

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.11
(231 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
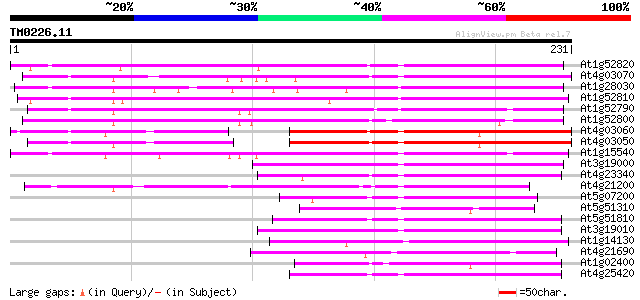
Score E
Sequences producing significant alignments: (bits) Value
At1g52820 oxidoreductase-like protein 231 2e-61
At4g03070 oxidoreductase - like protein (AOP1.1) 176 1e-44
At1g28030 hypothetical protein 155 2e-38
At1g52810 oxidoreductase like protein 146 1e-35
At1g52790 putative oxidoreductase 144 4e-35
At1g52800 putative oxidoreductase 144 5e-35
At4g03060 putative oxidoreductase 113 7e-26
At4g03050 2-oxoglutarate-dependent dioxygenase (AOP3) 110 5e-25
At1g15540 hypothetical protein 92 2e-19
At3g19000 unknown protein 80 7e-16
At4g23340 unknown protein 77 1e-14
At4g21200 gibberellin 20-oxidase - like protein 74 7e-14
At5g07200 gibberellin 20-oxidase 71 4e-13
At5g51310 gibberellin 20-oxidase-like protein 70 7e-13
At5g51810 gibberellin 20-oxidase (emb|CAA58294.1) 70 9e-13
At3g19010 oxidase like protein 68 4e-12
At1g14130 dioxygenase-like protein 68 4e-12
At4g21690 gibberellin 3 beta-hydroxylase - like protein 68 5e-12
At1g02400 unknown protein 67 8e-12
At4g25420 gibberellin 20-oxidase - Arabidopsis thaliana 65 2e-11
>At1g52820 oxidoreductase-like protein
Length = 317
Score = 231 bits (590), Expect = 2e-61
Identities = 131/295 (44%), Positives = 167/295 (56%), Gaps = 71/295 (24%)
Query: 1 MGSETTL---KLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEYMQL------------- 44
MGSET L +LPVIDF+N NL P W+ ++ V KAL +Y
Sbjct: 1 MGSETPLLPLRLPVIDFSN-KNLKPGEPEWDLTRADVQKALQDYGYFEASFDRIPFELRK 59
Query: 45 -------------------NVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKIL 85
NVSKKP+ GY GQ+P +PL++SMGIDD+++ EKV++ T+ L
Sbjct: 60 SVFGALEELFDLPLQTKLRNVSKKPFHGYVGQYPMVPLYESMGIDDSDIAEKVDAFTEKL 119
Query: 86 WPNGNPSFRYASQPIT--------------------------------YFLKTMKYKGPQ 113
WP GN SF Q + Y L+ MKYKGP
Sbjct: 120 WPQGNISFSTTIQSFSKKLSELDITIRRMIMESFGLDKYIDEHLHSTNYLLRVMKYKGPD 179
Query: 114 TSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHA 173
T +TKVGL+ HTD I+TIL+Q+ V+GLEV TKD + WI KP+ +S F VM+ D +A
Sbjct: 180 TEETKVGLNAHTDKNIVTILYQNHVEGLEVQTKD-KNWIKVKPTQDS--FTVMIGDSLYA 236
Query: 174 WSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
NGRLHSP+HRVMMTG E RYS LF +PK G I+ +P+ELVDEEHP L+KPFD
Sbjct: 237 LLNGRLHSPYHRVMMTGTETRYSLGLFSIPKAGHIVSSPDELVDEEHPRLFKPFD 291
>At4g03070 oxidoreductase - like protein (AOP1.1)
Length = 322
Score = 176 bits (445), Expect = 1e-44
Identities = 120/297 (40%), Positives = 155/297 (51%), Gaps = 79/297 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 11 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 69
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 70 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 125
Query: 93 FR-----YASQPI--------------------------TYFL-KTMKYKGPQTSD---- 116
++ Q + TY+L + MKY P D
Sbjct: 126 ISETIHLFSEQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDD 185
Query: 117 --TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAW 174
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A
