

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0218.20
(631 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
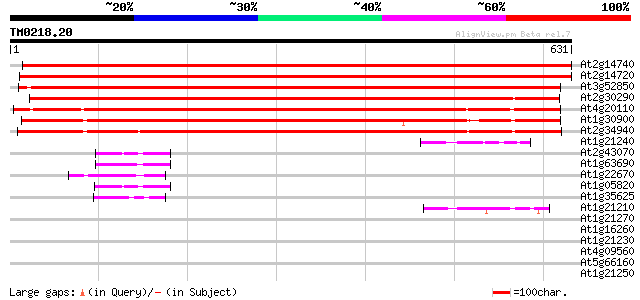
At2g14740 putative vacuolar sorting receptor 1124 0.0
At2g14720 putative vacuolar sorting receptor 1115 0.0
At3g52850 Spot 3 protein and vacuolar sorting receptor homolog/A... 969 0.0
At2g30290 putative vacuolar sorting receptor 917 0.0
At4g20110 vacuolar sorting receptor-like protein 830 0.0
At1g30900 F17F8.23 791 0.0
At2g34940 putative vacuolar sorting receptor 769 0.0
At1g21240 hypothetical protein 52 1e-06
At2g43070 unknown protein 50 3e-06
At1g63690 unknown protein 48 2e-05
At1g22670 putative RING zinc finger protein 48 2e-05
At1g05820 unknown protein 46 7e-05
At1g35625 integral membrane protein, putative 45 9e-05
At1g21210 hypothetical protein 44 3e-04
At1g21270 putative protein 42 0.001
At1g16260 putative wall-associated kinase 42 0.001
At1g21230 hypothetical protein 41 0.002
At4g09560 putative protein 41 0.002
At5g66160 ReMembR-H2 protein JR700 (gb|AAF32325.1) 40 0.004
At1g21250 wall-associated kinase 1 like protein 39 0.011
>At2g14740 putative vacuolar sorting receptor
Length = 628
Score = 1124 bits (2907), Expect = 0.0
Identities = 517/617 (83%), Positives = 569/617 (91%), Gaps = 1/617 (0%)
Query: 15 LLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVYPKDNH 74
LL L V+P + ARFVVEKNSL+VTSPE IKGTHDSAIGNFGIPQYGGSMAG VVYPK+N
Sbjct: 13 LLTLLVSPLNDARFVVEKNSLSVTSPESIKGTHDSAIGNFGIPQYGGSMAGTVVYPKENQ 72
Query: 75 KGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLIT 134
K CKEF + ISFKS+PGALPT +L+DRG+CFFALKVWNAQKAGASAVLVAD+++E LIT
Sbjct: 73 KSCKEFSDFSISFKSQPGALPTFLLVDRGDCFFALKVWNAQKAGASAVLVADNVDEPLIT 132
Query: 135 MDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWREAVPHPDDRVE 194
MDTPEED SSAKYIENITIPSAL+ K FGEKLKK+ISGG+MVN+NLDWREAVPHPDDRVE
Sbjct: 133 MDTPEEDVSSAKYIENITIPSALVTKGFGEKLKKAISGGDMVNLNLDWREAVPHPDDRVE 192
Query: 195 YELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCPKAFTLSKQCK 254
YELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGG+TQF PHYITWYCP AFTLS+QCK
Sbjct: 193 YELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGGFTQFRPHYITWYCPHAFTLSRQCK 252
Query: 255 SQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLWWDYVTDFQIR 314
SQCIN GRYCAPDPEQDFS+GYDGKDVV+ENLRQLCVYKV +E KPW+WWDYVTDFQIR
Sbjct: 253 SQCINKGRYCAPDPEQDFSSGYDGKDVVVENLRQLCVYKVANETGKPWVWWDYVTDFQIR 312
Query: 315 CPMKEKKYNKKCADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGSRGDVT 374
CPMKEKKYNK+CAD+VI+SLG+D+KK+++CMGDPDAD DNPVLKEEQDAQVGKGSRGDVT
Sbjct: 313 CPMKEKKYNKECADSVIKSLGIDSKKLDKCMGDPDADLDNPVLKEEQDAQVGKGSRGDVT 372
Query: 375 ILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKA 434
ILPTLVVNNRQYRGKLEK AV+KA+CSGFEETTEPA+CLS+DVE+NECL+NNGGCW+DK+
Sbjct: 373 ILPTLVVNNRQYRGKLEKSAVLKALCSGFEETTEPAICLSTDVESNECLDNNGGCWQDKS 432
Query: 435 ANITACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNGHAFSA 494
ANITACKDTFRGRVCECP VDGVQFKGDGY+ CE SGPGRC I NGGCWHE R+GHAFSA
Sbjct: 433 ANITACKDTFRGRVCECPTVDGVQFKGDGYSHCEPSGPGRCTINNGGCWHEERDGHAFSA 492
Query: 495 CVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYI 554
