+I+ G D +E+E+S+++ +G + GLII+GIE+RP+ N
Sbjct: 269 ELGKFYINNGGCGDDGSDEIEISIMETQNGNWKSGLIIQGIEIRPERSN 317
>At2g02360 putative phloem-specific lectin
Length = 272
Score = 184 bits (467), Expect = 4e-47
Identities = 107/259 (41%), Positives = 144/259 (55%), Gaps = 12/259 (4%)
Query: 5 PEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANASS 64
PE CI+ I+S T P DA + VSK F S SD +W++FLP+D S++ S SS
Sbjct: 18 PEDCISYIISFTNPRDACVAATVSKTFESTVKSDIIWEKFLPADYESLIPPSRV---FSS 74
Query: 65 KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESR 124
KK Y L + PV+ D+ KKS L+K SGK+C ML A L I W + Y W +PESR
Sbjct: 75 KKELYFSLCNDPVLFDDDKKSVWLEKASGKRCLMLSAMNLSIIWGDNPQYWQWIPIPESR 134
Query: 125 FPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIID-APNFHEVLVVLSVNNIGGH 183
F V LR + W EI G NT LSP T+Y A++VFK +D F V + +V +G
Sbjct: 135 FEKVAKLRDVCWFEIRGRTNTRVLSPRTRYSAYIVFKGVDKCYGFQNVAIEAAVGVVGQE 194
Query: 184 NITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISG--LEDEEVEMSVLQ 241
++ + G V P R DGW+EIE+GEFF G ++++E+EMS L+
Sbjct: 195 PSRRLICFSEAIRRGR-----RNVVKPKQREDGWMEIELGEFFNDGGIMDNDEIEMSALE 249
Query: 242 IDGC-FNVGLIIEGIEVRP 259
GLII+GIE+RP
Sbjct: 250 TKQLNRKCGLIIQGIEIRP 268
>At2g02250 lectin-like protein
Length = 305
Score = 184 bits (467), Expect = 4e-47
Identities = 110/260 (42%), Positives = 161/260 (61%), Gaps = 10/260 (3%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CI+ I+S T+P DA + VSK F SA +SD+VWD+FLPSD S+V S S
Sbjct: 45 LPEDCISNIISFTSPRDACVAASVSKTFESAVNSDSVWDKFLPSDYSSLVPPSRV---FS 101
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGAR-ALFIAWDNGDHYRCWTTLPE 122
SKK Y + D+PV++++G KSF L+K++GKKCFML + +++I W + Y W ++PE
Sbjct: 102 SKKELYFAICDNPVLVEDGGKSFWLEKENGKKCFMLSPKKSMWITWVSTPQYWRWISIPE 161
Query: 123 SRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIID-APNFHEVLVVLSVNNIG 181
+RF +V L ++ W E+ G +NT LSP T+Y A++VFK + PN +V V +V +G
Sbjct: 162 ARFEEVPELLNVCWFEVRGGMNTKELSPGTRYSAYIVFKTKNGCPNLGDVPVEATVGLVG 221
Query: 182 GHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFF-ISGLEDEEVEMSVL 240
+ + Y S + + D V P+ R DGW+E E+G+FF SG + V+ S+L
Sbjct: 222 QESSQRHIYFVGPSDQRRDRETRD-VTRPTKRKDGWMEAELGQFFNESGC--DVVDTSIL 278
Query: 241 QIDGCF-NVGLIIEGIEVRP 259
+I + GLII+GIE RP
Sbjct: 279 EIKTPYWKRGLIIQGIEFRP 298
>At2g02340 putative phloem-specific lectin
Length = 305
Score = 174 bits (440), Expect = 6e-44
Identities = 106/266 (39%), Positives = 152/266 (56%), Gaps = 9/266 (3%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE C++ I+S T+P DA L+ VSK F SA SD VW++F+P + S++SQS +