Sbjct: 186 EETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCAL 242
Query: 175 SNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 243 LNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 299
>At1g28030 hypothetical protein
Length = 322
Score = 155 bits (392), Expect = 2e-38
Identities = 112/295 (37%), Positives = 142/295 (47%), Gaps = 74/295 (25%)
Query: 3 SETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY--------------------- 41
S+ L LP+IDF+N +L +P W+ V+S+V KAL EY
Sbjct: 7 SQVPLSLPIIDFSN-PDLKPETPEWDLVRSQVRKALEEYGCFEALFDGASMELRKALFES 65
Query: 42 --------MQLNVSKKPYFGYAGQF--PNIPLFKSMG---IDDANVYEKVESLTKILWPN 88
++ +S K Y G P +P+ + MG ID+ NV V LT LWP
Sbjct: 66 SKEVFDLPLETKLSTKTDVHYEGYLTIPRVPIQEGMGFYGIDNPNV---VNDLTHKLWPQ 122
Query: 89 GN-------PSFRYASQPITYFLKTM-------------------------KYKGPQTSD 116
GN SF + ++TM KYKG +
Sbjct: 123 GNIFVGKNVQSFAEKLIELNLTVRTMTLESFGLEKYMEEHLNAANKHFQLLKYKGISDDN 182
Query: 117 T--KVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHA 173
T K+G H D LTIL Q+ VDGLE+ TKDG +WI KPS S SFIVM H
Sbjct: 183 TENKIGFYPHIDRHFLTILCQNDAVDGLEIKTKDGEEWIKVKPSQAS-SFIVMAGASLHV 241
Query: 174 WSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
NG + P HRV++TG + RY AALF +PK G II APEE+VD+EHP LYKPFD
Sbjct: 242 LLNGGVFPPLHRVVITGKKDRYVAALFTIPKEGVIINAPEEMVDDEHPRLYKPFD 296
>At1g52810 oxidoreductase like protein
Length = 289
Score = 146 bits (368), Expect = 1e-35
Identities = 102/263 (38%), Positives = 132/263 (49%), Gaps = 38/263 (14%)
Query: 4 ETTL--KLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY-----------MQLN----- 45
ETTL +LPVIDFT+ + GT W++V++ V +AL EY ++L
Sbjct: 5 ETTLALQLPVIDFTSRDLKPGTI-QWDSVRADVRRALEEYGCFEALFDKVPLELRKAVFD 63
Query: 46 ----------------VSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNG 89
VSK+ Y GY GQ P +PLF+ MG+D A +KV + T LWP G
Sbjct: 64 ASEEVFQLPLETKKRVVSKRKYRGYVGQIPTLPLFEVMGVDFAENPDKVNAFTHKLWPQG 123
Query: 90 NPSFRYASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAIL--TILFQSQVDGLEVLTKD 147
N +F A + + + + GLD L T DG+EV TKD
Sbjct: 124 NNNFSEAVMSFAEKVSELDFMTRRMIMESFGLDESYIKEHLNSTKCLNDVKDGIEVKTKD 183
Query: 148 GRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGC 207
+ WI PS +S SFIV+ HA NGR+ + HRVM G R+SA LF VPK
Sbjct: 184 DKHWIKANPSQDS-SFIVLGGAMLHALLNGRVLTGVHRVMRMGANIRFSAGLFSVPKTED 242
Query: 208 IIKAPEELVDEEHPLLYKPFDRE 230
+I APEELVD E+P LYKP D E
Sbjct: 243 LIYAPEELVDAEYPRLYKPVDFE 265
>At1g52790 putative oxidoreductase
Length = 310
Score = 144 bits (363), Expect = 4e-35
Identities = 93/286 (32%), Positives = 134/286 (46%), Gaps = 70/286 (24%)
Query: 8 KLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY-------------------------- 41
K+P +DF+ + GT WE+ + + +AL EY
Sbjct: 4 KIPTLDFSREDLKPGTK-YWESTRENIRQALEEYGCFIIDLKDKTPLDLLDRVFGSLVDL 62
Query: 42 -------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNGNPSF- 93
N KP GY GQ P +PL +S+GID+A E S T ++WP GN F
Sbjct: 63 FDLPTQTKMKNKYDKPLNGYVGQIPALPLHESLGIDNATSLEATRSFTGLMWPQGNEHFS 122
Query: 94 ----RYAS---------------------------QPITYFLKTMKYKGPQTSDTKVGLD 122
