

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.1
(305 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
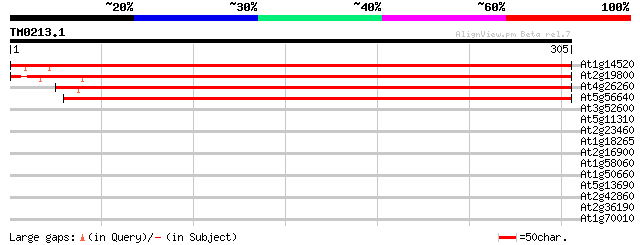
Score E
Sequences producing significant alignments: (bits) Value
At1g14520 unknown protein 504 e-143
At2g19800 unknown protein 479 e-136
At4g26260 unknown protein 471 e-133
At5g56640 unknown protein 454 e-128
At3g52600 cell wall invertase 33 0.19
At5g11310 putative protein 30 2.1
At2g23460 extra-large GTP-binding protein (XLG1) 29 3.6
At1g18265 unknown protein 28 4.7
At2g16900 unknown protein 28 6.2
At1g58060 hypothetical protein 28 6.2
At1g50660 unknown protein 28 6.2
At5g13690 alpha-N-acetylglucosaminidase 28 8.1
At2g42860 unknown protein 28 8.1
At2g36190 beta-fructofuranosidase like protein (invertase) 28 8.1
At1g70010 hypothetical protein 28 8.1
>At1g14520 unknown protein
Length = 311
Score = 504 bits (1298), Expect = e-143
Identities = 239/311 (76%), Positives = 269/311 (85%), Gaps = 6/311 (1%)
Query: 1 MTIIIDQ---PDHFGSDVEGKTV---PIDEKELVLDGGFVMPHTNSFGHTFRDYNAESER 54
MTI+ID+ + G ++ K +E ELVLD GF PHTNSFG TFRDY+AESER
Sbjct: 1 MTILIDRHSDQNDAGDEIVEKNQGNGKEEETELVLDAGFEAPHTNSFGRTFRDYDAESER 60
Query: 55 QEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQI 114
+ GVE FYR NHI Q+VDFV+KMREEY KLNR EMS+WECCELLNE +DESDPDLDEPQI
Sbjct: 61 RRGVEEFYRVNHIGQTVDFVRKMREEYEKLNRTEMSIWECCELLNEFIDESDPDLDEPQI 120
Query: 115 EHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDES 174
EHLLQTAEAIRKDYP+EDWLHLTGLIHDLGKVLL FG LPQWAVVGDTFPVGC FDES
Sbjct: 121 EHLLQTAEAIRKDYPDEDWLHLTGLIHDLGKVLLHSSFGELPQWAVVGDTFPVGCAFDES 180
Query: 175 IVHHKHFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAAL 234
IVHHK+FKENPDY+NP+YN++YG+Y+E CGL+NVLMSWGHDDYMYLVAKEN+TTLPSA L
Sbjct: 181 IVHHKYFKENPDYDNPSYNSKYGIYTEGCGLDNVLMSWGHDDYMYLVAKENQTTLPSAGL 240
Query: 235 FIIRYHSFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSL 294
FIIRYHSFYALH+ AYKHLMN ED EN++WL +FNKYDLYSKSKVR++VE+VKPYYLSL
Sbjct: 241 FIIRYHSFYALHKSEAYKHLMNNEDRENMKWLKVFNKYDLYSKSKVRVNVEEVKPYYLSL 300
Query: 295 IEKYFPAKLKW 305
KYFP+KLKW
Sbjct: 301 TNKYFPSKLKW 311
>At2g19800 unknown protein
Length = 317
Score = 479 bits (1233), Expect = e-136
Identities = 229/320 (71%), Positives = 264/320 (81%), Gaps = 18/320 (5%)
Query: 1 MTIIIDQPDHFGSDV---EGKTVPIDEKELVLDGGFVMPHT-----------NSFGHTFR 46
MTI+++ HF D E K + + ELVLDGGFV+P + N GH+FR
Sbjct: 1 MTILVE---HFVPDSRVDEKKVIEERDNELVLDGGFVVPKSKETDAFDAPDMNFLGHSFR 57
Query: 47 DY-NAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDES 105
DY N ESERQ+GVE FYR HI+Q+ DFVKKMR+EYGKLN++EMS+WECCELLN VVDES
Sbjct: 58 DYENGESERQQGVEEFYRMQHIHQTYDFVKKMRKEYGKLNKMEMSIWECCELLNNVVDES 117
Query: 106 DPDLDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTF 165
DPDLDEPQI+HLLQTAEAIR+DYP+EDWLHLT LIHDLGKVLLLP FGGLPQWAVVGDTF
Sbjct: 118 DPDLDEPQIQHLLQTAEAIRRDYPDEDWLHLTALIHDLGKVLLLPEFGGLPQWAVVGDTF 177
Query: 166 PVGCRFDESIVHHKHFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKEN 225
PVGC FD + +HHK+FK N D NNP YNT+ G+Y+E CGL+NVLMSWGHDDYMYLVAK+N
Sbjct: 178 PVGCTFDSANIHHKYFKGNHDINNPKYNTKNGVYTEGCGLDNVLMSWGHDDYMYLVAKKN 237
Query: 226 KTTLPSAALFIIRYHSFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVE 285
TTLP A LFIIRYHSFY LH+ GAY HLMN+ED ++L+WLH+FNKYDLYSKSKV +DVE
Sbjct: 238 GTTLPHAGLFIIRYHSFYPLHKAGAYTHLMNDEDRDDLKWLHVFNKYDLYSKSKVLVDVE 297
Query: 286 KVKPYYLSLIEKYFPAKLKW 305
+VKPYY+SLI KYFPAKLKW
Sbjct: 298 QVKPYYISLINKYFPAKLKW 317
>At4g26260 unknown protein
Length = 317
Score = 471 bits (1213), Expect = e-133
Identities = 221/293 (75%), Positives = 245/293 (83%), Gaps = 13/293 (4%)
Query: 26 ELVLDGGFVMP-------------HTNSFGHTFRDYNAESERQEGVENFYRKNHIYQSVD 72
EL LDGGF MP N+FG FRDY+ ESERQ+GVE FYR HI Q+VD
Sbjct: 25 ELKLDGGFSMPKMDTNDDEAFLAPEMNAFGRQFRDYDVESERQKGVEEFYRLQHINQTVD 84
Query: 73 FVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHLLQTAEAIRKDYPNED 132
FVKKMR EYGKL+++ MS+WECCELLNEVVDESDPDLDEPQI+HLLQ+AEAIRKDYPNED
Sbjct: 85 FVKKMRAEYGKLDKMVMSIWECCELLNEVVDESDPDLDEPQIQHLLQSAEAIRKDYPNED 144
Query: 133 WLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIVHHKHFKENPDYNNPAY 192
WLHLT LIHDLGKV+ LP FGGLPQWAVVGDTFPVGC FDES VHHK+F ENPD++N Y
Sbjct: 145 WLHLTALIHDLGKVITLPQFGGLPQWAVVGDTFPVGCAFDESNVHHKYFVENPDFHNETY 204
Query: 193 NTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYHSFYALHREGAYK 252
NT+ G+YSE CGLNNV+MSWGHDDYMYLVAKEN +TLPSA FIIRYHSFY LH G Y
Sbjct: 205 NTKNGIYSEGCGLNNVMMSWGHDDYMYLVAKENGSTLPSAGQFIIRYHSFYPLHTAGEYT 264
Query: 253 HLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFPAKLKW 305
HLMNEED ENL+WLH+FNKYDLYSKSKV +DVEKVKPYY+SLI+KYFP L+W
Sbjct: 265 HLMNEEDKENLKWLHVFNKYDLYSKSKVHVDVEKVKPYYMSLIKKYFPENLRW 317
>At5g56640 unknown protein
Length = 314
Score = 454 bits (1168), Expect = e-128
Identities = 209/277 (75%), Positives = 237/277 (85%), Gaps = 1/277 (0%)
Query: 30 DGGFVMPHTNSFGHTFRDY-NAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVE 88
D F+ P N+FG FRDY + SERQ+ VE+FY H Q++DFV+KMR EYGKL+++
Sbjct: 38 DEVFLAPEMNAFGRQFRDYTDTNSERQKSVEHFYATQHTNQTLDFVQKMRSEYGKLDKMV 97
Query: 89 MSMWECCELLNEVVDESDPDLDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLL 148
M++WECCEL EVVDESDPDLDEPQI+HLLQ+AEAIRKDYPNEDWLHLT LIHDLGKVL
Sbjct: 98 MNIWECCELSKEVVDESDPDLDEPQIQHLLQSAEAIRKDYPNEDWLHLTALIHDLGKVLT 157
Query: 149 LPVFGGLPQWAVVGDTFPVGCRFDESIVHHKHFKENPDYNNPAYNTRYGMYSEKCGLNNV 208
LP FGGLPQWAVVGDTFPVGC FDES VHHK+F ENPD+NNP YNT+ G+YSE CGL NV
Sbjct: 158 LPQFGGLPQWAVVGDTFPVGCAFDESNVHHKYFMENPDFNNPKYNTKAGIYSEGCGLENV 217
Query: 209 LMSWGHDDYMYLVAKENKTTLPSAALFIIRYHSFYALHREGAYKHLMNEEDFENLEWLHI 268
LMSWGHDDYMYLVAKEN +TLPS LFIIRYHSFY LH+ GAY HLMNEED ENL+WLH+
Sbjct: 218 LMSWGHDDYMYLVAKENGSTLPSPGLFIIRYHSFYPLHKAGAYTHLMNEEDKENLKWLHV 277
Query: 269 FNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFPAKLKW 305
