

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
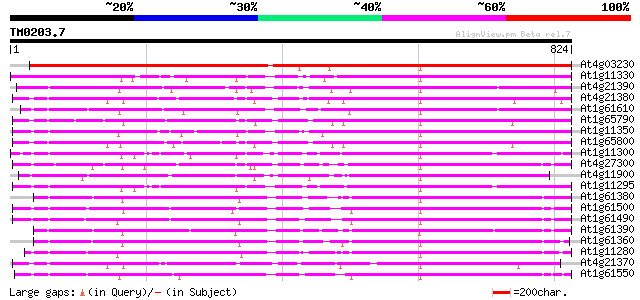
Query= TM0203.7
(824 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g03230 putative receptor kinase 1023 0.0
At1g11330 receptor-like protein kinase, putative 641 0.0
At4g21390 serine/threonine kinase - like protein 615 e-176
At4g21380 receptor-like serine/threonine protein kinase ARK3 612 e-175
At1g61610 604 e-173
At1g65790 receptor kinase, putative 595 e-170
At1g11350 putative receptor-like protein kinase gi|4008010; simi... 583 e-166
At1g65800 receptor kinase, putative 575 e-164
At1g11300 putative s-locus protein kinase (PPC:1.7.2) 572 e-163
At4g27300 putative receptor protein kinase 571 e-163
At4g11900 KI domain interacting kinase 1 -like protein 558 e-159
At1g11295 putative s-locus protein kinase (PPC:1.7.2) 555 e-158
At1g61380 S-like receptor protein kinase 551 e-157
At1g61500 549 e-156
At1g61490 S-like receptor protein kinase; member of gene cluster... 543 e-154
At1g61390 S-like receptor protein kinase; member of gene cluster... 540 e-153
At1g61360 S-like receptor protein kinase; member of gene cluster... 539 e-153
At1g11280 serine/threonine kinase like protein 536 e-152
At4g21370 receptor kinase - like protein 533 e-151
At1g61550 533 e-151
>At4g03230 putative receptor kinase
Length = 852
Score = 1023 bits (2645), Expect = 0.0
Identities = 506/823 (61%), Positives = 637/823 (76%), Gaps = 33/823 (4%)
Query: 29 LQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWVANRDNPLPDSGG 88
+ D +TL+SAG FELGFFTPNGSS+ RRY+GI ++ L P TVVWVANR++P+ D
Sbjct: 36 INDSHGETLVSAGQRFELGFFTPNGSSDERRYLGIWFYNLHPLTVVWVANRESPVLDRSC 95
Query: 89 AFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLI-VSDDKVKKILWQSF 147
F+I++DGNL V+D G+ +W T ++ +S S VKL+D+GNL+ +SD ++WQSF
Sbjct: 96 IFTISKDGNLEVIDSKGRVYWDTGVKPSSVSAERMVKLMDNGNLVLISDGNEANVVWQSF 155
Query: 148 ANPTDTFLPGMKMDDSITLTSWRSHDDPAPGNFSFEQDQGEN-QFVIWKRSMKYWKSSVS 206
NPTDTFLPGM+MD+++TL+SWRS +DP+ GNF+F+ DQ E+ QF+IWKRSM+YWKS +S
Sbjct: 156 QNPTDTFLPGMRMDENMTLSSWRSFNDPSHGNFTFQMDQEEDKQFIIWKRSMRYWKSGIS 215
Query: 207 GKFVGTGEMSSAISYLLSNFTLRISPNN-TIPFLTSSLYSNTRLVMTYWGQLQYLKMDSM 265
GKF+G+ EM AISY LSNFT ++ +N ++P L +SLY+NTR M+ GQ QY ++D
Sbjct: 216 GKFIGSDEMPYAISYFLSNFTETVTVHNASVPPLFTSLYTNTRFTMSSSGQAQYFRLDGE 275
Query: 266 KMWLMVWVEPRDRCSVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRK 325
+ W +W EPRD CSV+NACGNFGSCNSK + MCKCLPGFRPN +E W GDFSGGCSR+
Sbjct: 276 RFWAQIWAEPRDECSVYNACGNFGSCNSKNEEMCKCLPGFRPNFLEKWVKGDFSGGCSRE 335
Query: 326 TNVCSEDAK--SDTFLSLRMMKVGNPDAQFNAKNEEECKLECLNNCQCYAYSYEEYEKAR 383
+ +C +D D FL+L +++VG+PD+QF+A NE+EC+ ECLNNCQC AYSYEE
Sbjct: 336 SRICGKDGVVVGDMFLNLSVVEVGSPDSQFDAHNEKECRAECLNNCQCQAYSYEEV---- 391
Query: 384 QGDSVDPNAICWIWSQDLNNLEEEYEGGCDLHVRVAFSDI-----EGDSYQTERNLSSPT 438
D + N CWIW +DLNNL+E Y G ++ +RVA DI G E
Sbjct: 392 --DILQSNTKCWIWLEDLNNLKEGYLGSRNVFIRVAVPDIGSHVERGRGRYGEAKTPVVL 449
Query: 439 IILVTFTAVVFLIILSSTC--IYLRRRRQAKI-----RGIKLYGSERYVRDLIESGRLQE 491
II+VTFT+ L++LSST ++L+RR+ K RG+ L SER++++LIESGR ++