+YA + TY L+ +K + P + +G
Sbjct: 123 ECLYKYAEFAAELDQMVTRMVFQSYNVEKYYDPYIESTTYLLRVLKNRAPNNENPTLGFV 182
Query: 123 THTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSP 182
THTD + TIL Q QV+GLE+ T++G + I+ SS S F+V+ D AWSN R+ SP
Sbjct: 183 THTDKSFTTILHQDQVNGLEMETREGER-ININLSSPS-LFMVVAGDALMAWSNDRVWSP 240
Query: 183 FHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
H+V+++G RYS +F G ++ PEEL+D +HPL+YKPFD
Sbjct: 241 RHQVLVSGETDRYSLGMFAFNNG--TLQVPEELIDHQHPLMYKPFD 284
>At1g52800 putative oxidoreductase
Length = 314
Score = 144 bits (362), Expect = 5e-35
Identities = 95/288 (32%), Positives = 133/288 (45%), Gaps = 71/288 (24%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
T K+P++D T+ L + W +V + +AL EY
Sbjct: 7 TTKVPILDLTSQQELKPNTSTWRSVSREACEALEEYGCFLAVYDGVTQQLDDSIFAAAEE 66
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNGNPSF 93
+ NV++KPY GY GQ P IPL + +G+D E + T ++WP GN F
Sbjct: 67 LFDLPTETKKKNVNEKPYHGYVGQMPVIPLHEGLGVDYVTNKEIAQRFTHLMWPQGNDRF 126
Query: 94 ------------------------------RYASQ--PITYFLKTMKYKGPQTSDTKVGL 121
Y S TY LK +KY P S +
Sbjct: 127 CNTVHTFSNAVAELDRLVVRMIFENYGVEKHYESHVGSKTYLLKFLKYLAPPESISMPAF 186
Query: 122 DTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHS 181
HTD L+IL Q+ V+GLEV +KDG +WIS + +S++VM D WSN R+ S
Sbjct: 187 PQHTDKTFLSILHQNDVNGLEVKSKDG-EWISLQ--LPPKSYVVMAGDISMGWSNDRIRS 243
Query: 182 PFHRVMMTGNEARYSAALF-FVPKGGCIIKAPEELVDEEHPLLYKPFD 228
HRV M G++ RY+ LF F+ ++ PEELVD++HPL+YKPFD
Sbjct: 244 CEHRVTMEGDKTRYTLGLFSFLTD---LVSIPEELVDDKHPLMYKPFD 288
>At4g03060 putative oxidoreductase
Length = 399
Score = 113 bits (283), Expect = 7e-26
Identities = 58/117 (49%), Positives = 80/117 (67%), Gaps = 4/117 (3%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
+ K+GL HTD + T+LFQ +++GLEV TKD KWI KPS + FIV+ D A
Sbjct: 260 EKKLGLPCHTDKNLFTVLFQHEIEGLEVKTKD-EKWIRVKPSPNT--FIVIAGDSLCALM 316
Query: 176 NGRLHSPFHRVMMTGNE-ARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGR+ +P+HRV +T + RY+AA+F PK +I+AP+ELVDE+HP L++PFD D
Sbjct: 317 NGRIRAPYHRVRVTEKKRTRYTAAIFTCPKPDYVIEAPKELVDEKHPRLFRPFDYRD 373
Score = 46.6 bits (109), Expect = 1e-05
Identities = 38/122 (31%), Positives = 54/122 (44%), Gaps = 37/122 (30%)
Query: 1 MGSETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKAL---------------------- 38
MGS +L+LP+I+ + G+S W V+S V KAL
Sbjct: 1 MGS-CSLQLPLINLADKTLEPGSS-KWAEVRSDVRKALEDFGCFEASYDKVSLELQESIM 58
Query: 39 ----------VEYMQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN 88
VE Q NV KPY GY + L +S+GI +AN+ E + T+ LWP+
Sbjct: 59 KTMEELFALPVETKQRNVCPKPYVGYLN---HNNLSESLGISNANILENINEFTQQLWPH 115
Query: 89 GN 90
G+
Sbjct: 116 GD 117
>At4g03050 2-oxoglutarate-dependent dioxygenase (AOP3)
Length = 411
Score = 110 bits (276), Expect = 5e-25
Identities = 59/117 (50%), Positives = 80/117 (67%), Gaps = 4/117 (3%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
D ++GL +HTD ++ I++Q Q+DGLEV TK+G KWI KP+ + IV+ D A
Sbjct: 271 DAELGLPSHTDKSLTGIIYQHQIDGLEVKTKEG-KWIRVKPAPNT--VIVIAGDALCALM 327