Sbjct: 39 LPEECVSIIVSFTSPQDACVLASVSKTFASAVKSDIVWEKFIPPEYESLISQSRA-FKFL 97
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK Y L D V+ID+GKKS ++K + K+C M+ A L IAW N W P++
Sbjct: 98 SKKELYFALCDKSVLIDDGKKSLWIEKANAKRCIMISAMNLAIAWGNSPQSWRWIPDPQA 157
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGH 183
RF V L + EI G IN+ +SP T+Y A++V+K ++ E + V V + G
Sbjct: 158 RFETVAELLEVCLFEIRGRINSRVISPKTRYSAYIVYKKLNICYGFENVAVEVVVGVVGQ 217
Query: 184 NITKIA--YLNPDSLEGEVHDRWDRVQG---PSVRSDGWLEIEMGEFFISG--LEDEEVE 236
++ + Y+ D E R DR + P R DGW+EI++GEFF G L D+E+E
Sbjct: 218 DLEESCRRYICFDETMDEQFRRRDRGKNLVKPERRKDGWMEIKIGEFFNEGGLLNDDEIE 277
Query: 237 MSVLQI-DGCFNVGLIIEGIEVRPKN 261
M L+ + GLII+GIE+RP N
Sbjct: 278 MVALEAKQRHWKRGLIIQGIEIRPTN 303
>At1g09155 unknown protein
Length = 289
Score = 173 bits (438), Expect = 1e-43
Identities = 112/280 (40%), Positives = 154/280 (55%), Gaps = 23/280 (8%)
Query: 3 LLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANA 62
+LPE C+A ILS TTP D + VS +FR A DSD VW++FLP+D ++S+S
Sbjct: 2 MLPEACVATILSFTTPADTISSAAVSSVFRVAGDSDFVWEKFLPTDYCHVISRSTDPHRI 61
Query: 63 -SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
SSKK Y L + ++IDNG+K F+++K SGK ++L +R L I W + HY W+
Sbjct: 62 FSSKKELYRCLCE-SILIDNGRKIFKIEKLSGKISYILSSRDLSITWSDQRHYWSWSPRS 120
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIID-APNFHEVLVVLSVNNI 180
+SRF + V L WLEI G I T +LSPNT YGA+L+ K+ A V S+
Sbjct: 121 DSRFSEGVQLIMTDWLEIIGKIQTGALSPNTNYGAYLIMKVTSRAYGLDLVPAETSIKVG 180
Query: 181 GGHNITKIAYLN-----PDSLE----GEVHDRW-------DRVQGPSVRSDGWLEIEMGE 224
G K YL+ +E G+ R + P VR DGW+EIE+GE
Sbjct: 181 NGEKKIKSTYLSCLDNKKQQMERVFYGQREQRMATHEVVRSHRREPEVRDDGWMEIELGE 240
Query: 225 FFI---SGLEDEEVEMSVLQIDGC-FNVGLIIEGIEVRPK 260
F G +D+EV MS+ ++ G G+ I+GIEVRPK
Sbjct: 241 FETGSGEGDDDKEVVMSLTEVKGYQLKGGIAIDGIEVRPK 280
>At1g56240 hypothetical protein
Length = 284
Score = 167 bits (422), Expect = 7e-42
Identities = 110/281 (39%), Positives = 155/281 (55%), Gaps = 23/281 (8%)
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRA 60
M +LPE C+A IL+ T+P DA S VS +FR A DSD VW++FLPS S++SQS
Sbjct: 1 MMMLPEACVANILAFTSPADAFSSSEVSSVFRLAGDSDFVWEKFLPSHYKSLISQSTDHH 60
Query: 61 NA-SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTT 119
SSKK Y L D ++IDN +K F+++K SGK ++L AR + I + + Y W+
Sbjct: 61 RIFSSKKEIYRCLCDS-LLIDNARKLFKINKFSGKISYILSARDISITYSDHASYCSWSN 119
Query: 120 LPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLV----VL 175
+ +SRF + L LEI G I T LSPNT+YGA+L+ K+ + +++ V