CVD VKC+CP GFKGDG K C+D++ECKEKKACQCPECSCKNTWGSY+CSCSGDLLYI
Sbjct: 493 CVDKDSVKCECPPGFKGDGTKKCEDINECKEKKACQCPECSCKNTWGSYECSCSGDLLYI 552
Query: 555 RDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAIMAQYM 614
RDHDTCISKT +Q RSAWAA W+I+ L L+A G YL+YKYR+R YMDSEIRAIMAQYM
Sbjct: 553 RDHDTCISKTGAQ-VRSAWAAVWLIMLSLGLAAAGAYLVYKYRLRQYMDSEIRAIMAQYM 611
Query: 615 PLDSQAEVVNHVNDERA 631
PLDSQ E+ NHVNDERA
Sbjct: 612 PLDSQPEIPNHVNDERA 628
>At2g14720 putative vacuolar sorting receptor
Length = 628
Score = 1115 bits (2883), Expect = 0.0
Identities = 514/622 (82%), Positives = 572/622 (91%), Gaps = 3/622 (0%)
Query: 12 LGFLLLLS--VTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVY 69
L +LLLLS V+P + ARFVVEKNSL+VTSPE IKGTHDSAIGNFGIPQYGGSMAG VVY
Sbjct: 8 LPWLLLLSLVVSPFNEARFVVEKNSLSVTSPESIKGTHDSAIGNFGIPQYGGSMAGTVVY 67
Query: 70 PKDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVADDIE 129
PK+N K CKEF + ISFKS+PGALPT +L+DRG+CFFALKVWNAQKAGASAVLVAD+++
Sbjct: 68 PKENQKSCKEFSDFSISFKSQPGALPTFLLVDRGDCFFALKVWNAQKAGASAVLVADNVD 127
Query: 130 EKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWREAVPHP 189
E LITMDTPEED SSAKYIENITIPSAL+ K FGEKLKK+ISGG+MVN+NLDWREAVPHP
Sbjct: 128 EPLITMDTPEEDVSSAKYIENITIPSALVTKGFGEKLKKAISGGDMVNLNLDWREAVPHP 187
Query: 190 DDRVEYELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCPKAFTL 249
DDRVEYELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGG+TQF PHYITWYCP AFTL
Sbjct: 188 DDRVEYELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGGFTQFRPHYITWYCPHAFTL 247
Query: 250 SKQCKSQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLWWDYVT 309
S+QCKSQCIN GRYCAPDPEQDFS+GYDGKDVV+ENLRQLCVYKV +E KPW+WWDYVT
Sbjct: 248 SRQCKSQCINKGRYCAPDPEQDFSSGYDGKDVVVENLRQLCVYKVANETGKPWVWWDYVT 307
Query: 310 DFQIRCPMKEKKYNKKCADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGS 369
DFQIRCPMKEKKYNK CA++VI+SLG+D++KI++CMGDPDAD DNPVLKEEQDAQVGKG+
Sbjct: 308 DFQIRCPMKEKKYNKDCAESVIKSLGIDSRKIDKCMGDPDADLDNPVLKEEQDAQVGKGT 367
Query: 370 RGDVTILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGC 429
RGDVTILPTLVVNNRQYRGKLEK AV+KA+CSGFEE+TEPA+CLS+D+ETNECL+NNGGC
Sbjct: 368 RGDVTILPTLVVNNRQYRGKLEKSAVLKALCSGFEESTEPAICLSTDMETNECLDNNGGC 427
Query: 430 WKDKAANITACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNG 489
W+DK+ANITACKDTFRG+VC CP+VDGV+FKGDGY+ CE SGPGRC I NGGCWHE R+G
Sbjct: 428 WQDKSANITACKDTFRGKVCVCPIVDGVRFKGDGYSHCEPSGPGRCTINNGGCWHEERDG 487
Query: 490 HAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSG 549
HAFSACVD VKC+CP GFKGDGVK C+D++ECKEKKACQCPECSCKNTWGSY+CSCSG
Sbjct: 488 HAFSACVDKDSVKCECPPGFKGDGVKKCEDINECKEKKACQCPECSCKNTWGSYECSCSG 547
Query: 550 DLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAI 609
DLLY+RDHDTCISKT SQ +SAWAA W+I+ L L+A G YL+YKYR+R YMDSEIRAI
Sbjct: 548 DLLYMRDHDTCISKTGSQ-VKSAWAAVWLIMLSLGLAAAGAYLVYKYRLRQYMDSEIRAI 606
Query: 610 MAQYMPLDSQAEVVNHVNDERA 631
MAQYMPLDSQ EV NH NDERA
Sbjct: 607 MAQYMPLDSQPEVPNHTNDERA 628
>At3g52850 Spot 3 protein and vacuolar sorting receptor
homolog/AtELP1
Length = 623
Score = 969 bits (2506), Expect = 0.0
Identities = 432/610 (70%), Positives = 517/610 (83%), Gaps = 3/610 (0%)
Query: 10 FLLGFLLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVY 69
