

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.5
(295 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
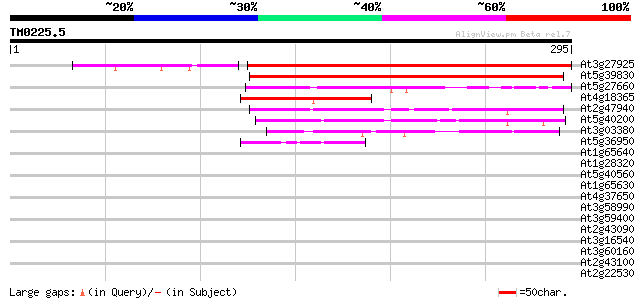
Score E
Sequences producing significant alignments: (bits) Value
At3g27925 DegP protease precursor, 5' partial 307 4e-84
At5g39830 DegP protease-like protein 162 2e-40
At5g27660 serine protease-like protein 72 4e-13
At4g18365 protease HhoA like precursor 70 1e-12
At2g47940 DegP2 protease (DEGP2) 57 9e-09
At5g40200 putative serine protease (MSN9.10) 49 3e-06
At3g03380 unknown protein 43 2e-04
At5g36950 unknown protein 42 5e-04
At1g65640 serine protease DO, putative 40 0.001
At1g28320 unknown protein 40 0.002
At5g40560 putative protein 38 0.006
At1g65630 serine protease DO, putative 33 0.24
At4g37650 SHORT-ROOT (SHR) 32 0.31
At3g58990 3-isopropylmalate dehydratase-like protein (small subu... 32 0.31
At3g59400 unknown protein (At3g59400) 32 0.53
At2g43090 3-isopropylmalate dehydratase, small subunit 32 0.53
At3g16540 putative serine protease 31 0.70
At3g60160 multi resistance protein homolog 31 0.91
At2g43100 3-isopropylmalate dehydratase, small subunit 30 1.2
At2g22530 unknown protein 30 1.5
>At3g27925 DegP protease precursor, 5' partial
Length = 386
Score = 307 bits (787), Expect = 4e-84
Identities = 153/170 (90%), Positives = 163/170 (95%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSG LIGINTAIYSPSGASSGVGFSIP+DTV GIVDQLV+F
Sbjct: 217 VIQTDAAINPGNSGGPLLDSSGTLIGINTAIYSPSGASSGVGFSIPVDTVGGIVDQLVRF 276
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAP +GPAGKAGLQSTKRD YGRL+LGDIIT
Sbjct: 277 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPPSGPAGKAGLQSTKRDGYGRLVLGDIIT 336
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
SVNGTKV++GSDLYRILDQCKVGD+VTVEVLRGDHKEKI V LEPKPDES
Sbjct: 337 SVNGTKVSNGSDLYRILDQCKVGDEVTVEVLRGDHKEKISVTLEPKPDES 386
Score = 73.2 bits (178), Expect = 2e-13
Identities = 51/116 (43%), Positives = 64/116 (54%), Gaps = 31/116 (26%)
Query: 34 TPKTCFNSILILCTSIALSFT------NADSAYAFVVTPPRKLQSDELATV--------- 78
TP + +LCTS+ALSF+ +SA AFVV+ P+KLQ+DELATV
Sbjct: 19 TPFSAVKPFFLLCTSVALSFSLFAASPAVESASAFVVSTPKKLQTDELATVRLFQENTPS 78
Query: 79 -VYITNLAVKMRLRWM-------------CWRFLKEGNIVTNYHVIPGASDLSLDL 120
VYITNLAV+ + W K+G+IVTNYHVI GASDL + L
Sbjct: 79 VVYITNLAVRQDAFTLDVLEVPQGSGSGFVWD--KQGHIVTNYHVIRGASDLRVTL 132
>At5g39830 DegP protease-like protein
Length = 448
Score = 162 bits (411), Expect = 2e-40
Identities = 86/166 (51%), Positives = 114/166 (67%), Gaps = 1/166 (0%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ +DAAINPGNSGGPLLDS GNLIGINTAI++ +G S+GVGF+IP TV IV QL++F
