

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.8
(124 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
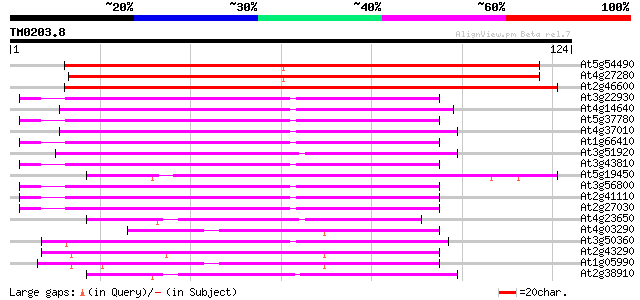
Score E
Sequences producing significant alignments: (bits) Value
At5g54490 unknown protein 136 3e-33
At4g27280 unknown protein 133 3e-32
At2g46600 caltractin like protein 130 2e-31
At3g22930 calmodulin, putative 47 2e-06
At4g14640 calmodulin 46 4e-06
At5g37780 calmodulin 1 (CAM1) 45 9e-06
At4g37010 caltractin-like protein 45 9e-06
At1g66410 calmodulin-4 45 9e-06
At3g51920 putative calmodulin 45 1e-05
At3g43810 calmodulin 7 44 2e-05
At5g19450 calcium-dependent protein kinase 43 3e-05
At3g56800 calmodulin-3 43 3e-05
At2g41110 calmodulin (cam2) 43 3e-05
At2g27030 calmodulin 43 3e-05
At4g23650 calcium-dependent protein kinase (CDPK6) 42 6e-05
At4g03290 putative calmodulin 42 1e-04
At3g50360 centrin 41 1e-04
At2g43290 putative calcium binding protein 41 1e-04
At1g05990 calcium-binding protein, putative 41 2e-04
At2g38910 putative calcium-dependent protein kinase 39 7e-04
>At5g54490 unknown protein
Length = 127
Score = 136 bits (342), Expect = 3e-33
Identities = 67/106 (63%), Positives = 83/106 (78%), Gaps = 1/106 (0%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQDMKEDEL 71
+F+D P MA KLGGEGLI+E+C GF++LMDKD+GVIT ESL+ NA+ VLGL D+ +D++
Sbjct: 18 QFQDFFPTMAGKLGGEGLIEEICKGFELLMDKDKGVITFESLRRNASTVLGLGDLTDDDV 77
Query: 72 VSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHE 117
MI EGD D DGAL QMEFCVLMFRLSPELME S + E ++ E
Sbjct: 78 RYMINEGDFDRDGALNQMEFCVLMFRLSPELMEASRCVVTEVIEEE 123
>At4g27280 unknown protein
Length = 130
Score = 133 bits (334), Expect = 3e-32
Identities = 66/105 (62%), Positives = 82/105 (77%), Gaps = 1/105 (0%)
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQDMKEDELV 72
F D LP MA LGGEGLI ELCNGF++LMD+++GVIT ESL+ NAA VLGL D+ ++++
Sbjct: 19 FHDFLPTMAGNLGGEGLIGELCNGFELLMDREKGVITFESLRRNAAAVLGLGDLTDEDVR 78
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHE 117
MI+EGD D DGAL QMEFCVLMFRLSP+LME S + E ++ E
Sbjct: 79 CMIKEGDFDCDGALNQMEFCVLMFRLSPDLMEASRCLVTEVIEEE 123
>At2g46600 caltractin like protein
Length = 135
Score = 130 bits (327), Expect = 2e-31
Identities = 60/109 (55%), Positives = 82/109 (75%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELV 72
++ED+LPVMA K+ E + ELC GF +L D +R +IT ESL+ N+ +LG++ M +++
Sbjct: 21 KYEDMLPVMAEKMDVEEFVSELCKGFSLLADPERHLITAESLRRNSGILGIEGMSKEDAQ 80
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNN 121
M+REGDLDGDGAL Q EFCVLM RLSPE+ME++ WLE+AL EL N+
Sbjct: 81 GMVREGDLDGDGALNQTEFCVLMVRLSPEMMEDAETWLEKALTQELCNH 129
>At3g22930 calmodulin, putative
Length = 173
Score = 47.0 bits (110), Expect = 2e-06
Identities = 31/93 (33%), Positives = 47/93 (50%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ EF + L +MAN+L +EL F + G I+ L+ LG
Sbjct: 83 GNGTI-----EFSEFLNLMANQLQETDADEELKEAFKVFDKDQNGYISASELRHVMINLG 137
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MI+E DLDGDG + EF +M
Sbjct: 138 -EKLTDEEVDQMIKEADLDGDGQVNYDEFVRMM 169
