

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0214.6
(398 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
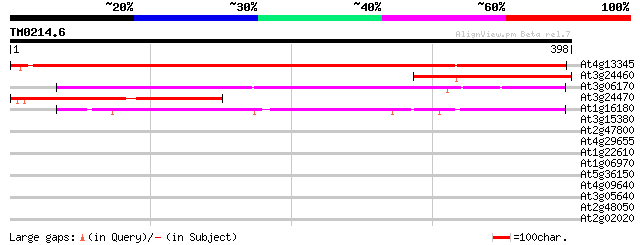
At4g13345 unknown protein 615 e-176
At3g24460 unknown protein 201 4e-52
At3g06170 unknown protein 199 2e-51
At3g24470 hypothetical protein 198 5e-51
At1g16180 194 9e-50
At3g15380 unknown protein 33 0.36
At2g47800 MRP-like ABC transporter 30 3.0
At4g29655 putative protein 29 4.0
At1g22610 Highly similar to phosphoribosylanthranilate transferase 29 4.0
At1g06970 hypothetical protein 29 5.2
At5g36150 putative protein 28 6.8
At4g09640 putative protein 28 6.8
At3g05640 putative protein phosphatase-2C 28 8.9
At2g48050 hypothetical protein 28 8.9
At2g02020 putative peptide/amino acid transporter 28 8.9
>At4g13345 unknown protein
Length = 394
Score = 615 bits (1586), Expect = e-176
Identities = 286/397 (72%), Positives = 335/397 (84%), Gaps = 6/397 (1%)
Query: 1 METGVS--NNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARDYGRG 58
METG S N G E K+ SW+ QFRN NP MARYVY L+FL+ANLLAWA RDYGRG
Sbjct: 1 METGTSIDNTGYEGI---KNGSWFIQFRNGCNPWMARYVYGLIFLLANLLAWALRDYGRG 57
Query: 59 ALTEMERLKGCNGGKDCLGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWS 118
ALTEM + K C G DCLG EGVLRVS GCF+FYFIMFL+T TSK + RD WHSGWW
Sbjct: 58 ALTEMRKFKNCKEGGDCLGTEGVLRVSFGCFLFYFIMFLSTVGTSKTHSSRDKWHSGWWF 117
Query: 119 VKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCESEKYA 178
K+ + +G+T+ PFLLPS IQ YGE+AHFGAGVFLLIQLISIISFITWLN+C +++K A
Sbjct: 118 AKLFMLLGLTIFPFLLPSSIIQFYGEIAHFGAGVFLLIQLISIISFITWLNECFQAQKDA 177
Query: 179 AKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPK 238
+C +HVML ATTAY VC++G+ILM+IWY P+PSCLLNIFFI WTL L+QLMTS+SLHPK
Sbjct: 178 ERCHVHVMLLATTAYTVCILGVILMYIWYVPEPSCLLNIFFITWTLFLIQLMTSISLHPK 237
Query: 239 VNAGILTPGLMGLYVVFLCWNAIRSEPDGGSCIRKSDSSTKTDWLSIISFIIAILAIVIA 298
+NAG LTP LMGLYVVF+CW AIRSEP G +C RK++ S++TDWL+IISF++A+LA+VIA
Sbjct: 238 INAGFLTPALMGLYVVFICWCAIRSEPVGETCNRKAEGSSRTDWLTIISFVVALLAMVIA 297
Query: 299 TFSTGIDSKCFQFRKDDDIPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWT 358
TFSTG+DS+CFQFRKD++ ED +PYGYGFFHFVFATGAMYFAMLL+GWN HHSM+KWT
Sbjct: 298 TFSTGVDSQCFQFRKDEN-HEEDAIPYGYGFFHFVFATGAMYFAMLLVGWNIHHSMKKWT 356
Query: 359 IDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSRQT 395
IDVGWTSTWVRIVNEWLAV VY+WMLVAP++ KSRQT
Sbjct: 357 IDVGWTSTWVRIVNEWLAVGVYIWMLVAPMVLKSRQT 393
>At3g24460 unknown protein
Length = 126
Score = 201 bits (512), Expect = 4e-52
Identities = 91/115 (79%), Positives = 105/115 (91%), Gaps = 3/115 (2%)
Query: 287 SFIIAILAIVIATFSTGIDSKCFQFRKDD---DIPAEDDVPYGYGFFHFVFATGAMYFAM 343
SF++A+LA+VIATFSTGIDS+CFQF+KD+ + AEDDVPYGYGFFHFVFATGAMYFAM
Sbjct: 10 SFVVALLAMVIATFSTGIDSQCFQFKKDENDQEEEAEDDVPYGYGFFHFVFATGAMYFAM 69
Query: 344 LLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSRQTDST 398
LLIGWN+HH M+KWTIDVGWTSTWVR+VNEWLAVCVY+WMLVAP+I KSR+ T
Sbjct: 70 LLIGWNTHHPMKKWTIDVGWTSTWVRVVNEWLAVCVYIWMLVAPLILKSRRQTPT 124
>At3g06170 unknown protein
Length = 409
Score = 199 bits (506), Expect = 2e-51
Identities = 112/382 (29%), Positives = 188/382 (48%), Gaps = 24/382 (6%)
