

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0207.3
(227 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
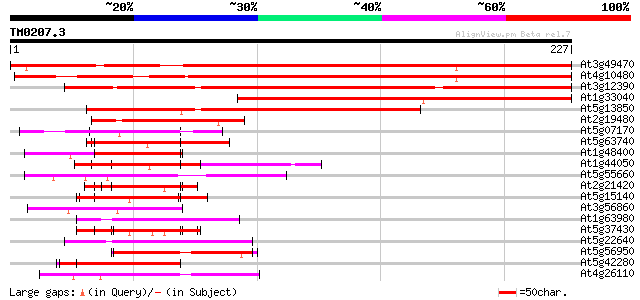
Score E
Sequences producing significant alignments: (bits) Value
At3g49470 alpha NAC-like protein 278 2e-75
At4g10480 putative alpha NAC 260 5e-70
At3g12390 unknown protein 243 5e-65
At1g33040 alpha NAC-like protein, putative 185 1e-47
At5g13850 unknown protein 171 2e-43
At2g19480 putative nucleosome assembly protein 55 4e-08
At5g07170 putative protein 54 9e-08
At5g63740 unknown protein 51 4e-07
At1g48400 50 1e-06
At1g44050 unknown protein 49 2e-06
At5g55660 putative protein 47 6e-06
At2g21420 Mutator-like transposase 47 6e-06
At5g15140 putative aldose 1-epimerase - like protein 47 8e-06
At3g56860 unknown protein 47 8e-06
At1g63980 unknown protein 45 2e-05
At5g37430 contains similarity to unknown protein (pir||C71432) 45 3e-05
At5g22640 unknown protein 45 3e-05
At5g56950 nucleosome assembly protein 45 4e-05
At5g42280 putative protein 45 4e-05
At4g26110 nucleosome assembly protein I-like protein 45 4e-05
>At3g49470 alpha NAC-like protein
Length = 217
Score = 278 bits (710), Expect = 2e-75
Identities = 156/229 (68%), Positives = 179/229 (78%), Gaps = 14/229 (6%)
Query: 1 MAPGP-VVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDED 59
M+P P VV S++ + + ++A L+KK Q E PIVEDVKDD+ D+ DD+E+ED
Sbjct: 1 MSPPPAVVTESADGQPEQPPVTAIAEELEKKLQTDE---PIVEDVKDDEDDDDDDEEEED 57
Query: 60 DDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISK 119
DD A G S SKQSRSEKKSRKAMLKLG+KPVTGVSRVTIKRTKN+LFFISK
Sbjct: 58 DD---------AQGVSGSSKQSRSEKKSRKAMLKLGMKPVTGVSRVTIKRTKNVLFFISK 108
Query: 120 PDVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQPEE 179
PDVFKSP+SETYVIFGEAKIEDLSSQLQTQAAQQFRMP++G+ + + TA Q EE
Sbjct: 109 PDVFKSPHSETYVIFGEAKIEDLSSQLQTQAAQQFRMPEIGATSQRAEASTATVEAQVEE 168
Query: 180 -EEEVDETGVEPHDIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELTT 227
EEE+DETGVE DIDLVMTQAGVSR KAVKALK+H+GDIV AIMELTT
Sbjct: 169 DEEEIDETGVEARDIDLVMTQAGVSRSKAVKALKSHDGDIVSAIMELTT 217
>At4g10480 putative alpha NAC
Length = 212
Score = 260 bits (664), Expect = 5e-70
Identities = 144/226 (63%), Positives = 174/226 (76%), Gaps = 16/226 (7%)
Query: 3 PGPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDD 62
PGPV++ +E ++ + K++ + Q+++ +VEDVKD D+D D+D DD
Sbjct: 2 PGPVIEEVNEEALMDAI--------KEQMKLQKENDVVVEDVKDGDED------DDDVDD 47
Query: 63 EDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKPDV 122
+DD+ DGA G +E SKQSRSEKKSRKAMLKLG+KPVT VSRVTIKR+KN+LF ISKPDV
Sbjct: 48 DDDEIADGA-GENEASKQSRSEKKSRKAMLKLGMKPVTDVSRVTIKRSKNVLFVISKPDV 106
Query: 123 FKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQPEE-EE 181
FKSPNSETYVIFGEAKI+D+SSQLQ QAAQ+F+MPD+ S++ D AA Q EE +E
Sbjct: 107 FKSPNSETYVIFGEAKIDDMSSQLQAQAAQRFKMPDVASMIPNTDGSEAATVAQEEEDDE 166
Query: 182 EVDETGVEPHDIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELTT 227
+VDETGVE DIDLVMTQAGVSR KAVKALK NGDIV AIMELTT
Sbjct: 167 DVDETGVEAKDIDLVMTQAGVSRPKAVKALKESNGDIVSAIMELTT 212
>At3g12390 unknown protein
Length = 203
Score = 243 bits (621), Expect = 5e-65
Identities = 132/205 (64%), Positives = 162/205 (78%), Gaps = 6/205 (2%)
Query: 23 EETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSR 82
E+ L K + Q+ D E V+DDD +E DD +D+D DD++ D DG GG SKQSR
Sbjct: 5 EKEILAAKLEEQKIDLDKPE-VEDDDDNEDDDSDDDDKDDDEADGLDGEAGGK--SKQSR 61
Query: 83 SEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKPDVFKSPNSETYVIFGEAKIEDL 142
SEKKSRKAMLKLG+KP+TGVSRVT+K++KNILF ISKPDVFKSP S+TYVIFGEAKIEDL
Sbjct: 62 SEKKSRKAMLKLGMKPITGVSRVTVKKSKNILFVISKPDVFKSPASDTYVIFGEAKIEDL 121
Query: 143 SSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQPEEEEEVDETGVEPHDIDLVMTQAGV 202
SSQ+Q+QAA+QF+ PD+ +V++K + +AA +++EEVDE GVEP DI+LVMTQAGV
