

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.2
(583 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
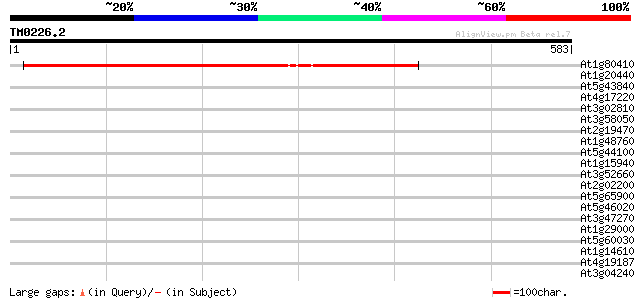
Score E
Sequences producing significant alignments: (bits) Value
At1g80410 putative N-terminal acetyltransferase (At1g80410) 653 0.0
At1g20440 unknown protein (At1g20440) 38 0.018
At5g43840 heat shock transcription factor-like protein 36 0.068
At4g17220 unknown protein 35 0.088
At3g02810 putative protein kinase 35 0.088
At3g58050 putative protein 35 0.15
At2g19470 putative casein kinase I 34 0.20
At1g48760 putative protein 34 0.20
At5g44100 casein kinase I 33 0.34
At1g15940 T24D18.4 33 0.34
At3g52660 putative RNA binding protein 33 0.44
At2g02200 putative protein on transposon FARE2.10 (cds2) 33 0.57
At5g65900 ATP-dependent RNA helicase-like 32 0.75
At5g46020 unknown protein 32 0.75
At3g47270 putative protein 32 0.75
At1g29000 hypothetical protein 32 0.75
At5g60030 KED - like protein 32 0.98
At1g14610 valyl tRNA synthetase (valRS) 32 0.98
At4g19187 unknown protein 32 1.3
At3g04240 O-linked GlcNAc transferase like protein 32 1.3
>At1g80410 putative N-terminal acetyltransferase (At1g80410)
Length = 714
Score = 653 bits (1685), Expect = 0.0
Identities = 325/411 (79%), Positives = 371/411 (90%), Gaps = 3/411 (0%)
Query: 15 QRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLEHSI 74
+RIPLDFLQ + F+EA YI+PLLTKGVPSLFSDLSSLY+HP K DILEQL+++++HSI
Sbjct: 302 KRIPLDFLQDENFKEAVAKYIKPLLTKGVPSLFSDLSSLYDHPRKPDILEQLVVEMKHSI 361
Query: 75 RTSGQYPGSMEKEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSVKS 134
T+G +PGS KEPPSTL+WTLF LAQHYDRRGQY++A+ KIDEAI HTPTVIDLYSVKS
Sbjct: 362 GTTGSFPGSDVKEPPSTLLWTLFFLAQHYDRRGQYDVALCKIDEAIAHTPTVIDLYSVKS 421
Query: 135 RILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKEGDQH 194
RI+KHAGD +AAAA ADEAR MDLADRY+NSECVKRMLQADQV LAEKTAVLFTKEGDQ
Sbjct: 422 RIMKHAGDLTAAAALADEARGMDLADRYINSECVKRMLQADQVPLAEKTAVLFTKEGDQL 481
Query: 195 NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCLRKMTLR 254
NNLHDMQCMWY+LASG+S+FRQGDLGRALKKFL VEKHYADI+EDQFDFHSYCLRKMTLR
Sbjct: 482 NNLHDMQCMWYDLASGDSYFRQGDLGRALKKFLAVEKHYADISEDQFDFHSYCLRKMTLR 541
Query: 255 TYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQ 314
+Y++MLKFQD+LHS YFHKAA AIRCY+KLHDS PKSTA ED EMSKL PAQKKK++
Sbjct: 542 SYVDMLKFQDRLHSFPYFHKAAIRAIRCYLKLHDS-PKSTAGED-EMSKLAPAQKKKIK- 598
Query: 315 KQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLK 374
KQ+KAEARAKK AE K+EE +ASG SKSGKR+VKPVDPDPHG+KL+QV++P++EA KYL+
Sbjct: 599 KQKKAEARAKKEAESKSEESTASGASKSGKRNVKPVDPDPHGQKLIQVEEPMAEASKYLR 658
Query: 375 LLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDSHRCLV 425
LLQK+SP+SLETHLLSFE+ RKQK LL FQAVKQLL+L AE+PDSHR LV
Sbjct: 659 LLQKHSPNSLETHLLSFEVNMRKQKFLLAFQAVKQLLKLGAENPDSHRSLV 709