Sbjct: 450 IIVVTFTSAAILVVLSSTASYVFLQRRKVNKELGSIPRGVHLCDSERHIKELIESGRFKQ 509
Query: 492 DDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQG 551
DD++ ID+P F LE+IL AT+NF+ ANKLGQGGFGPVYKG FPG QEIAVKRLS CSGQG
Sbjct: 510 DDSQGIDVPSFELETILYATSNFSNANKLGQGGFGPVYKGMFPGDQEIAVKRLSRCSGQG 569
Query: 552 LEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WD 603
LEEFKNEVVLIA+LQHRNLVRLLGYCV G+EK+L+YEYMP++SLD FIFD W
Sbjct: 570 LEEFKNEVVLIAKLQHRNLVRLLGYCVAGEEKLLLYEYMPHKSLDFFIFDRKLCQRLDWK 629
Query: 604 MRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVG 663
MR IILGIARGLLYLH+DSRLRIIHRDLK SNILLDEE NPKISDFGLARIFGG +T
Sbjct: 630 MRCNIILGIARGLLYLHQDSRLRIIHRDLKTSNILLDEEMNPKISDFGLARIFGGSETSA 689
Query: 664 NTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYA 723
NT+RVVGTYGYMSPEYAL+G FS KSDVFSFGVVV+E ISGKRNTGF++PE LSLLG+A
Sbjct: 690 NTNRVVGTYGYMSPEYALEGLFSFKSDVFSFGVVVIETISGKRNTGFHEPEKSLSLLGHA 749
Query: 724 WHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLG-SESN 782
W LWK R ++ +DQ L ++C+ E +KC+NVGLLC+QEDP +RPTMSNV+FMLG SE+
Sbjct: 750 WDLWKAERGIELLDQALQESCETEGFLKCLNVGLLCVQEDPNDRPTMSNVVFMLGSSEAA 809
Query: 783 TLPSPREPAFVIRRCP-SSRASTSSKMETFSRNDLTVTLENGR 824
TLP+P++PAFV+RRCP SS+AS+S+K ET S N+LT+TLE+GR
Sbjct: 810 TLPTPKQPAFVLRRCPSSSKASSSTKPETCSENELTITLEDGR 852
>At1g11330 receptor-like protein kinase, putative
Length = 840
Score = 641 bits (1654), Expect = 0.0
Identities = 365/866 (42%), Positives = 511/866 (58%), Gaps = 80/866 (9%)
Query: 2 VPIFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYV 61
V + L C+ + + LC +D IT ++ ++D +TL+ G F GFFTP S+ RYV
Sbjct: 12 VLLLLACTCLLSRRLCFGEDRITFSSPIKDSESETLLCKSGIFRFGFFTPVNSTTRLRYV 71
Query: 62 GIRYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKN 121
GI Y K+ QTVVWVAN+D+P+ D+ G SI +DGNL V D + W TN+ +
Sbjct: 72 GIWYEKIPIQTVVWVANKDSPINDTSGVISIYQDGNLAVTDGRNRLVWSTNVSVPVAPNA 131
Query: 122 MTVKLLDSGNLIVSDDKVK-KILWQSFANPTDTFLPGMKMDD------SITLTSWRSHDD 174
V+L+DSGNL++ D++ +ILW+SF +P D+F+P M + ++ LTSW SHDD
Sbjct: 132 TWVQLMDSGNLMLQDNRNNGEILWESFKHPYDSFMPRMTLGTDGRTGGNLKLTSWTSHDD 191
Query: 175 PAPGN-------FSFEQDQGENQFVIWKRSMKYWKSSV-SGK-FVGTGEMSSAISYLLSN 225
P+ GN F+F + +IWK ++ W+S +G+ F+G M S + L
Sbjct: 192 PSTGNYTAGIAPFTFPE------LLIWKNNVPTWRSGPWNGQVFIGLPNMDSLL--FLDG 243
Query: 226 FTLRISPNNTIPFLTSSLYSNTRLVMTYWGQLQ---YLK--MDSMKMWLMVWVEPRDRCS 280
F L TI S Y+N + + + Y K SM+ W + P C
Sbjct: 244 FNLNSDNQGTI----SMSYANDSFMYHFNLDPEGIIYQKDWSTSMRTWRIGVKFPYTDCD 299
Query: 281 VFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSE--------- 331
+ CG FGSC++ + CKC+ GF P + W+ G++S GC RK + E
Sbjct: 300 AYGRCGRFGSCHAGENPPCKCVKGFVPKNNTEWNGGNWSNGCMRKAPLQCERQRNVSNGG 359
Query: 332 -DAKSDTFLSLRMMKVGNPDAQFNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDP 390
K+D FL L+ MKV A+ + +E+ C CL+NC C AY+Y D
Sbjct: 360 GGGKADGFLKLQKMKVPI-SAERSEASEQVCPKVCLDNCSCTAYAY------------DR 406
Query: 391 NAICWIWSQDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFL 450
C +WS DL +++ G DL +RVA S+++ T NL+ ++ V+ +
Sbjct: 407 GIGCMLWSGDLVDMQSFLGSGIDLFIRVAHSELK-----THSNLA-----VMIAAPVIGV 456
