

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0221.5
(190 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
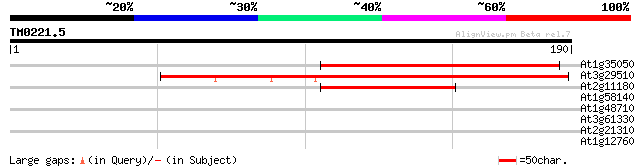
Sequences producing significant alignments: (bits) Value
At1g35050 putative protein 91 5e-19
At3g29510 hypothetical protein 90 6e-19
At2g11180 putative retroelement pol polyprotein 46 1e-05
At1g58140 hypothetical protein 30 0.59
At1g48710 hypothetical protein 30 0.59
At3g61330 copia-type polyprotein 29 1.7
At2g21310 putative retroelement pol polyprotein 28 3.8
At1g12760 unknown protein 28 3.8
>At1g35050 putative protein
Length = 450
Score = 90.5 bits (223), Expect = 5e-19
Identities = 44/81 (54%), Positives = 58/81 (71%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNE 165
FCL L IE+ +D +F++Q Y K+LK F MDK+ PL T ++ RSLNV DPFRP E NE
Sbjct: 254 FCLGLQIEHFQDGIFVHQSNYTKKILKRFNMDKANPLSTPMVNRSLNVENDPFRPCEDNE 313
Query: 166 ELLDPEVLYLSAIKALLYLAS 186
+ L P+V Y+SAI L+YLA+
Sbjct: 314 DFLGPKVPYMSAIGGLMYLAN 334
>At3g29510 hypothetical protein
Length = 1158
Score = 90.1 bits (222), Expect = 6e-19
Identities = 57/142 (40%), Positives = 87/142 (61%), Gaps = 4/142 (2%)
Query: 52 QNDPTNSCIFMMRYEIV-YVIINVYDNDLLRL*KSSQ--KL*IC*RKNLR*KS*-E*NFC 107
+NDP + CIF+ ++ +VII VY +DL L S + + +K K + FC
Sbjct: 804 KNDPISPCIFIKKFASKGFVIIAVYVDDLNILGTSGEIAQTVEYLKKEFEMKDLGKTKFC 863
Query: 108 LAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNEEL 167
L L +EY+ + ++Q+ Y VLK F MDK+ PL + + +RSL ++ DPF P + +EE+
Sbjct: 864 LGLQLEYIDKGILVHQKAYTETVLKRFNMDKAHPLTSPMQVRSLGLDSDPFGPKKDDEEI 923
Query: 168 LDPEVLYLSAIKALLYLASHCK 189
L PE+ YLSAI AL+YL+SH +
Sbjct: 924 LGPEMPYLSAIGALMYLSSHTR 945
>At2g11180 putative retroelement pol polyprotein
Length = 557
Score = 45.8 bits (107), Expect = 1e-05
Identities = 22/46 (47%), Positives = 30/46 (64%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSL 151
FCL L IE+LK+ + YQ TY VLK FYMD + PL + ++ + L
Sbjct: 401 FCLGLQIEHLKNVILEYQETYTKNVLKRFYMDGAYPLSSPMVGKRL 446
>At1g58140 hypothetical protein
Length = 1320
Score = 30.4 bits (67), Expect = 0.59
Identities = 14/40 (35%), Positives = 24/40 (60%)
Query: 105 NFCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCT 144
++ L + ++ + +F+ Q Y +VLK F MD S P+CT
Sbjct: 1033 SYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMDDSNPVCT 1072
>At1g48710 hypothetical protein
Length = 1352
Score = 30.4 bits (67), Expect = 0.59
Identities = 14/40 (35%), Positives = 24/40 (60%)
Query: 105 NFCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCT 144
++ L + ++ + +F+ Q Y +VLK F MD S P+CT
Sbjct: 1065 SYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMDDSNPVCT 1104
>At3g61330 copia-type polyprotein
Length = 1352
Score = 28.9 bits (63), Expect = 1.7
Identities = 13/40 (32%), Positives = 24/40 (59%)
Query: 105 NFCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCT 144
++ L + ++ + +F+ Q Y +VLK F +D S P+CT
Sbjct: 1065 SYYLGIEVKQEDNGIFITQEGYAKEVLKKFKIDDSNPVCT 1104
>At2g21310 putative retroelement pol polyprotein
Length = 838
Score = 27.7 bits (60), Expect = 3.8
Identities = 18/61 (29%), Positives = 32/61 (51%), Gaps = 2/61 (3%)
Query: 123 QRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNEELLDPEVLYLSAIKALL 182
QR+Y+ KVLK F MD+ P+ T L V E+ +++ + Y +A+ +++
Sbjct: 571 QRSYLQKVLKTFRMDECKPVKTPLAPHMKFVAATETEAEEQADQM--KSIPYANAVGSIM 628
Query: 183 Y 183
Y
Sbjct: 629 Y 629
>At1g12760 unknown protein
Length = 408
Score = 27.7 bits (60), Expect = 3.8
Identities = 10/25 (40%), Positives = 18/25 (72%)
Query: 132 KGFYMDKSCPLCTLLILRSLNVNKD 156
K Y++ +CPLC IL+S N++++
Sbjct: 382 KWLYINATCPLCKYNILKSSNLDRE 406
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.359 0.163 0.534
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,316,688
Number of Sequences: 26719
Number of extensions: 105074
Number of successful extensions: 463
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 454
Number of HSP's gapped (non-prelim): 8
length of query: 190
length of database: 11,318,596
effective HSP length: 94
effective length of query: 96
effective length of database: 8,807,010
effective search space: 845472960
effective search space used: 845472960
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0221.5