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNG-GKDCLGAEGVLRVSLGCFIFY 92
AR Y +F + +++W R+ G L ++ + + K+ + VLRVS G F+F+
Sbjct: 29 ARIAYCGLFGASLVVSWILRETGAPLLEKLPWINTSDSYTKEWYQQQAVLRVSFGNFLFF 88
Query: 93 FIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGV 152
I L N+ RD+WH G W +K+++W + V+ F +P+ + +YG ++ FGAG
Sbjct: 89 AIYALIMIGVKDQNDRRDSWHHGGWGLKMIVWFLLVVLMFFVPNVIVSLYGTLSKFGAGA 148
Query: 153 FLLIQLISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPK-P 211
FLL+Q++ ++ ND EK K I +++ + Y+ ++FIW+ P
Sbjct: 149 FLLVQVVLLLDATHNWNDSW-VEKDEKKWYIALLVISIVCYIATYTFSGILFIWFNPSGQ 207
Query: 212 SCLLNIFFIAWTLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEPDGGSCI 271
C LN+FFI ++L + ++LHP VN +L ++ +Y ++C+ + SEP C
Sbjct: 208 DCGLNVFFIVMPMILAFVFAIIALHPAVNGSLLPASVISVYCAYVCYTGLSSEPHDYVCN 267
Query: 272 RKSDSSTKTDWLSIISFIIAILAIVIATFSTGIDSKCF-------------------QFR 312
+ S I+ + +L+++ + G + +
Sbjct: 268 GLNKSKAVNASTLILGMLTTVLSVLYSALRAGSSTTFLSPPSSPRSGVKDALLGDPEDGK 327
Query: 313 KDDDIPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVN 372
K + A V Y Y FFH +FA +MY AMLL GW + S IDVGWTS WV+I
Sbjct: 328 KSGEAEAR-PVSYSYSFFHIIFALASMYAAMLLSGW-TDSSESATLIDVGWTSVWVKICT 385
Query: 373 EWLAVCVYLWMLVAPIIWKSRQ 394
W+ +Y+W L+AP+I R+
Sbjct: 386 GWVTAGLYIWTLIAPLILPDRE 407
>At3g24470 hypothetical protein
Length = 151
Score = 198 bits (503), Expect = 5e-51
Identities = 94/157 (59%), Positives = 116/157 (73%), Gaps = 12/157 (7%)
Query: 1 METG--VSNNG----NEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARD 54
METG VSNN N++ K+ SW+ QFRN NP MARYVY L+FL+ANLLAWAARD
Sbjct: 1 METGTSVSNNNQSIRNDSYEAIKNGSWFNQFRNGCNPWMARYVYGLIFLIANLLAWAARD 60
Query: 55 YGRGALTEMERLKGCNGGKDCLGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHS 114
YGRGAL ++ R K C GG++CLG +GVLR +FYF+MFL+T TSK + RD WHS
Sbjct: 61 YGRGALRKVTRFKNCKGGENCLGTDGVLR------LFYFVMFLSTLGTSKTHSSRDRWHS 114
Query: 115 GWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAG 151
GWW VK+++W +T+IPFLLPS I +YGE+AHFGAG
Sbjct: 115 GWWFVKLIMWPALTIIPFLLPSSIIHLYGEIAHFGAG 151
>At1g16180
Length = 412
Score = 194 bits (492), Expect = 9e-50
Identities = 118/393 (30%), Positives = 191/393 (48%), Gaps = 44/393 (11%)
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNG-----GKDCLGAEGVLRVSLGC 88
AR Y +F ++ +++W R+ A ME+L N ++ + VLRVSLG
Sbjct: 31 ARIAYCGLFALSLIVSWILREV---AAPLMEKLPWINHFHKTPDREWFETDAVLRVSLGN 87
Query: 89 FIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHF 148
F+F+ I+ + + RD H G W +KI+ W + + F LP+E I Y ++ F
Sbjct: 88 FLFFSILSVMMIGVKNQKDPRDGIHHGGWMMKIICWCILVIFMFFLPNEIISFYESMSKF 147
Query: 149 GAGVFLLIQLISIISFITWLNDCC----ESEKYAAKCQIHVMLFATTAYVVCLVGIILMF 204
GAG FLL+Q++ ++ F+ ND E YAA +++ + Y+ V +F
Sbjct: 148 GAGFFLLVQVVLLLDFVHGWNDTWVGYDEQFWYAA-----LLVVSLVCYLATFVFSGFLF 202
Query: 205 IWYAPKP-SCLLNIFFIAWTLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRS 263
W+ P C LN FFI TL+ + + V LHP V IL ++ LY ++LC++ + S
Sbjct: 203 HWFTPSGHDCGLNTFFIIMTLIFVFVFAIVVLHPTVGGSILPASVISLYCMYLCYSGLAS 262
Query: 264 EPDGGSC--IRKSDSSTKTDWLSIISFIIAILAIVIATFSTG------------------ 303
EP C + + T ++I + +L++V + G
