

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.15
(219 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
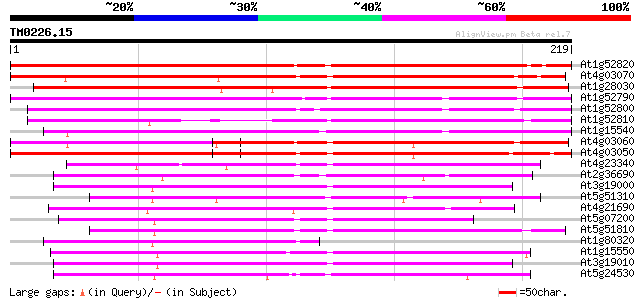
Score E
Sequences producing significant alignments: (bits) Value
At1g52820 oxidoreductase-like protein 249 8e-67
At4g03070 oxidoreductase - like protein (AOP1.1) 214 2e-56
At1g28030 hypothetical protein 187 4e-48
At1g52790 putative oxidoreductase 161 3e-40
At1g52800 putative oxidoreductase 157 3e-39
At1g52810 oxidoreductase like protein 147 3e-36
At1g15540 hypothetical protein 126 1e-29
At4g03060 putative oxidoreductase 124 3e-29
At4g03050 2-oxoglutarate-dependent dioxygenase (AOP3) 122 1e-28
At4g23340 unknown protein 92 2e-19
At2g36690 putative giberellin beta-hydroxylase 86 2e-17
At3g19000 unknown protein 85 3e-17
At5g51310 gibberellin 20-oxidase-like protein 84 6e-17
At4g21690 gibberellin 3 beta-hydroxylase - like protein 82 2e-16
At5g07200 gibberellin 20-oxidase 82 3e-16
At5g51810 gibberellin 20-oxidase (emb|CAA58294.1) 80 6e-16
At1g80320 oxidoreductase like protein 80 8e-16
At1g15550 Gibberillin 3 beta-hydroxylase (AtGA3ox1) 80 8e-16
At3g19010 oxidase like protein 80 1e-15
At5g24530 flavanone 3-hydroxylase-like protein 79 2e-15
>At1g52820 oxidoreductase-like protein
Length = 317
Score = 249 bits (636), Expect = 8e-67
Identities = 123/219 (56%), Positives = 163/219 (74%), Gaps = 5/219 (2%)
Query: 1 VKNTSVYEKVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEH 60
+ ++ + EKV+ T+ LWP GN SF + SFS+ LSELD +R+M++ES G++KY+DEH
Sbjct: 103 IDDSDIAEKVDAFTEKLWPQGNISFSTTIQSFSKKLSELDITIRRMIMESFGLDKYIDEH 162
Query: 61 LNSTNYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKP 120
L+STNY L+ MKYKGP T +TKVGL+ HTD I+TIL+Q+ V+GLEV TKD + WI KP
Sbjct: 163 LHSTNYLLRVMKYKGPDTEETKVGLNAHTDKNIVTILYQNHVEGLEVQTKD-KNWIKVKP 221
Query: 121 SSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELV 180
+ + SF VM+ D +A NGRL+SP+HRVMMTG E RYS LFS+PK G+I+ +P+ELV
Sbjct: 222 TQD--SFTVMIGDSLYALLNGRLHSPYHRVMMTGTETRYSLGLFSIPKAGHIVSSPDELV 279
Query: 181 DEEHPLLYKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
DEEHP L+KPFD EFL+ + E G+R Q AL +YCG
Sbjct: 280 DEEHPRLFKPFDHVEFLQFYYT-EAGQR-SQSALKTYCG 316
>At4g03070 oxidoreductase - like protein (AOP1.1)
Length = 322
Score = 214 bits (546), Expect = 2e-56
Identities = 118/224 (52%), Positives = 150/224 (66%), Gaps = 12/224 (5%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV T+ LWP+ GN S + FSE L ELD +VR+M++ES G+E Y+DE
Sbjct: 101 INDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLVELDLMVRRMIMESFGIENYIDE 160
Query: 60 HLNSTNYFLKTMKYKGPQTSD------TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGR 113
HLNST Y + MKY P D TK+GL +HTD I+TIL Q QVDGLEV TKD
Sbjct: 161 HLNSTYYLTRLMKYTSPPDDDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD- 219