FNKYDLYSKSKV ++VEKVKPYY+SLI+KYFP L+W
Sbjct: 278 FNKYDLYSKSKVHVNVEKVKPYYMSLIKKYFPENLRW 314
>At3g52600 cell wall invertase
Length = 590
Score = 33.1 bits (74), Expect = 0.19
Identities = 27/105 (25%), Positives = 44/105 (41%), Gaps = 13/105 (12%)
Query: 30 DGGFVMPHTNSFGHTFRD-----YNAESERQEGVENFYRKN----HIYQSVDFVKKMREE 80
D V P G FRD +N + + RKN ++Y+S DF K ++ +
Sbjct: 169 DNPIVKPDNGENGSAFRDPTTAWFNKKDGYWRMLVGSKRKNRGIAYMYKSRDFKKWVKSK 228
Query: 81 YGKLNRVEMSMWECCELLNEVVDES----DPDLDEPQIEHLLQTA 121
+R + MWEC + V + D D P +H+L+ +
Sbjct: 229 RPIHSRKKTGMWECPDFFPVSVTDKKNGLDFSYDGPNAKHVLKVS 273
>At5g11310 putative protein
Length = 602
Score = 29.6 bits (65), Expect = 2.1
Identities = 29/113 (25%), Positives = 52/113 (45%), Gaps = 12/113 (10%)
Query: 34 VMPHTNSFGHTFRDYNAESERQEGVENFYRKNHIYQSVD-----FVKKMREEYGKLNRVE 88
V P T ++ H F+ ++ ++ +EG+ +++ S D + KM E GKL+
Sbjct: 386 VDPTTTTYNHFFKYFSKHNKTEEGMNLYFKLIEAGHSPDRLTYHLILKMLCEDGKLS--- 442
Query: 89 MSMWECCELLNEVVDESDPD-LDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLI 140
++M E+ N + DPD L + HLL E + + + D G+I
Sbjct: 443 LAMQVNKEMKNRGI---DPDLLTTTMLIHLLCRLEMLEEAFEEFDNAVRRGII 492
>At2g23460 extra-large GTP-binding protein (XLG1)
Length = 888
Score = 28.9 bits (63), Expect = 3.6
Identities = 15/62 (24%), Positives = 31/62 (49%), Gaps = 9/62 (14%)
Query: 234 LFIIRYHSFYALHREGAYKHLMNEE---------DFENLEWLHIFNKYDLYSKSKVRIDV 284
+F++ + + +G K L+ ++ FEN+++L I NKYDL + R+ +
Sbjct: 729 VFVVSMSDYDQVSEDGTNKMLLTKKLFESIITHPIFENMDFLLILNKYDLLEEKVERVPL 788
Query: 285 EK 286
+
Sbjct: 789 AR 790
>At1g18265 unknown protein
Length = 280
Score = 28.5 bits (62), Expect = 4.7
Identities = 16/60 (26%), Positives = 26/60 (42%), Gaps = 1/60 (1%)
Query: 39 NSFGHTFRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELL 98
NSF +D N + F K+H+ +D K + + + +S WE C+LL
Sbjct: 79 NSFNREHKDCNNNNNNNNNNNGFISKSHLADGLDQKKPIILHWSSDSTNRLS-WENCDLL 137
>At2g16900 unknown protein
Length = 382
Score = 28.1 bits (61), Expect = 6.2
Identities = 25/99 (25%), Positives = 39/99 (39%), Gaps = 10/99 (10%)
Query: 12 GSDVEGKTVPIDEKELV----------LDGGFVMPHTNSFGHTFRDYNAESERQEGVENF 61
GS G+ + KE+V +D G++ F + ++E ERQEG+
Sbjct: 260 GSTPVGQLTELKVKEMVAVLKDLESVNIDVGWMRSVLEEFAQYQENTDSEKERQEGLVRS 319
Query: 62 YRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNE 100
++ Q D + +E RVE E EL E
Sbjct: 320 KKQEMEIQEADLARIEKEVAEARLRVEEMKAELAELETE 358
>At1g58060 hypothetical protein
Length = 1389
Score = 28.1 bits (61), Expect = 6.2
Identities = 22/73 (30%), Positives = 32/73 (43%), Gaps = 7/73 (9%)
Query: 152 FGGLPQWAVVGDTFPVGCRFDESIVHHKHFKENPD-----YNNPAYNTRYGMYSEKCGLN 206
FG P G T PV F E I ++ PD ++ + + G +++ G
Sbjct: 806 FGHCPVITAQGRTHPVTTHFLEEIYESINYLLAPDSPAALRSDTSIKDKLGSVNDRRGKK 865
Query: 207 N-VLMSWGHDDYM 218
N VL WG DDY+
Sbjct: 866 NLVLAGWG-DDYL 877
>At1g50660 unknown protein
Length = 725
Score = 28.1 bits (61), Expect = 6.2
