

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.8
(423 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
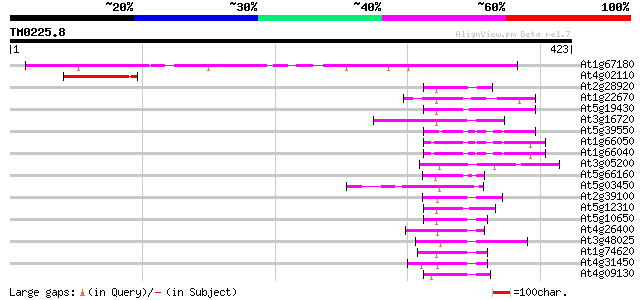
Score E
Sequences producing significant alignments: (bits) Value
At1g67180 hypothetical protein 205 4e-53
At4g02110 predicted protein of unknown function 48 1e-05
At2g28920 hypothetical protein 47 2e-05
At1g22670 putative RING zinc finger protein 47 3e-05
At5g19430 unknown protein 46 4e-05
At3g16720 putative RING zinc finger protein 46 4e-05
At5g39550 zinc finger -like protein 44 1e-04
At1g66050 hypothetical protein 44 1e-04
At1g66040 hypothetical protein 44 1e-04
At3g05200 RING-H2 zinc finger protein ATL6 (ATL6) 44 2e-04
At5g66160 ReMembR-H2 protein JR700 (gb|AAF32325.1) 44 2e-04
At5g03450 unknown protein 44 2e-04
At2g39100 RING zinc finger like protein 44 2e-04
At5g12310 RING finger -like protein 43 3e-04
At5g10650 Pspzf zinc finger protein - like 43 3e-04
At4g26400 unknown protein 43 3e-04
At3g48025 hypothetical protein 43 3e-04
At1g74620 putative RING zinc finger protein 43 3e-04
At4g31450 unknown protein 43 4e-04
At4g09130 putative protein 43 4e-04
>At1g67180 hypothetical protein
Length = 453
Score = 205 bits (521), Expect = 4e-53
Identities = 139/390 (35%), Positives = 211/390 (53%), Gaps = 42/390 (10%)
Query: 13 IVATVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHL---KFEGKKYDIARRLSIPVVN 69
+VATV+GYHG +RF LIKLI+++G SY+G+MS+SITHL KFEGKKYD+A++ VVN
Sbjct: 4 VVATVSGYHGSDRFKLIKLISHSGASYVGAMSRSITHLVCWKFEGKKYDLAKKFGTVVVN 63
Query: 70 HRWIEDCIREGKRLPIDSYTLQSGQEVGPLLLEVPASFGPNSWIKNKVVSDGLCDIESER 129
HRW+E+C++EG+R+ Y SG+EVGPL++E+PA + + KV + SE
Sbjct: 64 HRWVEECVKEGRRVSETPYMFDSGEEVGPLMIELPA-VSEEAKVTKKV------NKASET 116
Query: 130 QNTVISFGAPSYVLEDSCL---MKKYEETSLHSSRRLKKVKRNISCDNEVSTAAGPSRKE 186
+ S G + S L M+K E + HS R K +I + E S A SRK
Sbjct: 117 FDKYFSNGGENRSGSTSELATWMEKNVEANRHSVRLRTKRPSSILENKENSGVAESSRKG 176
Query: 187 KKFVKKVVDEVVSDPVILDLSPDDRLCRMDRDRLHTEAAATSSLSGGVDVNDLQENSSGP 246
KK V K S ++DL D+ + D H + + + N+ Q++
Sbjct: 177 KKRVVKQR----SYRNLIDLESDE-----ESDNNHHDNSDENQ-------NETQDHREPA 220
Query: 247 DARLCN------QNRAMNGSSDGIEQITDSNQIEDPQTVEQTSI----DLCSSDGKFIDG 296
D + + A+ D D ++IE+ + +++ S + K D
Sbjct: 221 DENVRGFVFEQGETSALRHPGDLATPNWDVDEIEESENWSHSAVFKRPRSFSPEIKPQDD 280
Query: 297 DQV---DNVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKIST 353
D+ + A + SC+IC+ +FSS+ G+LPC H FC +CIQKWA + +S K +T
Sbjct: 281 DESTREETEATEKAPAQVSCIICWTEFSSSRGILPCGHRFCYSCIQKWADRLVSERKKTT 340
Query: 354 CPLCKASFAWYKIVEHAATADKELNSQTVP 383
CPLCK++F +E A ++D+++ SQTVP