Query: 114 KWISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYII 173
KWI KPS +S +VM+ D A NGRL+SP+HRV+MTG + RYS LFS+PK G II
Sbjct: 220 KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVII 277
Query: 174 KAPEELVDEEHPLLYKPFDLEEFLKHSFNKEKGKRDDQFALHSY 217
+PEELVD+EHP ++KPF+ +FL H F E G R Q ALH++
Sbjct: 278 DSPEELVDKEHPRIFKPFEYTDFL-HFFQTEAG-RIAQSALHAF 319
>At1g28030 hypothetical protein
Length = 322
Score = 187 bits (475), Expect = 4e-48
Identities = 107/212 (50%), Positives = 130/212 (60%), Gaps = 6/212 (2%)
Query: 10 VETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHLNSTNYFLK 69
V LT LWP GN + SF+E L EL+ VR M LES G+EKY++EHLN+ N +
Sbjct: 112 VNDLTHKLWPQGNIFVGKNVQSFAEKLIELNLTVRTMTLESFGLEKYMEEHLNAANKHFQ 171
Query: 70 TMKYKGPQTSDT--KVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQS 126
+KYKG +T K+G H D LTIL Q+ VDGLE+ TKDG +WI KPS S S
Sbjct: 172 LLKYKGISDDNTENKIGFYPHIDRHFLTILCQNDAVDGLEIKTKDGEEWIKVKPSQAS-S 230
Query: 127 FIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPL 186
FIVM H NG + P HRV++TG + RY AALF++PK G II APEE+VD+EHP
Sbjct: 231 FIVMAGASLHVLLNGGVFPPLHRVVITGKKDRYVAALFTIPKEGVIINAPEEMVDDEHPR 290
Query: 187 LYKPFDLEEFLKHSFNKEKGKRDDQFALHSYC 218
LYKPFD FLK F+ R D L YC
Sbjct: 291 LYKPFDFWGFLK--FSNLPNARKDISDLTDYC 320
>At1g52790 putative oxidoreductase
Length = 310
Score = 161 bits (407), Expect = 3e-40
Identities = 85/219 (38%), Positives = 129/219 (58%), Gaps = 6/219 (2%)
Query: 1 VKNTSVYEKVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEH 60
+ N + E + T ++WP GN F L ++E +ELDQ+V +MV +S VEKY D +
Sbjct: 97 IDNATSLEATRSFTGLMWPQGNEHFSECLYKYAEFAAELDQMVTRMVFQSYNVEKYYDPY 156
Query: 61 LNSTNYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKP 120
+ ST Y L+ +K + P + +G THTD + TIL Q QV+GLE+ T++G + I+
Sbjct: 157 IESTTYLLRVLKNRAPNNENPTLGFVTHTDKSFTTILHQDQVNGLEMETREGER-ININL 215
Query: 121 SSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELV 180
SS S F+V+ D AWSN R+ SP H+V+++G RYS +F+ G ++ PEEL+
Sbjct: 216 SSPS-LFMVVAGDALMAWSNDRVWSPRHQVLVSGETDRYSLGMFAFNNG--TLQVPEELI 272
Query: 181 DEEHPLLYKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
D +HPL+YKPFD L+ F + + + +YCG
Sbjct: 273 DHQHPLMYKPFDHLGLLR--FYRTDTGYKSECPIKAYCG 309
>At1g52800 putative oxidoreductase
Length = 314
Score = 157 bits (398), Expect = 3e-39
Identities = 86/212 (40%), Positives = 125/212 (58%), Gaps = 7/212 (3%)
Query: 8 EKVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHLNSTNYF 67
E + T ++WP GN F + +FS ++ELD++V +M+ E+ GVEK+ + H+ S Y
Sbjct: 109 EIAQRFTHLMWPQGNDRFCNTVHTFSNAVAELDRLVVRMIFENYGVEKHYESHVGSKTYL 168
Query: 68 LKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSF 127
LK +KY P S + HTD L+IL Q+ V+GLEV +KDG +WIS + +S+