Sbjct: 122 SSQIQSQAAEQFKAPDLSNVISKGESSSAA---VVQDDEEVDEEGVEPKDIELVMTQAGV 178
Query: 203 SRGKAVKALKTHNGDIVGAIMELTT 227
SR AVKALK +GDIV AIMELTT
Sbjct: 179 SRPNAVKALKAADGDIVSAIMELTT 203
>At1g33040 alpha NAC-like protein, putative
Length = 136
Score = 185 bits (470), Expect = 1e-47
Identities = 94/136 (69%), Positives = 115/136 (84%), Gaps = 1/136 (0%)
Query: 93 KLGLKPVTGVSRVTIKRTKNILFFISKPDVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQ 152
KLG+KPV+ VSRVTIKR KN+LF ISKPDV+KSPN+ETYVIFGEAK++DLSSQLQTQAAQ
Sbjct: 1 KLGMKPVSDVSRVTIKRAKNVLFVISKPDVYKSPNAETYVIFGEAKVDDLSSQLQTQAAQ 60
Query: 153 QFRMPDMGSVMAKQ-DQGTAADGGQPEEEEEVDETGVEPHDIDLVMTQAGVSRGKAVKAL 211
+F+MPD+ S++ + T A + E+E++VD+TGVE DIDLVMTQAGVS+ KAV AL
Sbjct: 61 RFKMPDVTSMLPNAGSEATMAPLAEEEDEDDVDDTGVEARDIDLVMTQAGVSKAKAVSAL 120
Query: 212 KTHNGDIVGAIMELTT 227
K ++GDIV AIMELTT
Sbjct: 121 KANDGDIVSAIMELTT 136
>At5g13850 unknown protein
Length = 159
Score = 171 bits (434), Expect = 2e-43
Identities = 90/139 (64%), Positives = 111/139 (79%), Gaps = 6/139 (4%)
Query: 32 QPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKE----DGALGGSEGSKQSRSEKKS 87
Q E A + E D DK E +DD+D D+DD +DD E DG GG SKQSRSEKKS
Sbjct: 5 QKVELAAKLEEQKIDLDKPEVEDDDDNDEDDSEDDDEAEGHDGEAGGR--SKQSRSEKKS 62
Query: 88 RKAMLKLGLKPVTGVSRVTIKRTKNILFFISKPDVFKSPNSETYVIFGEAKIEDLSSQLQ 147
RKAMLKLG+KP+TGVSRVT+K++KNILF ISKPDVFKSP S+TYVIFGEAKIEDLSSQLQ
Sbjct: 63 RKAMLKLGMKPITGVSRVTVKKSKNILFVISKPDVFKSPASDTYVIFGEAKIEDLSSQLQ 122
Query: 148 TQAAQQFRMPDMGSVMAKQ 166
+QAA+QF+ P++ +V++++
Sbjct: 123 SQAAEQFKAPNLSNVISQE 141
>At2g19480 putative nucleosome assembly protein
Length = 379
Score = 54.7 bits (130), Expect = 4e-08
Identities = 28/63 (44%), Positives = 44/63 (69%), Gaps = 3/63 (4%)
Query: 34 QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRS-EKKSRKAML 92
++DD I ED DD++DE DD++DE++DDEDDD+E+ A G + K+S + KK+ ++ L
Sbjct: 307 EDDDDEIDED--DDEEDEEDDEDDEEEDDEDDDEEEEADQGKKSKKKSSAGHKKAGRSQL 364
Query: 93 KLG 95
G
Sbjct: 365 AEG 367
Score = 44.3 bits (103), Expect = 5e-05
Identities = 22/62 (35%), Positives = 37/62 (59%), Gaps = 1/62 (1%)
Query: 34 QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLK 93
+ DD I +D + D+D+ ++DE++D+DDE++D ED E + +S+KKS K
Sbjct: 300 EADDLDIEDDDDEIDEDDDEEDEEDDEDDEEEDDEDDD-EEEEADQGKKSKKKSSAGHKK 358
Query: 94 LG 95
G
Sbjct: 359 AG 360
Score = 37.0 bits (84), Expect = 0.009
Identities = 20/54 (37%), Positives = 30/54 (55%), Gaps = 6/54 (11%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSE------KKSRK 89
E V+ DD D DDD++ D+DD+++D+ED E + E KKS+K
Sbjct: 297 EAVEADDLDIEDDDDEIDEDDDEEDEEDDEDDEEEDDEDDDEEEEADQGKKSKK 350
Score = 33.9 bits (76), Expect = 0.072
Identities = 13/67 (19%), Positives = 35/67 (51%)
Query: 18 QVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEG 77
+ + A++ ++ ++D ++ D+D +E DD++D+++++ D K+ +
Sbjct: 297 EAVEADDLDIEDDDDEIDEDDDEEDEEDDEDDEEEDDEDDDEEEEADQGKKSKKKSSAGH 356
Query: 78 SKQSRSE 84
K RS+
Sbjct: 357 KKAGRSQ 363
>At5g07170 putative protein
Length = 542
Score = 53.5 bits (127), Expect = 9e-08
Identities = 30/82 (36%), Positives = 45/82 (54%), Gaps = 13/82 (15%)
Query: 5 PVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDED 64
P++D SS E E+ + +DDA +D +DDD D+ DDD+D+DDDD+D
Sbjct: 85 PLIDDSSSDE--------EDDSESTHCYAADDDADDTDDDEDDDDDDDDDDDDDDDDDDD 136
Query: 65 DDKEDGALGGSEGSKQSRSEKK 86
DD +D + SK S E++
Sbjct: 137 DDDDD-----DDESKDSEVEEE 153
Score = 42.0 bits (97), Expect = 3e-04
Identities = 21/52 (40%), Positives = 27/52 (51%), Gaps = 16/52 (30%)
Query: 33 PQEDDAPIVED---------------VKDDDKDEADDDEDEDDDDEDDDKED 69
P D P+++D DDD D+ DDDED+DDDD DDD +D
Sbjct: 79 PSSQDDPLIDDSSSDEEDDSESTHCYAADDDADDTDDDEDDDDDD-DDDDDD 129
>At5g63740 unknown protein
Length = 226
Score = 51.2 bits (121), Expect = 4e-07