Sbjct: 341 CPLCKSNFITITKIEDADSSDQKIYSQTVP 370
>At4g02110 predicted protein of unknown function
Length = 1293
Score = 47.8 bits (112), Expect = 1e-05
Identities = 23/57 (40%), Positives = 38/57 (66%), Gaps = 2/57 (3%)
Query: 41 GSMSKSITHLKFEGKKYDIARRLS-IPVVNHRWIEDCIREGKRLPIDSYTLQSGQEV 96
GS + + ++G+KY++A+R+ I +VNHRW+EDC++ K LP Y + SG E+
Sbjct: 108 GSKALVVCLTGYQGEKYELAKRIKRIKLVNHRWLEDCLKNWKLLPEVDYEI-SGYEL 163
>At2g28920 hypothetical protein
Length = 145
Score = 47.0 bits (110), Expect = 2e-05
Identities = 22/55 (40%), Positives = 31/55 (56%), Gaps = 9/55 (16%)
Query: 313 CVICFEDF--SSTLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWY 364
CVIC EDF + + VL C+H F + CI W ++ TCP+C+A F W+
Sbjct: 93 CVICLEDFKVNDVVRVLVRCKHVFHVDCIDSWCFYKL------TCPICRAPFQWF 141
>At1g22670 putative RING zinc finger protein
Length = 422
Score = 46.6 bits (109), Expect = 3e-05
Identities = 29/103 (28%), Positives = 49/103 (47%), Gaps = 23/103 (22%)
Query: 298 QVDNVAGSPISNEWSCVICFEDF--SSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCP 355
++DN G +SC IC ED+ L VLPC H F + C+ W + + CP
Sbjct: 223 KIDNTTG------FSCAICLEDYIVGDKLRVLPCSHKFHVACVDSWL-----ISWRTFCP 271
Query: 356 LCKASFAWYKIVEHAATADKELNSQTVP--CDNLASVTFISMD 396
+CK + TAD+ L +++ P ++A+ + + +D
Sbjct: 272 VCKR--------DARTTADEPLATESTPFLSSSIATSSLVCID 306
>At5g19430 unknown protein
Length = 255
Score = 45.8 bits (107), Expect = 4e-05
Identities = 26/86 (30%), Positives = 40/86 (46%), Gaps = 6/86 (6%)
Query: 313 CVICFEDF--SSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHA 370
C IC + T V CEHT+C+TCI +WA S + TCP CK F + +
Sbjct: 46 CAICLSEIPLQETAMVKGCEHTYCVTCILRWA----SCKESPTCPQCKHPFDFLSVHRAL 101
Query: 371 ATADKELNSQTVPCDNLASVTFISMD 396
+ ++ + C L + F+ +D
Sbjct: 102 DGSIEDFLFEESVCLLLRASWFVPLD 127
>At3g16720 putative RING zinc finger protein
Length = 304
Score = 45.8 bits (107), Expect = 4e-05
Identities = 31/102 (30%), Positives = 42/102 (40%), Gaps = 9/102 (8%)
Query: 275 DPQTVEQTSIDLCSSDGKFIDGDQVDNVAGSPISNEWSCVICFEDF--SSTLGVLP-CEH 331
DP T + + D I V + + C +C +F S T VLP C+H
Sbjct: 81 DPSTAATSVVASRGLDPNVIKSLPVFTFSDETHKDPIECAVCLSEFEESETGRVLPNCQH 140
Query: 332 TFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAATA 373
TF + CI W H STCPLC++ +E A A
Sbjct: 141 TFHVDCIDMWFHSH------STCPLCRSLVESLAGIESTAAA 176
>At5g39550 zinc finger -like protein
Length = 617
Score = 44.3 bits (103), Expect = 1e-04
Identities = 29/84 (34%), Positives = 42/84 (49%), Gaps = 11/84 (13%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC + + PC H FCL C +KWA + GK+ TC +C++ KI H A