Sbjct: 120 VSDSRFSESAELITTDRLEIKGKIQTTVLSPNTKYGAYLIMKVTNGAYGLDLVPAETSVK 179
Query: 176 SVNNIGGHNITKIAYLNPDSLE------GEVHDRW----DRVQG------PSVRSDGWLE 219
S N N T + L+ + G +R + V G P R DGWLE
Sbjct: 180 SKNGQNNKNTTYLCCLDEKKQQMKRLFYGNREERMAMTVEAVGGDGKRREPKARDDGWLE 239
Query: 220 IEMGEFFISGLEDEEVEMSVLQIDGC-FNVGLIIEGIEVRP 259
IE+GEF ED+EV MS+ ++ G G++I+GIEVRP
Sbjct: 240 IELGEFVTREGEDDEVNMSLTEVKGYQLKGGIVIDGIEVRP 280
>At1g56250 hypothetical protein
Length = 282
Score = 161 bits (407), Expect = 4e-40
Identities = 108/282 (38%), Positives = 154/282 (54%), Gaps = 23/282 (8%)
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRA 60
M +LPE CIA IL+ T+P DA S VS +FR A DSD VW++FLPSD S++SQS
Sbjct: 1 MMMLPEACIANILAFTSPADAFSSSEVSSVFRLAGDSDFVWEKFLPSDYKSLISQSTDHH 60
Query: 61 -NASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTT 119
N SSKK Y L D ++IDN +K F+++K SGK ++L AR + I + Y W+
Sbjct: 61 WNISSKKEIYRCLCD-SLLIDNARKLFKINKFSGKISYVLSARDISITHSDHASYWSWSN 119
Query: 120 LPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKII-DAPNFHEVLVVLSVN 178
+ +SRF + L LEI G I T LS NT+YGA+L+ K+ A V S+
Sbjct: 120 VSDSRFSESAELIITDRLEIEGKIQTRVLSANTRYGAYLIVKVTKGAYGLDLVPAETSIK 179
Query: 179 NIGG----------------HNITKIAYLNPD---SLEGEVHDRWDRVQGPSVRSDGWLE 219
+ G + ++ Y N + ++ E + + P R DGW+E
Sbjct: 180 SKNGQISKSATYLCCLDEKKQQMKRLFYGNREERMAMTVEAVGGDGKRREPKCRDDGWME 239
Query: 220 IEMGEFFISGLEDEEVEMSVLQIDGC-FNVGLIIEGIEVRPK 260
IE+GEF ED+EV M++ ++ G G++I+GIEVRPK
Sbjct: 240 IELGEFETREGEDDEVNMTLTEVKGYQLKGGILIDGIEVRPK 281
>At2g02350 SKP1 interacting partner 3 (SKIP3)
Length = 294
Score = 155 bits (393), Expect = 2e-38
Identities = 102/267 (38%), Positives = 148/267 (55%), Gaps = 16/267 (5%)
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRA 60
++ LPEGCI+ I+S T+P DA + VSKIF SA SD VW++FLP+D S+++ S
Sbjct: 24 LDSLPEGCISNIISFTSPEDACVAAAVSKIFESAVKSDIVWEKFLPTDYESLITPS---R 80
Query: 61 NASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIA--WDNGDHYRCWT 118
SSKK Y L + P++I++GK S L+K SGK+C ML A A+ ++ D + W
Sbjct: 81 VFSSKKELYFSLCNDPLLIEDGKMSLWLEKASGKRCIMLSATAMNLSSMADMSQRF-LWI 139
Query: 119 TLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDA-PNFHEVLVVLSV 177
PESRF V LR E + +NT LS T+Y ++VFK D F V + V
Sbjct: 140 PCPESRFETVAALREAYRFEFNCRMNTRVLSLRTRYSVYIVFKKADNWCGFKGVSIEAVV 199
Query: 178 NNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISG--LEDEEV 235
+G + + +G+ R V P +R DGW+E E+GEF+ G + +EV
Sbjct: 200 GIVGEESFRSFICFDTHG-KGQARKR-KVVAKPELREDGWMETEIGEFYNEGGLMSSDEV 257
Query: 236 EMSVLQIDGCF---NVGLIIEGIEVRP 259
E+S ++G + GL+I GIE+RP