Identities = 23/55 (41%), Positives = 34/55 (61%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRK 89
EDD +D DDD D+ADDDED+DD+D+D+D++D E ++ E S +
Sbjct: 89 EDDDDDDDDDDDDDADDADDDEDDDDEDDDEDEDDDDDDDDENDEECDDEYDSHR 143
Score = 46.6 bits (109), Expect = 1e-05
Identities = 20/39 (51%), Positives = 30/39 (76%), Gaps = 1/39 (2%)
Query: 32 QPQEDDAPIVEDVKDDDKDEADD-DEDEDDDDEDDDKED 69
+ +++D +D DDD D+ADD D+DEDDDDEDDD+++
Sbjct: 83 EDEDEDEDDDDDDDDDDDDDADDADDDEDDDDEDDDEDE 121
Score = 46.2 bits (108), Expect = 1e-05
Identities = 20/36 (55%), Positives = 25/36 (68%)
Query: 34 QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+++D ED DDD D+ DDD D+ DDDEDDD ED
Sbjct: 81 EDEDEDEDEDDDDDDDDDDDDDADDADDDEDDDDED 116
Score = 44.7 bits (104), Expect = 4e-05
Identities = 17/35 (48%), Positives = 27/35 (76%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
ED+ + +D+D+DE +DD+D+DDDD+DDD +D
Sbjct: 71 EDEDEDADADEDEDEDEDEDDDDDDDDDDDDDADD 105
Score = 43.9 bits (102), Expect = 7e-05
Identities = 16/38 (42%), Positives = 28/38 (73%)
Query: 32 QPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+ +++DA ED +D+ ++ DDD+D+DDDD+ DD +D
Sbjct: 71 EDEDEDADADEDEDEDEDEDDDDDDDDDDDDDADDADD 108
Score = 38.9 bits (89), Expect = 0.002
Identities = 16/27 (59%), Positives = 23/27 (84%), Gaps = 3/27 (11%)
Query: 46 DDDKDE---ADDDEDEDDDDEDDDKED 69
D+D+DE AD+DEDED+D++DDD +D
Sbjct: 70 DEDEDEDADADEDEDEDEDEDDDDDDD 96
Score = 38.9 bits (89), Expect = 0.002
Identities = 17/39 (43%), Positives = 26/39 (66%), Gaps = 4/39 (10%)
Query: 35 EDDAPIVEDVKDDDKDEADDDED----EDDDDEDDDKED 69
++D ED DDD D+ DDD+D +DD+D+DD+ +D
Sbjct: 80 DEDEDEDEDEDDDDDDDDDDDDDADDADDDEDDDDEDDD 118
Score = 37.0 bits (84), Expect = 0.009
Identities = 13/27 (48%), Positives = 22/27 (81%)
Query: 43 DVKDDDKDEADDDEDEDDDDEDDDKED 69
D +D+ ++AD DEDED+D+++DD +D
Sbjct: 68 DGDEDEDEDADADEDEDEDEDEDDDDD 94
Score = 35.4 bits (80), Expect = 0.025
Identities = 14/38 (36%), Positives = 26/38 (67%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGAL 72
+DD ++ D+D+D+ DDD+DE+D++ DD+ + L
Sbjct: 107 DDDEDDDDEDDDEDEDDDDDDDDENDEECDDEYDSHRL 144
Score = 33.1 bits (74), Expect = 0.12
Identities = 15/49 (30%), Positives = 28/49 (56%)
Query: 21 SAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+AE+ +K D + D D+DE +D + ++D+DED+D++D
Sbjct: 43 TAEDEYIKVYDDHNNDGEGDGDGDGDGDEDEDEDADADEDEDEDEDEDD 91
Score = 30.0 bits (66), Expect = 1.0
Identities = 14/32 (43%), Positives = 22/32 (68%), Gaps = 1/32 (3%)
Query: 34 QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDD 65
++DD ++ +DDD D+ DD+ DE+ DDE D
Sbjct: 110 EDDDDEDDDEDEDDDDDD-DDENDEECDDEYD 140
Score = 26.9 bits (58), Expect = 8.8
Identities = 11/41 (26%), Positives = 19/41 (45%)
Query: 44 VKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSE 84
V DD ++ + D D D D ++D+ ED E + +
Sbjct: 51 VYDDHNNDGEGDGDGDGDGDEDEDEDADADEDEDEDEDEDD 91
>At1g48400
Length = 513
Score = 49.7 bits (117), Expect = 1e-06
Identities = 20/36 (55%), Positives = 27/36 (74%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDG 70
+DD +D DDD D+ DDD+D+DDDD+DDD +DG
Sbjct: 291 DDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDG 326
Score = 47.4 bits (111), Expect = 6e-06
Identities = 19/35 (54%), Positives = 26/35 (74%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+DD +D DDD D+ DDD+D+DDDD+DDD +D
Sbjct: 290 DDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDD 324
Score = 47.4 bits (111), Expect = 6e-06
Identities = 19/35 (54%), Positives = 26/35 (74%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+DD +D DDD D+ DDD+D+DDDD+DDD +D
Sbjct: 285 DDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDD 319
Score = 47.4 bits (111), Expect = 6e-06
Identities = 19/35 (54%), Positives = 26/35 (74%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+DD +D DDD D+ DDD+D+DDDD+DDD +D