Sbjct: 169 LKFLKYLAPPESISMPAFPQHTDKTFLSILHQNDVNGLEVKSKDG-EWISLQ--LPPKSY 225
Query: 128 IVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLL 187
+VM D WSN R+ S HRV M G++ RY+ LFS ++ PEELVD++HPL+
Sbjct: 226 VVMAGDISMGWSNDRIRSCEHRVTMEGDKTRYTLGLFSFLTD--LVSIPEELVDDKHPLM 283
Query: 188 YKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
YKPFD + +F K R+ L +YCG
Sbjct: 284 YKPFDNIALI--NFYTTKEGREANSTLKAYCG 313
>At1g52810 oxidoreductase like protein
Length = 289
Score = 147 bits (372), Expect = 3e-36
Identities = 91/213 (42%), Positives = 117/213 (54%), Gaps = 35/213 (16%)
Query: 8 EKVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGV-EKYLDEHLNSTNY 66
+KV T LWP GN +F A+ SF+E +SELD + R+M++ES G+ E Y+ EHLNST
Sbjct: 110 DKVNAFTHKLWPQGNNNFSEAVMSFAEKVSELDFMTRRMIMESFGLDESYIKEHLNSTKC 169
Query: 67 FLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQS 126
+D K DG+EV TKD + WI PS +S S
Sbjct: 170 L-----------NDVK--------------------DGIEVKTKDDKHWIKANPSQDS-S 197
Query: 127 FIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPL 186
FIV+ HA NGR+ + HRVM G R+SA LFSVPK +I APEELVD E+P
Sbjct: 198 FIVLGGAMLHALLNGRVLTGVHRVMRMGANIRFSAGLFSVPKTEDLIYAPEELVDAEYPR 257
Query: 187 LYKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
LYKP D E + +++ E R D AL +YCG
Sbjct: 258 LYKPVDFEAYFRYTI--EGPGRRDLSALRTYCG 288
>At1g15540 hypothetical protein
Length = 320
Score = 126 bits (316), Expect = 1e-29
Identities = 66/209 (31%), Positives = 116/209 (54%), Gaps = 7/209 (3%)
Query: 14 TKILWPNG---NPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHLNSTNYFLKT 70
++++WP+ N F + ++++ +EL+Q+V +M+ ES VEKY ++++ T Y L+
Sbjct: 114 SRLMWPDDHDDNDRFCETVHAYAKMQAELEQLVIRMLFESYNVEKYTEKYIGGTRYLLRL 173
Query: 71 MKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVM 130
+KY+ + +HTD + ++IL Q+ + GL + ++ W + PS F+V+
Sbjct: 174 LKYRRLPNGEPNRKFISHTDKSFISILHQNHITGLMLKSEKEDVWYHFTPS--PTRFVVI 231
Query: 131 MNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKP 190
D AWSN R+ + +H+V M E RYS FS +G +I PEE+VD++HPL Y P
Sbjct: 232 AGDAIMAWSNDRIKACYHKVEMESVEMRYSLGFFSFQEG--MISTPEEMVDKDHPLAYNP 289
Query: 191 FDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
F + L++ E + + +YCG
Sbjct: 290 FHHDGLLEYYETLEVHLKAHRTMTKAYCG 318
>At4g03060 putative oxidoreductase
Length = 399
Score = 124 bits (312), Expect = 3e-29
Identities = 64/140 (45%), Positives = 92/140 (65%), Gaps = 6/140 (4%)
Query: 80 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 139
+ K+GL HTD + T+LFQ +++GLEV TKD KWI KPS + FIV+ D A
Sbjct: 260 EKKLGLPCHTDKNLFTVLFQHEIEGLEVKTKD-EKWIRVKPSPNT--FIVIAGDSLCALM 316
Query: 140 NGRLNSPFHRVMMTGNE-ARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEEFLK 198
NGR+ +P+HRV +T + RY+AA+F+ PK Y+I+AP+ELVDE+HP L++PFD +
Sbjct: 317 NGRIRAPYHRVRVTEKKRTRYTAAIFTCPKPDYVIEAPKELVDEKHPRLFRPFDYRDLF- 375