Sbjct: 258 EIST--VEGKYAQQKRGLVILGIEIRP 282
>At2g02320 unknown protein
Length = 307
Score = 146 bits (368), Expect = 1e-35
Identities = 99/263 (37%), Positives = 145/263 (54%), Gaps = 14/263 (5%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CI+ I+S T+P DA +LVSK F SA SD VW++F+P + S++S+S + S
Sbjct: 43 LPEECISLIISFTSPRDACVFALVSKTFESAVQSDIVWEKFIPPEYESLLSRS---QHFS 99
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK + L D V+I+ KK ++K +GK+C ML A AL + + H W T P S
Sbjct: 100 SKKELFFALCDESVLINVSKKDLWIEKATGKRCMMLSASALNL---STHHTWKWITNPVS 156
Query: 124 RFPDVV-VLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGG 182
+ + V L W EI NT LSP T+Y ++VF D + + +V + G
Sbjct: 157 AWLETVPELLTTRWFEIRCRTNTRFLSPRTRYSVYIVFLKADICYGFAYVAMEAVVRMVG 216
Query: 183 HNITKIA---YLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGL--EDEEVEM 237
H +++ +++E + R + V P R DGW+EIE+GEFF G ++E+EM
Sbjct: 217 HELSESCRRYVCFHEAMEWQFLTRKNLV-NPERREDGWMEIEIGEFFNEGAFRNNDEIEM 275
Query: 238 SVLQ-IDGCFNVGLIIEGIEVRP 259
SV + GLII+GIE+RP
Sbjct: 276 SVSETTQRNTKRGLIIQGIEIRP 298
>At2g02270 putative phloem-specific lectin
Length = 265
Score = 144 bits (364), Expect = 4e-35
Identities = 96/265 (36%), Positives = 135/265 (50%), Gaps = 47/265 (17%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CI+ I+S TTP DA + VSK F SA SD+VW++FLP D S+V +S
Sbjct: 16 LPENCISNIISFTTPRDACFAASVSKAFESAVQSDSVWEKFLPLDYSSLVPES---RVFL 72
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK L P++I+ GKKSF LDK SG+KC ML + + I+W N
Sbjct: 73 SKKELCFSLCRVPLLIEGGKKSFWLDKTSGEKCIMLSPKGMVISWVN-----------SP 121
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDA---------PNFHEVLVV 174
+F +V L + W E+ G ++T LSP T+Y ++VFK D P F
Sbjct: 122 QFEEVPELLYDSWFEVCGRLSTKYLSPRTRYSVYIVFKTNDLYPGVTLEPFPRF------ 175
Query: 175 LSVNNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEE 234
+ ++ P L+ E + V P R D W+E E+GEFF + +
Sbjct: 176 -------------VRFVGPTDLKYE----REYVTRPEKREDKWMEAELGEFF-NETSCGD 217
Query: 235 VEMSVLQIDGCFNVGLIIEGIEVRP 259
VE+SV+ + + GL+I+GIE RP
Sbjct: 218 VEVSVIDENSYWKSGLVIQGIEFRP 242
>At2g02310 putative phloem-specific lectin
Length = 307
Score = 141 bits (355), Expect = 4e-34
Identities = 91/261 (34%), Positives = 137/261 (51%), Gaps = 6/261 (2%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CI+ I+S T+P D + VSK F A D++W++FLPS+ S++ S
Sbjct: 48 LPEDCISNIISFTSPRDVCVSASVSKSFAHAVQCDSIWEKFLPSEYESLIPPWRV---FS 104
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK Y L PV++++GKKSF L+ SGKKC +L A+ L+I N Y W L ES