Query: 176 NGRLHSPFHRVMMTGNE-ARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGR+ SP+HRV +T + RY+AALF PK G II +P+ELVDE+HP +KPFD D
Sbjct: 328 NGRIPSPYHRVRVTEKKKTRYAAALFSNPKEGYIIDSPKELVDEKHPRAFKPFDFVD 384
Score = 45.1 bits (105), Expect = 3e-05
Identities = 36/117 (30%), Positives = 48/117 (40%), Gaps = 36/117 (30%)
Query: 8 KLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY-------------------------- 41
+LP+I ++ G+S W V+S V KAL +Y
Sbjct: 7 QLPLICLSDQTLKPGSS-KWVKVRSDVRKALEDYGCFEAKIDQVSMELQGSVLKAMQELF 65
Query: 42 ------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNGNPS 92
Q NV KP+ GY + L +S GI DAN+ EK T+ LWP GN S
Sbjct: 66 ALPTEAKQRNVCPKPFTGYLS---HNGLSESFGIKDANILEKAHEFTQQLWPEGNKS 119
>At1g15540 hypothetical protein
Length = 320
Score = 92.0 bits (227), Expect = 2e-19
Identities = 72/298 (24%), Positives = 121/298 (40%), Gaps = 73/298 (24%)
Query: 1 MGSETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKAL---------------------- 38
MG T +LP+I+ ++ NL + W + V +A+
Sbjct: 1 MGGNETRELPIINLSD-KNLKQNTELWSFTRDSVREAMEHHGWFVAEYNNFPTGLHQSIL 59
Query: 39 ----------VEYMQLNVSKKPYFGYAGQFPN-IPLFKSMGIDDANVYEKVESLTKILWP 87
VE N + K GY + P+ + +GID N ++ ++++WP
Sbjct: 60 EAAKELLDLPVEIKLKNENHKAGHGYITMMSDGQPVHEGLGIDQVNDVQQCRGFSRLMWP 119
Query: 88 NG---NPSF-----------------------------RYASQPI---TYFLKTMKYKGP 112
+ N F +Y + I Y L+ +KY+
Sbjct: 120 DDHDDNDRFCETVHAYAKMQAELEQLVIRMLFESYNVEKYTEKYIGGTRYLLRLLKYRRL 179
Query: 113 QTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFH 172
+ +HTD + ++IL Q+ + GL + ++ W + PS F+V+ D
Sbjct: 180 PNGEPNRKFISHTDKSFISILHQNHITGLMLKSEKEDVWYHFTPSPTR--FVVIAGDAIM 237
Query: 173 AWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRE 230
AWSN R+ + +H+V M E RYS F +G +I PEE+VD++HPL Y PF +
Sbjct: 238 AWSNDRIKACYHKVEMESVEMRYSLGFFSFQEG--MISTPEEMVDKDHPLAYNPFHHD 293
>At3g19000 unknown protein
Length = 352
Score = 80.5 bits (197), Expect = 7e-16
Identities = 43/128 (33%), Positives = 66/128 (50%), Gaps = 2/128 (1%)
Query: 101 TYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSES 160
T FL+ Y + +G+ H D LT+L Q V GL+V + +WI KP S++
Sbjct: 199 TSFLRFNHYPPCPNPELALGVGRHKDGGALTVLAQDSVGGLQVSRRSDGQWIPVKPISDA 258
Query: 161 QSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEH 220
I+ M +C W+N S HRV++ ++ R+S FF P I+ EEL+ EE+
Sbjct: 259 --LIINMGNCIQVWTNDEYWSAEHRVVVNTSKERFSIPFFFFPSHEANIEPLEELISEEN 316
Query: 221 PLLYKPFD 228
P YK ++
Sbjct: 317 PPCYKKYN 324
>At4g23340 unknown protein
Length = 277
Score = 76.6 bits (187), Expect = 1e-14
Identities = 46/129 (35%), Positives = 69/129 (52%), Gaps = 7/129 (5%)
Query: 103 FLKTMKYKGPQTSDTKV----GLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSS 158
+L+ + Y P + + GL HTD + +TI++Q V GL++ +K+G KWI P +
Sbjct: 114 YLRLVNYTPPHDVEKQEELVEGLGMHTDMSCITIVYQDSVGGLQMRSKEG-KWIDINPCN 172
Query: 159 ESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDE 218
+ +V + D AWSNGRL S HRV++ R S A F + +I AP+E+V E
Sbjct: 173 DF--LVVNIGDLMQAWSNGRLRSSEHRVVLRKLVNRVSLAFFLCFEDEKVILAPQEIVGE 230
Query: 219 EHPLLYKPF 227
YK F
Sbjct: 231 GKQRSYKSF 239
>At4g21200 gibberellin 20-oxidase - like protein
