

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.11
(579 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
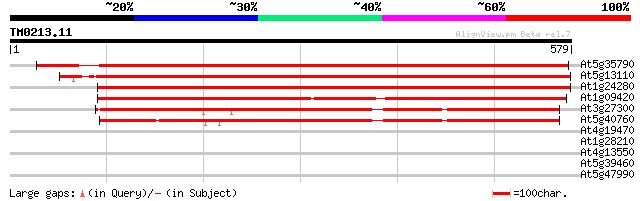
Score E
Sequences producing significant alignments: (bits) Value
At5g35790 glucose-6-phosphate dehydrogenase 855 0.0
At5g13110 glucose-6-phosphate 1-dehydrogenase 832 0.0
At1g24280 glucose-6-phosphate 1-dehydrogenase 820 0.0
At1g09420 putative Glucose-6-phosphate dehydrogenase (F14J9.8) 538 e-153
At3g27300 glucose-6-phosphate 1-dehydrogenase, putative 459 e-129
At5g40760 glucose-6-phosphate dehydrogenase 448 e-126
At4g19470 putative protein 32 1.3
At1g28210 AtJ1 (atj) 32 1.3
At4g13550 putative protein 30 4.8
At5g39460 hypothetical protein 29 6.3
At5g47990 cytochrome P450 29 8.2
>At5g35790 glucose-6-phosphate dehydrogenase
Length = 576
Score = 855 bits (2208), Expect = 0.0
Identities = 427/550 (77%), Positives = 472/550 (85%), Gaps = 20/550 (3%)
Query: 28 FSCIRPGNLPRNHFQLKSSNGHPLNAVS-SHSHDGLAGSSLTKEDSKPQPVDGPFLSSDS 86
FS +R H QL +SNG N S S D L +TK +S
Sbjct: 44 FSQVRLRFFAEKHSQLDTSNGCATNFASLQDSGDQLTEEHVTKGES-------------- 89
Query: 87 ECTGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMIS 146
LSITVVGASGDLAKKKIFPALFALFYE CLP++F VFG+ARTK+T EELR+MIS
Sbjct: 90 -----TLSITVVGASGDLAKKKIFPALFALFYEGCLPQDFSVFGYARTKLTHEELRDMIS 144
Query: 147 RTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLS 206
TLTCRID+R C DKM+QFLKRCFYHSG YNSE F++L+ K+KEKE GK SNRL+YLS
Sbjct: 145 STLTCRIDQREKCGDKMEQFLKRCFYHSGQYNSEEDFAELNKKLKEKEAGKISNRLYYLS 204
Query: 207 IPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDH 266
IPPNIFV+VVRCASL+ASS+ GWTRVIVEKPFGRDSESS ELTRCLKQYLTE+QIFRIDH
Sbjct: 205 IPPNIFVDVVRCASLRASSENGWTRVIVEKPFGRDSESSGELTRCLKQYLTEEQIFRIDH 264
Query: 267 YLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQ 326
YLGKELVENLSVLRFSNLVFEPLWSR YIRNVQ IFSEDFGTEGRGGYFD YGIIRDIMQ
Sbjct: 265 YLGKELVENLSVLRFSNLVFEPLWSRNYIRNVQLIFSEDFGTEGRGGYFDQYGIIRDIMQ 324
Query: 327 NHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGY 386
NHLLQILALFAME PVSLDAEDIR+EKVKVLRSM+PL+LE+VV GQYKGH+KGGK+YPGY
Sbjct: 325 NHLLQILALFAMETPVSLDAEDIRSEKVKVLRSMKPLRLEDVVVGQYKGHNKGGKTYPGY 384
Query: 387 ADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYK 446
DDP+VP SLTPTFAAAA+FI+NARWDGVPFLMKAGKALHT+ AEIRVQFRHVPGNLYK
Sbjct: 385 TDDPTVPNHSLTPTFAAAAMFINNARWDGVPFLMKAGKALHTRGAEIRVQFRHVPGNLYK 444
Query: 447 RNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAY 506
++F ++D ATNELV+RVQPDE IYL++NNK+PGLGMRLDRSDLNLLYR+RYP EIPDAY
Sbjct: 445 KSFATNLDNATNELVIRVQPDEGIYLRINNKVPGLGMRLDRSDLNLLYRSRYPREIPDAY 504