Identities = 21/87 (24%), Positives = 43/87 (49%), Gaps = 7/87 (8%)
Query: 47 DYNAESERQEGVENFYRK--NHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDE 104
D N E + ++ +E K N + S VK+ ++Y K + + E C+ L + + E
Sbjct: 299 DMNREKKTRQRLEIVNHKLVNELADSKLAVKRYMQDYEKERKARELIEEVCDELAKEIGE 358
Query: 105 SDPDLDEPQIEHLLQTAEAIRKDYPNE 131
D+ +IE L + + ++R++ +E
Sbjct: 359 -----DKAEIEALKRESMSLREEVDDE 380
>At5g13690 alpha-N-acetylglucosaminidase
Length = 806
Score = 27.7 bits (60), Expect = 8.1
Identities = 26/107 (24%), Positives = 46/107 (42%), Gaps = 30/107 (28%)
Query: 73 FVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHLLQTA--EAIRKDYPN 130
F+K+ EEYG++ + + C + +E+ P EP+ L A +A+ K N
Sbjct: 324 FIKQQTEEYGEITNI----YNC-----DTFNENTPPTSEPEYISSLGAAVYKAMSKGNKN 374
Query: 131 EDWL----------------HLTGLIHDL--GKVLLLPVFGGL-PQW 158
WL L L+H + GK+++L ++ + P W
Sbjct: 375 AVWLMQGWLFSSDSKFWKPPQLKALLHSVPFGKMIVLDLYAEVKPIW 421
>At2g42860 unknown protein
Length = 224
Score = 27.7 bits (60), Expect = 8.1
Identities = 21/63 (33%), Positives = 27/63 (42%), Gaps = 13/63 (20%)
Query: 4 IIDQPDHFGSDVEGKTVPIDEKELVLDGGFVMPHTNSFGHTFRDYNAESERQEGVENFYR 63
I DQ D S EG T P + +V S H F + + E E Q V F+R
Sbjct: 35 IADQVDPVDSTEEGSTGPSNSHTVV-----------SKDHDFSESDMEDEEQ--VLEFFR 81
Query: 64 KNH 66
+NH
Sbjct: 82 RNH 84
>At2g36190 beta-fructofuranosidase like protein (invertase)
Length = 591
Score = 27.7 bits (60), Expect = 8.1
Identities = 25/104 (24%), Positives = 43/104 (41%), Gaps = 12/104 (11%)
Query: 30 DGGFVMPHTNSFGHTFRDYNAESERQEG----VENFYRKN----HIYQSVDFVKKMREEY 81
D +P G FRD ++G V RK +IY+S DF ++ ++
Sbjct: 171 DNPIAIPDYTMNGSAFRDPTTAWFSKDGHWRTVVGSKRKRRGIAYIYRSRDFKHWVKAKH 230
Query: 82 GKLNRVEMSMWECCEL----LNEVVDESDPDLDEPQIEHLLQTA 121
++ MWEC + L + + D D P +H+L+ +
Sbjct: 231 PVHSKQSTGMWECPDFFPVSLTDFRNGLDLDYVGPNTKHVLKVS 274
>At1g70010 hypothetical protein
Length = 1315
Score = 27.7 bits (60), Expect = 8.1
Identities = 20/88 (22%), Positives = 40/88 (44%), Gaps = 9/88 (10%)
Query: 217 YMYLVAKENKTTLPSAALFIIRYHSFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYS 276
++ V+ NK++ P + I + F +G L +D+ W+++ L +
Sbjct: 449 HLPFVSHNNKSSRPFDLIHIDTWGPFSVQTHDGYRYFLTIVDDYSRATWVYL-----LRN 503
Query: 277 KSKVRIDVEKVKPYYLSLIEKYFPAKLK 304
KS DV V P +++++E F +K
Sbjct: 504 KS----DVLTVIPTFVTMVENQFETTIK 527
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.139 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,842,554
Number of Sequences: 26719
Number of extensions: 364076
Number of successful extensions: 836
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 826
Number of HSP's gapped (non-prelim): 15
length of query: 305
length of database: 11,318,596
effective HSP length: 99
effective length of query: 206
effective length of database: 8,673,415
effective search space: 1786723490
effective search space used: 1786723490
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0213.1