>At1g20440 unknown protein (At1g20440)
Length = 265
Score = 37.7 bits (86), Expect = 0.018
Identities = 37/150 (24%), Positives = 66/150 (43%), Gaps = 11/150 (7%)
Query: 285 KLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGK 344
KLH S S++ DEE ++KK ++K+ KKG EK +E K+ +
Sbjct: 103 KLHRSNSSSSSSSDEE------GEEKKEKKKKIVEGEEDKKGLVEKIKEKLPGHHDKTAE 156
Query: 345 RHVKPVD---PDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVL 401
V PV P P E +++ D P E ++ +++ P + T ++
Sbjct: 157 DDV-PVSTTIPVPVSESVVEHDHPEEEKKGLVEKIKEKLPGHHDEKAEDSPAVT-STPLV 214
Query: 402 LTFQAVKQLLRLDAEHPDSHRCLVCQLHEK 431
+T V+ L EHP+ + ++ ++ EK
Sbjct: 215 VTEHPVEPTTELPVEHPEEKKGILEKIKEK 244
>At5g43840 heat shock transcription factor-like protein
Length = 251
Score = 35.8 bits (81), Expect = 0.068
Identities = 24/99 (24%), Positives = 44/99 (44%), Gaps = 2/99 (2%)
Query: 279 AIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKK--MRQKQRKAEARAKKGAEEKNEELSA 336
+++ Y+ L + K T + E M L + KK Q RK + K E+K E +S+
Sbjct: 147 SVKTYLHLMEEKLKVTEVKQEMMMNFLLKKIKKPSFLQSLRKRNLQGIKNREQKQEVISS 206
Query: 337 SGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLKL 375
G + H+ ++ GE +++D + I +L
Sbjct: 207 HGFDYGDELHIASMEHQGQGEDEIEMDSESTPLIVQRRL 245
>At4g17220 unknown protein
Length = 513
Score = 35.4 bits (80), Expect = 0.088
Identities = 44/174 (25%), Positives = 72/174 (41%), Gaps = 14/174 (8%)
Query: 274 KAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEE 333
KA A+R +L D K E ++ + L A + ++ QK E KK EEK +
Sbjct: 30 KAEVEALRTNEELKDRVFKELRENVRKLEEKLGATENQVDQK----ELERKKLEEEKEDA 85
Query: 334 LSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFEL 393
L+A ++ R V D L + PL IK + K+ +L+ + E
Sbjct: 86 LAAQDAAEEALRRVYTHQQDDDSLPLESIIAPLESQIK----IHKHEISALQEDKKALER 141
Query: 394 YTR-KQKVLLTFQAVKQLL---RLDAEHPDSHRCLVCQLHEKTLFEANNSFLEK 443
T+ K+ LL + + + L E +H + + E + + N FLEK
Sbjct: 142 LTKSKESALLEAERILRSALERALIVEEVQNHNFELRRQIE--ICQDENKFLEK 193
>At3g02810 putative protein kinase
Length = 558
Score = 35.4 bits (80), Expect = 0.088
Identities = 20/70 (28%), Positives = 31/70 (43%)
Query: 289 SPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVK 348
SPP A ED++ S + +K+ ++ KK EE++ + + S S H K
Sbjct: 378 SPPPELATEDDKSSTSSGEESSLESEKESVSKNEYKKKHEEEDSSMESDDESDSNSEHEK 437
Query: 349 PVDPDPHGEK 358
P P EK
Sbjct: 438 DQPPKPIDEK 447
>At3g58050 putative protein
Length = 1209
Score = 34.7 bits (78), Expect = 0.15
Identities = 32/108 (29%), Positives = 49/108 (44%), Gaps = 5/108 (4%)
Query: 284 IKLHDSPPKSTAEEDEEMSKLLPAQK-KKMRQKQRKAEARAKKGAEEKNEELSASGVSKS 342
+KL + K EE+E K ++ KK+R+K+R E KG E+KN E S + +
Sbjct: 529 VKLLEEEEKEKREEEERKEKKRSKEREKKLRKKERLKE--KDKGKEKKNPECSDKDMLLN 586
Query: 343 GKRHVKPVDPDPHGEKLLQVDDPLSE-AIKYLKLLQKNSPDSLETHLL 389
R + + P+ + E ++ SE Y L SPD E L