Sbjct: 105 SKKDLYFTLCYDPVLVEDGKKSFWLETASGKKCVLLAAKELWITGGNNPEYWQWIELCES 164
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGH 183
F V L + ++ G ++T LS T Y ++V+KI D + L + G
Sbjct: 165 SFEKVPELLNNRSFQMGGSMSTQILSLGTHYSVYIVYKIKDERHGLRDLPIQVGVGFKGQ 224
Query: 184 NITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISG--LEDEEVEMSVLQ 241
+ K +S + ++ R DGW+E E+G+FF G + +EVE+S++
Sbjct: 225 EMPKQFICFDESTDKTKEWPKKKLMKSKKRGDGWMEAEIGDFFNDGGLMGFDEVEVSIVD 284
Query: 242 IDG-CFNVGLIIEGIEVRPKN 261
+ G++IEGIE RPK+
Sbjct: 285 VTSPNLKCGVMIEGIEFRPKD 305
>At2g02300 putative phloem-specific lectin
Length = 284
Score = 135 bits (341), Expect = 2e-32
Identities = 97/265 (36%), Positives = 139/265 (51%), Gaps = 26/265 (9%)
Query: 4 LPEGCIAAILSRT-TPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANA 62
LP+ C+A I S T TP DA +LVSK F +SD+VW++FLP + VS
Sbjct: 37 LPDDCLAIISSFTSTPRDAFLAALVSKSFGLQFNSDSVWEKFLPPPDY--VSLLPKSRVF 94
Query: 63 SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPE 122
SSKK Y L D P NGK SF+LDK SGKKC ML A+ L I+ Y W ++PE
Sbjct: 95 SSKKELYFALCD-PFPNHNGKMSFRLDKASGKKCVMLSAKKLLISRVVNPKYWKWISIPE 153
Query: 123 SRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGG 182
SRF +V L ++ +I G++NT +SP T Y A++V+ N + + +
Sbjct: 154 SRFDEVPELLNIDSFDIRGVLNTRIISPGTHYSAYIVYTKTSHFNGFQTSPIQAGVGFQR 213
Query: 183 HNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISG--LEDEEVEMSVL 240
H ++K ++ DS + R DGW+E ++G+F+ G + +E+SV+
Sbjct: 214 HGMSK-TFIRFDSKK---------------RQDGWMEAKIGDFYNEGGLIGFNLIEVSVV 257
Query: 241 QIDGC----FNVGLIIEGIEVRPKN 261
+ GLIIEGIE RPK+
Sbjct: 258 DVARYPHMNMKSGLIIEGIEFRPKD 282
>At3g53000 unknown protein
Length = 300
Score = 92.8 bits (229), Expect = 2e-19
Identities = 74/277 (26%), Positives = 124/277 (44%), Gaps = 31/277 (11%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVS-QSLSRANA 62
+PE C+A + TP + L+ +++ FR AA SD+VW++ LP + ++ R ++
Sbjct: 24 IPESCVACVFMYLTPPEICNLAGLNRSFRGAASSDSVWEKKLPENYQDLLDLLPPERYHS 83
Query: 63 SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPE 122
SKK + LS P+ D+ K +D+ +G+ C + AR + I Y W E
Sbjct: 84 LSKKDIFAVLS-RPIPFDDDNKEVWIDRVTGRVCMAISARGMSITGIEDRRYWNWIPTEE 142
Query: 123 SRFPDVVVLRHLLWLEIHGMI------NTFSLSPNTQYGAFLV--------FKIIDAPNF 168
SRF V L+ + W E+ G + +SLS G F F++ +
Sbjct: 143 SRFHVVAYLQQIWWFEVDGTVRFHLPPGVYSLSFRIHLGRFTKRLGRRVCHFELTHGWDL 202
Query: 169 HEVLVVLSVNNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDG-WLEIEMGEFFI 227
V LS ++ G + YL D +R + G W+E +GEF +
Sbjct: 203 KPVRFSLSTSD--GQEASCEYYL----------DDVERNEALGKHKRGYWIEYRVGEFIV 250