Sbjct: 129 CSICIQLPERPI-TTPCGHNFCLKCFEKWA---VGQGKL-TCMICRS-----KIPRHVA- 177
Query: 373 ADKELNSQTVPCDNLASVTFISMD 396
+ +N V LA+VT S++
Sbjct: 178 KNPRINLALVSAIRLANVTKCSVE 201
Score = 34.3 bits (77), Expect = 0.13
Identities = 34/136 (25%), Positives = 54/136 (39%), Gaps = 27/136 (19%)
Query: 301 NVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQ-KWA-----HQRISMGK---- 350
N + + E+SC IC E S + PC H FC C++ K+A +R + G+
Sbjct: 483 NTMKARLLKEFSCQICREVLSLPV-TTPCAHNFCKACLEAKFAGITQLRERSNGGRKLRA 541
Query: 351 ---ISTCPLCKASFAWYKIVEHAATADKELNSQTVPCDNLASVTFISMDQELPDNSF-ER 406
I TCP C + + L + V + + + +E D S E
Sbjct: 542 KKNIMTCPCCTTDLSEF------------LQNPQVNREMMEIIENFKKSEEEADASISEE 589
Query: 407 REEPRVPVTFQWRLSN 422
EE P T + ++ N
Sbjct: 590 EEEESEPPTKKIKMDN 605
>At1g66050 hypothetical protein
Length = 598
Score = 44.3 bits (103), Expect = 1e-04
Identities = 33/104 (31%), Positives = 46/104 (43%), Gaps = 23/104 (22%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC + PC H FCL C +KWA + GK+ TC +C++ KI H A
Sbjct: 129 CSICIQ-LPERPVTTPCGHNFCLKCFEKWA---VGQGKL-TCMICRS-----KIPRHVA- 177
Query: 373 ADKELNSQTVPCDNLASVT------------FISMDQELPDNSF 404
+ +N V LA+VT I +Q+ PD +F
Sbjct: 178 KNPRINLALVSAIRLANVTKCSGEATAAKVHHIIRNQDRPDKAF 221
Score = 30.4 bits (67), Expect = 1.9
Identities = 20/71 (28%), Positives = 31/71 (43%), Gaps = 14/71 (19%)
Query: 300 DNVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQ-KWA------------HQRI 346
+N + + E+SC IC + S + PC H FC C++ K+A +
Sbjct: 460 NNAMKARLLKEFSCQICRKVLSLPV-TTPCAHNFCKACLEAKFAGITQLRDRSNGVRKLR 518
Query: 347 SMGKISTCPLC 357
+ I TCP C
Sbjct: 519 AKKNIMTCPCC 529
>At1g66040 hypothetical protein
Length = 622
Score = 44.3 bits (103), Expect = 1e-04
Identities = 33/104 (31%), Positives = 46/104 (43%), Gaps = 23/104 (22%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC + PC H FCL C +KWA + GK+ TC +C++ KI H A
Sbjct: 129 CSICIQ-LPERPVTTPCGHNFCLKCFEKWA---VGQGKL-TCMICRS-----KIPRHVA- 177
Query: 373 ADKELNSQTVPCDNLASVT------------FISMDQELPDNSF 404
+ +N V LA+VT I +Q+ PD +F
Sbjct: 178 KNPRINLALVSAIRLANVTKCSGEATAAKVHHIIRNQDRPDKAF 221
Score = 30.4 bits (67), Expect = 1.9
Identities = 20/71 (28%), Positives = 31/71 (43%), Gaps = 14/71 (19%)
Query: 300 DNVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQ-KWA------------HQRI 346
+N + + E+SC IC + S + PC H FC C++ K+A +
Sbjct: 485 NNAMKARLLKEFSCQICRKVLSLPV-TTPCAHNFCKACLEAKFAGITQLRDRSNGVRKLR 543
Query: 347 SMGKISTCPLC 357
+ I TCP C
Sbjct: 544 AKKNIMTCPCC 554
>At3g05200 RING-H2 zinc finger protein ATL6 (ATL6)
Length = 398
Score = 43.9 bits (102), Expect = 2e-04