Length = 293
Score = 73.9 bits (180), Expect = 7e-14
Identities = 65/228 (28%), Positives = 100/228 (43%), Gaps = 31/228 (13%)
Query: 7 LKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY--------------------MQLNV 46
++LPVID + L + G E K + +A E+ Q+ V
Sbjct: 40 VELPVIDVSRL--IDGAEEEREKCKEAIARASREWGFFQVINHGISMDVLEKMRQEQIRV 97
Query: 47 SKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNGNPSFRYASQPITYFLKT 106
++P+ + N + K +A Y E L + N + F+ T +L+
Sbjct: 98 FREPF----DKKSNSTMEKFASESEALAYMLAEVLAEKSGQNSS-FFKENCVRNTCYLRM 152
Query: 107 MKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVM 166
+Y GL HTD+ LTIL+Q QV GL+++ KD R WI+ KP+ ++ I+
Sbjct: 153 NRYPPCPKPSEVYGLMPHTDSDFLTILYQDQVGGLQLI-KDNR-WIAVKPNPKA--LIIN 208
Query: 167 MNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEE 214
+ D F AWSNG S HRVM R+S A F P +I+ +
Sbjct: 209 IGDLFQAWSNGMYKSVEHRVMTNPKVERFSTAYFMCPSYDAVIECSSD 256
>At5g07200 gibberellin 20-oxidase
Length = 380
Score = 71.2 bits (173), Expect = 4e-13
Identities = 43/108 (39%), Positives = 58/108 (52%), Gaps = 6/108 (5%)
Query: 112 PQTSDTKVGLDT--HTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 169
PQ ++ L T H D LTIL Q QV GL+V + KW S P+ + F+V + D
Sbjct: 232 PQCKQPELALGTGPHCDPTSLTILHQDQVGGLQVFVDN--KWQSIPPNPHA--FVVNIGD 287
Query: 170 CFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVD 217
F A +NGR S HR ++ R + A F PKG ++K PEELV+
Sbjct: 288 TFMALTNGRYKSCLHRAVVNSERERKTFAFFLCPKGEKVVKPPEELVN 335
>At5g51310 gibberellin 20-oxidase-like protein
Length = 333
Score = 70.5 bits (171), Expect = 7e-13
Identities = 40/102 (39%), Positives = 59/102 (57%), Gaps = 9/102 (8%)
Query: 120 GLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRL 179
GL HTD + +TI+ Q + GL+V T+DG + P E + +V + D HAW+NGRL
Sbjct: 190 GLGMHTDMSCITIVDQDDIGGLQVRTRDGIGLMDINPKDE--ALVVNVGDLLHAWTNGRL 247
Query: 180 HSPFHRVMM-----TGNEARYSAALFFVPKGGCIIKAPEELV 216
S HRV++ GN R+S A F+ G ++ AP+E+V
Sbjct: 248 RSSQHRVILKRRGFVGN--RFSLAFFWCFDDGKVVFAPDEVV 287
>At5g51810 gibberellin 20-oxidase (emb|CAA58294.1)
Length = 378
Score = 70.1 bits (170), Expect = 9e-13
Identities = 41/119 (34%), Positives = 62/119 (51%), Gaps = 4/119 (3%)
Query: 109 YKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMN 168
Y QT D +G H D + LTIL Q V+GL+V + +W S +P+ ++ F+V +
Sbjct: 230 YPPCQTPDLTLGTGPHCDPSSLTILHQDHVNGLQVFVDN--QWQSIRPNPKA--FVVNIG 285
Query: 169 DCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
D F A SNG S HR ++ AR S A F PK ++K P +++++ Y F
Sbjct: 286 DTFMALSNGIFKSCLHRAVVNRESARKSMAFFLCPKKDKVVKPPSDILEKMKTRKYPDF 344
>At3g19010 oxidase like protein
Length = 349
Score = 68.2 bits (165), Expect = 4e-12
Identities = 35/125 (28%), Positives = 59/125 (47%), Gaps = 2/125 (1%)
Query: 103 FLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQS 162
F + +Y D +G+ H D ++++L Q V GL+V + W +P +
Sbjct: 196 FFRINRYPPCPRPDLALGVGHHKDADVISLLAQDDVGGLQVSRRSDGVWFPIRPVPNA-- 253
Query: 163 FIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPL 222
++ + +C W+N + S HRV++ RYS F +P +K EELV E+P