Sbjct: 281 IQTDAAINPGNSGGPLLDSKGNLIGINTAIFTQTGTSAGVGFAIPSSTVLKIVPQLIQFS 340
Query: 187 KVTRPILGIKFAPDQSVEQLGV-SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
KV R + I+ APD QL V +G LVL P A KAGL T R G ++LGDII
Sbjct: 341 KVLRAGINIELAPDPVANQLNVRNGALVLQVPGKSLAEKAGLHPTSRGFAGNIVLGDIIV 400
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPK 291
+V+ V + ++L +ILD+ VGDKVT+++ RG+ ++ + LE K
Sbjct: 401 AVDDKPVKNKAELMKILDEYSVGDKVTLKIKRGNEDLELKISLEEK 446
>At5g27660 serine protease-like protein
Length = 459
Score = 72.0 bits (175), Expect = 4e-13
Identities = 59/182 (32%), Positives = 92/182 (50%), Gaps = 35/182 (19%)
Query: 125 QLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVK 184
+ + +D +IN GNSGGPL++ G +IG+N A+ G+GFS+P+D+V+ I++ K
Sbjct: 298 EYLQTDCSINAGNSGGPLVNLDGEVIGVNIMKVL---AADGLGFSVPIDSVSKIIEHFKK 354
Query: 185 FGKVTRPILGIKFAP--DQSVEQLG---------VSGVLVLDAPANGPAGKAGLQSTKRD 233
G+V RP +G+K + V QL GVLV PA +AG +
Sbjct: 355 SGRVIRPWIGLKMVELNNLIVAQLKERDPMFPDVERGVLVPTVIPGSPADRAGFKP---- 410
Query: 234 SYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPD 293
GD++ +G V I+D +VG ++ V V R +KE+ V LE P+
Sbjct: 411 -------GDVVVRFDGKPV------IEIMDD-RVGKRMQVVVER-SNKER--VTLEVIPE 453
Query: 294 ES 295
E+
Sbjct: 454 EA 455
>At4g18365 protease HhoA like precursor
Length = 323
Score = 70.1 bits (170), Expect = 1e-12
Identities = 36/71 (50%), Positives = 49/71 (68%), Gaps = 2/71 (2%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYS--PSGASSGVGFSIPLDTVNGIV 179
++ + + +DA IN GNSGGPLLDS G+ IG+NTA ++ SG SSGV F+IP+DTV V
Sbjct: 250 SISEAIQTDADINSGNSGGPLLDSYGHTIGVNTATFTRKGSGMSSGVNFAIPIDTVVRTV 309
Query: 180 DQLVKFGKVTR 190
L+ +G R
Sbjct: 310 PYLIVYGTAYR 320
>At2g47940 DegP2 protease (DEGP2)
Length = 607
Score = 57.4 bits (137), Expect = 9e-09
Identities = 50/176 (28%), Positives = 79/176 (44%), Gaps = 18/176 (10%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ DAAINPGNSGGP + G IG+ +Y S + +G+ IP V+ + + G
Sbjct: 257 IQIDAAINPGNSGGPAFNDQGECIGVAFQVYR-SEETENIGYVIPTTVVSHFLTDYERNG 315
Query: 187 KVT-RPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
K T P LG+ Q +E + L P N ++ T D+ L GD+I
Sbjct: 316 KYTGYPCLGVLL---QKLENPALRE--CLKVPTNEGVLVRRVEPTS-DASKVLKEGDVIV 369
Query: 246 SVNGTKV-TSGSDLYR---------ILDQCKVGDKVTVEVLRGDHKEKIPVILEPK 291
S + V G+ +R ++ Q GD + ++R +K+ V+L P+
Sbjct: 370 SFDDLHVGCEGTVPFRSSERIAFRYLISQKFAGDIAEIGIIRAGEHKKVQVVLRPR 425
>At5g40200 putative serine protease (MSN9.10)
Length = 592
Score = 48.9 bits (115), Expect = 3e-06
Identities = 51/185 (27%), Positives = 80/185 (42%), Gaps = 28/185 (15%)
Query: 130 DAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFGKVT 189