Query: 199 HSFNKEKGKRDDQFALHSYC 218
+F + R Q+ L +YC
Sbjct: 376 -TFYHSEAGRKIQYTLQAYC 394
Score = 66.2 bits (160), Expect = 1e-11
Identities = 34/93 (36%), Positives = 53/93 (56%), Gaps = 3/93 (3%)
Query: 1 VKNTSVYEKVETLTKILWPNG--NPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLD 58
+ N ++ E + T+ LWP+G N + + F+E L E+D +VR+MV+ES G+EKY+D
Sbjct: 96 ISNANILENINEFTQQLWPHGDGNENISKTIQLFAEKLVEIDVMVRRMVMESFGIEKYID 155
Query: 59 EHLNSTNYFLKTMKYKGPQTS-DTKVGLDTHTD 90
+HL ST Y + G + + KV D D
Sbjct: 156 DHLKSTAYATDELNIVGVEPNVGVKVNADISDD 188
>At4g03050 2-oxoglutarate-dependent dioxygenase (AOP3)
Length = 411
Score = 122 bits (306), Expect = 1e-28
Identities = 67/141 (47%), Positives = 95/141 (66%), Gaps = 6/141 (4%)
Query: 80 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 139
D ++GL +HTD ++ I++Q Q+DGLEV TK+G KWI KP+ + IV+ D A
Sbjct: 271 DAELGLPSHTDKSLTGIIYQHQIDGLEVKTKEG-KWIRVKPAPNT--VIVIAGDALCALM 327
Query: 140 NGRLNSPFHRVMMTGNE-ARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEEFLK 198
NGR+ SP+HRV +T + RY+AALFS PK GYII +P+ELVDE+HP +KPFD + L
Sbjct: 328 NGRIPSPYHRVRVTEKKKTRYAAALFSNPKEGYIIDSPKELVDEKHPRAFKPFDFVD-LF 386
Query: 199 HSFNKEKGKRDDQFALHSYCG 219
+ ++ E G+R L ++CG
Sbjct: 387 NFYHTEAGRRAPS-TLQAFCG 406
Score = 79.0 bits (193), Expect = 2e-15
Identities = 35/90 (38%), Positives = 56/90 (61%)
Query: 1 VKNTSVYEKVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEH 60
+K+ ++ EK T+ LWP GN S + ++E L+ELD +VR+++LES G+E ++DEH
Sbjct: 96 IKDANILEKAHEFTQQLWPEGNKSISKMIQLYAEKLAELDMMVRRLILESYGIEYFIDEH 155
Query: 61 LNSTNYFLKTMKYKGPQTSDTKVGLDTHTD 90
LNST Y ++ MKY +D + + D
Sbjct: 156 LNSTYYRMRLMKYIARPDNDITAAVGANVD 185
>At4g23340 unknown protein
Length = 277
Score = 92.4 bits (228), Expect = 2e-19
Identities = 62/194 (31%), Positives = 97/194 (49%), Gaps = 13/194 (6%)
Query: 23 PSFRYALCSFSEHLSELDQIVRKMVL-----ESLGVEKYLDEHLNSTNYFLKTMKYKGPQ 77
P R + + ++EL + + K++L + G Y + N Y L+ + Y P
Sbjct: 66 PELRETMQEYGAKMAELSKRLIKILLMMTLGDETGKRLYQTDFSNCHGY-LRLVNYTPPH 124
Query: 78 TSDTKV----GLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 133
+ + GL HTD + +TI++Q V GL++ +K+G KWI P ++ +V + D
Sbjct: 125 DVEKQEELVEGLGMHTDMSCITIVYQDSVGGLQMRSKEG-KWIDINPCNDF--LVVNIGD 181
Query: 134 CFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDL 193
AWSNGRL S HRV++ R S A F + +I AP+E+V E YK F
Sbjct: 182 LMQAWSNGRLRSSEHRVVLRKLVNRVSLAFFLCFEDEKVILAPQEIVGEGKQRSYKSFKC 241
Query: 194 EEFLKHSFNKEKGK 207
E+LK + E+GK
Sbjct: 242 SEYLKFRQSNEEGK 255
>At2g36690 putative giberellin beta-hydroxylase
Length = 392
Score = 85.9 bits (211), Expect = 2e-17
Identities = 54/190 (28%), Positives = 100/190 (52%), Gaps = 8/190 (4%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLD--EHLNSTNYFLKTMKYKG 75