Sbjct: 254 LVINIGNCMEIWTNDKYWSAEHRVVVNTTRERYSIPFFLLPSHDVEVKPLEELVSPENPP 313
Query: 223 LYKPF 227
YK +
Sbjct: 314 KYKGY 318
>At1g14130 dioxygenase-like protein
Length = 308
Score = 68.2 bits (165), Expect = 4e-12
Identities = 43/124 (34%), Positives = 58/124 (46%), Gaps = 3/124 (2%)
Query: 108 KYKGPQTSDTKVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQSFIVM 166
KY + K+G+ HTD+ LTIL + V GLE + + P + + +
Sbjct: 160 KYHFKPETVGKLGVQLHTDSGFLTILQDDENVGGLEAMDNSSGTFFPIDPLPNTLA--IN 217
Query: 167 MNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
+ D WSNGRL + HRV RYS A F + ++ P E VD EHP LYKP
Sbjct: 218 LGDMATIWSNGRLCNVKHRVQCKEATMRYSIASFLLGPMDTDLEPPSEFVDAEHPRLYKP 277
Query: 227 FDRE 230
E
Sbjct: 278 ISHE 281
>At4g21690 gibberellin 3 beta-hydroxylase - like protein
Length = 349
Score = 67.8 bits (164), Expect = 5e-12
Identities = 42/128 (32%), Positives = 65/128 (49%), Gaps = 6/128 (4%)
Query: 100 ITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLT--KDGRKWISYKPS 157
I FL+ Y +GL HTD+++LTIL+Q + GLE+ + ++G +WI +P
Sbjct: 202 IQSFLQLNSYPVCPDPHLAMGLAPHTDSSLLTILYQGNIPGLEIESPQEEGSRWIGVEPI 261
Query: 158 SESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVD 217
S +V+M D H SNG+ S HR ++ R SAA F P ++ D
Sbjct: 262 --EGSLVVIMGDLSHIISNGQFRSTMHRAVVNKTHHRVSAAYFAGPPKN--LQIGPLTSD 317
Query: 218 EEHPLLYK 225
+ HP +Y+
Sbjct: 318 KNHPPIYR 325
>At1g02400 unknown protein
Length = 329
Score = 67.0 bits (162), Expect = 8e-12
Identities = 41/111 (36%), Positives = 58/111 (51%), Gaps = 4/111 (3%)
Query: 118 KVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNG 177
++G H+D ILT+L + VDGLE+ ++DG WI S+ F V++ DC A +NG
Sbjct: 197 QIGFGEHSDPQILTVLRSNDVDGLEICSRDG-LWIPI--PSDPTCFFVLVGDCLQALTNG 253
Query: 178 RLHSPFHRVMM-TGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
R S HRV+ T + R SA F P I ++V E+P Y F
Sbjct: 254 RFTSVRHRVLANTAKKPRMSAMYFAAPPLEAKISPLPKMVSPENPRRYNSF 304
>At4g25420 gibberellin 20-oxidase - Arabidopsis thaliana
Length = 377
Score = 65.5 bits (158), Expect = 2e-11
Identities = 39/112 (34%), Positives = 56/112 (49%), Gaps = 4/112 (3%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
D +G H D LTIL Q V+GL+V ++ +W S +P+ ++ F+V + D F A S
Sbjct: 239 DLTLGTGPHCDPTSLTILHQDHVNGLQVFVEN--QWRSIRPNPKA--FVVNIGDTFMALS 294
Query: 176 NGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
N R S HR ++ R S A F PK ++ P EL+D Y F
Sbjct: 295 NDRYKSCLHRAVVNSESERKSLAFFLCPKKDRVVTPPRELLDSITSRRYPDF 346
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,659,026
Number of Sequences: 26719
Number of extensions: 240660
Number of successful extensions: 688
Number of sequences better than 10.0: 103
Number of HSP's better than 10.0 without gapping: 84
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 500
Number of HSP's gapped (non-prelim): 120
length of query: 231
length of database: 11,318,596
effective HSP length: 96
effective length of query: 135
effective length of database: 8,753,572
effective search space: 1181732220
effective search space used: 1181732220
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0226.11


