

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.7
(451 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
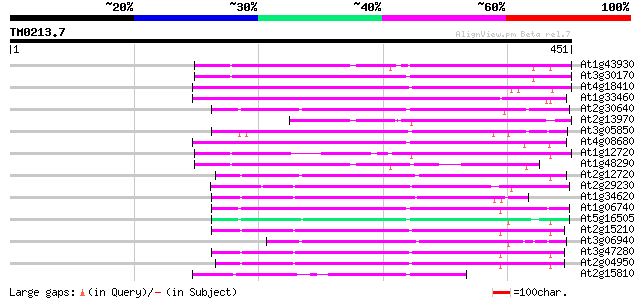
Score E
Sequences producing significant alignments: (bits) Value
At1g43930 hypothetical protein 145 6e-35
At3g30170 unknown protein 141 8e-34
At4g18410 MuDR transposable element - like protein 140 1e-33
At1g33460 mutator transposase MUDRA, putative 135 5e-32
At2g30640 Mutator-like transposase 133 2e-31
At2g13970 Mutator-like transposase 122 3e-28
At3g05850 unknown protein 116 3e-26
At4g08680 putative MuDR-A-like transposon protein 112 3e-25
At1g12720 hypothetical protein 111 7e-25
At1g48290 hypothetical protein 108 6e-24
At2g12720 putative protein 104 9e-23
At2g29230 Mutator-like transposase 99 6e-21
At1g34620 Mutator-like protein 98 8e-21
At1g06740 mudrA-like protein 96 3e-20
At5g16505 mutator-like transposase-like protein (MQK4.25) 95 7e-20
At2g15210 putaive transposase of transposon FARE2.7 (cds1) 95 9e-20
At3g06940 putative mudrA protein 94 2e-19
At3g47280 putative protein 94 2e-19
At2g04950 putative transposase of transposon FARE2.9 (cds1) 93 3e-19
At2g15810 Mutator-like transposase 91 1e-18
>At1g43930 hypothetical protein
Length = 946
Score = 145 bits (365), Expect = 6e-35
Identities = 100/323 (30%), Positives = 152/323 (46%), Gaps = 29/323 (8%)
Query: 149 GLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQ 208
GL+ ++ EH C RH+ N+K+ + ++ L ++ + M L+
Sbjct: 508 GLVKAIHSVIPQAEHRQCARHIMENWKRN-SHDMELQRLFWKIVRSYTEGEFGAHMRALK 566
Query: 209 ALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIR 268
+ + A+ L++ W + F C+ +NNLSESFN TI R KP+L M++ IR
Sbjct: 567 SYNASAFELLLKTLPVTWSRAFFKIGSCCNDNLNNLSESFNRTIREARRKPLLDMLEDIR 626
Query: 269 SYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMV-------RYWVATRCGEDKYEVKH 321
M R NEK + + KR EIE + + W+A +K+E+K
Sbjct: 627 RQCMVR----NEKRYVIADRLRTRFTKRAHAEIEKMIAGSQVCQRWMARH---NKHEIK- 678
Query: 322 MVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYG 381
+ D F V++ +TC C W++ GIPC HA + I Q+ ED+V +Y + TY
Sbjct: 679 VGPVDSFTVDMNNNTCGCMKWQMTGIPCIHAASVIIGKRQKVEDYVSDWYTTSMWKQTYN 738
Query: 382 HVITPINGQKLWPTTNDPYILPPLYKRA-PGRPKKLRRR-------DPHEDTTDNQTRLR 433
I P+ G+ LWPT N +LPP ++R PGRPK RR +T +RL
Sbjct: 739 DGIGPVQGKLLWPTVNKVGVLPPPWRRGNPGRPKNHARRKGVFESSTASSSSTTELSRLH 798
Query: 434 -----SRCGQFGHNSRSCRNPVV 451
S C GHN + C+N V
Sbjct: 799 RVMTCSNCQGEGHNKQGCKNETV 821
>At3g30170 unknown protein
Length = 995
Score = 141 bits (355), Expect = 8e-34
Identities = 94/314 (29%), Positives = 147/314 (45%), Gaps = 14/314 (4%)
Query: 149 GLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQ 208
GL+ ++ EH C +H+ N+K+ + ++ L A++ + + MMELQ
Sbjct: 480 GLVKAIHTILPHAEHRQCSKHIMDNWKRD-SHDLELQRLFWKIARSYTVEEFNNHMMELQ 538
Query: 209 ALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIR 268
+ +AY L W + F C+ +NNLSESFN TI R KP+L M++ IR
Sbjct: 539 QYNRQAYESLQLTSPVTWSRAFFRIGTCCNDNLNNLSESFNRTIRQARRKPILDMLEDIR 598
Query: 269 SYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQF 328