Sbjct: 263 EPRDYECNGLHNHSKAVSTGTMTI-GLLTTVLSVVYSAVRAGSSTTLLSPPDSPRAEKPL 321
Query: 304 --IDSKCFQFRKDDDIPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDV 361
ID K + + ++ + V Y Y FFH +F+ +MY AMLL GW++ +DV
Sbjct: 322 LPIDGKAEEKEEKEN---KKPVSYSYAFFHIIFSLASMYSAMLLTGWSTSVGESGKLVDV 378
Query: 362 GWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSRQ 394
GW S WVR+V W +++W LVAPI++ R+
Sbjct: 379 GWPSVWVRVVTSWATAGLFIWSLVAPILFPDRE 411
>At3g15380 unknown protein
Length = 700
Score = 32.7 bits (73), Expect = 0.36
Identities = 25/113 (22%), Positives = 48/113 (42%), Gaps = 16/113 (14%)
Query: 124 WVGMTVIPFLLPSE--FIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCESEKYAAKC 181
W+G + ++ + +YG GV +L+ IS+++ +T +
Sbjct: 294 WIGNDAVTPIIGEHDPYFHVYGRELTHVRGVAILMTFISVVAILTSI------------A 341
Query: 182 QIHVMLFATTAYVVC--LVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTS 232
I +L AT+ V ++G + I + P +L IF++ W L L +S
Sbjct: 342 IIRRILMATSVLKVAAKVIGEVQALIIFPAIPFAMLAIFYMFWISAALHLFSS 394
>At2g47800 MRP-like ABC transporter
Length = 1516
Score = 29.6 bits (65), Expect = 3.0
Identities = 19/56 (33%), Positives = 29/56 (50%), Gaps = 8/56 (14%)
Query: 115 GWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVA----HFGAGVFLL----IQLISII 162
GWW + +VL+ +T L+ S++ Y A F A VF+L I L+SI+
Sbjct: 953 GWWGIVLVLFFSLTWQGSLMASDYWLAYETSAKNAISFDASVFILGYVIIALVSIV 1008
>At4g29655 putative protein
Length = 190
Score = 29.3 bits (64), Expect = 4.0
Identities = 25/96 (26%), Positives = 38/96 (39%), Gaps = 24/96 (25%)
Query: 184 HVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAW--------------------- 222
++M+F T V+ +V I + +W P + I +AW
Sbjct: 62 NIMIFRTNYIVIFIVSIFISMLW-QPVHLSVFVILIVAWLYVYSRDNEPWVIFGSVIDDS 120
Query: 223 --TLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFL 256
LVLL L + L V+ GI+ L GL VV +
Sbjct: 121 TLVLVLLVLTIGIFLLTDVSRGIVIGVLAGLPVVLV 156
>At1g22610 Highly similar to phosphoribosylanthranilate transferase
Length = 1029
Score = 29.3 bits (64), Expect = 4.0
Identities = 11/54 (20%), Positives = 26/54 (47%), Gaps = 4/54 (7%)
Query: 157 QLISIISFIT----WLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIW 206
+++S++S +T W ND C C +HV+ Y ++ + ++++
Sbjct: 829 RIMSLLSSVTLVCKWFNDICTWRNPITTCLVHVLFLILVCYPELILPTVFLYLF 882
>At1g06970 hypothetical protein
Length = 829
Score = 28.9 bits (63), Expect = 5.2
Identities = 35/156 (22%), Positives = 65/156 (41%), Gaps = 11/156 (7%)
Query: 89 FIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIV----LWVGMTV------IPFLLPSEF 138
F ++ LT + TS++ V + + W V + L G+T +LP F
Sbjct: 283 FFPIIMVLLTISLTSEVLGVHAAFGAFWLGVSLPDGPPLGTGLTTKLEMFATSLMLPC-F 341
Query: 139 IQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLV 198
I I G +F +++I + IT+ + +A C I + + A ++C
Sbjct: 342 ISISGLQTNFFIIGESHVKIIEAVILITYGCKFLGTAAASAYCNIQIGDAFSLALLMCCQ 401
Query: 199 GIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVS 234
G+I ++ K +LN ++ L L+T +S
Sbjct: 402 GVIEIYTCVMWKDEKVLNTECFNLLIITLLLVTGIS 437
>At5g36150 putative protein
Length = 729
Score = 28.5 bits (62), Expect = 6.8
Identities = 20/65 (30%), Positives = 26/65 (39%), Gaps = 8/65 (12%)