Sbjct: 286 DDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDD 320
Score = 47.4 bits (111), Expect = 6e-06
Identities = 19/35 (54%), Positives = 26/35 (74%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+DD +D DDD D+ DDD+D+DDDD+DDD +D
Sbjct: 287 DDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDD 321
Score = 47.4 bits (111), Expect = 6e-06
Identities = 19/35 (54%), Positives = 26/35 (74%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+DD +D DDD D+ DDD+D+DDDD+DDD +D
Sbjct: 289 DDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDD 323
Score = 47.4 bits (111), Expect = 6e-06
Identities = 19/35 (54%), Positives = 26/35 (74%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+DD +D DDD D+ DDD+D+DDDD+DDD +D
Sbjct: 288 DDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDD 322
Score = 46.6 bits (109), Expect = 1e-05
Identities = 25/66 (37%), Positives = 36/66 (53%), Gaps = 3/66 (4%)
Query: 7 VDASSEAEALEQVLSAE---ETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDE 63
+D SS A V+ + E L + +D +D DDD D+ DDD+D+DDDD+
Sbjct: 247 LDYSSYVSAKYDVVDFDLLVEARLSLRLWVSTNDYDYSDDDDDDDDDDDDDDDDDDDDDD 306
Query: 64 DDDKED 69
DDD +D
Sbjct: 307 DDDDDD 312
Score = 45.8 bits (107), Expect = 2e-05
Identities = 19/35 (54%), Positives = 25/35 (71%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+DD +D DDD D+ DDD+D+DDDD+DDD D
Sbjct: 293 DDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDDGD 327
>At1g44050 unknown protein
Length = 734
Score = 49.3 bits (116), Expect = 2e-06
Identities = 21/44 (47%), Positives = 28/44 (62%)
Query: 34 QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEG 77
++D+ D DDD D+ DDD+D+DDDD DDD EDG +G
Sbjct: 101 EDDNEDDDNDDDDDDDDDDDDDDDDDDDDGDDDNEDGDCDDDDG 144
Score = 46.6 bits (109), Expect = 1e-05
Identities = 25/85 (29%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTG 101
+D +DD++D+ +DD+D+DDDD+DDD +D G + ++ + +L P
Sbjct: 98 DDNEDDNEDDDNDDDDDDDDDDDDDDDDDDDDGDDDNEDGDCDDDDGDLLLPFDSTPHFP 157
Query: 102 VSRVTIKRTKNILFFISKPDVFKSP 126
+R ++ +++L + PDV K P
Sbjct: 158 STRSGDQQGESLL-DCNHPDVCKLP 181
Score = 40.8 bits (94), Expect = 6e-04
Identities = 19/46 (41%), Positives = 29/46 (62%), Gaps = 3/46 (6%)
Query: 27 LKKKPQPQEDDAPIVEDVKDDDKDEADDD---EDEDDDDEDDDKED 69
L+ K +DD +D DDK++ DDD +D DDD+EDD+++D
Sbjct: 62 LRLKMHNHDDDDDAEDDDNGDDKEDGDDDNKGDDGDDDNEDDNEDD 107
Score = 39.7 bits (91), Expect = 0.001
Identities = 18/37 (48%), Positives = 24/37 (64%)
Query: 36 DDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGAL 72
DD +D DDD D+ DDD D+D++D D D +DG L
Sbjct: 110 DDDDDDDDDDDDDDDDDDDDGDDDNEDGDCDDDDGDL 146
Score = 35.8 bits (81), Expect = 0.019
Identities = 16/35 (45%), Positives = 21/35 (59%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+DD +D DDD D+ D D+D +D D DDD D
Sbjct: 111 DDDDDDDDDDDDDDDDDDDGDDDNEDGDCDDDDGD 145
Score = 32.0 bits (71), Expect = 0.27
Identities = 12/38 (31%), Positives = 21/38 (54%)
Query: 47 DDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSE 84
DD D+A+DD++ DD ++ DD G G + + +
Sbjct: 70 DDDDDAEDDDNGDDKEDGDDDNKGDDGDDDNEDDNEDD 107
Score = 27.3 bits (59), Expect = 6.8
Identities = 11/25 (44%), Positives = 15/25 (60%)
Query: 55 DEDEDDDDEDDDKEDGALGGSEGSK 79
+ D+DDD EDDD D G + +K
Sbjct: 68 NHDDDDDAEDDDNGDDKEDGDDDNK 92
>At5g55660 putative protein
Length = 759
Score = 47.4 bits (111), Expect = 6e-06
Identities = 39/121 (32%), Positives = 58/121 (47%), Gaps = 26/121 (21%)
Query: 7 VDASSEAEAL----EQVLSAEETTLKK---KPQPQEDDA--------PIVEDVKDDDKDE 51
VD + EAL E L+ EE T K K +EDD P VED K + KDE
Sbjct: 186 VDEDDKEEALKEKNEAELAEEEETNKGEEVKEANKEDDVEADTKVAEPEVEDKKTESKDE 245
Query: 52 ADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTK 111
+D E+E +D+++D+KE +S +K+ +K +K K G + T +TK
Sbjct: 246 NEDKEEEKEDEKEDEKE-----------ESNDDKEDKKEDIKKSNKRGKGKTEKTRGKTK 294