Query: 228 SGLE-DEEVEMSVLQIDGCFNV-GLIIEGIEVRPKND 262
+G E E++ S+ QID + GL ++ + + P D
Sbjct: 251 NGSEPSTEIQWSMKQIDCTHSKGGLCVDSVFINPIGD 287
>At1g80110 unknown protein
Length = 163
Score = 92.0 bits (227), Expect = 3e-19
Identities = 58/163 (35%), Positives = 84/163 (50%), Gaps = 3/163 (1%)
Query: 98 MLGARALFIAWDNGDHYRCWTTLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAF 157
M+ ARAL I W + Y W +LP +RF +V L + WLEI G IN LS +T Y A+
Sbjct: 1 MMAARALNIVWGHEQRYWHWISLPNTRFGEVAELIMVWWLEITGKINITLLSDDTLYAAY 60
Query: 158 LVFKIIDAPNFHEVLVVLSVNNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGW 217
VFK +P V S+ + + + P + + Q P +R DGW
Sbjct: 61 FVFKWNHSPYGFRQPVETSLVLADTESTDNV--VQPSMISLMQDSGGEEGQSPVLRRDGW 118
Query: 218 LEIEMGEFFISGLEDEEVEMSVLQIDGCF-NVGLIIEGIEVRP 259
E+E+G+FF + E+EMS+ + G + GLI+ GIE+RP
Sbjct: 119 YEVELGQFFKRRGDLGEIEMSLKETKGPYEKKGLIVYGIEIRP 161
>At3g61060 unknown protein
Length = 290
Score = 80.9 bits (198), Expect = 7e-16
Identities = 45/142 (31%), Positives = 76/142 (52%), Gaps = 7/142 (4%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE C+A I++R P + RL+ ++++FR A+ +D +W+ LP++ I + +
Sbjct: 28 LPENCVALIMTRLDPPEICRLARLNRMFRRASSADFIWESKLPANYRVIAHKVFDEITLT 87
Query: 64 S--KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
KK Y LS P + D+G K +DK +G+ C + ++AL I D R W+ +P
Sbjct: 88 KLIKKDLYAKLS-QPNLFDDGTKELWIDKNTGRLCLSISSKALRIT--GIDDRRYWSHIP 144
Query: 122 --ESRFPDVVVLRHLLWLEIHG 141
ESRF ++ + W E+ G
Sbjct: 145 TDESRFQSAAYVQQIWWFEVGG 166
>At1g63090 unknown protein
Length = 289
Score = 78.2 bits (191), Expect = 4e-15
Identities = 69/264 (26%), Positives = 112/264 (42%), Gaps = 17/264 (6%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE C+A IL PV+ R S ++ F A+ +D VW+ LP D I+ + L +
Sbjct: 29 LPESCVALILQNLDPVEICRFSKLNTAFHGASWADFVWESKLPPDYKLILEKILGSFPDN 88
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP-- 121
+KR D G K +DK++G C A+ L I D R W+ +P
Sbjct: 89 LRKRDIFTFLSRVNSFDEGNKKAWVDKRTGGLCLCTSAKGLSIT--GIDDRRYWSHIPSD 146
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIG 181
+SRF V ++ + W ++ G I+ F T Y + + + P V+ +
Sbjct: 147 DSRFASVAYVQQIWWFQVDGEID-FPFPAGT-YSVYFRLQ-LGKPGKRFGWKVVDTEQVH 203
Query: 182 GHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFI--SGLEDEEVEMSV 239
G NI + + S E H Q + W G+F + S +++ S+
Sbjct: 204 GWNIKPVRF--QLSTEDGQH---SSSQCMLTEAGNWSHYHAGDFVVGKSKSSSTKIKFSM 258
Query: 240 LQIDGCFNV--GLIIEGIEVRPKN 261
QID C + GL ++ + V P +
Sbjct: 259 TQID-CTHTKGGLCVDSVVVYPSS 281
>At1g12710 unknown protein
Length = 291
Score = 75.5 bits (184), Expect = 3e-14