M R A EK + + + +E I +R E+ +E+ V+ +
Sbjct: 599 RQCMVRSAKRFLIAEKLQTRFTKRAHEEIEKMIVGLRQCERYMARENLHEI--YVNNVSY 656
Query: 329 VVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPIN 388
V++ + TC C ++VGIPC HA I ++ ED+V YY + + TY I P+
Sbjct: 657 FVDMEQKTCDCRKCQMVGIPCIHATCVIIGKKEKVEDYVSDYYTKVRWRETYLKGIRPVQ 716
Query: 389 GQKLWPTTNDPYILPPLYKRA-PGRPKKLRRR----------DPHEDTTDNQTRLRSRCG 437
G LW TN +LPP ++R GRP RR +P++ + + + S C
Sbjct: 717 GMPLWCRTNRLPVLPPPWRRGNAGRPSNYARRKGRNEAAAPSNPNKMSREKRIMTCSNCH 776
Query: 438 QFGHNSRSCRNPVV 451
Q GHN + C P V
Sbjct: 777 QEGHNKQRCNIPTV 790
>At4g18410 MuDR transposable element - like protein
Length = 633
Score = 140 bits (353), Expect = 1e-33
Identities = 88/315 (27%), Positives = 148/315 (46%), Gaps = 13/315 (4%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ + +EH C++H+Y N KK +G +I+ L+ A + + + + ++
Sbjct: 303 KGLIKAVQLELPKIEHRMCVQHIYGNLKKTYGSKTMIKPLLWNLAWSYNETEYKQHLEKI 362
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+ + Y +M+ R+W + C+ + NN ESFN ++ R+K + M++ I
Sbjct: 363 RCYDTKVYESVMKTNPRSWVRAFQKIGSFCEDVDNNSVESFNGSLNKAREKQFVAMLETI 422
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R M R A + + + G P +K L E ++ YEV+H D
Sbjct: 423 RRMAMVRIAKRSVESHTHTGVCTPYVMKFLAGEHKVASTAKVAPGTNGMYEVRH--GGDT 480
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI 387
V+LA +TC+C W++ GIPC HA I H +PEDFV ++R + Y + P
Sbjct: 481 HRVDLAAYTCTCIKWQICGIPCEHAYGVILHKKLQPEDFVCQWFRTAMWKKNYTDGLFPQ 540
Query: 388 NGQKLWPTTNDPYIL---PPLYK--RAPGRPKKLRRRDPHEDTTDNQTRLRSR------C 436
G K WP +N P + PP + + + +K R++ +E T Q + + R C
Sbjct: 541 RGPKFWPESNGPRVFAAEPPEGEEDKKMTKAEKKRKKGVNESPTKKQPKAKKRIMHCGVC 600
Query: 437 GQFGHNSRSCRNPVV 451
G HNSR +N V
Sbjct: 601 GAADHNSRHHKNDKV 615
>At1g33460 mutator transposase MUDRA, putative
Length = 826
Score = 135 bits (340), Expect = 5e-32
Identities = 87/317 (27%), Positives = 147/317 (45%), Gaps = 18/317 (5%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+G+++ + + EH C++H+ N KK+ G +++ + A + + + E+
Sbjct: 442 KGIISAVKNALPNAEHRPCVKHIVENLKKRHGSLDLLKKFVWNLAWSYSDTQYKANLNEM 501
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+A Y +M+ WC+ F C+ + NN +ESFNATI+ R K ++ M++ I
Sbjct: 502 RAYIMSLYEDVMKEEPNTWCRSWFRIGSYCEDVDNNATESFNATIVKARAKALVPMMETI 561
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R M R + K+ K+K ++ + L+ E E+ T+ +K+EV + +
Sbjct: 562 RRQAMARISKRKAKIGKWKKKISEYVSEILKEEWELAVKCEVTKGTHEKFEVWFDGNSNA 621
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI 387
+ E CSC W++ GIPC HA AAI+ + EDFV P + A+ Y P+
Sbjct: 622 VSLKTEEWDCSCCKWQVTGIPCEHAYAAINDVGKDVEDFVIPMFYTIAWREQYDTGPDPV 681
Query: 388 NGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQT-------RLR------- 433
GQ WP T I PL P KK +++ H N++ RL+
Sbjct: 682 RGQMYWP-TGLGLITAPLQDPVPPGRKKGEKKNYHRIKGPNESPKKKGPQRLKVGKKGVV 740
Query: 434 ---SRCGQFGHNSRSCR 447
CG+ GHN+ C+
Sbjct: 741 MHCKSCGEAGHNAAGCK 757
>At2g30640 Mutator-like transposase
Length = 754
Score = 133 bits (334), Expect = 2e-31
Identities = 90/299 (30%), Positives = 139/299 (46%), Gaps = 17/299 (5%)