Query: 507 ERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAA 566
ERLLLDAIEGERRLFIRSDELDAAW LFTP LKE+E+KKI PELYP+GSRGP+GAHYLA+
Sbjct: 505 ERLLLDAIEGERRLFIRSDELDAAWDLFTPALKELEEKKIIPELYPYGSRGPVGAHYLAS 564
Query: 567 KHNVRWGDTG 576
K+NVRWGD G
Sbjct: 565 KYNVRWGDLG 574
>At5g13110 glucose-6-phosphate 1-dehydrogenase
Length = 596
Score = 832 bits (2149), Expect = 0.0
Identities = 413/533 (77%), Positives = 465/533 (86%), Gaps = 13/533 (2%)
Query: 52 NAVSSHSHDGLAG------SSLTKEDSKPQPVDGPFLSSDSECTGSNLSITVVGASGDLA 105
N+ SS S L G S+L++E +K + SD + + S +SITVVGASGDLA
Sbjct: 70 NSNSSESKTSLKGLKDEVLSALSQEAAKVG------VESDGQ-SQSTVSITVVGASGDLA 122
Query: 106 KKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCRIDKRANCEDKMDQ 165
KKKIFPALFAL+YE CLPE+F +FG++R+KMT ELRNM+S+TLTCRIDKRANC +KM++
Sbjct: 123 KKKIFPALFALYYEGCLPEHFTIFGYSRSKMTDVELRNMVSKTLTCRIDKRANCGEKMEE 182
Query: 166 FLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLKASS 225
FLKRCFYHSG Y+S+ F++LD K+KE E G+ SNRLFYLSIPPNIFV+ V+CAS ASS
Sbjct: 183 FLKRCFYHSGQYDSQEHFTELDKKLKEHEAGRISNRLFYLSIPPNIFVDAVKCASTSASS 242
Query: 226 KKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGKELVENLSVLRFSNLV 285
GWTRVIVEKPFGRDSE+S LT+ LKQYL EDQIFRIDHYLGKELVENLSVLRFSNL+
Sbjct: 243 VNGWTRVIVEKPFGRDSETSAALTKSLKQYLEEDQIFRIDHYLGKELVENLSVLRFSNLI 302
Query: 286 FEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMEPPVSLD 345
FEPLWSR YIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAME PVSLD
Sbjct: 303 FEPLWSRQYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMETPVSLD 362
Query: 346 AEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGSLTPTFAAAA 405
AEDIRNEKVKVLRSMRP+++E+VV GQYK H+KGG +YP Y DD +VPKGSLTPTFAAAA
Sbjct: 363 AEDIRNEKVKVLRSMRPIRVEDVVIGQYKSHTKGGVTYPAYTDDKTVPKGSLTPTFAAAA 422
Query: 406 LFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKATNELVLRVQ 465
LFIDNARWDGVPFLMKAGKALHT+ AEIRVQFRHVPGNLY RN G+D+D+ATNELV+RVQ
Sbjct: 423 LFIDNARWDGVPFLMKAGKALHTRSAEIRVQFRHVPGNLYNRNTGSDLDQATNELVIRVQ 482
Query: 466 PDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFIRSD 525
PDEAIYLK+NNK+PGLGMRLDRS+LNLLY ARY EIPDAYERLLLDAIEGERRLFIRSD
Sbjct: 483 PDEAIYLKINNKVPGLGMRLDRSNLNLLYSARYSKEIPDAYERLLLDAIEGERRLFIRSD 542
Query: 526 ELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTGND 578
ELDAAW+LFTPLLKE+E+KK PE YP+GSRGP+GAHYLAAKH V+WGD D
Sbjct: 543 ELDAAWSLFTPLLKEIEEKKRIPEYYPYGSRGPVGAHYLAAKHKVQWGDVSID 595
>At1g24280 glucose-6-phosphate 1-dehydrogenase
Length = 599
Score = 820 bits (2118), Expect = 0.0
Identities = 400/488 (81%), Positives = 439/488 (88%)
Query: 91 SNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLT 150