DAAIN GNSGGP + G +GI A + +G+ IP + + K K T
Sbjct: 270 DAAINSGNSGGPAFNDKGKCVGIAFQSLKHEDAEN-IGYVIPTPVIVHFIQDYEKHDKYT 328
Query: 190 R-PILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIITSVN 248
P+LGI++ Q +E + + +++ G + ++ T +S L DII S +
Sbjct: 329 GFPVLGIEW---QKMENPDLRKSMGMESHQKGVRIRR-IEPTAPESQ-VLKPSDIILSFD 383
Query: 249 GTKVTS-GSDLYR---------ILDQCKVGDKVTVEVLRGD-----------HKEKIPVI 287
G + + G+ +R ++ Q GD V+VLR HK IP
Sbjct: 384 GVNIANDGTVPFRHGERIGFSYLISQKYTGDSALVKVLRNKEILEFNIKLAIHKRLIPAH 443
Query: 288 LEPKP 292
+ KP
Sbjct: 444 ISGKP 448
>At3g03380 unknown protein
Length = 627
Score = 42.7 bits (99), Expect = 2e-04
Identities = 48/185 (25%), Positives = 77/185 (40%), Gaps = 50/185 (27%)
Query: 136 GNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVK----------- 184
G+SG P++D G + +N S +SS F +PL V + L K
Sbjct: 191 GSSGSPVIDWQGRAVALNAG----SKSSSASAFFLPLQRVVRALSFLQKSIDSRTDKPKA 246
Query: 185 ----------------FGKVTRPILGIKFAPDQSVEQL---GVSGVLVLDAPA-NGPAGK 224
F ++ R LG++ +Q V G +G+LV+D+ +GPA K
Sbjct: 247 VHIPRGTLQMTFLHKGFDEIRR--LGLRSETEQVVRHASPTGETGMLVVDSVVPSGPADK 304
Query: 225 AGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKI 284
L GD++ VNGT +T +L +LD VG + +E+ RG +
Sbjct: 305 ------------HLEPGDVLVRVNGTVLTQFLNLENLLDD-GVGQILELEIERGGQPLSV 351
Query: 285 PVILE 289
V ++
Sbjct: 352 SVSVQ 356
>At5g36950 unknown protein
Length = 586
Score = 41.6 bits (96), Expect = 5e-04
Identities = 25/66 (37%), Positives = 37/66 (55%), Gaps = 4/66 (6%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQ 181
T L + DAAINPGNSGGP + GN + A + SGA + +G+ IP + ++
Sbjct: 249 TQLMAIQIDAAINPGNSGGPAI--MGNKVA-GVAFQNLSGAEN-IGYIIPTPVIKHFING 304
Query: 182 LVKFGK 187
+ + GK
Sbjct: 305 VEECGK 310
>At1g65640 serine protease DO, putative
Length = 518
Score = 40.4 bits (93), Expect = 0.001
Identities = 41/189 (21%), Positives = 84/189 (43%), Gaps = 34/189 (17%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGI--- 178
T L + +DAAINPGNSGGP + + G+ + ++ +G+ IP +
Sbjct: 209 TTLLAIQTDAAINPGNSGGPAI-IGNKMAGV---AFQKDPSADNIGYIIPTPVIKHFLTA 264
Query: 179 VDQLVKFGKVTRPILGIKFAPDQSVE---QLG--VSGVLVLDAPANGPAGKAGLQSTKRD 233
V++ ++G + + + + ++G ++G+L+ + + D
Sbjct: 265 VEENGQYGGFCTLDISYQLMENSQLRNHFKMGPEMTGILINEI------------NPLSD 312
Query: 234 SYGRLILGDIITSV------NGTKVTSGS----DLYRILDQCKVGDKVTVEVLRGDHKEK 283
+Y RL DII ++ N KVT + + + K+ + V ++VLR + +
Sbjct: 313 AYKRLRKDDIILAIDDVLIGNDAKVTFRNKERINFNHFVSMKKLDETVLLQVLRDGKEHE 372
Query: 284 IPVILEPKP 292
++++P P
Sbjct: 373 FHIMVKPVP 381
>At1g28320 unknown protein
Length = 709
Score = 39.7 bits (91), Expect = 0.002