Query: 163 HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIP 222
H C+ L + + +F +++ NL+ AAK + KM E+ + PEA +W+ I
Sbjct: 462 HGLCVHCLTESVRTQFNNSILV-NLVWEAAKCLTDFEFEGKMGEIAQISPEAASWIRNIQ 520
Query: 223 TRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKL 282
W + F + L N+SES N+ + P++ M++ IR +M F E
Sbjct: 521 HSQWATYCFEG-TRFGHLTANVSESLNSWVQDASGLPIIQMLESIRRQLMTLFNERRETS 579
Query: 283 EKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFW 342
++ G ++P + + IE R + + E ++EV M S+ +++V++ TC C W
Sbjct: 580 MQWSGMLVPSAERHVLEAIEECRLYPVHKANEAQFEV--MTSEGKWIVDIRCRTCYCRGW 637
Query: 343 ELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLWPTT------ 396
EL G+PC HAVAA+ Q F Y+ Y TY I P+ + W TT
Sbjct: 638 ELYGLPCSHAVAALLACRQNVYRFTESYFTVANYRRTYAETIHPVPDKTEWKTTEPAGES 697
Query: 397 ----NDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLR-SRCGQFGHNSRSCRNPV 450
+ I PP R P RPKK RR ED + +R SRC Q GH +C P+
Sbjct: 698 GEDGEEIVIRPPRDLRQPPRPKK--RRSQGEDRGRQKRVVRCSRCNQAGHFRTTCTAPI 754
>At2g13970 Mutator-like transposase
Length = 597
Score = 122 bits (307), Expect = 3e-28
Identities = 83/234 (35%), Positives = 116/234 (49%), Gaps = 22/234 (9%)
Query: 226 WCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKLEKY 285
W + F C+ +NNLSESFN TI R KP+L M++ IR M R NEK
Sbjct: 236 WSRAFFRIGSCCNDNLNNLSESFNRTIRNARRKPLLDMLEDIRRQCMVR----NEKRFVI 291
Query: 286 KGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKH------MVSQDQFVVNLAEHTCSC 339
+ + KR EIE + A C D++ +H + D F V++ + TC C
Sbjct: 292 AYRLKSRFTKRAHMEIEKMIDGAAV-C--DRWMARHNRIEVRVGLDDSFSVDMNDRTCGC 348
Query: 340 NFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLWPTTNDP 399
W++ GIPC HA + I + Q+ EDFV +Y + TY I P+ G+ LWP N
Sbjct: 349 RKWQMTGIPCIHAASVIIGNRQKVEDFVSDWYTTYMWKQTYYDGIMPVQGKMLWPIVNRV 408
Query: 400 YILPPLYKRA-PGRPKKL-RRRDPHEDTTDNQTRLRSRCGQFGHNSRSCRNPVV 451
+LPP ++R PGRPK RR+ +E T + +R GHN C+N V
Sbjct: 409 GVLPPPWRRGNPGRPKNHDRRKGIYESATASTSR-------EGHNKAGCKNETV 455
>At3g05850 unknown protein
Length = 777
Score = 116 bits (290), Expect = 3e-26
Identities = 84/298 (28%), Positives = 137/298 (45%), Gaps = 18/298 (6%)
Query: 163 HIFCLRHLYANFKKKFGGGVV--IRNLMM----AAAKATYHQLWTEKMMELQALHPEAYT 216
H +CLR+L K G I+ L++ +AA A + + ++ L PEAY
Sbjct: 481 HAYCLRYLTDELIKDLKGPFSHEIKRLIVDDFYSAAYAPRADSFERHVENIKGLSPEAYD 540
Query: 217 WLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFA 276
W++Q +A+ + + + ++ E F + D P+ MVD IR IMG
Sbjct: 541 WIVQKSQPDHWANAYFRGARYNHMTSHSGEPFFSWASDANDLPITQMVDVIRGKIMGLIH 600
Query: 277 TMNEKLEKYKGEVMPKPLKRLEHE-IEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEH 335
+ G + P +LE E + VA + ++V+ + +VN+AE
Sbjct: 601 VRRISANEANGNLTPSMEVKLEKESLRAQTVHVAPSADNNLFQVR---GETYELVNMAEC 657
Query: 336 TCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI---NGQKL 392
CSC W+L G+PC HAVA I++ + P D+ Y+ Y +TY I P+ G+
Sbjct: 658 DCSCKGWQLTGLPCHHAVAVINYYGRNPYDYCSKYFTVAYYRSTYAQSINPVPLLEGEMC 717
Query: 393 WPTTNDP--YILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGHNSRSCRN 448