Query: 451 IILSSTCIYLR----RRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLES 506
+++++ C+ L ++R AK R +L + L + K ++P F +
Sbjct: 457 MLIAAVCVLLACRKYKKRPAKDRSAELMFKR--MEALTSDNESASNQIKLKELPLFEFQV 514
Query: 507 ILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQ 566
+ +T++F++ NKLGQGGFGPVYKGK P GQEIAVKRLS SGQGLEE NEVV+I++LQ
Sbjct: 515 LATSTDSFSLRNKLGQGGFGPVYKGKLPEGQEIAVKRLSRKSGQGLEELMNEVVVISKLQ 574
Query: 567 HRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLY 618
HRNLV+LLG C+EG+E+MLVYEYMP +SLDA++FD W RF I+ GI RGLLY
Sbjct: 575 HRNLVKLLGCCIEGEERMLVYEYMPKKSLDAYLFDPMKQKILDWKTRFNIMEGICRGLLY 634
Query: 619 LHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPE 678
LH DSRL+IIHRDLKASNILLDE NPKISDFGLARIF + NT RVVGTYGYMSPE
Sbjct: 635 LHRDSRLKIIHRDLKASNILLDENLNPKISDFGLARIFRANEDEANTRRVVGTYGYMSPE 694
Query: 679 YALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQ 738
YA++G FS KSDVFS GV+ LEIISG+RN+ ++ E+ L+LL YAW LW G D
Sbjct: 695 YAMEGFFSEKSDVFSLGVIFLEIISGRRNSSSHKEENNLNLLAYAWKLWNDGEAASLADP 754
Query: 739 TLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCP 798
+ C +E KCV++GLLC+QE +RP +SNV++ML +E+ +L P++PAF++RR
Sbjct: 755 AVFDKCFEKEIEKCVHIGLLCVQEVANDRPNVSNVIWMLTTENMSLADPKQPAFIVRRGA 814
Query: 799 SSRASTSSKMETFSRNDLTVTLENGR 824
S S+ + S ND+++T GR
Sbjct: 815 SEAESSDQSSQKVSINDVSLTAVTGR 840
>At4g21390 serine/threonine kinase - like protein
Length = 849
Score = 615 bits (1586), Expect = e-176
Identities = 368/866 (42%), Positives = 508/866 (58%), Gaps = 83/866 (9%)
Query: 10 FIFTFNLCSAKDTITINNNLQDG-GEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKL 68
+ F + A +TI +L+DG L+S FELGFF+P S++ R++GI Y +
Sbjct: 16 YFFLYESSMAANTIRRGESLRDGINHKPLVSPQKTFELGFFSPGSSTH--RFLGIWYGNI 73
Query: 69 APQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLER-TSSSKNMTVKLL 127
+ VVWVANR P+ D G I+ DGNL +LD + W +N+E T+++ N V +
Sbjct: 74 EDKAVVWVANRATPISDQSGVLMISNDGNLVLLDGKNITVWSSNIESSTTNNNNRVVSIH 133
Query: 128 DSGNLIVSDDKVKKILWQSFANPTDTFLPGMKM------DDSITLTSWRSHDDPAPGNFS 181
D+GN ++S+ + +W+SF +PTDTFLP M++ D+ SWRS DP+PGN+S
Sbjct: 134 DTGNFVLSETDTDRPIWESFNHPTDTFLPQMRVRVNPQTGDNHAFVSWRSETDPSPGNYS 193
Query: 182 FEQD-QGENQFVIWK-RSMKYWKSSV--SGKFVGTGEMSSAISYLLSNFTLRISPNNT-- 235
D G + V+W+ + W+S S F G MS +YL F L P+ T
Sbjct: 194 LGVDPSGAPEIVLWEGNKTRKWRSGQWNSAIFTGIPNMSLLTNYLYG-FKLSSPPDETGS 252
Query: 236 --IPFLTSSLYSNTRLVMTYWGQLQYLKM-DSMKMWLMVWVEPRDRCSVFNACGNFGSCN 292
++ S R + Y G + L+ +++K W EP C +N CG FG C+
Sbjct: 253 VYFTYVPSDPSVLLRFKVLYNGTEEELRWNETLKKWTKFQSEPDSECDQYNRCGKFGICD 312
Query: 293 SK-YDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSE---DAKSDTFLSLRMMKVGN 348
K + +C C+ G+ SV NWS G C R+T + E D FL+L+ +K+
Sbjct: 313 MKGSNGICSCIHGYEQVSVGNWSRG-----CRRRTPLKCERNISVGEDEFLTLKSVKL-- 365
Query: 349 PDAQF---NAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLE 405
PD + N + E+C+ CL NC C AYS + C IW+QDL +L+
Sbjct: 366 PDFEIPEHNLVDPEDCRERCLRNCSCNAYS------------LVGGIGCMIWNQDLVDLQ 413
Query: 406 EEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRR-RR 464
+ GG LH+R+A S++ G++ +T+ I V +V +I++ + L R +R
Sbjct: 414 QFEAGGSSLHIRLADSEV-GENRKTK--------IAVIVAVLVGVILIGIFALLLWRFKR 464