Query: 332 FVFATGAMYFAMLLIGWNSHHS-------MRKWTIDVGWTSTWVRIVNEWLAVC-VYLWM 383
F+F T Y + + G + H RKW ID G + WL+V VY W
Sbjct: 175 FMFCTVINYICLRIFGVDPDHDGESACARARKWIIDHGGATYTPLFGKAWLSVLGVYEWS 234
Query: 384 LVAPI 388
PI
Sbjct: 235 GCKPI 239
>At4g09640 putative protein
Length = 339
Score = 28.5 bits (62), Expect = 6.8
Identities = 22/101 (21%), Positives = 48/101 (46%), Gaps = 20/101 (19%)
Query: 124 WVGMTVIPFLLPSEFIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCESEKYAAKCQI 183
W+GM + I GE+A+F A F L++ + ++ + C +++K +
Sbjct: 68 WIGMITM----------IVGEIANFAAYAFAPAILVTPLGALSIIIRCEQTQK------L 111
Query: 184 HVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTL 224
H F +C+VG + + + +AP+ ++++ + W L
Sbjct: 112 HT--FGILGCALCIVGSVTI-VLHAPQEQDIVSVLEV-WNL 148
>At3g05640 putative protein phosphatase-2C
Length = 358
Score = 28.1 bits (61), Expect = 8.9
Identities = 28/80 (35%), Positives = 37/80 (46%), Gaps = 6/80 (7%)
Query: 7 NNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAAR-DYGRGALTEMER 65
N+G A TI + G +N G +R V A + +L+A D+ E ER
Sbjct: 170 NSGTTALTIVRQ----GDVIYIANVGDSRAVLATVSDEGSLVAVQLTVDFKPNLPQEEER 225
Query: 66 LKGCNGGKDCLGAE-GVLRV 84
+ GCNG CL E GV RV
Sbjct: 226 IIGCNGRVFCLQDEPGVHRV 245
>At2g48050 hypothetical protein
Length = 1500
Score = 28.1 bits (61), Expect = 8.9
Identities = 27/135 (20%), Positives = 53/135 (39%), Gaps = 25/135 (18%)
Query: 193 YVVCLVGIILMFIW------------YAPKPSCLLNIFFIAWTLVLLQLMT----SVSLH 236
++ CL + + +W Y K L N++ L+++QL + +
Sbjct: 92 FIYCLSSFVSLQLWLSGFIDLYFYLGYNSKAPLLDNVWESLAVLIVMQLYSYERRQSGHY 151
Query: 237 PKVNAGILTPGLMGLYVVFLCWNAIRSEPDGGSCIRKSDSSTKTDWLSIISFIIAILAIV 296
+ +L PG+ G + FL W+ G + + +S+ F+ + ++
Sbjct: 152 IPGQSSLLHPGVFGFFERFLAWH-------GQKILFAALFYASLSPISVFGFVYLLGLVI 204
Query: 297 IATF--STGIDSKCF 309
TF S+ I SK F
Sbjct: 205 CTTFPKSSSIPSKSF 219
>At2g02020 putative peptide/amino acid transporter
Length = 545
Score = 28.1 bits (61), Expect = 8.9
Identities = 42/157 (26%), Positives = 66/157 (41%), Gaps = 24/157 (15%)
Query: 160 SIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAP--KPSCLLNI 217
++I++ T N+ E+ AA+ HVM + T Y+ L+G ++ ++ +C I
Sbjct: 69 NLITYFT--NELHETNVSAAR---HVMTWQGTCYITPLIGALIADAYWGRYWTIACFSAI 123
Query: 218 FFIAWTLVLLQLMTSV-SLHPKVNAGILTPG---------LMGLYVVFLCWNAIRS--EP 265
+F +V L L SV L P G L P GLY++ L I+
Sbjct: 124 YFTG--MVALTLSASVPGLKPAECIGSLCPPATMVQSTVLFSGLYLIALGTGGIKPCVSS 181
Query: 266 DGGSCIRKSDSS---TKTDWLSIISFIIAILAIVIAT 299
G K+D S K + + F I I A V +T
Sbjct: 182 FGADQFDKTDPSERVRKASFFNWFYFTINIGAFVSST 218
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.140 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,412,737
Number of Sequences: 26719
Number of extensions: 394273
Number of successful extensions: 1043
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1028
Number of HSP's gapped (non-prelim): 15
length of query: 398
length of database: 11,318,596
effective HSP length: 101
effective length of query: 297
effective length of database: 8,619,977
effective search space: 2560133169
effective search space used: 2560133169
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0214.6