F L FLL+L++ +M RFVVEKN+L VTSP+ IKG ++ AIGNFG+PQYGG++ G VVY
Sbjct: 6 FTLSFLLILNL---AMGRFVVEKNNLKVTSPDSIKGIYECAIGNFGVPQYGGTLVGTVVY 62
Query: 70 PKDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVADDIE 129
PK N K CK + + ISFKSKPG LPT VL+DRG+C+F LK W AQ+AGA+A+LVAD
Sbjct: 63 PKSNQKACKSYSDFDISFKSKPGRLPTFVLIDRGDCYFTLKAWIAQQAGAAAILVADSKA 122
Query: 130 EKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWREAVPHP 189
E LITMDTPEED S A Y++NITIPSALI K+ G+ +K ++SGG+MVN+ LDW E+VPHP
Sbjct: 123 EPLITMDTPEEDKSDADYLQNITIPSALITKTLGDSIKSALSGGDMVNMKLDWTESVPHP 182
Query: 190 DDRVEYELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCPKAFTL 249
D+RVEYELWTNSNDECG KCD +EF+K+FKGAAQILEKGG+TQFTPHYITWYCP+AFTL
Sbjct: 183 DERVEYELWTNSNDECGKKCDTQIEFLKNFKGAAQILEKGGHTQFTPHYITWYCPEAFTL 242
Query: 250 SKQCKSQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLWWDYVT 309
SKQCKSQCINHGRYCAPDPEQDF+ GYDGKDVV++NLRQ CVY+V ++ KPW+WWDYVT
Sbjct: 243 SKQCKSQCINHGRYCAPDPEQDFTKGYDGKDVVVQNLRQACVYRVMNDTGKPWVWWDYVT 302
Query: 310 DFQIRCPMKEKKYNKKCADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGS 369
DF IRCPMKEKKY K+CAD +I+SLG+D KK+++C+GDP+AD +NPVLK EQ++Q+GKGS
Sbjct: 303 DFAIRCPMKEKKYTKECADGIIKSLGIDLKKVDKCIGDPEADVENPVLKAEQESQIGKGS 362
Query: 370 RGDVTILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGC 429
RGDVTILPTLVVNNRQYRGKLEKGAV+KA+CSGF+E+TEPA+CL+ D+ETNECLENNGGC
Sbjct: 363 RGDVTILPTLVVNNRQYRGKLEKGAVLKAMCSGFQESTEPAICLTEDLETNECLENNGGC 422
Query: 430 WKDKAANITACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNG 489
W+DKAANITAC+DTFRGR+CECP V GV+F GDGYT C+ASG C I NGGCW E+R G
Sbjct: 423 WQDKAANITACRDTFRGRLCECPTVQGVKFVGDGYTHCKASGALHCGINNGGCWRESRGG 482
Query: 490 HAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSG 549
+SACVDD C+CP GFKGDGVKNC+DVDECKEK CQCPEC CKNTWGSY+CSCS
Sbjct: 483 FTYSACVDDHSKDCKCPLGFKGDGVKNCEDVDECKEKTVCQCPECKCKNTWGSYECSCSN 542
Query: 550 DLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAI 609
LLY+R+HDTCI + +W+ W++I G+ ++ GY +YKYRIRSYMD+EIR I
Sbjct: 543 GLLYMREHDTCIGSGKVGTTKLSWSFLWILIIGVGVAGLSGYAVYKYRIRSYMDAEIRGI 602
Query: 610 MAQYMPLDSQ 619
MAQYMPL+SQ
Sbjct: 603 MAQYMPLESQ 612
>At2g30290 putative vacuolar sorting receptor
Length = 625
Score = 917 bits (2370), Expect = 0.0
Identities = 409/596 (68%), Positives = 498/596 (82%), Gaps = 2/596 (0%)
Query: 23 SSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVYPKDNHKGCKEFDE 82
S RFVVEKN+L VTSPE I+G ++ A+GNFG+PQYGGSM+G VVYPK N K CK FD+
Sbjct: 20 SCTGRFVVEKNNLRVTSPESIRGVYECALGNFGVPQYGGSMSGAVVYPKTNQKACKNFDD 79
Query: 83 SGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLITMDTPEEDG 142
ISF+S+ LPT VL+DRG+C+F LK WNAQ+AGA+ +LVAD+ E+LITMD PE++
Sbjct: 80 FEISFRSRVAGLPTFVLVDRGDCYFTLKAWNAQRAGAATILVADNRPEQLITMDAPEDET 139
Query: 143 SSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWREAVPHPDDRVEYELWTNSN 202
S A Y++NITIPSAL+ +S G +K +I+ G+ V+++LDWREA+PHP+DRV YELWTNSN
Sbjct: 140 SDADYLQNITIPSALVSRSLGSAIKTAIAHGDPVHISLDWREALPHPNDRVAYELWTNSN 199
Query: 203 DECGVKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCPKAFTLSKQCKSQCINHGR 262
DECG KCD + F+K FKGAAQILEKGGYT+FTPHYITWYCP+AF S+QCK+QCIN GR