Sbjct: 587 SSREEEDL-PNLYDETNNTINSEESEIETGYADLSPPGSPDVQERQCL 633
>At2g19470 putative casein kinase I
Length = 433
Score = 34.3 bits (77), Expect = 0.20
Identities = 23/68 (33%), Positives = 33/68 (47%), Gaps = 9/68 (13%)
Query: 216 QGDLGRALKKFLGVEKHYADINEDQF-----DFHSYCLRKMTLRTYL----EMLKFQDQL 266
QG G K+ GVE Y + D D SYC R+ +L+T L +M+ + +
Sbjct: 60 QGGTGIPNMKWYGVEGDYNVLVMDLLGPSLEDLFSYCKRQFSLKTVLMLADQMINRLEFI 119
Query: 267 HSHSYFHK 274
HS SY H+
Sbjct: 120 HSKSYLHR 127
>At1g48760 putative protein
Length = 869
Score = 34.3 bits (77), Expect = 0.20
Identities = 18/54 (33%), Positives = 32/54 (58%), Gaps = 3/54 (5%)
Query: 287 HDSPPKSTAEEDEEMSKL---LPAQKKKMRQKQRKAEARAKKGAEEKNEELSAS 337
++ P S +E EE S++ ++KKK ++K++K E +K + +NE SAS
Sbjct: 806 YEGNPNSGQQEKEESSRIENHQNSEKKKKKKKKKKGEGSSKHKSRRQNEVASAS 859
>At5g44100 casein kinase I
Length = 476
Score = 33.5 bits (75), Expect = 0.34
Identities = 29/107 (27%), Positives = 46/107 (42%), Gaps = 17/107 (15%)
Query: 177 VALAEKTAVLFTKEGDQHNNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADI 236
V E+ AV +H LH ++ L QG G K+ GVE Y+ +
Sbjct: 29 VQTGEEVAVKLENVKTKHPQLHYESKLYMLL--------QGGSGIPNIKWFGVEGDYSVM 80
Query: 237 NEDQF-----DFHSYCLRKMTLRTYL----EMLKFQDQLHSHSYFHK 274
D D +YC RK+TL+T L ++L + +H+ + H+
Sbjct: 81 VIDLLGPSLEDLFNYCNRKLTLKTVLMLADQLLNRVEFMHTRGFLHR 127
>At1g15940 T24D18.4
Length = 1012
Score = 33.5 bits (75), Expect = 0.34
Identities = 28/152 (18%), Positives = 70/152 (45%), Gaps = 5/152 (3%)
Query: 284 IKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSG 343
+K ++ PKS EE E + K ++ E A++ +E+ +++ KS
Sbjct: 849 LKEPNAEPKSDGEEQEAAKEPNAELKTDGENQEAAKELTAERKTDEEEHKVADEVEQKSQ 908
Query: 344 KRHVKPVDPDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLL- 402
K V+P+ GE+ V++P +E ++ + + +T L+ E ++ + +
Sbjct: 909 KE--TNVEPEAEGEEQKSVEEPNAEPKTKVEEKESAKEQTADTKLIEKEDMSKTKGEEID 966
Query: 403 --TFQAVKQLLRLDAEHPDSHRCLVCQLHEKT 432
T+ ++ + ++ E + + ++ +L E++
Sbjct: 967 KETYSSIPETGKVGNEAEEDDQRVIKELEEES 998
>At3g52660 putative RNA binding protein
Length = 471
Score = 33.1 bits (74), Expect = 0.44
Identities = 17/72 (23%), Positives = 39/72 (53%), Gaps = 3/72 (4%)
Query: 288 DSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHV 347
D+ P+ EE+ E ++ +++++ + + + E + EE+ E+ A+ + KRHV
Sbjct: 26 DNDPEEILEEEVEYEEV---EEEEIEEIEEEIEEEVEVEEEEEEEDAVATEEEEEKKRHV 82
Query: 348 KPVDPDPHGEKL 359
+ + PHG ++
Sbjct: 83 ELLALPPHGSEV 94
>At2g02200 putative protein on transposon FARE2.10 (cds2)
Length = 586
Score = 32.7 bits (73), Expect = 0.57
Identities = 24/96 (25%), Positives = 45/96 (46%), Gaps = 11/96 (11%)
Query: 274 KAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEE 333
K G + Y+ + P T EE QK++ +K+++ E + ++G EEK E+
Sbjct: 182 KKKVGRLEGYLGIERVPASYTREE----------QKEEDEKKEQEEEKQEEEGKEEKLEK 231
Query: 334 LSASGVSKSGKRHV-KPVDPDPHGEKLLQVDDPLSE 368