Query: 112 N 112
+
Sbjct: 295 S 295
Score = 38.5 bits (88), Expect = 0.003
Identities = 25/88 (28%), Positives = 44/88 (49%), Gaps = 3/88 (3%)
Query: 9 ASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDED---D 65
+SS+ A Q + E T KK DD E D+++++ + E+E++++E+ D
Sbjct: 480 SSSKRSAKSQKKTEEATRTNKKSVAHSDDESEEEKEDDEEEEKEQEVEEEEEENENGIPD 539
Query: 66 DKEDGALGGSEGSKQSRSEKKSRKAMLK 93
ED A SE + SE++S + K
Sbjct: 540 KSEDEAPQLSESEENVESEEESEEETKK 567
Score = 37.4 bits (85), Expect = 0.007
Identities = 20/81 (24%), Positives = 45/81 (54%), Gaps = 5/81 (6%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDD--EDD 65
D ++ + E + ++T K + + +E++ ED K+D+K+E++DD+++ +D + +
Sbjct: 223 DVEADTKVAEPEVEDKKTESKDENEDKEEEK---EDEKEDEKEESNDDKEDKKEDIKKSN 279
Query: 66 DKEDGALGGSEGSKQSRSEKK 86
+ G + G +S EKK
Sbjct: 280 KRGKGKTEKTRGKTKSDEEKK 300
Score = 32.3 bits (72), Expect = 0.21
Identities = 21/85 (24%), Positives = 41/85 (47%), Gaps = 6/85 (7%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDD-- 65
+ E E+ E+ L + ++ +E +V K DD DEA+ E+ D+DD+++
Sbjct: 139 EGEQEKESKEEKLEGGKANGNEEGDTEEK---LVGGDKGDDVDEAEKVENVDEDDKEEAL 195
Query: 66 -DKEDGALGGSEGSKQSRSEKKSRK 89
+K + L E + + K++ K
Sbjct: 196 KEKNEAELAEEEETNKGEEVKEANK 220
Score = 30.8 bits (68), Expect = 0.61
Identities = 21/87 (24%), Positives = 40/87 (45%), Gaps = 7/87 (8%)
Query: 12 EAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGA 71
E E ++V EE P ED+AP + + E + + +E+ ++E K+ G+
Sbjct: 519 EEEKEQEVEEEEEENENGIPDKSEDEAPQLSE------SEENVESEEESEEETKKKKRGS 572
Query: 72 LGGSEGSKQSRSEKKSRKAMLKLGLKP 98
S+ K+S + +S+K + P
Sbjct: 573 RTSSD-KKESAGKSRSKKTAVPTKSSP 598
Score = 29.6 bits (65), Expect = 1.4
Identities = 17/65 (26%), Positives = 30/65 (46%), Gaps = 14/65 (21%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDD--------------EDDDKEDGALGGSEGSKQ 80
E+DA + E V+ D +A++ E E + + E+ D E+ +GG +G
Sbjct: 118 EEDAVMKESVESADNKDAENPEGEQEKESKEEKLEGGKANGNEEGDTEEKLVGGDKGDDV 177
Query: 81 SRSEK 85
+EK
Sbjct: 178 DEAEK 182
Score = 28.9 bits (63), Expect = 2.3
Identities = 15/58 (25%), Positives = 36/58 (61%), Gaps = 7/58 (12%)
Query: 20 LSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDED-EDDDDEDDDKEDGALGGSE 76
L+++++++K+ Q + + + D+ +DE ++ED E +++E+ +K G+ GG E
Sbjct: 705 LASKKSSIKRMIQDE------LTKLADEAEDEEGEEEDAEHEEEEEKEKAKGSGGGEE 756
Score = 28.1 bits (61), Expect = 4.0
Identities = 20/86 (23%), Positives = 41/86 (47%), Gaps = 16/86 (18%)
Query: 4 GPVVDASSEAEALEQVLSAEETTLKK------------KPQPQEDDAPIVEDVKDD---D 48
G V ++ + +++ E+T L+K + + + + + E+ K D D
Sbjct: 29 GKEVQENASGKEVQESKKEEDTGLEKMEIDDEGKQHEGESETGDKEVEVTEEEKKDVGED 88
Query: 49 KDEADDDE-DEDDDDEDDDKEDGALG 73
K++ + D+ DED DD++ +DG G
Sbjct: 89 KEQPEADKMDEDTDDKNLKADDGVSG 114
>At2g21420 Mutator-like transposase
Length = 468
Score = 47.4 bits (111), Expect = 6e-06
Identities = 17/35 (48%), Positives = 27/35 (76%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSE 76
+D DDD D+ DDD+D+DDDD++DD++DG + +
Sbjct: 396 DDDDDDDDDDDDDDDDDDDDDDEDDEDDGYIDSDD 430
Score = 46.6 bits (109), Expect = 1e-05
Identities = 18/28 (64%), Positives = 23/28 (81%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKED 69
+D DDD D+ DDD+D+DDDD+DDD ED
Sbjct: 392 DDDDDDDDDDDDDDDDDDDDDDDDDDED 419
Score = 46.2 bits (108), Expect = 1e-05
Identities = 17/28 (60%), Positives = 24/28 (85%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKED 69
+D DDD D+ DDD+D+DDDD+DDD++D
Sbjct: 393 DDDDDDDDDDDDDDDDDDDDDDDDDEDD 420
Score = 45.4 bits (106), Expect = 2e-05
Identities = 20/39 (51%), Positives = 27/39 (68%), Gaps = 4/39 (10%)
Query: 31 PQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
P +DD +D DDD D+ DDD+D+DDDD+D+D ED
Sbjct: 388 PGCDDDD----DDDDDDDDDDDDDDDDDDDDDDDEDDED 422
Score = 45.1 bits (105), Expect = 3e-05