Sbjct: 200 DECGSKCDAQIRFLKRFKGAAQILEKGGYTRFTPHYITWYCPEAFLASRQCKTQCINGGR 259
Query: 263 YCAPDPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLWWDYVTDFQIRCPMKEKKY 322
YCAPDPEQDFS GY+GKDV+I+NLRQ C ++VT+E+ KPWLWWDYVTDF IRCPMKE+KY
Sbjct: 260 YCAPDPEQDFSRGYNGKDVIIQNLRQACFFRVTNESGKPWLWWDYVTDFAIRCPMKEEKY 319
Query: 323 NKKCADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGSRGDVTILPTLVVN 382
NKKCAD VI+SLG+D KKI++C+GD DA+++NPVLKEEQ AQVGKGSRGDVTILPT+V+N
Sbjct: 320 NKKCADQVIQSLGVDVKKIDKCIGDIDANAENPVLKEEQVAQVGKGSRGDVTILPTIVIN 379
Query: 383 NRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKAANITACKD 442
NRQYRGKL++ AV+KA+CSGF ETTEP +CL+ D+ETNECL+NNGGCW+DK NITAC+D
Sbjct: 380 NRQYRGKLQRSAVLKALCSGFRETTEPPICLTEDIETNECLQNNGGCWEDKTTNITACRD 439
Query: 443 TFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNGHAFSACVDDGEVK 502
TFRGRVC+CP+V GV+F GDGYT CEASG RC I NGGCW + + G +SAC DD
Sbjct: 440 TFRGRVCQCPIVQGVKFLGDGYTHCEASGALRCGINNGGCWKQTQMGKTYSACRDDHSKG 499
Query: 503 CQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCIS 562
C+CP GF GDG+K C+DV+EC+EK ACQC +C CKNTWGSY+CSCSG LLYIR+HD CI+
Sbjct: 500 CKCPPGFIGDGLKECKDVNECEEKTACQCRDCKCKNTWGSYECSCSGSLLYIREHDICIN 559
Query: 563 KTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAIMAQYMPLDS 618
+ A G +W W+II GL +A G Y +YKYRIR+YMDSEIRAIMAQYMPLD+
Sbjct: 560 RDAR--GDFSWGVIWIIIMGLGAAALGAYTVYKYRIRTYMDSEIRAIMAQYMPLDN 613
>At4g20110 vacuolar sorting receptor-like protein
Length = 625
Score = 830 bits (2144), Expect = 0.0
Identities = 377/615 (61%), Positives = 480/615 (77%), Gaps = 10/615 (1%)
Query: 5 RFSLGFLLGFLLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMA 64
R SL FLL L ++++ ARFVVEK S++V +PE+++ HD +I NFG+P YGG +
Sbjct: 7 RASLTFLLAALTIIAMVVE--ARFVVEKESISVLNPEEMRSKHDGSIANFGLPDYGGFLI 64
Query: 65 GNVVYPKDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLV 124
G+VVYP GC F G +FK K PTI+LLDRG C+FALK W+AQ+AGA+AVLV
Sbjct: 65 GSVVYPDSKTDGCSAF---GKTFKPK-FPRPTILLLDRGGCYFALKAWHAQQAGAAAVLV 120
Query: 125 ADDIEEKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWRE 184
AD+++E L+TMD+PEE + +IE +TIPS LI+KSFG+ L++ G+ + + LDWRE
Sbjct: 121 ADNVDEPLLTMDSPEESKDADGFIEKLTIPSVLIDKSFGDDLRQGFQKGKNIVIKLDWRE 180
Query: 185 AVPHPDDRVEYELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCP 244
+VPHPD RVEYELWTNSNDECG +CD M+FVK+FKG AQILEKGGYT FTPHYITW+CP
Sbjct: 181 SVPHPDKRVEYELWTNSNDECGARCDEQMDFVKNFKGHAQILEKGGYTAFTPHYITWFCP 240
Query: 245 KAFTLSKQCKSQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLW 304
F S CKSQCINHGRYCAPDPE +F GY+GKDVV+ENLRQLCV++V +E+ +PW+W
Sbjct: 241 FQFINSPHCKSQCINHGRYCAPDPEDNFREGYEGKDVVLENLRQLCVHRVANESSRPWVW 300
Query: 305 WDYVTDFQIRCPMKEKKYNKKCADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQ 364
WDYVTDF RC MKEKKY+ CA++VI+SL L +KI++C+GDP+AD++N VL+ EQ +Q
Sbjct: 301 WDYVTDFHSRCSMKEKKYSIDCAESVIKSLNLPIEKIKKCIGDPEADTENQVLRTEQVSQ 360
Query: 365 VGKGSRGDVTILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLE 424
+G+G+RGDVTILPTLV+NN QYRG+LE+ AV+KAIC+GF ET+EPA+CL++ +ETNECLE
Sbjct: 361 IGRGNRGDVTILPTLVINNAQYRGRLERTAVLKAICAGFNETSEPAICLNTGLETNECLE 420
Query: 425 NNGGCWKDKAANITACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWH 484