Query: 465 QAKIRGI---KLYGSERYVRDLIESGRLQEDDAKAIDI------------PHFHLESILD 509
+ + G K + V DL +S + ++DI P F L +I
Sbjct: 465 KKDVSGAYCGKNTDTSVVVADLTKSKETTSAFSGSVDIMIEGKAVNTSELPVFSLNAIAI 524
Query: 510 ATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRN 569
ATN+F N+LG+GGFGPVYKG G+EIAVKRLS SGQG++EFKNE++LIA+LQHRN
Sbjct: 525 ATNDFCKENELGRGGFGPVYKGVLEDGREIAVKRLSGKSGQGVDEFKNEIILIAKLQHRN 584
Query: 570 LVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHE 621
LVRLLG C EG+EKMLVYEYMPN+SLD F+FD W +RF II GIARGLLYLH
Sbjct: 585 LVRLLGCCFEGEEKMLVYEYMPNKSLDFFLFDETKQALIDWKLRFSIIEGIARGLLYLHR 644
Query: 622 DSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYAL 681
DSRLRIIHRDLK SN+LLD E NPKISDFG+ARIFGG NT RVVGTYGYMSPEYA+
Sbjct: 645 DSRLRIIHRDLKVSNVLLDAEMNPKISDFGMARIFGGNQNEANTVRVVGTYGYMSPEYAM 704
Query: 682 DGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLS 741
+G FSVKSDV+SFGV++LEI+SGKRNT EH SL+GYAW+L+ GR+ + +D +
Sbjct: 705 EGLFSVKSDVYSFGVLLLEIVSGKRNTSLRSSEHG-SLIGYAWYLYTHGRSEELVDPKIR 763
Query: 742 QTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPS-- 799
TC E ++C++V +LC+Q+ ERP M++VL ML S++ TL +PR+P F R S
Sbjct: 764 VTCSKREALRCIHVAMLCVQDSAAERPNMASVLLMLESDTATLAAPRQPTFTSTRRNSID 823
Query: 800 -SRASTSSKMETFSRNDLTVTLENGR 824
+ A SS+ S N++T T+ GR
Sbjct: 824 VNFALDSSQQYIVSSNEITSTVVLGR 849
>At4g21380 receptor-like serine/threonine protein kinase ARK3
Length = 850
Score = 612 bits (1579), Expect = e-175
Identities = 365/863 (42%), Positives = 516/863 (59%), Gaps = 73/863 (8%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L ++ + N SA +++TI++N +T++S G FELGFF P S R Y+GI
Sbjct: 19 LILFPAYSISANTLSASESLTISSN------NTIVSPGNVFELGFFKPGLDS--RWYLGI 70
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y ++ +T VWVANRD PL S G I+ D NL VLD++ W TNL +
Sbjct: 71 WYKAISKRTYVWVANRDTPLSSSIGTLKIS-DSNLVVLDQSDTPVWSTNLTGGDVRSPLV 129
Query: 124 VKLLDSGNLIVSDDKVKK---ILWQSFANPTDTFLPGMKMD-DSIT-----LTSWRSHDD 174
+LLD+GN ++ D K +LWQSF PTDT LP MK+ D+ T + SW+S DD
Sbjct: 130 AELLDNGNFVLRDSKNSAPDGVLWQSFDFPTDTLLPEMKLGWDAKTGFNRFIRSWKSPDD 189
Query: 175 PAPGNFSFE-QDQGENQFVIWKRSMKYWKSSVSG--KFVGTGEMSSAISYLLSNFTL-RI 230
P+ G+FSF+ + +G + +W R + ++S +F G EM Y++ NFT +
Sbjct: 190 PSSGDFSFKLETEGFPEIFLWNRESRMYRSGPWNGIRFSGVPEMQP-FEYMVFNFTTSKE 248
Query: 231 SPNNTIPFLTSSLYSNTRLVMTYWGQLQ-YLKMDSMKMWLMVWVEPRDRCSVFNACGNFG 289
+ S +YS RL ++ G LQ + +++ + W W P+D+C + CG +G
Sbjct: 249 EVTYSFRITKSDVYS--RLSISSSGLLQRFTWIETAQNWNQFWYAPKDQCDEYKECGVYG 306
Query: 290 SCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNP 349
C+S +C C+ GF+P + + W D S GC RKT + D F+ L+ MK+ P
Sbjct: 307 YCDSNTSPVCNCIKGFKPRNPQVWGLRDGSDGCVRKTLLSC--GGGDGFVRLKKMKL--P 362
Query: 350 DAQFNAKNE----EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLE 405
D + + +EC+ +CL +C C A++ + + G C W+ +L ++
Sbjct: 363 DTTTASVDRGIGVKECEQKCLRDCNCTAFANTDIRGSGSG--------CVTWTGELFDIR 414
Query: 406 EEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQ 465
+GG DL+VR+A +D+E +RN S+ I+ + V L++LS +L +R+Q
Sbjct: 415 NYAKGGQDLYVRLAATDLED-----KRNRSAK--IIGSSIGVSVLLLLSFIIFFLWKRKQ 467