Identities = 17/32 (53%), Positives = 25/32 (78%)
Query: 38 APIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+P +D DDD D+ DDD+D+DDDD+DDD ++
Sbjct: 387 SPGCDDDDDDDDDDDDDDDDDDDDDDDDDDDE 418
Score = 41.6 bits (96), Expect = 3e-04
Identities = 20/39 (51%), Positives = 25/39 (63%), Gaps = 3/39 (7%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDD---DEDDDKEDG 70
+DD +D DDD D+ DDDED++DD D DDD DG
Sbjct: 397 DDDDDDDDDDDDDDDDDDDDDEDDEDDGYIDSDDDDVDG 435
Score = 37.7 bits (86), Expect = 0.005
Identities = 20/41 (48%), Positives = 26/41 (62%), Gaps = 4/41 (9%)
Query: 35 EDDAPIVEDVKDDDKDEADDDE---DEDDDDED-DDKEDGA 71
+DD +D DDD DE D+D+ D DDDD D DD +DG+
Sbjct: 402 DDDDDDDDDDDDDDDDEDDEDDGYIDSDDDDVDGDDNDDGS 442
Score = 35.8 bits (81), Expect = 0.019
Identities = 18/43 (41%), Positives = 25/43 (57%), Gaps = 3/43 (6%)
Query: 35 EDDAPIVEDVKDDDKDEADD---DEDEDDDDEDDDKEDGALGG 74
+DD +D DDD+D+ DD D D+DD D DD+ + L G
Sbjct: 404 DDDDDDDDDDDDDDEDDEDDGYIDSDDDDVDGDDNDDGSELDG 446
>At5g15140 putative aldose 1-epimerase - like protein
Length = 490
Score = 47.0 bits (110), Expect = 8e-06
Identities = 21/49 (42%), Positives = 31/49 (62%), Gaps = 3/49 (6%)
Query: 35 EDDAPIVEDVKDD---DKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQ 80
+DD +D K D D D++DDD D+DDDD+DDD +D + G + K+
Sbjct: 110 DDDEKKHKDKKKDGHNDDDDSDDDTDDDDDDDDDDDDDDEVDGDDNEKE 158
Score = 43.5 bits (101), Expect = 9e-05
Identities = 18/42 (42%), Positives = 29/42 (68%)
Query: 28 KKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+KK + ++ D +D DDD D+ DDD+D+DDDD++ D +D
Sbjct: 113 EKKHKDKKKDGHNDDDDSDDDTDDDDDDDDDDDDDDEVDGDD 154
Score = 40.8 bits (94), Expect = 6e-04
Identities = 17/40 (42%), Positives = 25/40 (62%)
Query: 29 KKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKE 68
KK DD +D DDD D+ DDD+D++ D +D++KE
Sbjct: 119 KKKDGHNDDDDSDDDTDDDDDDDDDDDDDDEVDGDDNEKE 158
Score = 38.9 bits (89), Expect = 0.002
Identities = 18/42 (42%), Positives = 28/42 (65%), Gaps = 2/42 (4%)
Query: 28 KKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
KKK +DD +D DDD D+ DDD+D+D+ D DD++++
Sbjct: 119 KKKDGHNDDDDS--DDDTDDDDDDDDDDDDDDEVDGDDNEKE 158
Score = 36.6 bits (83), Expect = 0.011
Identities = 24/69 (34%), Positives = 32/69 (45%), Gaps = 1/69 (1%)
Query: 28 KKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDD-DEDDDKEDGALGGSEGSKQSRSEKK 86
KKK + D + KD KD +DD+D DDD D+DDD +D E ++K
Sbjct: 100 KKKSGGHDKDDDDEKKHKDKKKDGHNDDDDSDDDTDDDDDDDDDDDDDDEVDGDDNEKEK 159
Query: 87 SRKAMLKLG 95
LK G
Sbjct: 160 IGLYELKKG 168
Score = 27.7 bits (60), Expect = 5.2
Identities = 19/66 (28%), Positives = 32/66 (47%), Gaps = 1/66 (1%)
Query: 4 GPVVD-ASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDD 62
G VD A S+ E ++ +E ++KK + E+ E D K D+D+DD+
Sbjct: 55 GEKVDGAGSDDEDNDKKEKKKEHDVQKKDKQHENKDKDDEKKHVDKKKSGGHDKDDDDEK 114
Query: 63 EDDDKE 68
+ DK+
Sbjct: 115 KHKDKK 120
>At3g56860 unknown protein
Length = 478
Score = 47.0 bits (110), Expect = 8e-06
Identities = 27/68 (39%), Positives = 41/68 (59%), Gaps = 5/68 (7%)
Query: 8 DASSEAEALEQVLSA---EETTLKKKPQPQEDDAPIVE--DVKDDDKDEADDDEDEDDDD 62
+ S+EAE Q L EE L K Q + D VE +V+++ ++E +DD+DEDD D
Sbjct: 10 EESNEAEEPSQKLKQTPEEEQQLVIKNQDNQGDVEEVEYEEVEEEQEEEVEDDDDEDDGD 69
Query: 63 EDDDKEDG 70
E++D+ DG
Sbjct: 70 ENEDQTDG 77
Score = 34.7 bits (78), Expect = 0.042
Identities = 18/69 (26%), Positives = 39/69 (56%), Gaps = 10/69 (14%)
Query: 17 EQVLSAEETTLKKKPQPQEDDAPIVE------DVKDDDKDEADDDEDE----DDDDEDDD 66
E+ AEE + K K P+E+ +++ DV++ + +E +++++E DDD++D D
Sbjct: 10 EESNEAEEPSQKLKQTPEEEQQLVIKNQDNQGDVEEVEYEEVEEEQEEEVEDDDDEDDGD 69
Query: 67 KEDGALGGS 75
+ + G+
Sbjct: 70 ENEDQTDGN 78
>At1g63980 unknown protein
Length = 391
Score = 45.4 bits (106), Expect = 2e-05
Identities = 25/66 (37%), Positives = 34/66 (50%), Gaps = 4/66 (6%)