NNGGCW+D ANITAC+DTFRGR+CECP+V GVQ+KGDGYT+C GP RC + NGGCW
Sbjct: 421 NNGGCWQDTKANITACQDTFRGRLCECPVVKGVQYKGDGYTSCTPYGPARCTMNNGGCWS 480
Query: 485 EARNGHAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYD 544
+ RNG FSAC D C+CP GF+GDG+ C+D++ECKE+ CQC C CKN+WG Y
Sbjct: 481 DTRNGLTFSACSDSVSTGCKCPEGFQGDGL-TCEDINECKERSVCQCSGCRCKNSWGGYK 539
Query: 545 CSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDS 604
CSCSGD LYI D DTCI + G ++AW ++I+A + ++ GY+ YKYR RSYMDS
Sbjct: 540 CSCSGDRLYINDQDTCIER---YGSKTAWWLTFLILAIVAVAGLAGYIFYKYRFRSYMDS 596
Query: 605 EIRAIMAQYMPLDSQ 619
EI IM+QYMPL+SQ
Sbjct: 597 EIMTIMSQYMPLESQ 611
>At1g30900 F17F8.23
Length = 649
Score = 791 bits (2044), Expect = 0.0
Identities = 371/633 (58%), Positives = 460/633 (72%), Gaps = 44/633 (6%)
Query: 14 FLLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVYPKDN 73
FL L V RF+VEK+S+T+ +P ++ HD+AI NFG+P YGG M G+VVY
Sbjct: 13 FLALTMVVNGVFGRFIVEKSSVTILNPLAMRSKHDAAIANFGVPNYGGYMIGSVVYAGQG 72
Query: 74 HKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLI 133
GC FD++ FK K PTI+++DRG C+FALKVWN Q++G +AVLVAD+++E LI
Sbjct: 73 AYGCDSFDKT---FKPK-FPRPTILIIDRGECYFALKVWNGQQSGVAAVLVADNVDEPLI 128
Query: 134 TMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWREAVPHPDDRV 193
TMD+PEE + +IE + IPSALI+ SF LK+++ GE V + +DW E++PHPD+RV
Sbjct: 129 TMDSPEESKEADDFIEKLNIPSALIDFSFANTLKQALKKGEEVVLKIDWSESLPHPDERV 188
Query: 194 EYELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCPKAFTLSKQC 253
EYELWTN+NDECG +CD M FVK+FKG AQILEKGGY+ FTPHYITW+CPK + S QC
Sbjct: 189 EYELWTNTNDECGARCDEQMNFVKNFKGHAQILEKGGYSLFTPHYITWFCPKDYVSSNQC 248
Query: 254 KSQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLWWDYVTDFQI 313
KSQCIN GRYCAPDPEQDF GYDGKD+V ENLRQLCV+KV E + W+WWDYVTDF I
Sbjct: 249 KSQCINQGRYCAPDPEQDFGDGYDGKDIVFENLRQLCVHKVAKENNRSWVWWDYVTDFHI 308
Query: 314 RCPMKEKKYNKKCADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGSRGDV 373
RC MKEKKY+K+CA+ V+ESLGL KI++C+GDPDA+ +N VLK EQ QVG+G RGDV
Sbjct: 309 RCSMKEKKYSKECAERVVESLGLPLDKIKKCIGDPDANVENEVLKAEQALQVGQGDRGDV 368
Query: 374 TILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDK 433
TILPTL+VNN QYRGKLE+ AV+KAICSGF+E TEP +CLS D+ETNECLE NGGCW+DK
Sbjct: 369 TILPTLIVNNAQYRGKLERNAVLKAICSGFKERTEPGICLSGDIETNECLEANGGCWEDK 428
Query: 434 AANITACK---------------------------DTFRGRVCECPLVDGVQFKGDGYTT 466
+N+TACK DTFRGRVCECP+V+GVQ+KGDGYT+
Sbjct: 429 KSNVTACKVLRTDELKGLHFYRYLVSFIPKNGFYQDTFRGRVCECPVVNGVQYKGDGYTS 488
Query: 467 CEASGPGRCKIKNGGCWHEARNGHAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEK 526
CE GP RC I GGCW E + G FSAC + C+CP GFKGDG+K C+
Sbjct: 489 CEPYGPARCSINQGGCWSETKKGLTFSACSNLETSGCRCPPGFKGDGLK-CE-------- 539
Query: 527 KACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLS 586
ACQC C+CKN WG ++C CSG+ LY+++ DTCI ++ G R W +VI+A +
Sbjct: 540 -ACQCDGCNCKNKWGGFECKCSGNRLYMKEQDTCIERS---GSRIGWFPTFVILAAVASI 595
Query: 587 AGGGYLIYKYRIRSYMDSEIRAIMAQYMPLDSQ 619
GGY+ YKYR+RSYMDSEI AIM+QYMPL+SQ
Sbjct: 596 CVGGYVFYKYRLRSYMDSEIMAIMSQYMPLESQ 628
>At2g34940 putative vacuolar sorting receptor
Length = 618
Score = 770 bits (1987), Expect = 0.0
Identities = 353/614 (57%), Positives = 464/614 (75%), Gaps = 11/614 (1%)