Identities = 21/56 (37%), Positives = 34/56 (60%), Gaps = 5/56 (8%)
Query: 116 LSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIP 171
+S ++ ++ + AA++PG SGG +L+SSG++IG+ T S A G G IP
Sbjct: 558 ISQEVAEFPAMLETTAAVHPGGSGGAVLNSSGHMIGLVT-----SNARHGAGTVIP 608
>At5g40560 putative protein
Length = 410
Score = 38.1 bits (87), Expect = 0.006
Identities = 21/64 (32%), Positives = 34/64 (52%), Gaps = 4/64 (6%)
Query: 124 LQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLV 183
L + +DAA+NPGNSGGP+ ++G+ + G S+ +G IP V + +
Sbjct: 102 LPAIQTDAAMNPGNSGGPVC-IGNKVVGV---AFQTLGHSNNIGCLIPAPVVKHFITGVE 157
Query: 184 KFGK 187
K G+
Sbjct: 158 KTGQ 161
>At1g65630 serine protease DO, putative
Length = 559
Score = 32.7 bits (73), Expect = 0.24
Identities = 20/66 (30%), Positives = 33/66 (49%), Gaps = 4/66 (6%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQ 181
T L + DAAIN GNSGGP++ GN + + S +G+ IP + ++
Sbjct: 231 TELLAIQIDAAINNGNSGGPVI--MGNKVA--GVAFESLCYSDSIGYIIPTPVIRHFLNA 286
Query: 182 LVKFGK 187
+ + G+
Sbjct: 287 IEESGE 292
>At4g37650 SHORT-ROOT (SHR)
Length = 531
Score = 32.3 bits (72), Expect = 0.31
Identities = 18/64 (28%), Positives = 29/64 (45%), Gaps = 1/64 (1%)
Query: 4 HYKHHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSILILCTSIALSFTNADSAYAFV 63
H+ HH + +++YSP TP SSTP + + + + N SA++
Sbjct: 70 HHNHHNHN-NPNTYYSPFTTPTQYHPATSSTPSSTAAAAALASPYSSSGHHNDPSAFSIP 128
Query: 64 VTPP 67
TPP
Sbjct: 129 QTPP 132
>At3g58990 3-isopropylmalate dehydratase-like protein (small
subunit)
Length = 253
Score = 32.3 bits (72), Expect = 0.31
Identities = 21/60 (35%), Positives = 32/60 (53%), Gaps = 8/60 (13%)
Query: 223 GKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKE 282
G AG ++ +SY R+ + + + G S++ RI D+CK GD VT+E HKE
Sbjct: 157 GAAGAKAVVAESYARIFFRNCVAT--GEIFPLESEV-RICDECKTGDVVTIE-----HKE 208
>At3g59400 unknown protein (At3g59400)
Length = 265
Score = 31.6 bits (70), Expect = 0.53
Identities = 23/74 (31%), Positives = 37/74 (49%), Gaps = 10/74 (13%)
Query: 5 YKHHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSILILCTSIALSFTNADSAYAFVV 64
Y HH+ LH S HF P SLSL + ++ T ++C+ A S +++ +A + V
Sbjct: 17 YTHHRNNLHCQS----HFGPTSLSLKQPTSAAT----FSLICS--ASSTSSSTTAVSAVS 66
Query: 65 TPPRKLQSDELATV 78
T + E AT+
Sbjct: 67 TTNASATTAETATI 80
>At2g43090 3-isopropylmalate dehydratase, small subunit
Length = 251
Score = 31.6 bits (70), Expect = 0.53
Identities = 18/57 (31%), Positives = 29/57 (50%), Gaps = 3/57 (5%)
Query: 223 GKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGD 279
G AG ++ SY R+ + + + G S++ R+ D+C GD TVE+ GD
Sbjct: 154 GAAGAKAVVAQSYARIFFRNSVAT--GEVYPLDSEV-RVCDECTTGDVATVELREGD 207