Query: 28 KKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
KK DD +D DDD DE D++EDED+ + DDD +D + S +K+ E
Sbjct: 259 KKTSFDNSDD----DDDDDDDDDEEDEEEDEDESEADDDDKDSVIESSLPAKRKHDEIIE 314
Query: 88 RKAMLK 93
K LK
Sbjct: 315 PKIKLK 320
Score = 36.6 bits (83), Expect = 0.011
Identities = 16/40 (40%), Positives = 25/40 (62%)
Query: 30 KPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
K +P++ E K + DDD+D+DDDDE+D++ED
Sbjct: 245 KDRPKKIAGVRYEGKKTSFDNSDDDDDDDDDDDEEDEEED 284
Score = 34.3 bits (77), Expect = 0.055
Identities = 18/65 (27%), Positives = 33/65 (50%)
Query: 23 EETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSR 82
++ T K+ +D + V+ + K + D+ D+DDDD+DDD E+ + S+
Sbjct: 233 DKATAGKQGLGIKDRPKKIAGVRYEGKKTSFDNSDDDDDDDDDDDEEDEEEDEDESEADD 292
Query: 83 SEKKS 87
+K S
Sbjct: 293 DDKDS 297
>At5g37430 contains similarity to unknown protein (pir||C71432)
Length = 607
Score = 45.1 bits (105), Expect = 3e-05
Identities = 19/36 (52%), Positives = 25/36 (68%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDG 70
+DD +D DDD D+ DDD D+ DDD+DDD +DG
Sbjct: 562 DDDGDDDDDDGDDDGDDDDDDGDDGDDDDDDDGDDG 597
Score = 43.1 bits (100), Expect = 1e-04
Identities = 21/50 (42%), Positives = 31/50 (62%), Gaps = 1/50 (2%)
Query: 28 KKKPQPQEDDAPIVEDVKDD-DKDEADDDEDEDDDDEDDDKEDGALGGSE 76
K+ P+ + D + +D DD D D+ DDD+D+ DDD DDD +DG G +
Sbjct: 540 KQLPETEFDWLNLHDDDGDDGDDDDGDDDDDDGDDDGDDDDDDGDDGDDD 589
Score = 42.7 bits (99), Expect = 2e-04
Identities = 18/35 (51%), Positives = 23/35 (65%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+DD D DDD D+ DDD+D+D DD DDD +D
Sbjct: 569 DDDGDDDGDDDDDDGDDGDDDDDDDGDDGDDDDDD 603
Score = 42.4 bits (98), Expect = 2e-04
Identities = 18/36 (50%), Positives = 23/36 (63%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEG 77
+D DDD D+ DDD D+DDDD DD +D G +G
Sbjct: 562 DDDGDDDDDDGDDDGDDDDDDGDDGDDDDDDDGDDG 597
Score = 41.2 bits (95), Expect = 5e-04
Identities = 19/41 (46%), Positives = 26/41 (63%), Gaps = 5/41 (12%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDD-----DEDDDKEDG 70
+DD +D DDD D+ DD +D+DDD D+DDD +DG
Sbjct: 566 DDDDDDGDDDGDDDDDDGDDGDDDDDDDGDDGDDDDDDDDG 606
Score = 40.4 bits (93), Expect = 8e-04
Identities = 18/36 (50%), Positives = 25/36 (69%), Gaps = 1/36 (2%)
Query: 43 DVKDDDKDEADDDE-DEDDDDEDDDKEDGALGGSEG 77
++ DDD D+ DDD+ D+DDDD DDD +D G +G
Sbjct: 551 NLHDDDGDDGDDDDGDDDDDDGDDDGDDDDDDGDDG 586
>At5g22640 unknown protein
Length = 871
Score = 45.1 bits (105), Expect = 3e-05
Identities = 24/76 (31%), Positives = 39/76 (50%), Gaps = 2/76 (2%)
Query: 23 EETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSR 82
EE + K + QE+ E KDDD ++ DD +D+DDDD+DDD + G ++ +++
Sbjct: 662 EEKVVTAKEKIQENKQE--EKYKDDDDEDDDDGDDDDDDDDDDDLGPSSFGSADKGRRNS 719
Query: 83 SEKKSRKAMLKLGLKP 98
S + L P
Sbjct: 720 PFSSSSLSFASCTLFP 735
Score = 37.4 bits (85), Expect = 0.007
Identities = 17/47 (36%), Positives = 30/47 (63%), Gaps = 6/47 (12%)
Query: 17 EQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDE 63
E+V++A+E + K + + +D D+D D+ DDD+D+DDDD+
Sbjct: 663 EKVVTAKEKIQENKQEEK------YKDDDDEDDDDGDDDDDDDDDDD 703
Score = 35.4 bits (80), Expect = 0.025
Identities = 28/103 (27%), Positives = 47/103 (45%), Gaps = 27/103 (26%)
Query: 12 EAE-ALEQVLSAEETTLKKKPQPQE----------DDAPIV------------EDVKDDD 48
EAE LE+ + + LKKK Q +E D+ +V E ++++
Sbjct: 617 EAEICLEEAIEDMDEELKKKEQEEEKKTEMGLTEEDEDVLVPVYKEEKVVTAKEKIQENK 676
Query: 49 KDEADDDEDEDD----DDEDDDKEDGALGGSEGSKQSRSEKKS 87
++E D+D++D DD+DDD +D LG S + + S
Sbjct: 677 QEEKYKDDDDEDDDDGDDDDDDDDDDDLGPSSFGSADKGRRNS 719
>At5g56950 nucleosome assembly protein
Length = 374
Score = 44.7 bits (104), Expect = 4e-05
Identities = 21/57 (36%), Positives = 39/57 (67%), Gaps = 5/57 (8%)
Query: 43 DVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAML-KLGLKP 98