S +SITVVGASGDLAKKKIFPALFAL+YE CLPE+F +FG+AR+KMT ELR M+S+TLT
Sbjct: 111 STVSITVVGASGDLAKKKIFPALFALYYEGCLPEHFTIFGYARSKMTDAELRVMVSKTLT 170
Query: 151 CRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPN 210
CRIDKRANC +KM++FLKRCFYHSG Y+S+ F LD K+KE EGG+ SNRLFYLSIPPN
Sbjct: 171 CRIDKRANCGEKMEEFLKRCFYHSGQYDSQEHFVALDEKLKEHEGGRLSNRLFYLSIPPN 230
Query: 211 IFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGK 270
IFV+ V+CAS ASS GWTRVIVEKPFGRDS++S LT+ LKQYL EDQIFRIDHYLGK
Sbjct: 231 IFVDAVKCASSSASSVNGWTRVIVEKPFGRDSKTSAALTKSLKQYLEEDQIFRIDHYLGK 290
Query: 271 ELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLL 330
ELVENLSVLRFSNL+FEPLWSR YIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLL
Sbjct: 291 ELVENLSVLRFSNLIFEPLWSRQYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLL 350
Query: 331 QILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDP 390
QILALFAME PVSLDAEDIRNEKVKVLRSMRP++LE+VV GQYK HS GG +YP Y DD
Sbjct: 351 QILALFAMETPVSLDAEDIRNEKVKVLRSMRPIKLEDVVIGQYKSHSIGGVTYPSYTDDK 410
Query: 391 SVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFG 450
+VPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKAL+T+ AEIRVQFRHVPGNLY RN G
Sbjct: 411 TVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALNTRSAEIRVQFRHVPGNLYNRNSG 470
Query: 451 ADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLL 510
D D+ TNELV+RVQPDEAIYLK+NNK+PGLGMRLD+S+LNLLY ARY EIPDAYERLL
Sbjct: 471 TDRDQTTNELVIRVQPDEAIYLKINNKVPGLGMRLDQSNLNLLYSARYSKEIPDAYERLL 530
Query: 511 LDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNV 570
LDAIEGERRLFIRSDELDAAWALFTPLLKE+E+KK PE YP+GSRGP+GAHYLAAKH V
Sbjct: 531 LDAIEGERRLFIRSDELDAAWALFTPLLKEIEEKKTTPEFYPYGSRGPVGAHYLAAKHKV 590
Query: 571 RWGDTGND 578
+WGD D
Sbjct: 591 QWGDLSLD 598
>At1g09420 putative Glucose-6-phosphate dehydrogenase (F14J9.8)
Length = 625
Score = 538 bits (1387), Expect = e-153
Identities = 264/484 (54%), Positives = 346/484 (70%), Gaps = 11/484 (2%)
Query: 91 SNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLT 150
++L I VVGA+G+LA+ KIFPALFAL+Y LPE+ +FG +R +T E+LR++I+ TLT
Sbjct: 152 ASLCIAVVGATGELARGKIFPALFALYYSGYLPEDVAIFGVSRKNLTDEDLRSIIASTLT 211
Query: 151 CRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPN 210
CR+D + NC KMD F R +Y +G YN+ S L +MK+ EG +NR+FYLS+P
Sbjct: 212 CRVDHQENCGGKMDAFQSRTYYINGGYNNRDGMSRLAERMKQIEGESEANRIFYLSVPQE 271
Query: 211 IFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGK 270
V+V A + +GWTR+IVEKPFG +S SS++LT+ L E QI+RIDH LG+
Sbjct: 272 ALVDVACTIGDNAQAPRGWTRIIVEKPFGFNSHSSHQLTKSLLSKFEEKQIYRIDHMLGR 331
Query: 271 ELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLL 330