Query: 466 AK-------IRGIKLYGSERYVRDLIESGRL---QEDDAKAIDIPHFHLESILDATNNFA 515
+ I +L + + +++ S R +E++ +++P E + ATNNF+
Sbjct: 468 KRSILIETPIVDHQLRSRDLLMNEVVISSRRHISRENNTDDLELPLMEFEEVAMATNNFS 527
Query: 516 IANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLG 575
ANKLGQGGFG VYKGK GQE+AVKRLS S QG +EFKNEV LIARLQH NLVRLL
Sbjct: 528 NANKLGQGGFGIVYKGKLLDGQEMAVKRLSKTSVQGTDEFKNEVKLIARLQHINLVRLLA 587
Query: 576 YCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRI 627
CV+ EKML+YEY+ N SLD+ +FD W MRF II GIARGLLYLH+DSR RI
Sbjct: 588 CCVDAGEKMLIYEYLENLSLDSHLFDKSRNSKLNWQMRFDIINGIARGLLYLHQDSRFRI 647
Query: 628 IHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSV 687
IHRDLKASNILLD+ PKISDFG+ARIFG +T NT +VVGTYGYMSPEYA+DG FS+
Sbjct: 648 IHRDLKASNILLDKYMTPKISDFGMARIFGRDETEANTRKVVGTYGYMSPEYAMDGIFSM 707
Query: 688 KSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTL---SQTC 744
KSDVFSFGV++LEIIS KRN GFY + +L+LLG W WK G+ L+ ID + S T
Sbjct: 708 KSDVFSFGVLLLEIISSKRNKGFYNSDRDLNLLGCVWRNWKEGKGLEIIDPIITDSSSTF 767
Query: 745 KAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRAST 804
+ E ++C+ +GLLC+QE +RPTMS V+ MLGSES T+P P+ P + + R S+
Sbjct: 768 RQHEILRCIQIGLLCVQERAEDRPTMSLVILMLGSESTTIPQPKAPGYCLERSLLDTDSS 827
Query: 805 SSKM---ETFSRNDLTVTLENGR 824
SSK E+++ N +TV++ + R
Sbjct: 828 SSKQRDDESWTVNQITVSVLDAR 850
>At1g61610
Length = 842
Score = 604 bits (1558), Expect = e-173
Identities = 351/847 (41%), Positives = 493/847 (57%), Gaps = 69/847 (8%)
Query: 17 CSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWV 76
CS ++ T N+ +++G D+LIS FELGFFTP S+ RYVGI Y + PQTVVWV
Sbjct: 26 CSTSNSFTRNHTIREG--DSLISEDESFELGFFTPKNST--LRYVGIWYKNIEPQTVVWV 81
Query: 77 ANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIV-S 135
ANR+ PL D GA I +DGNL +++ ++ W TN+E S N L +G+L++ S
Sbjct: 82 ANREKPLLDHKGALKIADDGNLVIVNGQNETIWSTNVE--PESNNTVAVLFKTGDLVLCS 139
Query: 136 DDKVKKILWQSFANPTDTFLPGMK------MDDSITLTSWRSHDDPAPGNFSFEQDQ-GE 188
D +K W+SF NPTDTFLPGM+ + ++ W+S DP+PG +S D G
Sbjct: 140 DSDRRKWYWESFNNPTDTFLPGMRVRVNPSLGENRAFIPWKSESDPSPGKYSMGIDPVGA 199
Query: 189 NQFVIWKRSMKYWKSSV--SGKFVGTGEMSSAISYLLSNFTLRISPNNTIPFLTSSLYSN 246
+ VIW+ + W+S S F G +M +Y+ F L P+ + + S+
Sbjct: 200 LEIVIWEGEKRKWRSGPWNSAIFTGIPDMLRFTNYIYG-FKLSSPPDRDGSVYFTYVASD 258
Query: 247 TRLVMTYW-----GQLQYLKMDSMKMWLMVWVEPRDRCSVFNACGNFGSCNS--KYDS-M 298
+ + +W + Q+ ++ W ++ +P C +N CGN+ C+ ++DS
Sbjct: 259 SSDFLRFWIRPDGVEEQFRWNKDIRNWNLLQWKPSTECEKYNRCGNYSVCDDSKEFDSGK 318
Query: 299 CKCLPGFRPNSVENWSAGDFSGGCSRKTNV-CSED---AKSDTFLSLRMMKVGNPDAQFN 354
C C+ GF P + W+ DFSGGC R+ + C++ + D F L+ +KV + +
Sbjct: 319 CSCIDGFEPVHQDQWNNRDFSGGCQRRVPLNCNQSLVAGQEDGFTVLKGIKVPDFGSVVL 378
Query: 355 AKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYEGGCDL 414
N E CK C +C C AY+ + C IW++DL ++E GG +
Sbjct: 379 HNNSETCKDVCARDCSCKAYA------------LVVGIGCMIWTRDLIDMEHFERGGNSI 426
Query: 415 HVRVAFSDIEGDSYQTERNLSSPTIILVTFTAV-VFLIILSSTCIYLRRRRQAKIRGIKL 473
++R+A S + G + T+ ++ F+ + FL+ L CI++ + + ++
Sbjct: 427 NIRLAGSKLGGGK-------ENSTLWIIVFSVIGAFLLGL---CIWILWKFKKSLKAFLW 476