Query: 9 GFLLGFLLLLS--VTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGN 66
G +L +L L+ V +RF VEK+SLTV + ++ HD+AI NFG+P+YGG M G+
Sbjct: 7 GTVLALILALTMVVVNGFSSRFFVEKSSLTVLNSWEMGAKHDAAIANFGLPKYGGFMIGS 66
Query: 67 VVYPKDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVAD 126
VVY + GC F+++ F +K P I+L+DRG C FALK+WN Q++GA+AVL+AD
Sbjct: 67 VVYAGQDAYGCNSFNKT---FNTK-SPYPKILLIDRGVCNFALKIWNGQQSGAAAVLLAD 122
Query: 127 DIEEKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWREAV 186
+I E LITMDTP+++ +I+ + IPSALI +SFG+ LKK++ GE V + +DW E++
Sbjct: 123 NIVEPLITMDTPQDEDPD--FIDKVKIPSALILRSFGDSLKKALKRGEEVILKMDWSESI 180
Query: 187 PHPDDRVEYELWTNSNDECGVKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCPKA 246
P+PD+RVEYELW N+NDECGV CD ++F+K+FKG AQILEKGGYT F PHYI+W CPK
Sbjct: 181 PNPDERVEYELWANTNDECGVHCDKQIDFIKNFKGMAQILEKGGYTLFRPHYISWVCPKE 240
Query: 247 FTLSKQCKSQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLWWD 306
LSKQC++QCIN GRYCA D +Q+F GY+GKDVV ENLRQLCV+KV E W+WWD
Sbjct: 241 LLLSKQCRTQCINQGRYCALDTKQEFEDGYNGKDVVYENLRQLCVHKVAKEKNTSWVWWD 300
Query: 307 YVTDFQIRCPMKEKKYNKKCADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVG 366
YVTDF IRC MKEKKY+++CA+ ++ESLGL +KI++C+GDPDAD +N VLK E+ Q+G
Sbjct: 301 YVTDFNIRCSMKEKKYSRECAETIVESLGLSLEKIKKCIGDPDADVENEVLKAEEAFQLG 360
Query: 367 KGSRGDVTILPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENN 426
+ +RG VTI PTL++NN QYRGKLE+ AV+KAICSGF+E TEP++CL+SD+ETNECL N
Sbjct: 361 QENRGIVTIFPTLMINNAQYRGKLERTAVLKAICSGFKERTEPSICLNSDIETNECLIEN 420
Query: 427 GGCWKDKAANITACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEA 486
GGCW+DK +N+TACKDTFRGRVCECP+VDGVQ+KGDGYT+C+ GP RC + NG CW E
Sbjct: 421 GGCWQDKRSNVTACKDTFRGRVCECPVVDGVQYKGDGYTSCKPYGPARCSMNNGDCWSET 480
Query: 487 RNGHAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCS 546
R G FS+C D C+CP GF GDG+K C+D+DECKEK AC+C C CKN WG Y+C
Sbjct: 481 RKGLTFSSCSDSETSGCRCPLGFLGDGLK-CEDIDECKEKSACKCDGCKCKNNWGGYECK 539
Query: 547 CSGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEI 606
CS + +Y+++ DTCI + + G RS V++ + + G Y+ YKY ++SYMDSEI
Sbjct: 540 CSNNSIYMKEEDTCIERRS--GSRSRGLFTIVVLTAIAGISLGAYIFYKYHLQSYMDSEI 597
Query: 607 RAIMAQYMPLDSQA 620
+IM+QY+PLDSQ+
Sbjct: 598 VSIMSQYIPLDSQS 611
>At1g21240 hypothetical protein
Length = 741
Score = 52.0 bits (123), Expect = 1e-06
Identities = 38/127 (29%), Positives = 62/127 (47%), Gaps = 22/127 (17%)
Query: 463 GYTTCEASGPGRCKIKNGGCWHE-ARNGHAFSACVDDGEVKCQCPAGFKGDGVKN--CQD 519
G TCE +G R KN C++ RNG+ C+C G+ G+ ++ C+D
Sbjct: 245 GNQTCEQAGSTRICGKNSSCYNSTTRNGYI-----------CKCNEGYDGNPYRSEGCKD 293
Query: 520 VDEC-KEKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWV 578
+DEC + C P+ +C+N G +DC C D ++ +S T + R+ F V
Sbjct: 294 IDECISDTHNCSDPK-TCRNRDGGFDCKCPSGY----DLNSSMSCTRPEYKRT--RIFLV 346
Query: 579 IIAGLVL 585
II G+++
Sbjct: 347 IIIGVLV 353
>At2g43070 unknown protein
Length = 543
Score = 50.4 bits (119), Expect = 3e-06
Identities = 35/84 (41%), Positives = 48/84 (56%), Gaps = 6/84 (7%)
Query: 97 IVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLITMDTPEEDGSSAKYIENITIPSA 156
I L RGNC F K +A+ AGASA+LV +D +E L M E+D S N++IP
Sbjct: 108 IALSIRGNCAFTEKAKHAEAAGASALLVIND-KEDLDEMGCMEKDTSL-----NVSIPVL 161