Score = 33.1 bits (74), Expect = 0.036
Identities = 25/56 (44%), Positives = 29/56 (51%), Gaps = 2/56 (3%)
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
L DKD G IT + L T L Q+ E EL MI E D DG+G + EF LM
Sbjct: 42 LFDKDGDGCITADELATVIRSLD-QNPTEQELQDMITEIDSDGNGTIEFSEFLNLM 96
>At4g14640 calmodulin
Length = 151
Score = 46.2 bits (108), Expect = 4e-06
Identities = 28/87 (32%), Positives = 43/87 (49%), Gaps = 1/87 (1%)
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
+EF + L +MA KL +EL F + G I+ L LG + + ++E+
Sbjct: 65 IEFAEFLNLMAKKLQESDAEEELKEAFKVFDKDQNGYISASELSHVMINLG-EKLTDEEV 123
Query: 72 VSMIREGDLDGDGALTQMEFCVLMFRL 98
MI+E DLDGDG + EF +M +
Sbjct: 124 EQMIKEADLDGDGQVNYDEFVKMMINI 150
>At5g37780 calmodulin 1 (CAM1)
Length = 149
Score = 45.1 bits (105), Expect = 9e-06
Identities = 29/93 (31%), Positives = 46/93 (49%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 60 GNGTI-----DFPEFLNLMAKKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 114
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 115 -EKLTDEEVEEMIREADVDGDGQINYEEFVKIM 146
Score = 36.2 bits (82), Expect = 0.004
Identities = 33/76 (43%), Positives = 39/76 (50%), Gaps = 4/76 (5%)
Query: 21 MANKLGGEGLIKELCNGFDMLMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGD 79
MA++L E I E F L DKD G IT + L T LG Q+ E EL MI E D
Sbjct: 1 MADQLTDEQ-ISEFKEAFS-LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINEVD 57
Query: 80 LDGDGALTQMEFCVLM 95
DG+G + EF LM
Sbjct: 58 ADGNGTIDFPEFLNLM 73
>At4g37010 caltractin-like protein
Length = 167
Score = 45.1 bits (105), Expect = 9e-06
Identities = 26/88 (29%), Positives = 45/88 (50%), Gaps = 1/88 (1%)
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
++F++ + +M K G I EL F ++ + G I+ +K A LG ++ ++++
Sbjct: 79 IDFDEFVHMMTTKFGERDSIDELSKAFKIIDHDNNGKISPRDIKMIAKELG-ENFTDNDI 137
Query: 72 VSMIREGDLDGDGALTQMEFCVLMFRLS 99
MI E D D DG + EF +M R S
Sbjct: 138 EEMIEEADRDKDGEVNLEEFMKMMKRTS 165
Score = 27.3 bits (59), Expect = 2.0
Identities = 20/70 (28%), Positives = 29/70 (40%), Gaps = 4/70 (5%)
Query: 54 LKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEA 113
LK GL + K E+ + D+DG G++ E V M L E+ +
Sbjct: 11 LKPKGKTYGLTNQKRREIREIFDLFDIDGSGSIDASELNVAMRSLGFEMNNQQ----INE 66
Query: 114 LQHELDNNYS 123
L E+D N S
Sbjct: 67 LMAEVDKNQS 76
>At1g66410 calmodulin-4
Length = 149
Score = 45.1 bits (105), Expect = 9e-06
Identities = 29/93 (31%), Positives = 46/93 (49%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 60 GNGTI-----DFPEFLNLMAKKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 114
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 115 -EKLTDEEVEEMIREADVDGDGQINYEEFVKIM 146
Score = 36.2 bits (82), Expect = 0.004
Identities = 33/76 (43%), Positives = 39/76 (50%), Gaps = 4/76 (5%)
Query: 21 MANKLGGEGLIKELCNGFDMLMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGD 79
MA++L E I E F L DKD G IT + L T LG Q+ E EL MI E D
Sbjct: 1 MADQLTDEQ-ISEFKEAFS-LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINEVD 57
Query: 80 LDGDGALTQMEFCVLM 95
DG+G + EF LM