Identities = 36/113 (31%), Positives = 49/113 (42%), Gaps = 17/113 (15%)
Query: 310 EWSCVICFEDFSS--TLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWY-- 364
E C IC +F TL +LP C+H F CI W + TCP+C+A+ A
Sbjct: 125 ELECAICLNEFEDDETLRLLPKCDHVFHPHCIDAWLEAHV------TCPVCRANLAEQVA 178
Query: 365 --KIVEHAAT-ADKELNSQTVPCDNLASVTFISMDQELPDNSFERREEPRVPV 414
+ VE T D EL V N V + ++L + + R P VPV
Sbjct: 179 EGESVEPGGTEPDLELQQVVV---NPEPVVTAPVPEQLVTSEVDSRRLPGVPV 228
>At5g66160 ReMembR-H2 protein JR700 (gb|AAF32325.1)
Length = 310
Score = 43.5 bits (101), Expect = 2e-04
Identities = 21/49 (42%), Positives = 28/49 (56%), Gaps = 7/49 (14%)
Query: 312 SCVICFED--FSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
+C IC ED F +L +LPC+H F L CI W G ++CP+CK
Sbjct: 231 TCAICLEDYRFGESLRLLPCQHAFHLNCIDSWL---TKWG--TSCPVCK 274
>At5g03450 unknown protein
Length = 630
Score = 43.5 bits (101), Expect = 2e-04
Identities = 35/107 (32%), Positives = 44/107 (40%), Gaps = 24/107 (22%)
Query: 255 RAMNGSSDGIEQITDSNQIEDPQTVEQTSIDLCSSDGKFIDGDQVDNVAGSPISNEWSCV 314
R S+GIE + S +CSS G Q D + SC
Sbjct: 79 RVSQNESEGIEN-------------QNRSQGVCSSSS----GSQEDVEWKQGDTEGLSCS 121
Query: 315 ICFEDFSS----TLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLC 357
IC E ++S + LPC H + +CI KW QR S GK CPLC
Sbjct: 122 ICMEVWTSGGQHQVCCLPCGHLYGYSCINKWFQQRRSGGK---CPLC 165
>At2g39100 RING zinc finger like protein
Length = 296
Score = 43.5 bits (101), Expect = 2e-04
Identities = 21/64 (32%), Positives = 34/64 (52%), Gaps = 10/64 (15%)
Query: 312 SCVICFEDFS---STLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASF-AWYKIV 367
SC IC E+ + S + C+H +CL CI+KW+ + CPLC F +W+ +
Sbjct: 38 SCPICLENLTERRSAAVITVCKHGYCLACIRKWSSFK------RNCPLCNTRFDSWFIVS 91
Query: 368 EHAA 371
+ A+
Sbjct: 92 DFAS 95
>At5g12310 RING finger -like protein
Length = 254
Score = 43.1 bits (100), Expect = 3e-04
Identities = 21/56 (37%), Positives = 28/56 (49%), Gaps = 6/56 (10%)
Query: 313 CVICFEDF--SSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKI 366
C IC + T V CEH +C+TCI +WA S + TCP CK F + +
Sbjct: 43 CAICLDTIPLQETAMVKGCEHAYCVTCILRWA----SYKEKPTCPQCKLPFDFLNV 94
>At5g10650 Pspzf zinc finger protein - like
Length = 525
Score = 43.1 bits (100), Expect = 3e-04
Identities = 17/50 (34%), Positives = 29/50 (58%), Gaps = 8/50 (16%)
Query: 313 CVICFEDF--SSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKAS 360
C IC E++ LG +PC+H + ++C+Q+W + + CP+CK S
Sbjct: 475 CSICQEEYVDGDELGTIPCQHMYHVSCVQQWLRMK------NWCPICKTS 518
>At4g26400 unknown protein
Length = 356
Score = 43.1 bits (100), Expect = 3e-04
Identities = 22/62 (35%), Positives = 30/62 (47%), Gaps = 8/62 (12%)