L+ENL+VLRFSNLVFEPLW+R YIRN+Q I SE + + D YGIIRDI+ +H+L
Sbjct: 332 NLIENLTVLRFSNLVFEPLWNRTYIRNIQVIISESIAQTEK--FSDGYGIIRDIVHSHIL 389
Query: 331 QILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDP 390
Q +AL AMEPP+SLD EDIRNEKVKVLRS+R + +V+ GQYK S+ D
Sbjct: 390 QTIALLAMEPPISLDGEDIRNEKVKVLRSIRKIDPRDVILGQYKSSSR---------DKN 440
Query: 391 SVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFG 450
V + PT+ AAAL+IDNARWDGVPFL++ G L R EI VQFRHVPGNLY+ N G
Sbjct: 441 GVILNGVDPTYCAAALYIDNARWDGVPFLVRVGTGLIKHRVEIHVQFRHVPGNLYRENIG 500
Query: 451 ADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLL 510
++D TNEL+LR +PDEAI +K+NNK+PGLG++LD S+LNLLY+ RY TE+PD+YE L+
Sbjct: 501 INIDLGTNELILRDEPDEAILVKINNKVPGLGLQLDASELNLLYKDRYKTEVPDSYEHLI 560
Query: 511 LDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNV 570
D I+G+ LF+RSDE+ AAW + +P+L+E++ APELY G RGP+ A+YL AKH V
Sbjct: 561 HDVIDGDNHLFMRSDEVAAAWNILSPVLEEIDKHHTAPELYEFGGRGPVAAYYLWAKHGV 620
Query: 571 RWGD 574
W D
Sbjct: 621 PWAD 624
>At3g27300 glucose-6-phosphate 1-dehydrogenase, putative
Length = 516
Score = 459 bits (1180), Expect = e-129
Identities = 250/489 (51%), Positives = 333/489 (67%), Gaps = 27/489 (5%)
Query: 89 TGSNLSITVVGASGDLAKKKIFPALFALFYEDCL-PENFLVFGFARTKMTSEELRNMISR 147
TGS LSI V+GASGDLAKKK FPALF LF++ L P+ +FG+AR+K+T EELR+ I
Sbjct: 29 TGS-LSIIVLGASGDLAKKKTFPALFNLFHQGFLNPDEVHIFGYARSKITDEELRDKIRG 87
Query: 148 TLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKR-----SNRL 202
L + E + +FLK Y SG Y+SE F LD + E E K+ S RL
Sbjct: 88 YLVDEKNASKKTE-ALSKFLKLIKYVSGPYDSEEGFKRLDKAILEHEISKKTAEGSSRRL 146
Query: 203 FYLSIPPNIFVNVVRCASLKASSKK---GWTRVIVEKPFGRDSESSNELTRCLKQYLTED 259
FYL++PP+++ V + ++K GWTR++VEKPFG+D ES+ +L+ + E
Sbjct: 147 FYLALPPSVYPPVSKMIKAWCTNKSDLGGWTRIVVEKPFGKDLESAEQLSSQIGALFEEP 206
Query: 260 QIFRIDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYG 319
QI+RIDHYLGKELV+N+ VLRF+N +F PLW+R I NVQ +F EDFGTEGRGGYFD YG
Sbjct: 207 QIYRIDHYLGKELVQNMLVLRFANRLFLPLWNRDNIANVQIVFREDFGTEGRGGYFDEYG 266
Query: 320 IIRDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKG 379
IIRDI+QNHLLQ+L L AME P+SL E IR+EKVKVL+S+ P++ E VV GQY+
Sbjct: 267 IIRDIIQNHLLQVLCLVAMEKPISLKPEHIRDEKVKVLQSVIPIKDEEVVLGQYE----- 321
Query: 380 GKSYPGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRH 439
GY DDP+VP S TPTFA L I+N RW+GVPF++KAGKA+ +K+A+IR+QF+
Sbjct: 322 -----GYRDDPTVPNDSNTPTFATTILRINNERWEGVPFILKAGKAMSSKKADIRIQFKD 376
Query: 440 VPGNLYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARY- 498
VPG+++K ++ NE V+R+QP EA+Y+K+ K PGL M+ +S+L+L Y+ RY
Sbjct: 377 VPGDIFK-----CQNQGRNEFVIRLQPSEAMYMKLTVKQPGLEMQTVQSELDLSYKQRYQ 431