++ + PP +R PGRP K ++ P E+ Q + SRC GHN +C++
Sbjct: 718 RESSGGSAVTVTPPPTRRPPGRPPK--KKTPAEEVMKRQLQC-SRCKGLGHNKSTCKD 772
>At4g08680 putative MuDR-A-like transposon protein
Length = 761
Score = 112 bits (281), Expect = 3e-25
Identities = 73/321 (22%), Positives = 138/321 (42%), Gaps = 23/321 (7%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GLL+ + EH C++H+ N KK +++ L+ A + + + + + L
Sbjct: 376 KGLLSAVKQELPNAEHRMCVKHIVENLKKNHAKKDMLKTLVWKLAWSYNEKEYGKNLNNL 435
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+ Y ++ W + + C+ + NN +ESFN+TI R K ++ M++ I
Sbjct: 436 RCYDEALYNDVLNEEPHTWSRCFYKLGSCCEDVDNNATESFNSTITKARAKSLIPMLETI 495
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R M R N+K +++G LK L E RC +EV + ++
Sbjct: 496 RRQGMTRIVKRNKKSLRHEGRFTKYALKMLALEKTDADRSKVYRCTHGVFEV--YIDENG 553
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI 387
V++ + CSC W++ GIPC H+ A+ + E+++ ++ + + +Y P+
Sbjct: 554 HRVDIPKTQCSCGKWQISGIPCEHSYGAMIEAGLDAENYISEFFSTDLWRDSYETATMPL 613
Query: 388 NGQKLWPTTNDPYILPPLYKRAPGR-------------------PKKLRRRDPHEDTTDN 428
G K W ++ + P PGR PKK +R+ +
Sbjct: 614 RGPKYWLNSSYGLVTAPPEPILPGRKKEKSKKEKFARIKGKNESPKKKKRKKNEVEKLGK 673
Query: 429 QTRL--RSRCGQFGHNSRSCR 447
+ ++ CG+ GHN+ C+
Sbjct: 674 KGKIIHCKSCGEAGHNALRCK 694
>At1g12720 hypothetical protein
Length = 585
Score = 111 bits (278), Expect = 7e-25
Identities = 83/319 (26%), Positives = 136/319 (42%), Gaps = 48/319 (15%)
Query: 149 GLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQ 208
GL+ +++ EH C +H+ N+K+ + ++ L +++ + + M L+
Sbjct: 190 GLVKAIHNVLPQAEHRQCSKHIMDNWKRD-SHDMELQRLFWKISRSYTIEEFNTHMANLK 248
Query: 209 ALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIR 268
+ +P+AY L W TI R KP+L M++ IR
Sbjct: 249 SYNPQAYASLQLTSPMTW------------------------TIRQARRKPLLDMLEDIR 284
Query: 269 SYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKH-----MV 323
M R A E+ K P R EIE + +A G +++ ++ V
Sbjct: 285 RQCMVRTAKRFIIAERLKSRFTP----RAHAEIEKM---IAGSAGCERHLARNNLHEIYV 337
Query: 324 SQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHV 383
+ + V++ + TC C WE+VGIPC H I ++ ED+V YY + + TY
Sbjct: 338 NDVGYFVDMDKKTCGCRKWEMVGIPCVHTPCVIIGRKEKVEDYVSDYYTKVRWRETYRDG 397
Query: 384 ITPINGQKLWPTTNDPYILPPLYKRA-PGRPKKLRRRDPHEDTTDNQTRLR--------- 433
I P+ G LWP + +LPP ++R PGR R+ +T + + +
Sbjct: 398 IRPVQGMPLWPRMSRLPVLPPPWRRGNPGRQSNYARKKGRYETASSSNKNKMSRANRIMT 457
Query: 434 -SRCGQFGHNSRSCRNPVV 451
S C Q GHN SC+N V
Sbjct: 458 CSNCKQEGHNKSSCKNATV 476
>At1g48290 hypothetical protein
Length = 444
Score = 108 bits (270), Expect = 6e-24
Identities = 81/287 (28%), Positives = 128/287 (44%), Gaps = 33/287 (11%)
Query: 149 GLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQ 208
GL+ + + EH C +H+ N+K+ + ++ L A++ +T + EL+
Sbjct: 164 GLVKAIHNRIPAAEHRQCAKHIMDNWKRN-SHDMELQRLFWKIARSYTIGEYTANLEELK 222
Query: 209 ALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIR 268
+P A LM W + F C+ +NNLSESFN TI R KP+L M++ IR
Sbjct: 223 TYNPGAAASLMNTKPMEWSRAFFRIGSCCNDNLNNLSESFNRTIRQARRKPLLDMLEDIR 282