WP+ FR + ++++ E+ +++ K +LESL ++ + + L + + Y
Sbjct: 193 WPSSPSDFRSSAATYAKETKEMFEMMVKAILESLEIDGSDEAAKELEEGSQVVVVNCYPP 252
Query: 76 PQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCF 135
+ +G+ H+D LT+L Q +V+GL++L +D +W++ P SF+V + D
Sbjct: 253 CPEPELTLGMPPHSDYGFLTLLLQDEVEGLQILYRD--EWVTVDPI--PGSFVVNVGDHL 308
Query: 136 HAWSNGRLNSPFHRVMMTGNEARYS-AALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLE 194
+SNGR S HRV++ + R S A+L S P ++K +LVD+ +P Y D
Sbjct: 309 EIFSNGRYKSVLHRVLVNSTKPRISVASLHSFPLTS-VVKPSPKLVDKHNPSQYMDTDFT 367
Query: 195 EFLKHSFNKE 204
FL++ ++E
Sbjct: 368 TFLQYITSRE 377
>At3g19000 unknown protein
Length = 352
Score = 85.1 bits (209), Expect = 3e-17
Identities = 53/180 (29%), Positives = 85/180 (46%), Gaps = 3/180 (1%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVE-KYLDEHLNSTNYFLKTMKYKGP 76
WP FR ++ + +L + ++V SLG+ L N FL+ Y
Sbjct: 151 WPQNPSHFREVCQEYAREVEKLAFRLLELVSISLGLPGDRLTGFFNEQTSFLRFNHYPPC 210
Query: 77 QTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFH 136
+ +G+ H D LT+L Q V GL+V + +WI KP S++ I+ M +C
Sbjct: 211 PNPELALGVGRHKDGGALTVLAQDSVGGLQVSRRSDGQWIPVKPISDA--LIINMGNCIQ 268
Query: 137 AWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEEF 196
W+N S HRV++ ++ R+S F P I+ EEL+ EE+P YK ++ +F
Sbjct: 269 VWTNDEYWSAEHRVVVNTSKERFSIPFFFFPSHEANIEPLEELISEENPPCYKKYNWGKF 328
>At5g51310 gibberellin 20-oxidase-like protein
Length = 333
Score = 84.0 bits (206), Expect = 6e-17
Identities = 58/195 (29%), Positives = 96/195 (48%), Gaps = 23/195 (11%)
Query: 32 FSEHLSELDQIVRKMVLESLGVE---KYLDEHLNSTNYFLKTMKYKGPQTS--------- 79
+ E +++L + + K +L S G + KY + + + + + Y P
Sbjct: 125 YGEKMTKLCEKIMKAILSSFGDDLHHKYYESEFGNCHGYFRINNYTIPSDQEDDHHNGDE 184
Query: 80 -DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAW 138
D GL HTD + +TI+ Q + GL+V T+DG + P E + +V + D HAW
Sbjct: 185 QDLIEGLGMHTDMSCITIVDQDDIGGLQVRTRDGIGLMDINPKDE--ALVVNVGDLLHAW 242
Query: 139 SNGRLNSPFHRVMM-----TGNEARYSAALFSVPKGGYIIKAPEELVDE-EHPLLYKPFD 192
+NGRL S HRV++ GN R+S A F G ++ AP+E+V E +++ F
Sbjct: 243 TNGRLRSSQHRVILKRRGFVGN--RFSLAFFWCFDDGKVVFAPDEVVGGCEGMRVFRSFK 300
Query: 193 LEEFLKHSFNKEKGK 207
++L+ + EKGK
Sbjct: 301 CGDYLRFRESNEKGK 315
>At4g21690 gibberellin 3 beta-hydroxylase - like protein
Length = 349
Score = 82.0 bits (201), Expect = 2e-16
Identities = 55/193 (28%), Positives = 92/193 (47%), Gaps = 15/193 (7%)
Query: 16 ILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLG---------VEKYLDEHLNSTNY 66
+LWP+ + F + + + + +L + M++ SLG V +S
Sbjct: 145 LLWPDDHAEFCNVMEEYQKAMDDLSHRLISMLMGSLGLTHEDLGWLVPDKTGSGTDSIQS 204
Query: 67 FLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLT--KDGRKWISYKPSSES 124
FL+ Y +GL HTD+++LTIL+Q + GLE+ + ++G +WI +P S
Sbjct: 205 FLQLNSYPVCPDPHLAMGLAPHTDSSLLTILYQGNIPGLEIESPQEEGSRWIGVEPIEGS 264