Identities = 43/143 (30%), Positives = 70/143 (48%), Gaps = 4/143 (2%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE C+A I+ PV+ R S +++ FR A+ +D VW+ LP + ++ + L +
Sbjct: 31 LPEACVAIIVENLDPVEICRFSKLNRAFRGASWADCVWESKLPQNYRDVLEKILGGFPEN 90
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP-- 121
+KR D+ K +DK++ C + A+ L I D R W+ +P
Sbjct: 91 LQKRHLYAFLSRINSFDDATKKVWIDKRTSGVCLSISAKGLSIT--GIDDRRYWSHIPTD 148
Query: 122 ESRFPDVVVLRHLLWLEIHGMIN 144
ESRF V L+ + W E+ G I+
Sbjct: 149 ESRFSSVAYLQQIWWFEVDGEID 171
>At5g52120 unknown protein
Length = 291
Score = 73.2 bits (178), Expect = 1e-13
Identities = 57/170 (33%), Positives = 80/170 (46%), Gaps = 13/170 (7%)
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRA 60
+E +PE CI A+ P + L+ V+K F A+ SDAVW+ LPS+ +V + L
Sbjct: 21 LEDVPENCITAMFMYMEPPEICLLARVNKSFHRASRSDAVWEDKLPSNYKFLVRRILEDQ 80
Query: 61 NASS--------KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGD 112
KK Y L P + D G K LDK+SGK + +A+ I +
Sbjct: 81 QQVGVKDKLIYRKKEIYARLC-RPNLFDTGTKEAWLDKRSGKVFLAISPKAMKITGIDDR 139
Query: 113 HYRCWTTLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKI 162
Y + ESRF + LR + WLE G I F +P +Y L+FKI
Sbjct: 140 RYWEHISSDESRFGSITYLRQIWWLEAVGKIR-FEFAPG-KYS--LLFKI 185
>At1g65400 hypothetical protein
Length = 172
Score = 58.5 bits (140), Expect = 3e-09
Identities = 46/179 (25%), Positives = 77/179 (42%), Gaps = 37/179 (20%)
Query: 97 FMLGARALFIAWDNGDHYRCWTTLPESR---------FPDVVVLRHLLWLEIHGMINTFS 147
FM+ AR L IAW ++ W LP ++ L+ WL++ G +T
Sbjct: 15 FMIDARDLSIAWSEDSNHWTWLPLPNQNSNTILSSESVMEIAFLKSASWLDVAGKFDTRY 74
Query: 148 LSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGHNITKIAYLNPDSLEGEVHDRWDRV 207
L+P T+Y V K ++ E LV L + ++ + W++
Sbjct: 75 LTPRTRYEVVFVVK-LEYTFEWETLVKLKL---------------------DLPNTWEKP 112
Query: 208 QGPSVR-----SDGWLEIEMGEFFISGLEDEEVEMSVLQID-GCFNVGLIIEGIEVRPK 260
Q SV SD WL+I +GEF S E+ ++ + + + GL ++G+ +RPK
Sbjct: 113 QEQSVDMFDYISDQWLDIPVGEFTTSKKNVGEISFAMYEHECQLWKSGLFVKGVTIRPK 171
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,126,755
Number of Sequences: 26719
Number of extensions: 260105
Number of successful extensions: 751
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 656
Number of HSP's gapped (non-prelim): 51
length of query: 263
length of database: 11,318,596
effective HSP length: 97
effective length of query: 166
effective length of database: 8,726,853
effective search space: 1448657598
effective search space used: 1448657598
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0226.6