++ +DD+D+ D+DEDED++DED+D+E+ E ++ S+ K + ++L K G +P
Sbjct: 305 EIDNDDEDDIDEDEDEDEEDEDEDEEE----DDEDEEEEVSKTKKKPSVLHKKGGRP 357
Score = 42.0 bits (97), Expect = 3e-04
Identities = 22/59 (37%), Positives = 35/59 (59%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVT 100
E+ + D+ DE D DEDED+D+ED+D+++ E + S+++KK K G VT
Sbjct: 302 EEFEIDNDDEDDIDEDEDEDEEDEDEDEEEDDEDEEEEVSKTKKKPSVLHKKGGRPQVT 360
Score = 38.1 bits (87), Expect = 0.004
Identities = 12/31 (38%), Positives = 26/31 (83%)
Query: 36 DDAPIVEDVKDDDKDEADDDEDEDDDDEDDD 66
DD +++ +D+D+++ D+DE+EDD+DE+++
Sbjct: 309 DDEDDIDEDEDEDEEDEDEDEEEDDEDEEEE 339
Score = 35.4 bits (80), Expect = 0.025
Identities = 12/31 (38%), Positives = 25/31 (79%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDD 65
+D+ I ED +D++DE +D+E++D+D+E++
Sbjct: 309 DDEDDIDEDEDEDEEDEDEDEEEDDEDEEEE 339
Score = 35.4 bits (80), Expect = 0.025
Identities = 15/48 (31%), Positives = 27/48 (56%)
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRK 89
ED +D+D+DE +DDEDE+++ K+ L G Q +++ +
Sbjct: 320 EDEEDEDEDEEEDDEDEEEEVSKTKKKPSVLHKKGGRPQVTDDQQGER 367
>At5g42280 putative protein
Length = 694
Score = 44.7 bits (104), Expect = 4e-05
Identities = 18/42 (42%), Positives = 28/42 (65%)
Query: 28 KKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
K K +E + +D +DDD D+ DDD+D+DDD++DD +D
Sbjct: 141 KDKSSTEESIYEVDDDCEDDDNDDDDDDDDDDDDNDDDVVDD 182
Score = 44.3 bits (103), Expect = 5e-05
Identities = 21/49 (42%), Positives = 30/49 (60%)
Query: 21 SAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
S+ E ++ + EDD +D DDD D+ DDD +DDDD+DDD +D
Sbjct: 144 SSTEESIYEVDDDCEDDDNDDDDDDDDDDDDNDDDVVDDDDDDDDDDDD 192
Score = 40.4 bits (93), Expect = 8e-04
Identities = 18/50 (36%), Positives = 29/50 (58%)
Query: 20 LSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
L + + T P+ +D + E + + D D DDD D+DDDD+DDD ++
Sbjct: 126 LISVDPTFDFSPRNWKDKSSTEESIYEVDDDCEDDDNDDDDDDDDDDDDN 175
Score = 38.9 bits (89), Expect = 0.002
Identities = 24/65 (36%), Positives = 34/65 (51%), Gaps = 5/65 (7%)
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDD-EDDD 66
D SS E++ +V + + +DD +D DDD + DDD+D+DDDD DDD
Sbjct: 142 DKSSTEESIYEV----DDDCEDDDNDDDDDDDDDDDDNDDDVVDDDDDDDDDDDDVVDDD 197
Query: 67 KEDGA 71
D A
Sbjct: 198 YGDSA 202
Score = 28.1 bits (61), Expect = 4.0
Identities = 14/36 (38%), Positives = 21/36 (57%), Gaps = 2/36 (5%)
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDG 70
++D +V+D DDD D+ DDD +DD + DG
Sbjct: 174 DNDDDVVDD--DDDDDDDDDDVVDDDYGDSACLSDG 207
>At4g26110 nucleosome assembly protein I-like protein
Length = 372
Score = 44.7 bits (104), Expect = 4e-05
Identities = 29/97 (29%), Positives = 53/97 (53%), Gaps = 12/97 (12%)
Query: 13 AEALEQVLSAEE---TTLKKKPQPQE-----DDAPIVEDVKDDDKDEADDDEDEDDDDED 64
AE L+ ++ + +T+++K P+ +A ED + DD +E D DEDED++DE+
Sbjct: 266 AEDLQNLMEQDYDIGSTIREKIIPRAVSWFTGEAMEAEDFEIDDDEEDDIDEDEDEEDEE 325
Query: 65 DDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTG 101
D+++D E ++S+++KK K G + G
Sbjct: 326 DEEDD----DDEDEEESKTKKKPSIGNKKGGRSQIVG 358
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.305 0.127 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,469,886
Number of Sequences: 26719
Number of extensions: 280261
Number of successful extensions: 11899
Number of sequences better than 10.0: 823
Number of HSP's better than 10.0 without gapping: 580
Number of HSP's successfully gapped in prelim test: 253
Number of HSP's that attempted gapping in prelim test: 3242
Number of HSP's gapped (non-prelim): 3622
length of query: 227
length of database: 11,318,596
effective HSP length: 96
effective length of query: 131
effective length of database: 8,753,572
effective search space: 1146717932
effective search space used: 1146717932
T: 11
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0207.3