Query: 157 LIEKSFGEKLKKSISGGEMVNVNL 180
+I KS G+ L KS+ + V + L
Sbjct: 162 MISKSSGDALNKSMVDNKNVELLL 185
>At1g63690 unknown protein
Length = 540
Score = 47.8 bits (112), Expect = 2e-05
Identities = 26/84 (30%), Positives = 47/84 (55%), Gaps = 6/84 (7%)
Query: 97 IVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLITMDTPEEDGSSAKYIENITIPSA 156
+V+++RGNC F K NA+ AGASA+L+ ++ +E + P+E +I IP+
Sbjct: 107 VVIVERGNCRFTAKANNAEAAGASALLIINNQKELYKMVCEPDETDL------DIQIPAV 160
Query: 157 LIEKSFGEKLKKSISGGEMVNVNL 180
++ + G L+K ++ V+ L
Sbjct: 161 MLPQDAGASLQKMLANSSKVSAQL 184
>At1g22670 putative RING zinc finger protein
Length = 422
Score = 47.8 bits (112), Expect = 2e-05
Identities = 32/109 (29%), Positives = 48/109 (43%), Gaps = 11/109 (10%)
Query: 67 VVYPKDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVAD 126
VVY + C+ + P +VL+ RG C F KV NAQ++G A +V D
Sbjct: 54 VVYVAEPLNACRNLRNKP---EQSPYGTSPLVLIIRGGCSFEYKVRNAQRSGFKAAIVYD 110
Query: 127 DIEEKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEM 175
+++ ++ + DG I I + + K GE LKK EM
Sbjct: 111 NVDRNFLSAMGGDSDG--------IKIQAVFVMKRAGEMLKKYAGSEEM 151
>At1g05820 unknown protein
Length = 441
Score = 45.8 bits (107), Expect = 7e-05
Identities = 32/85 (37%), Positives = 48/85 (55%), Gaps = 6/85 (7%)
Query: 96 TIVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLITMDTPEEDGSSAKYIENITIPS 155
+I L RG C F +K AQ GA+A+++ +D +E+L M E+D S N++IP
Sbjct: 103 SIALSVRGECAFTVKAQVAQAGGAAALVLIND-KEELDEMVCGEKDTSL-----NVSIPI 156
Query: 156 ALIEKSFGEKLKKSISGGEMVNVNL 180
+I S G+ LKKSI + V + L
Sbjct: 157 LMITTSSGDALKKSIMQNKKVELLL 181
>At1g35625 integral membrane protein, putative
Length = 279
Score = 45.4 bits (106), Expect = 9e-05
Identities = 30/81 (37%), Positives = 43/81 (53%), Gaps = 10/81 (12%)
Query: 95 PTIVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLITMDTPEEDGSSAKYIENITIP 154
P+ VL+ RG C F K+ NAQ+AG A +V +D E+L+ SS YI +
Sbjct: 40 PSYVLIVRGGCSFEEKIRNAQEAGYKAAIVYNDRYEELLV-----RRNSSGVYIHGV--- 91
Query: 155 SALIEKSFGEKLKKSISGGEM 175
L+ ++ GE LK+ S EM
Sbjct: 92 --LVTRTSGEVLKEYTSRAEM 110
>At1g21210 hypothetical protein
Length = 738
Score = 43.9 bits (102), Expect = 3e-04
Identities = 37/155 (23%), Positives = 66/155 (41%), Gaps = 33/155 (21%)
Query: 466 TCEASGPGRCKIKNGGCWHEARNGHAFSACVDDGEVKCQCPAGFKGDGV--KNCQDVDEC 523
TC G +C + NG C + A +G ++ C+C GF+G+ CQD++EC
Sbjct: 235 TCGQVGEKKCGV-NGICSNSA-SGIGYT---------CKCKGGFQGNPYLQNGCQDINEC 283
Query: 524 KEKKACQCPECS----CKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWVI 579
CS C+N G + C+C +TC K G + + I
Sbjct: 284 TTANPIHKHNCSGDSTCENKLGHFRCNCRSRYELNTTTNTCKPK-----GNPEYVEWTTI 338
Query: 580 IAGLVLSAGGGYLI-------YKYRIRSYMDSEIR 607
+ G + G+L+ ++++++ D+E+R
Sbjct: 339 VLGTTI----GFLVILLAISCIEHKMKNTKDTELR 369
>At1g21270 putative protein
Length = 732
Score = 42.0 bits (97), Expect = 0.001
Identities = 17/47 (36%), Positives = 28/47 (59%), Gaps = 2/47 (4%)
Query: 503 CQCPAGFKGDGV--KNCQDVDECKEKKACQCPECSCKNTWGSYDCSC 547
C+C GF+G+ CQD++EC + +C+NT GS++C+C
Sbjct: 260 CKCLEGFEGNPYLPNGCQDINECISSRHNCSEHSTCENTKGSFNCNC 306
Score = 33.5 bits (75), Expect = 0.37
Identities = 22/81 (27%), Positives = 35/81 (43%), Gaps = 10/81 (12%)