Query: 474 YGSERYVRDLIESGRLQEDDAKAI--------DIPHFHLESILDATNNFAIANKLGQGGF 525
+ V D+IE+ K + D+P F +S+ AT +FA NKLGQGGF
Sbjct: 477 KKKDITVSDIIENRDYSSSPIKVLVGDQVDTPDLPIFSFDSVASATGDFAEENKLGQGGF 536
Query: 526 GPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKML 585
G VYKG F G+EIAVKRLS S QGLEEFKNE++LIA+LQHRNLVRLLG C+E +EKML
Sbjct: 537 GTVYKGNFSEGREIAVKRLSGKSKQGLEEFKNEILLIAKLQHRNLVRLLGCCIEDNEKML 596
Query: 586 VYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNI 637
+YEYMPN+SLD F+FD W R+++I GIARGLLYLH DSRL+IIHRDLKASNI
Sbjct: 597 LYEYMPNKSLDRFLFDESKQGSLDWRKRWEVIGGIARGLLYLHRDSRLKIIHRDLKASNI 656
Query: 638 LLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVV 697
LLD E NPKISDFG+ARIF + NT RVVGTYGYM+PEYA++G FS KSDV+SFGV+
Sbjct: 657 LLDTEMNPKISDFGMARIFNYRQDHANTIRVVGTYGYMAPEYAMEGIFSEKSDVYSFGVL 716
Query: 698 VLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGL 757
+LEI+SG++N F +H SL+GYAWHLW G+T + ID + T E M+C++VG+
Sbjct: 717 ILEIVSGRKNVSFRGTDHG-SLIGYAWHLWSQGKTKEMIDPIVKDTRDVTEAMRCIHVGM 775
Query: 758 LCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLT 817
LC Q+ I RP M +VL ML S+++ LP PR+P F + S ND+T
Sbjct: 776 LCTQDSVIHRPNMGSVLLMLESQTSQLPPPRQPTFHSFLNSGDIELNFDGHDVASVNDVT 835
Query: 818 VTLENGR 824
T GR
Sbjct: 836 FTTIVGR 842
>At1g65790 receptor kinase, putative
Length = 843
Score = 595 bits (1535), Expect = e-170
Identities = 350/857 (40%), Positives = 496/857 (57%), Gaps = 66/857 (7%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L +F + N SA +++TI++N T+IS FELGFF P SS R Y+GI
Sbjct: 17 LILFLAFSVSPNTLSATESLTISSN------KTIISPSQIFELGFFNPASSS--RWYLGI 68
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y + +T VWVANRDNPL S G I+ + NL + D++ + W TN+ +
Sbjct: 69 WYKIIPIRTYVWVANRDNPLSSSNGTLKISGN-NLVIFDQSDRPVWSTNITGGDVRSPVA 127
Query: 124 VKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKMD-DSIT-----LTSWRSHDDPAP 177
+LLD+GN ++ D ++LWQSF PTDT L MK+ D T L SW++ DDP+
Sbjct: 128 AELLDNGNFLLRDSN-NRLLWQSFDFPTDTLLAEMKLGWDQKTGFNRILRSWKTTDDPSS 186
Query: 178 GNFSFEQDQGE--NQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSNFTLRISPNNT 235
G FS + + E ++ K S+ Y +G + + + Y++ NFT T
Sbjct: 187 GEFSTKLETSEFPEFYICSKESILYRSGPWNGMRFSSVPGTIQVDYMVYNFTAS-KEEVT 245
Query: 236 IPFLTSSLYSNTRLVMTYWGQLQYLK-MDSMKMWLMVWVEPRDRCSVFNACGNFGSCNSK 294
+ + +RL + G LQ L ++ + W +W P+D C + CGNFG C+S
Sbjct: 246 YSYRINKTNLYSRLYLNSAGLLQRLTWFETTQSWKQLWYSPKDLCDNYKVCGNFGYCDSN 305
Query: 295 YDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNPDAQFN 354
C C+ GF+P + + W D S GC RKT + + D F L+ MK+ PD
Sbjct: 306 SLPNCYCIKGFKPVNEQAWDLRDGSAGCMRKTRLSCDGR--DGFTRLKRMKL--PDTTAT 361
Query: 355 AKNEE----ECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYEG 410
+ E CK CL +C C A++ + G C IW++++ ++ +G
Sbjct: 362 IVDREIGLKVCKERCLEDCNCTAFANADIRNGGSG--------CVIWTREILDMRNYAKG 413
Query: 411 GCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRG 470
G DL+VR+A +++E + E+ + S V L++LS + +R+Q +
Sbjct: 414 GQDLYVRLAAAELEDKRIKNEKIIGSSI-------GVSILLLLSFVIFHFWKRKQKRSIT 466
Query: 471 IK------LYGSERYVRDLIESGR---LQEDDAKAIDIPHFHLESILDATNNFAIANKLG 521