Query: 125 QSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEH 184
+V+M D H SNG+ S HR ++ R SAA F+ P ++ D+ H
Sbjct: 265 --LVVIMGDLSHIISNGQFRSTMHRAVVNKTHHRVSAAYFAGPPKN--LQIGPLTSDKNH 320
Query: 185 PLLYKPFDLEEFL 197
P +Y+ EE+L
Sbjct: 321 PPIYRRLIWEEYL 333
>At5g07200 gibberellin 20-oxidase
Length = 380
Score = 81.6 bits (200), Expect = 3e-16
Identities = 52/163 (31%), Positives = 81/163 (48%), Gaps = 5/163 (3%)
Query: 20 NGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEK-YLDEHLNSTNYFLKTMKYKGPQT 78
+G F ++E ++ L + +++ SLGVE+ Y E ++ + Y +
Sbjct: 177 DGYEDFGKVYQEYAEAMNTLSLKIMELLGMSLGVERRYFKEFFEDSDSIFRLNYYPQCKQ 236
Query: 79 SDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAW 138
+ +G H D LTIL Q QV GL+V + KW S P+ + F+V + D F A
Sbjct: 237 PELALGTGPHCDPTSLTILHQDQVGGLQVFVDN--KWQSIPPNPHA--FVVNIGDTFMAL 292
Query: 139 SNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVD 181
+NGR S HR ++ R + A F PKG ++K PEELV+
Sbjct: 293 TNGRYKSCLHRAVVNSERERKTFAFFLCPKGEKVVKPPEELVN 335
>At5g51810 gibberellin 20-oxidase (emb|CAA58294.1)
Length = 378
Score = 80.5 bits (197), Expect = 6e-16
Identities = 56/187 (29%), Positives = 91/187 (47%), Gaps = 8/187 (4%)
Query: 32 FSEHLSELDQIVRKMVLESLGVEK-YLDEHLNSTNYFLKTMKYKGPQTSDTKVGLDTHTD 90
+ E +S L + +++ SLGV + Y + ++ Y QT D +G H D
Sbjct: 188 YCEAMSSLSLKIMELLGLSLGVNRDYFRGFFEENDSIMRLNHYPPCQTPDLTLGTGPHCD 247
Query: 91 TAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRV 150
+ LTIL Q V+GL+V + +W S +P+ ++ F+V + D F A SNG S HR
Sbjct: 248 PSSLTILHQDHVNGLQVFVDN--QWQSIRPNPKA--FVVNIGDTFMALSNGIFKSCLHRA 303
Query: 151 MMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEEFLKHSFNKEKGKRDD 210
++ AR S A F PK ++K P +++++ Y F FL+ + +K R D
Sbjct: 304 VVNRESARKSMAFFLCPKKDKVVKPPSDILEKMKTRKYPDFTWSMFLEFT---QKHYRAD 360
Query: 211 QFALHSY 217
L S+
Sbjct: 361 VNTLDSF 367
>At1g80320 oxidoreductase like protein
Length = 249
Score = 80.1 bits (196), Expect = 8e-16
Identities = 39/110 (35%), Positives = 64/110 (57%), Gaps = 3/110 (2%)
Query: 14 TKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVE--KYLDEHLNSTNYFLKTM 71
+K+LWP GN F ++ ++ELDQ V +M+ ES G++ K+ H ST Y L+ +
Sbjct: 114 SKLLWPQGNDPFCQTTHMYAMTMAELDQTVMRMLYESYGMDEKKHSVSHSESTRYLLRML 173
Query: 72 KYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPS 121
Y+ Q + G +HTD + ++IL Q+ V GL++ T G +W+ + PS
Sbjct: 174 SYRRQQNGEANTGFVSHTDKSFMSILHQNHVGGLQLKTMTG-QWVGFNPS 222
>At1g15550 Gibberillin 3 beta-hydroxylase (AtGA3ox1)
Length = 358
Score = 80.1 bits (196), Expect = 8e-16
Identities = 59/196 (30%), Positives = 88/196 (44%), Gaps = 12/196 (6%)
Query: 17 LWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKY------LDEHLNSTNYFLKT 70
LWP + ++ + + EH+ +L + + L SLGV + L LN L+