Sbjct: 58 ADGNGTIDFPEFLNLM 73
>At3g51920 putative calmodulin
Length = 151
Score = 44.7 bits (104), Expect = 1e-05
Identities = 29/89 (32%), Positives = 43/89 (47%), Gaps = 1/89 (1%)
Query: 11 GLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDE 70
G+ F+D L +MA E EL F + G+I+ L +G++ + +E
Sbjct: 63 GITFDDFLYIMAQNTSQESASDELIEVFRVFDRDGDGLISQLELGEGMKDMGMK-ITAEE 121
Query: 71 LVSMIREGDLDGDGALTQMEFCVLMFRLS 99
M+RE DLDGDG L+ EF +M S
Sbjct: 122 AEHMVREADLDGDGFLSFHEFSKMMIAAS 150
Score = 31.2 bits (69), Expect = 0.14
Identities = 19/54 (35%), Positives = 27/54 (49%), Gaps = 3/54 (5%)
Query: 40 MLMDKD---RGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQME 90
M+ D D G IT + A Q+ DEL+ + R D DGDG ++Q+E
Sbjct: 52 MMSDVDIFGNGGITFDDFLYIMAQNTSQESASDELIEVFRVFDRDGDGLISQLE 105
>At3g43810 calmodulin 7
Length = 149
Score = 44.3 bits (103), Expect = 2e-05
Identities = 29/93 (31%), Positives = 46/93 (49%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 60 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 114
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 115 -EKLTDEEVDEMIREADVDGDGQINYEEFVKVM 146
Score = 36.2 bits (82), Expect = 0.004
Identities = 27/58 (46%), Positives = 31/58 (52%), Gaps = 2/58 (3%)
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFR 97
L DKD G IT + L T LG Q+ E EL MI E D DG+G + EF LM R
Sbjct: 19 LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 75
>At5g19450 calcium-dependent protein kinase
Length = 533
Score = 43.1 bits (100), Expect = 3e-05
Identities = 39/123 (31%), Positives = 54/123 (43%), Gaps = 22/123 (17%)
Query: 18 LPVMANKLGGEGL--IKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMI 75
L V+A L E + IKE F+M+ K G I LE LK LG Q + + +L ++
Sbjct: 348 LRVIAEHLSVEEVAGIKE---AFEMMDSKKTGKINLEELKFGLHKLGQQQIPDTDLQILM 404
Query: 76 REGDLDGDGALTQMEFCVLMFRLSPELMEE--------------SWIWLE---EALQHEL 118
D+DGDG L EF + L +E +I +E EAL E+
Sbjct: 405 EAADVDGDGTLNYGEFVAVSVHLKKMANDEHLHKAFSFFDQNQSDYIEIEELREALNDEV 464
Query: 119 DNN 121
D N
Sbjct: 465 DTN 467
>At3g56800 calmodulin-3
Length = 149
Score = 43.1 bits (100), Expect = 3e-05
Identities = 28/93 (30%), Positives = 46/93 (49%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 60 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 114
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MI+E D+DGDG + EF +M
Sbjct: 115 -EKLTDEEVDEMIKEADVDGDGQINYEEFVKVM 146
Score = 36.2 bits (82), Expect = 0.004
Identities = 27/58 (46%), Positives = 31/58 (52%), Gaps = 2/58 (3%)
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFR 97
L DKD G IT + L T LG Q+ E EL MI E D DG+G + EF LM R
Sbjct: 19 LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 75
>At2g41110 calmodulin (cam2)
Length = 149
Score = 43.1 bits (100), Expect = 3e-05
Identities = 28/93 (30%), Positives = 46/93 (49%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 60 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 114
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MI+E D+DGDG + EF +M
Sbjct: 115 -EKLTDEEVDEMIKEADVDGDGQINYEEFVKVM 146