I+ + + + D++ S R +E ++ +++P LE++ ATNNF+ NKLG
Sbjct: 467 IQTPNVDQVRSQDSLINDVVVSRRGYTSKEKKSEYLELPLLELEALATATNNFSNDNKLG 526
Query: 522 QGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGD 581
QGGFG VYKG+ G+EIAVKRLS S QG +EF NEV LIA+LQH NLVRLLG CV+
Sbjct: 527 QGGFGIVYKGRLLDGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRLLGCCVDKG 586
Query: 582 EKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLK 633
EKML+YEY+ N SLD+ +FD W RF II GIARGLLYLH+DSR RIIHRDLK
Sbjct: 587 EKMLIYEYLENLSLDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRCRIIHRDLK 646
Query: 634 ASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFS 693
ASN+LLD+ PKISDFG+ARIFG ++T NT RVVGTYGYMSPEYA+DG FS+KSDVFS
Sbjct: 647 ASNVLLDKNMTPKISDFGMARIFGREETEANTRRVVGTYGYMSPEYAMDGIFSMKSDVFS 706
Query: 694 FGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFID----QTLSQTCKAEEC 749
FGV++LEIISGKRN GFY +L+LLG+ W WK G L+ +D +LS E
Sbjct: 707 FGVLLLEIISGKRNKGFYNSNRDLNLLGFVWRHWKEGNELEIVDPINIDSLSSKFPTHEI 766
Query: 750 MKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSS--K 807
++C+ +GLLC+QE +RP MS+V+ MLGSE+ +P P+ P F I R P S+SS +
Sbjct: 767 LRCIQIGLLCVQERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCIGRSPLEADSSSSTQR 826
Query: 808 METFSRNDLTVTLENGR 824
+ + N +T+++ + R
Sbjct: 827 DDECTVNQITLSVIDAR 843
>At1g11350 putative receptor-like protein kinase gi|4008010; similar
to EST gb|AI999419.1
Length = 830
Score = 583 bits (1502), Expect = e-166
Identities = 343/859 (39%), Positives = 497/859 (56%), Gaps = 72/859 (8%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
I LL F+ LC A D IT ++ +D +T++S F GFF+P S+ RY GI
Sbjct: 6 ILLLTLICFSLRLCLATDVITFSSEFRDS--ETVVSNHSTFRFGFFSPVNSTG--RYAGI 61
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
++ + QTVVWVAN ++P+ DS G SI+++GNL V+D G+ W TN+ ++
Sbjct: 62 WFNNIPVQTVVWVANSNSPINDSSGMVSISKEGNLVVMDGRGQVHWSTNVLVPVAANTFY 121
Query: 124 VKLLDSGNLIV--SDDKVKKILWQSFANPTDTFLPGM------KMDDSITLTSWRSHDDP 175
+LL++GNL++ + + +ILW+SF +P + +LP M K S+ L SW+S DP
Sbjct: 122 ARLLNTGNLVLLGTTNTGDEILWESFEHPQNIYLPTMSLATDTKTGRSLKLRSWKSPFDP 181
Query: 176 APGNFSFEQ-DQGENQFVIWKRSMKYWKSSV-SGK-FVGTGEMSSAISYLLSNFTLRISP 232
+PG +S + V+WK + W+S +G+ F+G M Y ++ F L +S
Sbjct: 182 SPGRYSAGLIPLPFPELVVWKDDLLMWRSGPWNGQYFIGLPNMD----YRINLFELTLSS 237
Query: 233 NNTIPFLTSSLYSNTRLVM-------TYWGQLQYLKMDSMKMWLMVWVEPRDRCSVFNAC 285
+N ++ S NT L + + + + + K WL V P +C + C
Sbjct: 238 DNR-GSVSMSYAGNTLLYHFLLDSEGSVFQRDWNVAIQEWKTWLKV---PSTKCDTYATC 293
Query: 286 GNFGSCNSKYDSM--CKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDA-------KSD 336
G F SC S C C+ GF+P S W+ G+++ GC RK + E KSD
Sbjct: 294 GQFASCRFNPGSTPPCMCIRGFKPQSYAEWNNGNWTQGCVRKAPLQCESRDNNDGSRKSD 353
Query: 337 TFLSLRMMKVGNPDAQFNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWI 396
F+ ++ MKV + + Q + NE++C CL NC C AYS+ D C +
Sbjct: 354 GFVRVQKMKVPH-NPQRSGANEQDCPESCLKNCSCTAYSF------------DRGIGCLL 400
Query: 397 WSQDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSST 456
WS +L +++E G ++R+A S+ + +T R+ I++T T +V + + T
Sbjct: 401 WSGNLMDMQEFSGTGVVFYIRLADSEFKK---RTNRS------IVITVTLLVGAFLFAGT 451