Query: 269 SYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMV----RYWVATRCGEDKYEVKHMVS 324
S M R NEK G + KR EIE + ++ + +K+E+ H
Sbjct: 283 SQCMVR----NEKRYIIAGRWKSRFTKRAHEEIEKMIAGSQFCERSMARHNKHEISHF-- 336
Query: 325 QDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVI 384
++ V++ ++TC C W++ ED+V +Y + TY I
Sbjct: 337 GRKYSVDMNDNTCGCRKWQMT-----------------VEDYVSDWYTTRMWQLTYNDGI 379
Query: 385 TPINGQKLWPTTNDPYILPPLYKRA-PGRPK----KLRRRDPHEDTT 426
P+ GQ LWP N +LPP ++R PG K+ PH+ T
Sbjct: 380 APVQGQLLWPRVNRLGVLPPPWRRGLPGLSTLFDVKIVTSSPHQHFT 426
>At2g12720 putative protein
Length = 819
Score = 104 bits (260), Expect = 9e-23
Identities = 75/287 (26%), Positives = 124/287 (43%), Gaps = 9/287 (3%)
Query: 166 CLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIPTRA 225
C HLY N KF + L+ AA+ +T E++ L P + +L +
Sbjct: 536 CTYHLYKNILVKFKRKDLFP-LVKKAARCYRLNDFTNAFNEIEELDPLLHAYLQRAGVEM 594
Query: 226 WCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKLEKY 285
W + F + +++ N++ES N + ++ P++ M++ IR + FA + K
Sbjct: 595 WARAHFPG-DRYNLMTTNIAESMNRALSQAKNLPIVRMLEAIRQMMTRWFAERRDDASKQ 653
Query: 286 KGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFWELV 345
++ P K L+ + R + +V + S VVN+ E C+C +
Sbjct: 654 HTQLTPGVEKLLQTRVTSSRLLDVQTIDASRVQVAYEASLH--VVNVDEKQCTCRLFNKE 711
Query: 346 GIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLWP-TTNDPYILPP 404
+PC HA+AA H PYY+ + Y I P + + P D LPP
Sbjct: 712 KLPCIHAIAAAEHMGVSRISLCSPYYKSSHLVNVYACAIMPSDTEVPVPQIVIDQPCLPP 771
Query: 405 LYKRAPGRPKKLRRRDPHEDTTDNQTRLR----SRCGQFGHNSRSCR 447
+ PGRPKKLR + E + + + SRC + GHN ++CR
Sbjct: 772 IVANQPGRPKKLRMKSALEVAVETKRPRKEHACSRCKETGHNVKTCR 818
>At2g29230 Mutator-like transposase
Length = 915
Score = 98.6 bits (244), Expect = 6e-21
Identities = 76/300 (25%), Positives = 131/300 (43%), Gaps = 20/300 (6%)
Query: 162 EHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQI 221
+H C+ HL N + ++ ++ AA + T E+ L P +L +
Sbjct: 625 KHGACIVHLQRNIATSYKKKHLLFHVSRAARAFRICEFHTY-FNEVIRLDPACARYLESV 683
Query: 222 PTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEK 281
W + A+ + +V+ +N++ES NA + R+ P+++++++IR+ ++ FA E
Sbjct: 684 GFCHWTR-AYFLGKRYNVMTSNVAESLNAVLKEARELPIISLLEFIRTTLISWFAMRREA 742
Query: 282 LEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNF 341
+ PK + + E + R YE++ + V L E TCSC
Sbjct: 743 ARTEASPLPPKMREVVHRNFEKSVRFAVHRLDRYDYEIREE-GASVYHVKLMERTCSCRA 801
Query: 342 WELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLWPTTNDPYI 401
++L+ +PC HA+AA + + P Y E++ +Y I P+ P D +
Sbjct: 802 FDLLHLPCPHAIAAAVAEGVPIQGLMAPEYSVESWRMSYLGTIKPV------PEVGDVFA 855
Query: 402 L----------PPLYKRAPGRPKKLRRRDPHEDTTDNQTRLR-SRCGQFGHNSRSCRNPV 450
L PP +R GRPKK R E T + R SRC GHN +C+ P+
Sbjct: 856 LPEPIASLRLFPPATRRPSGRPKKKRITSRGEFTCPQRQVTRCSRCTGAGHNRATCKMPI 915
>At1g34620 Mutator-like protein
Length = 1034
Score = 98.2 bits (243), Expect = 8e-21
Identities = 70/262 (26%), Positives = 128/262 (48%), Gaps = 12/262 (4%)
Query: 163 HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIP 222