Score = 36.2 bits (82), Expect = 0.004
Identities = 27/58 (46%), Positives = 31/58 (52%), Gaps = 2/58 (3%)
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFR 97
L DKD G IT + L T LG Q+ E EL MI E D DG+G + EF LM R
Sbjct: 19 LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 75
>At2g27030 calmodulin
Length = 149
Score = 43.1 bits (100), Expect = 3e-05
Identities = 28/93 (30%), Positives = 46/93 (49%), Gaps = 6/93 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 60 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 114
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MI+E D+DGDG + EF +M
Sbjct: 115 -EKLTDEEVDEMIKEADVDGDGQINYEEFVKVM 146
Score = 36.2 bits (82), Expect = 0.004
Identities = 27/58 (46%), Positives = 31/58 (52%), Gaps = 2/58 (3%)
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFR 97
L DKD G IT + L T LG Q+ E EL MI E D DG+G + EF LM R
Sbjct: 19 LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 75
>At4g23650 calcium-dependent protein kinase (CDPK6)
Length = 529
Score = 42.4 bits (98), Expect = 6e-05
Identities = 27/76 (35%), Positives = 43/76 (56%), Gaps = 6/76 (7%)
Query: 18 LPVMANKLGGEGLI--KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMI 75
L V+A L E +I KE+ F L + G++TLE L+T LG + + E E+ ++
Sbjct: 369 LKVIAENLSEEEIIGLKEM---FKSLDTDNNGIVTLEELRTGLPKLGSK-ISEAEIRQLM 424
Query: 76 REGDLDGDGALTQMEF 91
D+DGDG++ +EF
Sbjct: 425 EAADMDGDGSIDYLEF 440
Score = 32.7 bits (73), Expect = 0.047
Identities = 21/70 (30%), Positives = 33/70 (47%), Gaps = 2/70 (2%)
Query: 34 LCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCV 93
L F + + G IT+E L+ + D K + +I E D D DG + EF
Sbjct: 456 LYTAFQFFDNDNSGYITMEELELAMKKYNMGDDKS--IKEIIAEVDTDRDGKINYEEFVA 513
Query: 94 LMFRLSPELM 103
+M + +PEL+
Sbjct: 514 MMKKGNPELV 523
>At4g03290 putative calmodulin
Length = 154
Score = 41.6 bits (96), Expect = 1e-04
Identities = 29/70 (41%), Positives = 39/70 (55%), Gaps = 4/70 (5%)
Query: 27 GEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKE-DELVSMIREGDLDGDGA 85
GE +KE N FD D G IT++ LK + LGL+ K +E MI + D+DGDG
Sbjct: 78 GEEDMKEAFNVFDRNGD---GFITVDELKAVLSSLGLKQGKTLEECRKMIMQVDVDGDGR 134
Query: 86 LTQMEFCVLM 95
+ MEF +M
Sbjct: 135 VNYMEFRQMM 144
Score = 31.2 bits (69), Expect = 0.14
Identities = 21/59 (35%), Positives = 31/59 (51%), Gaps = 1/59 (1%)
Query: 33 ELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
EL F M G IT + L + LG+ + EDEL +I++ D++GDG + EF
Sbjct: 5 ELNRVFQMFDKDGDGKITTKELNESFKNLGII-IPEDELTQIIQKIDVNGDGCVDIEEF 62
>At3g50360 centrin
Length = 169
Score = 41.2 bits (95), Expect = 1e-04
Identities = 26/91 (28%), Positives = 47/91 (51%), Gaps = 2/91 (2%)
Query: 8 NGKG-LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDM 66
+G G ++F++ + +M K+G +EL F ++ G I+ + +K A LG ++
Sbjct: 74 DGSGAIDFDEFVHMMTAKIGERDTKEELTKAFQIIDLDKNGKISPDDIKRMAKDLG-ENF 132
Query: 67 KEDELVSMIREGDLDGDGALTQMEFCVLMFR 97
+ E+ M+ E D D DG + EF +M R
Sbjct: 133 TDAEIREMVEEADRDRDGEVNMDEFMRMMRR 163
Score = 32.0 bits (71), Expect = 0.080