Sbjct: 155 LWPQHHLNYCDIVEEYEEHMKKLASKLMWLALNSLGVSEEDIEWASLSSDLNWAQAALQL 214
Query: 71 MKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVM 130
Y D +GL HTD+ +LTIL+Q+ GL+V +D W++ P S +V
Sbjct: 215 NHYPVCPEPDRAMGLAAHTDSTLLTILYQNNTAGLQVF-RDDLGWVTVPPFPGS--LVVN 271
Query: 131 MNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKP 190
+ D FH SNG S HR + AR S A P+ I +LV LY+
Sbjct: 272 VGDLFHILSNGLFKSVLHRARVNQTRARLSVAFLWGPQSDIKISPVPKLVSPVESPLYQS 331
Query: 191 FDLEEFLKHS---FNK 203
+E+L+ FNK
Sbjct: 332 VTWKEYLRTKATHFNK 347
>At3g19010 oxidase like protein
Length = 349
Score = 79.7 bits (195), Expect = 1e-15
Identities = 47/180 (26%), Positives = 82/180 (45%), Gaps = 3/180 (1%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKY-LDEHLNSTNYFLKTMKYKGP 76
WP FR A ++ H +L + +++ SLG+ K ++ F + +Y
Sbjct: 146 WPQSPSDFREACEVYARHAEKLAFKLLELISLSLGLPKERFHDYFKEQMSFFRINRYPPC 205
Query: 77 QTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFH 136
D +G+ H D ++++L Q V GL+V + W +P + ++ + +C
Sbjct: 206 PRPDLALGVGHHKDADVISLLAQDDVGGLQVSRRSDGVWFPIRPVPNA--LVINIGNCME 263
Query: 137 AWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEEF 196
W+N + S HRV++ RYS F +P +K EELV E+P YK + +F
Sbjct: 264 IWTNDKYWSAEHRVVVNTTRERYSIPFFLLPSHDVEVKPLEELVSPENPPKYKGYKWGKF 323
>At5g24530 flavanone 3-hydroxylase-like protein
Length = 341
Score = 79.0 bits (193), Expect = 2e-15
Identities = 56/191 (29%), Positives = 92/191 (47%), Gaps = 9/191 (4%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEK-YLDEHLNSTNYFLKTMKYKGP 76
WP+ PSF+ + +S + E+ + +++ ESLG+EK Y+ + L + Y
Sbjct: 141 WPSNPPSFKEIVSKYSREVREVGFKIEELISESLGLEKDYMKKVLGEQGQHMAVNYYPPC 200
Query: 77 QTSDTKVGLDTHTDTAILTILFQ-SQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCF 135
+ GL HTD LTIL Q + V GL++L DG +W + P + +F++ + D
Sbjct: 201 PEPELTYGLPAHTDPNALTILLQDTTVCGLQILI-DG-QWFAVNPHPD--AFVINIGDQL 256
Query: 136 HAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPE---ELVDEEHPLLYKPFD 192
A SNG S +HR + R S A F P ++ + E D+E +YK F
Sbjct: 257 QALSNGVYKSVWHRAVTNTENPRLSVASFLCPADCAVMSPAKPLWEAEDDETKPVYKDFT 316
Query: 193 LEEFLKHSFNK 203
E+ K +++
Sbjct: 317 YAEYYKKFWSR 327
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,295,387
Number of Sequences: 26719
Number of extensions: 226668
Number of successful extensions: 769
Number of sequences better than 10.0: 105
Number of HSP's better than 10.0 without gapping: 85
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 563
Number of HSP's gapped (non-prelim): 109
length of query: 219
length of database: 11,318,596
effective HSP length: 95
effective length of query: 124
effective length of database: 8,780,291
effective search space: 1088756084
effective search space used: 1088756084
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0226.15