H C HL+ N + + + R LM AA+A + + +++Q L+P +L+ +
Sbjct: 647 HAACTVHLWRNIRHLYKPKSLCR-LMSEAAQAFHVTDFNRIFLKIQKLNPGCAAYLVDLG 705
Query: 223 TRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKL 282
W + + + +++ +N+ ES+N I R+ P++ M+++IR+ +M FAT +
Sbjct: 706 FSEWTR-VHSKGRRFNIMDSNICESWNNVIREAREYPLICMLEYIRTTLMDWFATRRAQA 764
Query: 283 EKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFW 342
E + P+ +R+E + +++V+ + F+V L E TCSC +
Sbjct: 765 EDCPTTLAPRVQERVEENYQSAMSMSVKPICNFEFQVQERTGEC-FIVKLDESTCSCLEF 823
Query: 343 ELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPIN--GQKLWP-----T 395
+ +GIPC HA+AA + + Y E +Y I PI+ G ++ P T
Sbjct: 824 QGLGIPCAHAIAAAARIGVPTDSLAANGYFNELVKLSYEEKIYPIHSVGGEVAPEIAAGT 883
Query: 396 TNDPYILPPLYKRAPGRPKKLR 417
T + + PP +R PGRP+K+R
Sbjct: 884 TGE--LQPPFVRRPPGRPRKIR 903
>At1g06740 mudrA-like protein
Length = 726
Score = 96.3 bits (238), Expect = 3e-20
Identities = 68/291 (23%), Positives = 119/291 (40%), Gaps = 7/291 (2%)
Query: 163 HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIP 222
H FCL +L F+++F V++ +L AA + K+ +++ + PEA W+
Sbjct: 440 HGFCLHYLTERFQREFQSSVLV-DLFWEAAHCLTVLEFKSKINKIEQISPEASLWIQNKS 498
Query: 223 TRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKL 282
W F + N ++ES + + P++ ++ I +++ E
Sbjct: 499 PARWASSYFEGTRFGQLTANVITESLSNWVEDTSGLPIIQTMECIHRHLINMLKERRETS 558
Query: 283 EKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFW 342
+ ++P K++ IE R R E ++EV M + VVN+ +C C W
Sbjct: 559 LHWSNVLVPSAEKQMLAAIEQSRAHRVYRANEAEFEV--MTCEGNVVVNIENCSCLCGRW 616
Query: 343 ELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLW---PTTNDP 399
++ G+PC HAV A+ + + + E Y Y + PI+ + W + D
Sbjct: 617 QVYGLPCSHAVGALLSCEEDVYRYTESCFTVENYRRAYAETLEPISDKVQWKENDSERDS 676
Query: 400 YILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGHNSRSCRNPV 450
+ K G P+K R R D RC Q GH +C P+
Sbjct: 677 ENVIKTPKAMKGAPRKRRVRAEDRDRVRRVVHC-GRCNQTGHFRTTCTAPM 726
>At5g16505 mutator-like transposase-like protein (MQK4.25)
Length = 597
Score = 95.1 bits (235), Expect = 7e-20
Identities = 71/301 (23%), Positives = 122/301 (39%), Gaps = 23/301 (7%)
Query: 163 HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIP 222
H FCLR++ NF+ F ++ N+ +A A + K E+ + + W
Sbjct: 307 HGFCLRYVSENFRDTFKNTKLV-NIFWSAVYALTPAEFETKSNEMIEISQDVVQWFELYL 365
Query: 223 TRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKL 282
W F + ++E L + P++ M++ IR I F E
Sbjct: 366 PHLWAVAYFQGV-RYGHFGLGITEVLYNWALECHELPIIQMMEHIRHQISSWFDNRRELS 424
Query: 283 EKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFW 342
+ ++P +R+ + R + R E ++E+ + ++ +V++ CSC W
Sbjct: 425 MGWNSILVPSAERRITEAVADARCYQVLRANEVEFEI--VSTERTNIVDIRTRDCSCRRW 482
Query: 343 ELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLWPTTNDP--- 399
++ G+PC HA AA+ + F P + +Y TY +I I + LW +
Sbjct: 483 QIYGLPCAHAAAALISCGRNVHLFAEPCFTVSSYQQTYSQMIGEIPDRSLWKDEGEEAGG 542
Query: 400 -----YILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLR-----SRCGQFGHNSRSCRNP 449
I PP +R PGRPKK R +N R + RC GH+ + C P