Query: 299 VDNVAGSPISNEWSCVICFEDFS--STLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPL 356
VDN+ IS C IC +DF S +PC+H F + CI W S+CP+
Sbjct: 227 VDNLPTVKISESLQCSICLDDFDKGSEAKEMPCKHKFHIRCIVPWLELH------SSCPV 280
Query: 357 CK 358
C+
Sbjct: 281 CR 282
>At3g48025 hypothetical protein
Length = 268
Score = 43.1 bits (100), Expect = 3e-04
Identities = 28/88 (31%), Positives = 39/88 (43%), Gaps = 10/88 (11%)
Query: 307 ISNEWSCVICFEDFSST--LGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAW 363
+ + C +C +FS T L +LP C H F L CI W STCPLC+ S +
Sbjct: 120 LEQPFDCAVCLNEFSDTDKLRLLPVCSHAFHLHCIDTWLLSN------STCPLCRRSLST 173
Query: 364 YKI-VEHAATADKELNSQTVPCDNLASV 390
+ H+ T L+ D AS+
Sbjct: 174 SNVCYNHSETLVAPLSGHQQVDDGKASL 201
>At1g74620 putative RING zinc finger protein
Length = 249
Score = 43.1 bits (100), Expect = 3e-04
Identities = 22/56 (39%), Positives = 28/56 (49%), Gaps = 9/56 (16%)
Query: 308 SNEWSCVICFEDF---SSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKAS 360
+ E C IC ED+ SS + LPC+H F CI KW CPLC++S
Sbjct: 178 TEENGCAICMEDYIEGSSIVAKLPCDHEFHGDCINKWLQLN------HMCPLCRSS 227
>At4g31450 unknown protein
Length = 497
Score = 42.7 bits (99), Expect = 4e-04
Identities = 22/64 (34%), Positives = 36/64 (55%), Gaps = 10/64 (15%)
Query: 301 NVAGSPISN--EWSCVICFEDFS--STLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPL 356
++ SP N + C IC E+++ +G L CEHT+ + C+Q+W + S CP+
Sbjct: 434 SITKSPSDNKEDAKCSICQEEYTIGDEVGRLHCEHTYHVKCVQEWLRIK------SWCPI 487
Query: 357 CKAS 360
CKA+
Sbjct: 488 CKAT 491
>At4g09130 putative protein
Length = 357
Score = 42.7 bits (99), Expect = 4e-04
Identities = 21/53 (39%), Positives = 27/53 (50%), Gaps = 9/53 (16%)
Query: 313 CVIC---FEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFA 362
C IC FED + PC HTF CI +W R STCP+C+A+ +
Sbjct: 120 CAICLCEFEDEEPLRWMPPCSHTFHANCIDEWLSSR------STCPVCRANLS 166
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,982,523
Number of Sequences: 26719
Number of extensions: 434672
Number of successful extensions: 1606
Number of sequences better than 10.0: 305
Number of HSP's better than 10.0 without gapping: 50
Number of HSP's successfully gapped in prelim test: 255
Number of HSP's that attempted gapping in prelim test: 1480
Number of HSP's gapped (non-prelim): 340
length of query: 423
length of database: 11,318,596
effective HSP length: 102
effective length of query: 321
effective length of database: 8,593,258
effective search space: 2758435818
effective search space used: 2758435818
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0225.8