Query: 438 TACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNGHAFSACVD 497
+ C D+ G C ++G F+G+ Y P C+ N C N S C +
Sbjct: 247 STCFDSTGGTGYNCKCLEG--FEGNPYL------PNGCQDINE-CISSRHNCSEHSTCEN 297
Query: 498 D-GEVKCQCPAGFKGDGVKNC 517
G C CP+G++ D + +C
Sbjct: 298 TKGSFNCNCPSGYRKDSLNSC 318
>At1g16260 putative wall-associated kinase
Length = 720
Score = 41.6 bits (96), Expect = 0.001
Identities = 17/46 (36%), Positives = 26/46 (55%), Gaps = 2/46 (4%)
Query: 502 KCQCPAGFKGDGV--KNCQDVDECKEKKACQCPECSCKNTWGSYDC 545
+C C G++G+ CQD+DEC++ +C + C N GSY C
Sbjct: 265 QCSCHNGYEGNPYIPGGCQDIDECRDPHLNKCGKRKCVNVLGSYRC 310
>At1g21230 hypothetical protein
Length = 733
Score = 41.2 bits (95), Expect = 0.002
Identities = 32/120 (26%), Positives = 49/120 (40%), Gaps = 13/120 (10%)
Query: 493 SACVDDGEVK---CQCPAGFKGDGVKN--CQDVDECKEKKACQCPECSCKNTWGSYDCSC 547
S C D K C+C GF G+ + CQD++EC + +C+NT GS+ C C
Sbjct: 248 STCFDSTRGKGYNCKCLQGFDGNPYLSDGCQDINECTTRIHNCSDTSTCENTLGSFHCQC 307
Query: 548 SGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIR 607
+CI + W ++L G+LI I SY+ ++R
Sbjct: 308 PSGSDLNTTTMSCIDTPKEEPKYLGWTT-------VLLGTTIGFLIILLTI-SYIQQKMR 359
>At4g09560 putative protein
Length = 431
Score = 40.8 bits (94), Expect = 0.002
Identities = 33/112 (29%), Positives = 50/112 (44%), Gaps = 12/112 (10%)
Query: 15 LLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVYPKDNH 74
LLL+S S+ + SL+ +D++ T P S G V+Y +
Sbjct: 11 LLLISHLVSAKVLLIGNSTSLSF---DDVEATFT--------PMIKRSDQGGVLYVAEPL 59
Query: 75 KGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVAD 126
C + + ++ K+ P VL+ RG C F K+ NAQKAG A +V D
Sbjct: 60 DACSDLVNT-VNVKNGTTVSPPYVLIIRGGCSFEDKIRNAQKAGYKAAIVYD 110
>At5g66160 ReMembR-H2 protein JR700 (gb|AAF32325.1)
Length = 310
Score = 40.0 bits (92), Expect = 0.004
Identities = 27/75 (36%), Positives = 38/75 (50%), Gaps = 10/75 (13%)
Query: 99 LLDRGNCFFALKVWNAQKAGASAVLVADDIE-EKLITMDTPEEDGSSAKYIENITIPSAL 157
L+ RG C F K+ NAQ +G AV+V D+I+ E LI M +D IT+ +
Sbjct: 86 LIIRGECSFEDKLLNAQNSGFQAVIVYDNIDNEDLIVMKVNPQD---------ITVDAVF 136
Query: 158 IEKSFGEKLKKSISG 172
+ GE L+K G
Sbjct: 137 VSNVAGEILRKYARG 151
>At1g21250 wall-associated kinase 1 like protein
Length = 735
Score = 38.5 bits (88), Expect = 0.011
Identities = 28/92 (30%), Positives = 35/92 (37%), Gaps = 18/92 (19%)
Query: 463 GYTTCEASGPGRCKIKNGGCWHEA-RNGHAFSACVDDGEVKCQCPAGFKGDGVKN--CQD 519
G TCE G N C RNG+ C+C GF G+ + CQD
Sbjct: 234 GNQTCEQVGSTSICGGNSTCLDSTPRNGYI-----------CRCNEGFDGNPYLSAGCQD 282
Query: 520 VDECKEKKACQCPECS----CKNTWGSYDCSC 547
V+EC CS C+N G + C C
Sbjct: 283 VNECTTSSTIHRHNCSDPKTCRNKVGGFYCKC 314
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,030,623
Number of Sequences: 26719
Number of extensions: 771759
Number of successful extensions: 1748
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1679
Number of HSP's gapped (non-prelim): 50
length of query: 631
length of database: 11,318,596
effective HSP length: 105
effective length of query: 526
effective length of database: 8,513,101
effective search space: 4477891126
effective search space used: 4477891126
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0218.20