Sbjct: 543 GVESLRIRPPKTRRPPGRPKKKVVR------VENLKRPKRIVQCGRCHLLGHSQKKCTQP 596
Query: 450 V 450
+
Sbjct: 597 I 597
>At2g15210 putaive transposase of transposon FARE2.7 (cds1)
Length = 580
Score = 94.7 bits (234), Expect = 9e-20
Identities = 71/291 (24%), Positives = 132/291 (44%), Gaps = 11/291 (3%)
Query: 163 HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIP 222
H C HL N + F G +I + AA++ + E+ + +L ++
Sbjct: 290 HGLCTFHLQKNLETHFRGSSLIP-VYYAASRVYTKTEFDSLFWEITNSDKKLAQYLWEVH 348
Query: 223 TRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKL 282
R W + A++ + +++ +NL+ES NA + R+ P++ + + IRS + F E+
Sbjct: 349 VRKWSR-AYSPSNRYNIMTSNLAESVNALLKQNREYPIVCLFESIRSIMTRWFNERREES 407
Query: 283 EKYKGEVMPKPLKRLEHEIEMVRYWVAT-RCGEDKYEVKHMVSQDQFVVNLAEHTCSCNF 341
++ V K+++ + W+ + ++++EVK +VNL + TC+C
Sbjct: 408 SQHPSAVTINVGKKMKASYDTSTRWLEVCQVNQEEFEVKG--DTKTHLVNLDKRTCTCCM 465
Query: 342 WELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLW---PTTND 398
+++ PC H +A+ +H + FV ++ + Y I P + W T ++
Sbjct: 466 FDIDKFPCAHGIASANHINLNENMFVDEFHSTYRWRQAYSESIHPNGDMEYWEIPETISE 525
Query: 399 PYILPPLYKRAPGRPKKLRRRDPHE-DTTDNQTRLR--SRCGQFGHNSRSC 446
LPP + GR KK R E + + +L SRCGQ GHN +C
Sbjct: 526 VICLPPSTRVPSGRRKKKRIPSVWEHGRSQPKPKLHKCSRCGQSGHNKSTC 576
>At3g06940 putative mudrA protein
Length = 749
Score = 94.0 bits (232), Expect = 2e-19
Identities = 58/245 (23%), Positives = 113/245 (45%), Gaps = 9/245 (3%)
Query: 207 LQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDW 266
++++ P+AY W+++ W F + + + +N + F + + + P+ M+D
Sbjct: 508 IKSISPDAYNWVIESEPHHWANALFQG-ERYNKMNSNFGKDFYSWVSEAHEFPITQMIDE 566
Query: 267 IRSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQD 326
R+ +M T K ++ + P ++L+ EIE+ + + ++V S
Sbjct: 567 FRAKMMQSIYTRQVKSREWVTTLTPSNEEKLQKEIELAQLLQVSSPEGSLFQVNGGESSV 626
Query: 327 QFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITP 386
+V++ + C C W L G+PC HA+A I + P ++ Y E++ Y I P
Sbjct: 627 S-IVDINQCDCDCKTWRLTGLPCSHAIAVIGCIEKSPYEYCSTYLTVESHRLMYAESIQP 685
Query: 387 INGQKLWPTTNDP----YILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGHN 442
+ + P + PP +R PGRP K+++ +P D Q + S+C GHN
Sbjct: 686 VPNMDRMMLDDPPEGLVCVTPPPTRRTPGRP-KIKKVEP-LDMMKRQLQC-SKCKGLGHN 742
Query: 443 SRSCR 447
++C+
Sbjct: 743 KKTCK 747
>At3g47280 putative protein
Length = 739
Score = 93.6 bits (231), Expect = 2e-19
Identities = 71/291 (24%), Positives = 131/291 (44%), Gaps = 11/291 (3%)
Query: 163 HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIP 222
H C HL N + F G +I + AA++ + E+ + +L ++
Sbjct: 449 HGLCTFHLQKNLETHFRGSSLIP-VYYAASRIYTKTEFDSLFWEITNSDKKLAQYLCEVD 507
Query: 223 TRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKL 282
R W + A++ + +++ +NL+ES NA + R+ PV+ + + IRS + F E+
Sbjct: 508 VRKWSR-AYSPSNRYNIMTSNLAESVNALLKQNREYPVVCLFESIRSIMTRWFNERREES 566
Query: 283 EKYKGEVMPKPLKRLEHEIEMVRYWVAT-RCGEDKYEVKHMVSQDQFVVNLAEHTCSCNF 341
++ V K+++ + W+ + ++++EVK +VNL + TC+C
Sbjct: 567 SQHPSAVTINVGKKMKASYDTSTRWLEVCQVNQEEFEVKG--DTKTHLVNLDKRTCTCCM 624
Query: 342 WELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLW---PTTND 398