+ G + K+ + K D + GE+ Q ++ E
Sbjct: 232 IEYRGDEGTEKQEIPKQGDEEMEGEEEKQEEEGKEE 267
>At5g65900 ATP-dependent RNA helicase-like
Length = 633
Score = 32.3 bits (72), Expect = 0.75
Identities = 26/111 (23%), Positives = 52/111 (46%), Gaps = 6/111 (5%)
Query: 295 AEEDEEMSKLLPAQK-----KKMRQKQRKAEARAKKG-AEEKNEELSASGVSKSGKRHVK 348
+ E+EE+ K ++ KK++Q + E + G A+E N + K K++ K
Sbjct: 10 SSENEEIKKKKHKKRARDEAKKLKQPAMEEEPDHEDGDAKENNALIDEEPKKKKKKKNKK 69
Query: 349 PVDPDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQK 399
D D ++ + ++P + K KL Q+ + E +++ E +K+K
Sbjct: 70 RGDTDDGEDEAVAEEEPKKKKKKNKKLQQRGDTNDEEDEVIAEEEEPKKKK 120
>At5g46020 unknown protein
Length = 164
Score = 32.3 bits (72), Expect = 0.75
Identities = 32/110 (29%), Positives = 51/110 (46%), Gaps = 7/110 (6%)
Query: 307 AQKKKMRQKQRKAEARAKKGAEEKNEELS--ASGVSKSGKRHVKPVD-PDPHGEKLLQVD 363
A+ + +QK+ + E ++ +EE++EE S + V K G V VD P+ +K L+
Sbjct: 29 ARPRSFKQKEAEYEEDVEEESEEESEEESEDEADVKKKGAEAVIEVDNPNRVRQKTLKAK 88
Query: 364 DPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRL 413
D + L ++ + H E Y R Q+ T QA K L RL
Sbjct: 89 DLDASKTTELSRREREELEKQRAH----ERYMRLQEQGKTEQARKDLDRL 134
>At3g47270 putative protein
Length = 671
Score = 32.3 bits (72), Expect = 0.75
Identities = 23/96 (23%), Positives = 46/96 (46%), Gaps = 11/96 (11%)
Query: 274 KAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEE 333
K G + Y+ + P T EE QK++ +K+++ E + ++G EE+ E+
Sbjct: 271 KKKVGRLEGYLGIQRVPASYTREE----------QKEEDEKKEQEEEKQEEEGKEEELEK 320
Query: 334 LSASGVSKSGKRHV-KPVDPDPHGEKLLQVDDPLSE 368
+ G ++ K+ + K D + GE+ Q ++ E
Sbjct: 321 VEYRGDERTEKQEIPKQGDEEMEGEEEKQKEEGKEE 356
>At1g29000 hypothetical protein
Length = 287
Score = 32.3 bits (72), Expect = 0.75
Identities = 19/64 (29%), Positives = 31/64 (47%)
Query: 285 KLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGK 344
K+H +++ +EE K +KKK + ++K E KK E+K EE + + K
Sbjct: 170 KVHKHAEIISSKTEEEKKKEEEDKKKKEEEDKKKKEDEKKKEEEKKKEEENKKKEGEKKK 229
Query: 345 RHVK 348
VK
Sbjct: 230 EEVK 233
Score = 30.8 bits (68), Expect = 2.2
Identities = 18/55 (32%), Positives = 27/55 (48%), Gaps = 4/55 (7%)
Query: 292 KSTAEED----EEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKS 342
K EED EE K +KK ++++K E KK E+K EE+ +K+
Sbjct: 186 KKKEEEDKKKKEEEDKKKKEDEKKKEEEKKKEEENKKKEGEKKKEEVKVEVTTKT 240
>At5g60030 KED - like protein
Length = 292
Score = 32.0 bits (71), Expect = 0.98
Identities = 29/120 (24%), Positives = 53/120 (44%), Gaps = 18/120 (15%)
Query: 298 DEEMSKLLPAQKKKMRQKQRKAEARAKKGAEE------------KNEELSASGVSKSGKR 345
DE++++ L A+++ +++RK E + KK ++ ++E+ SA + K+
Sbjct: 130 DEKVNEKLEAEQRSEERRERKKEKKKKKNNKDEDVVDEKVKEKLEDEQKSADRKERKKKK 189
Query: 346 HVKPVDPDPHGEKLLQVDDPLSEAIKYLK------LLQKNSPDSLETHLLSFELYTRKQK 399
K D D EK D+ S IK K ++ + + LE S E K+K