>At3g16540 putative serine protease
Length = 555
Score = 31.2 bits (69), Expect = 0.70
Identities = 38/178 (21%), Positives = 73/178 (40%), Gaps = 26/178 (14%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ DA IN NSGGP++ ++G+ +Y +GF IP + + + +
Sbjct: 247 IQIDATINDENSGGPVI-MGNKVVGV---VYE-------IGFVIPTPIIKHFITSVQESR 295
Query: 187 KVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIITS 246
+ + G QS+E + + + G ++ +Y L DII +
Sbjct: 296 QYS--CFGSLDLSYQSLENVQIRNHFKMSHEMTGIL--INKINSSSGAYKILRKDDIILA 351
Query: 247 VNGTKVTSGS----------DLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEP-KPD 293
++G + + D ++ K G+K V+VLR + + + L+P KP+
Sbjct: 352 IDGVPIGNDEKVPFQNKRRIDFSYLVSMKKPGEKALVKVLRNGKEYEYNISLKPVKPN 409
>At3g60160 multi resistance protein homolog
Length = 1490
Score = 30.8 bits (68), Expect = 0.91
Identities = 33/125 (26%), Positives = 53/125 (42%), Gaps = 15/125 (12%)
Query: 27 LSLLRSSTPKTCFNSILILCTSIALSFTNADSAYAFVVTPPRKLQSDELATVVYITNLAV 86
L L R S C +S+ + ++ SF+ + FV K++ L ++
Sbjct: 101 LLLFRDSVVSRCDSSVSVFSAEVSQSFS-----WLFVSVVVVKIRERRLVKFPWM----- 150
Query: 87 KMRLRWMCWRFLK---EGNIVTNYHVIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLL 143
+R W+C L + + +T H D + DLT LL + A G +G LL
Sbjct: 151 -LRSWWLCSFILSFSFDAHFITAKHEPLEFQDYA-DLTGLLASLFLLAVSIRGKTGFHLL 208
Query: 144 DSSGN 148
+SSGN
Sbjct: 209 ESSGN 213
>At2g43100 3-isopropylmalate dehydratase, small subunit
Length = 256
Score = 30.4 bits (67), Expect = 1.2
Identities = 17/53 (32%), Positives = 28/53 (52%), Gaps = 3/53 (5%)
Query: 223 GKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEV 275
G AG ++ +SY R+ + SV +V R+ ++CK GD VT+E+
Sbjct: 160 GAAGAKAIVAESYARIFFRN---SVATGEVFPLESEVRVCEECKTGDTVTIEL 209
>At2g22530 unknown protein
Length = 897
Score = 30.0 bits (66), Expect = 1.5
Identities = 18/54 (33%), Positives = 28/54 (51%), Gaps = 3/54 (5%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNL---IGINTAIYSPSGASSGVGFSIPL 172
TLL +VS GN GG + + +L IG+N+ I + A++ V F + L
Sbjct: 273 TLLIIVSDHGMTENGNHGGSSYEETDSLMLFIGLNSNISDYASATNNVAFQVDL 326
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,749,685
Number of Sequences: 26719
Number of extensions: 284543
Number of successful extensions: 891
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 858
Number of HSP's gapped (non-prelim): 40
length of query: 295
length of database: 11,318,596
effective HSP length: 99
effective length of query: 196
effective length of database: 8,673,415
effective search space: 1699989340
effective search space used: 1699989340
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0225.5