Query: 499 PTEIPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGP 558
IP+AYERL+LD I G+++ F+R DEL AAW +FTPLL ++ ++ Y GSRGP
Sbjct: 432 DVSIPEAYERLILDTIRGDQQHFVRRDELKAAWEIFTPLLHRIDKGEVKSVPYKQGSRGP 491
Query: 559 IGAHYLAAK 567
A L K
Sbjct: 492 AEADQLLKK 500
>At5g40760 glucose-6-phosphate dehydrogenase
Length = 515
Score = 448 bits (1153), Expect = e-126
Identities = 242/485 (49%), Positives = 326/485 (66%), Gaps = 27/485 (5%)
Query: 93 LSITVVGASGDLAKKKIFPALFALFYEDCL-PENFLVFGFARTKMTSEELRNMISRTLTC 151
LSI V+GASGDLAKKK FPALF L+ + L P+ +FG+ARTK++ EELR+ I L
Sbjct: 32 LSIIVLGASGDLAKKKTFPALFNLYRQGFLNPDEVHIFGYARTKISDEELRDRIRGYLVD 91
Query: 152 RIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSN-----RLFYLS 206
+K A + + +FL+ Y SG Y++E F LD + E E K S RLFYL+
Sbjct: 92 --EKNAEQAEALSKFLQLIKYVSGPYDAEEGFQRLDKAISEHEISKNSTEGSSRRLFYLA 149
Query: 207 IPPNIFVNV---VRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFR 263
+PP+++ +V ++ + S GWTR++VEKPFG+D ES+ +L+ + + E QI+R
Sbjct: 150 LPPSVYPSVCKMIKTCCMNKSDLGGWTRIVVEKPFGKDLESAEQLSSQIGELFDESQIYR 209
Query: 264 IDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRD 323
IDHYLGKELV+N+ VLRF+N F PLW+R I NVQ +F EDFGTEGRGGYFD YGIIRD
Sbjct: 210 IDHYLGKELVQNMLVLRFANRFFLPLWNRDNIENVQIVFREDFGTEGRGGYFDEYGIIRD 269
Query: 324 IMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSY 383
I+QNHLLQ+L L AME P+SL E IR+EKVKVL+S+ P+ + VV GQY+
Sbjct: 270 IIQNHLLQVLCLVAMEKPISLKPEHIRDEKVKVLQSVVPISDDEVVLGQYE--------- 320
Query: 384 PGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGN 443
GY DD +VP S TPTFA L I N RW+GVPF++KAGKAL++++AEIR+QF+ VPG+
Sbjct: 321 -GYRDDDTVPNDSNTPTFATTILRIHNERWEGVPFILKAGKALNSRKAEIRIQFKDVPGD 379
Query: 444 LYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYP-TEI 502
+++ + NE V+R+QP EA+Y+K+ K PGL M +S+L+L Y RY I
Sbjct: 380 IFR-----CQKQGRNEFVIRLQPSEAMYMKLTVKQPGLDMNTVQSELDLSYGQRYQGVAI 434
Query: 503 PDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAH 562
P+AYERL+LD I+G+++ F+R DEL AW +FTPLL ++ ++ Y GSRGP A
Sbjct: 435 PEAYERLILDTIKGDQQHFVRRDELKVAWEIFTPLLHRIDKGEVKSIPYKPGSRGPKEAD 494
Query: 563 YLAAK 567
L K
Sbjct: 495 QLLEK 499
>At4g19470 putative protein
Length = 417
Score = 31.6 bits (70), Expect = 1.3
Identities = 44/181 (24%), Positives = 72/181 (39%), Gaps = 19/181 (10%)
Query: 125 NFLVFGFARTKMTSEELRNMISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFS 184
N +F F +K+ ++ R+M C + K + + C + + N GS
Sbjct: 17 NLKLFSFGGSKV--QDFRDMY--LTDCNLYKFPDNFSCLSSLQSLCLSRNSIENLPGSIK 72
Query: 185 DLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLKASSKKGWTRVIVEK-------- 236
L +K N + +P N +++V C SL+ SK VI EK