+++ PC H +A+ H + FV ++ + Y I P + W T ++
Sbjct: 625 FDIDKFPCAHGIASAKHINLNENMFVDEFHSTYRWRQAYSESIHPNGDMEYWEIPETISE 684
Query: 399 PYILPPLYKRAPGRPKKLRRRDPHE-DTTDNQTRLR--SRCGQFGHNSRSC 446
LPP + G+ KK R E + + +L SRCGQ GHN +C
Sbjct: 685 VICLPPSTRVPTGKRKKKRIPSVWEHGRSQPKPKLHKCSRCGQSGHNKSTC 735
>At2g04950 putative transposase of transposon FARE2.9 (cds1)
Length = 616
Score = 93.2 bits (230), Expect = 3e-19
Identities = 70/288 (24%), Positives = 131/288 (45%), Gaps = 11/288 (3%)
Query: 166 CLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIPTRA 225
C HL N + F G +I + AA++ + E+ + +L ++ R
Sbjct: 329 CTFHLQKNLETHFRGSSLIP-VYYAASRVYTKTEFDSLFWEITNSDKKLAQYLWEVDVRK 387
Query: 226 WCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKLEKY 285
W + A++ + +++ +NL+ES NA + R+ P++ + + IRS + F E+ ++
Sbjct: 388 WSR-AYSPSNRYNIMTSNLAESVNALLKQNREYPIVCLFESIRSIMTRWFNERREESSQH 446
Query: 286 KGEVMPKPLKRLEHEIEMVRYWVAT-RCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFWEL 344
V K+++ + W+ + ++++EVK +VNL + TC+C +++
Sbjct: 447 PSAVTINVGKKMKASYDTSTRWLEVCQVNQEEFEVKG--DTKTHLVNLDKRTCTCCMFDI 504
Query: 345 VGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLW---PTTNDPYI 401
PC H +A+ H + FV ++ + Y I P + W T ++
Sbjct: 505 DKFPCAHGIASAKHINLNENMFVDEFHSTYKWRQAYSKSIHPNGDMEYWEIPETISEVIC 564
Query: 402 LPPLYKRAPGRPKKLRRRDPHE-DTTDNQTRLR--SRCGQFGHNSRSC 446
LPP + GR KK R E + + +L SRCGQ GHN+ +C
Sbjct: 565 LPPSTRVPSGRRKKKRIPSVWEHGRSQPKPKLHKCSRCGQSGHNNSTC 612
>At2g15810 Mutator-like transposase
Length = 545
Score = 90.9 bits (224), Expect = 1e-18
Identities = 61/220 (27%), Positives = 100/220 (44%), Gaps = 21/220 (9%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GLL DL+ EH C RH+Y+N+KK +G + A A ++ +T M L
Sbjct: 126 KGLLNAVADLLPEAEHRHCARHIYSNWKKVYGD-YCHESYFWAIAYSSTDGTYTYNMRAL 184
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
++ P AY L++ + + N L ++ R KPV++M++ +
Sbjct: 185 KSYDPLAYEDLLKTEPKTEFQQGH----------NGLRDA--------RKKPVISMLEDV 226
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R M R + ++ K K + P + L+ + + + E YEV M
Sbjct: 227 RRQAMKRISKRRDQTAKCKSKFPPHIMGVLKANRKSAKLCTVLKNSEHVYEV--MEGNGG 284
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFV 367
F N+ TC+CN W L GIPC HA+ I+ ++ PED +
Sbjct: 285 FTANILHRTCACNQWNLTGIPCPHAIYVINEHNRDPEDTI 324
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.341 0.149 0.525
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,877,443
Number of Sequences: 26719
Number of extensions: 405308
Number of successful extensions: 1470
Number of sequences better than 10.0: 75
Number of HSP's better than 10.0 without gapping: 70
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 1283
Number of HSP's gapped (non-prelim): 88
length of query: 451
length of database: 11,318,596
effective HSP length: 103
effective length of query: 348
effective length of database: 8,566,539
effective search space: 2981155572
effective search space used: 2981155572
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0213.7