Sbjct: 190 SKKNNDEDVVDEKEKLEDEQKSAEIKEKKKNKDEDVVDEKEKEKLEDEQRSGERKKEKKK 249
Score = 29.3 bits (64), Expect = 6.3
Identities = 17/57 (29%), Positives = 28/57 (48%), Gaps = 2/57 (3%)
Query: 297 EDEEMSKLLPAQKKKMRQKQRK--AEARAKKGAEEKNEELSASGVSKSGKRHVKPVD 351
EDE+ S +KKK R+ + +E R K + +EE+ + KR +K +D
Sbjct: 235 EDEQRSGERKKEKKKKRKSDEEIVSEERKSKKKRKSDEEMGSEERKSKKKRKLKEID 291
>At1g14610 valyl tRNA synthetase (valRS)
Length = 1108
Score = 32.0 bits (71), Expect = 0.98
Identities = 25/91 (27%), Positives = 43/91 (46%), Gaps = 7/91 (7%)
Query: 296 EEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH 355
++ EE +K +K+K +K+R AE +AK+ + N + KS KR +P
Sbjct: 62 KKKEEKAKEKELKKQKALEKERLAELKAKQAKDGTN--VPKKSAKKSSKRDASEENP--- 116
Query: 356 GEKLLQVDDPLSEAIKYLKLLQKN-SPDSLE 385
E + + PL E + + K SP ++E
Sbjct: 117 -EDFVDPETPLGERKRLSSQMAKQYSPATVE 146
>At4g19187 unknown protein
Length = 595
Score = 31.6 bits (70), Expect = 1.3
Identities = 21/72 (29%), Positives = 35/72 (48%), Gaps = 4/72 (5%)
Query: 285 KLHDSPPKSTAEE-DEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSK-- 341
K H T EE DE + +++K+ +K++K + ++KK + + SG
Sbjct: 324 KRHKKHSSRTVEETDESSTGSEDSREKRGSKKRKKLKKKSKKQYDSDSLSFEGSGSDSYR 383
Query: 342 -SGKRHVKPVDP 352
S +RH K VDP
Sbjct: 384 LSRRRHTKHVDP 395
>At3g04240 O-linked GlcNAc transferase like protein
Length = 977
Score = 31.6 bits (70), Expect = 1.3
Identities = 48/220 (21%), Positives = 80/220 (35%), Gaps = 53/220 (24%)
Query: 48 SDLSSLYNHPGKADILEQLILDLEHSIRTSGQYPGSMEKEPPSTLMW--------TLFLL 99
S L +N + D + L L H + G + ++E S +++ L L+
Sbjct: 39 SSLLQQFNKTHEGD--DDARLALAHQLYKGGDFKQALEH---SNMVYQRNPLRTDNLLLI 93
Query: 100 AQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSVKSRILKHAGDFSAAAAF---ADEAR-- 154
Y + +Y++ IA+ +EA+ P + Y + K GD A + A E R
Sbjct: 94 GAIYYQLQEYDMCIARNEEALRIQPQFAECYGNMANAWKEKGDTDRAIRYYLIAIELRPN 153
Query: 155 ----CMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKEGDQHNNLHDMQ--------- 201
+LA Y+ + Q Q AL+ ++ D H+NL ++
Sbjct: 154 FADAWSNLASAYMRKGRLSEATQCCQQALSLNPLLV-----DAHSNLGNLMKAQGLIHEA 208
Query: 202 ---------------CMWYELASGESFFRQGDLGRALKKF 226
W LA F GDL RAL+ +
Sbjct: 209 YSCYLEAVRIQPTFAIAWSNLAG--LFMESGDLNRALQYY 246
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,029,981
Number of Sequences: 26719
Number of extensions: 560509
Number of successful extensions: 2703
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 57
Number of HSP's that attempted gapping in prelim test: 2592
Number of HSP's gapped (non-prelim): 134
length of query: 583
length of database: 11,318,596
effective HSP length: 105
effective length of query: 478
effective length of database: 8,513,101
effective search space: 4069262278
effective search space used: 4069262278
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0226.2