Identities = 19/64 (29%), Positives = 31/64 (47%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ F++ G I + L LG + M E+++ MI + D DG GA+ EF
Sbjct: 26 QEIKEAFELFDTDGSGTIDAKELNVAMRALGFE-MTEEQINKMIADVDKDGSGAIDFDEF 84
Query: 92 CVLM 95
+M
Sbjct: 85 VHMM 88
Score = 26.2 bits (56), Expect = 4.4
Identities = 15/44 (34%), Positives = 19/44 (43%)
Query: 62 GLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEE 105
GL K+ E+ D DG G + E V M L E+ EE
Sbjct: 19 GLTTQKKQEIKEAFELFDTDGSGTIDAKELNVAMRALGFEMTEE 62
>At2g43290 putative calcium binding protein
Length = 215
Score = 41.2 bits (95), Expect = 1e-04
Identities = 32/95 (33%), Positives = 47/95 (48%), Gaps = 7/95 (7%)
Query: 8 NGKGL----EFEDLLPVMANKLGGEGLIKE--LCNGFDMLMDKDRGVITLESLKTNAAVL 61
NG G EFE L + ++ +G +E + + F++ G IT+E LK+ A L
Sbjct: 112 NGDGCVDIDEFESLYSSIVDEHHNDGETEEEDMKDAFNVFDQDGDGFITVEELKSVMASL 171
Query: 62 GLQDMKE-DELVSMIREGDLDGDGALTQMEFCVLM 95
GL+ K D MI + D DGDG + EF +M
Sbjct: 172 GLKQGKTLDGCKKMIMQVDADGDGRVNYKEFLQMM 206
>At1g05990 calcium-binding protein, putative
Length = 150
Score = 40.8 bits (94), Expect = 2e-04
Identities = 35/95 (36%), Positives = 49/95 (50%), Gaps = 9/95 (9%)
Query: 7 VNGKGL----EFEDLLP-VMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVL 61
VNG G EF +L +M + E +KE N FD D G IT++ LK + L
Sbjct: 51 VNGDGCVDIDEFGELYKTIMDEEDEEEEDMKEAFNVFDQNGD---GFITVDELKAVLSSL 107
Query: 62 GLQDMKE-DELVSMIREGDLDGDGALTQMEFCVLM 95
GL+ K D+ MI++ D+DGDG + EF +M
Sbjct: 108 GLKQGKTLDDCKKMIKKVDVDGDGRVNYKEFRQMM 142
>At2g38910 putative calcium-dependent protein kinase
Length = 583
Score = 38.9 bits (89), Expect = 7e-04
Identities = 26/84 (30%), Positives = 44/84 (51%), Gaps = 6/84 (7%)
Query: 18 LPVMANKLGGEGL--IKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMI 75
+ V+A L E + +KE+ F M+ + G ITLE LK +G D+K+ E++ ++
Sbjct: 425 IKVIAESLSEEEIAGLKEM---FKMIDTDNSGHITLEELKKGLDRVGA-DLKDSEILGLM 480
Query: 76 REGDLDGDGALTQMEFCVLMFRLS 99
+ D+D G + EF M L+
Sbjct: 481 QAADIDNSGTIDYGEFIAAMVHLN 504
Score = 30.0 bits (66), Expect = 0.30
Identities = 24/90 (26%), Positives = 38/90 (41%), Gaps = 8/90 (8%)
Query: 8 NGKGLEFEDLLPVMA--NKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQD 65
N +++ + + M NK+ E L F G IT + L+ GL D
Sbjct: 487 NSGTIDYGEFIAAMVHLNKIEKED---HLFTAFSYFDQDGSGYITRDELQQACKQFGLAD 543
Query: 66 MKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ D++ +RE D D DG + EF +M
Sbjct: 544 VHLDDI---LREVDKDNDGRIDYSEFVDMM 570
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,959,812
Number of Sequences: 26719
Number of extensions: 119519
Number of successful extensions: 447
Number of sequences better than 10.0: 92
Number of HSP's better than 10.0 without gapping: 57
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 314
Number of HSP's gapped (non-prelim): 158
length of query: 124
length of database: 11,318,596
effective HSP length: 87
effective length of query: 37
effective length of database: 8,994,043
effective search space: 332779591
effective search space used: 332779591
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0203.8