Sbjct: 73 KLH-HLKSLYLKNCKNLISLPVLPSNQYLDVHGCISLETVSKPMTLLVIAEKTHSTFVFT 131
Query: 237 ---PFGRDSESSNELTRCLKQYLTEDQIFRIDHYL-GKELV-ENLSVLRFSNLVFEPLWS 291
RD++ LK + ++ F+++H + ELV E LS + F PLW
Sbjct: 132 DCYKLNRDAQEKIVAHTQLKSQILANRSFQLNHKVQSLELVLEPLSAVSFPGNDL-PLWF 190
Query: 292 R 292
R
Sbjct: 191 R 191
>At1g28210 AtJ1 (atj)
Length = 427
Score = 31.6 bits (70), Expect = 1.3
Identities = 18/67 (26%), Positives = 33/67 (48%), Gaps = 2/67 (2%)
Query: 465 QPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFIRS 524
QPD+ + L+ +P G +D D + +R +PTE+ + +R +L+ E S
Sbjct: 336 QPDQLLVLR-GKGLPKQGFFVDHGDQYVRFRVNFPTEVNER-QRAILEEFAKEEINNELS 393
Query: 525 DELDAAW 531
D + +W
Sbjct: 394 DSAEGSW 400
>At4g13550 putative protein
Length = 805
Score = 29.6 bits (65), Expect = 4.8
Identities = 28/127 (22%), Positives = 51/127 (40%), Gaps = 5/127 (3%)
Query: 126 FLVFGFARTKMTSEE-LRNMISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSE---- 180
F+ + F + K ++ L+N T + A+ ED DQ SG + +
Sbjct: 254 FVEYAFGQLKSLNDAPLKNTELLNNTAEDSEGASSEDSSDQHRSTNLSSSGKLSKDKDGD 313
Query: 181 GSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGR 240
G + + + G +S F+ +IP + N+V+ L + K W + + FG
Sbjct: 314 GDGHGNELEDDNESGSIQSESNFWDNIPDIVGQNIVQKLGLPSPEKLKWNGTELLENFGL 373
Query: 241 DSESSNE 247
S + E
Sbjct: 374 QSRKTAE 380
>At5g39460 hypothetical protein
Length = 571
Score = 29.3 bits (64), Expect = 6.3
Identities = 23/81 (28%), Positives = 36/81 (44%), Gaps = 5/81 (6%)
Query: 153 IDKRANCEDKMDQFLK----RCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIP 208
ID + ED+++ + R F SG+ + G FS +RSNR +LS
Sbjct: 295 IDNMEDIEDQIEVTPRKKSFRRFLRSGIKHILGKFSSSKINSPSSSETRRSNRQSFLS-S 353
Query: 209 PNIFVNVVRCASLKASSKKGW 229
N F ++ + SS +GW
Sbjct: 354 GNTFCLSLKASCTLMSSYEGW 374
>At5g47990 cytochrome P450
Length = 511
Score = 28.9 bits (63), Expect = 8.2
Identities = 18/41 (43%), Positives = 24/41 (57%), Gaps = 2/41 (4%)
Query: 62 LAGSSLTKEDSKPQPVDG--PFLSSDSECTGSNLSITVVGA 100
LA S L +ED K + + PF S C GS+L+ TVVG+
Sbjct: 426 LASSRLGEEDEKREDMLKYIPFGSGRRACPGSHLAYTVVGS 466
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,624,987
Number of Sequences: 26719
Number of extensions: 611495
Number of successful extensions: 1362
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 1341
Number of HSP's gapped (non-prelim): 11
length of query: 579
length of database: 11,318,596
effective HSP length: 105
effective length of query: 474
effective length of database: 8,513,101
effective search space: 4035209874
effective search space used: 4035209874
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0213.11