Query: 457 CI---YLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILDATNN 513
+ + + + K R +L + G + + K ++P F + + ATNN
Sbjct: 452 VVLALWKIAKHREKNRNTRLLNERMEALSSNDVGAILVNQYKLKELPLFEFQVLAVATNN 511
Query: 514 FAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRL 573
F+I NKLGQGGFG VYKG+ G +IAVKRLS SGQG+EEF NEVV+I++LQHRNLVRL
Sbjct: 512 FSITNKLGQGGFGAVYKGRLQEGLDIAVKRLSRTSGQGVEEFVNEVVVISKLQHRNLVRL 571
Query: 574 LGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRL 625
LG+C+EG+E+MLVYE+MP LDA++FD W RF II GI RGL+YLH DSRL
Sbjct: 572 LGFCIEGEERMLVYEFMPENCLDAYLFDPVKQRLLDWKTRFNIIDGICRGLMYLHRDSRL 631
Query: 626 RIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHF 685
+IIHRDLKASNILLDE NPKISDFGLARIF G + +T RVVGTYGYM+PEYA+ G F
Sbjct: 632 KIIHRDLKASNILLDENLNPKISDFGLARIFQGNEDEVSTVRVVGTYGYMAPEYAMGGLF 691
Query: 686 SVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCK 745
S KSDVFS GV++LEI+SG+RN+ FY +L YAW LW G + +D + + C
Sbjct: 692 SEKSDVFSLGVILLEIVSGRRNSSFYNDGQNPNLSAYAWKLWNTGEDIALVDPVIFEECF 751
Query: 746 AEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTS 805
E +CV+VGLLC+Q+ +RP+++ V++ML SE++ LP P++PAF+ RR S S+
Sbjct: 752 ENEIRRCVHVGLLCVQDHANDRPSVATVIWMLSSENSNLPEPKQPAFIPRRGTSEVESSG 811
Query: 806 SKMETFSRNDLTVTLENGR 824
S N++++T GR
Sbjct: 812 QSDPRASINNVSLTKITGR 830
>At1g65800 receptor kinase, putative
Length = 847
Score = 575 bits (1481), Expect = e-164
Identities = 350/865 (40%), Positives = 497/865 (56%), Gaps = 78/865 (9%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
I L +F + SA +++TI++N T+IS FELGFF P+ SS R Y+GI
Sbjct: 17 IILFLAFSVYASNFSATESLTISSN------KTIISPSQIFELGFFNPDSSS--RWYLGI 68
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y + +T VWVANRDNPL S G I+ D NL + D++ + W TN+ +
Sbjct: 69 WYKIIPIRTYVWVANRDNPLSSSNGTLKIS-DNNLVIFDQSDRPVWSTNITGGDVRSPVA 127
Query: 124 VKLLDSGNLIVSDDKVKK---ILWQSFANPTDTFLPGMKMD-------DSITLTSWRSHD 173
+LLD GN ++ D K K LWQSF PTDT L MKM + L SW++ D
Sbjct: 128 AELLDYGNFVLRDSKNNKPSGFLWQSFDFPTDTLLSDMKMGWDNKSGGFNRILRSWKTTD 187
Query: 174 DPAPGNFSFE-QDQGENQFVIW-KRSMKYWKSSVSGKFVGTGEMSSAISYLLSNFTLRIS 231
DP+ G+FS + + G +F I+ K S+ Y G + + Y+ ++FT
Sbjct: 188 DPSSGDFSTKLRTSGFPEFYIYNKESITYRSGPWLGNRFSSVPGMKPVDYIDNSFT---E 244
Query: 232 PNNTIPFL----TSSLYSNTRLVMTYWGQLQYLK-MDSMKMWLMVWVEPRDRCSVFNACG 286
N + + +++YS L T G LQ L M++ + W +W P+D C + CG
Sbjct: 245 NNQQVVYSYRVNKTNIYSILSLSST--GLLQRLTWMEAAQSWKQLWYSPKDLCDNYKECG 302
Query: 287 NFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKV 346
N+G C++ +C C+ GF P + E + D S GC RKT + + D F+ L+ M++
Sbjct: 303 NYGYCDANTSPICNCIKGFEPMN-EQAALRDDSVGCVRKTKLSCDGR--DGFVRLKKMRL 359
Query: 347 GNPDAQFNAKNE----EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLN 402
PD + ++ +EC+ CL C C A++ + G C IWS L
Sbjct: 360 --PDTTETSVDKGIGLKECEERCLKGCNCTAFANTDIRNGGSG--------CVIWSGGLF 409
Query: 403 NLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRR 462
++ +GG DL+VRVA D+E ++++ + S V L++LS + +
Sbjct: 410 DIRNYAKGGQDLYVRVAAGDLEDKRIKSKKIIGSSI-------GVSILLLLSFIIFHFWK 462
Query: 463 RRQAKIRGIK------LYGSERYVRDLIESGRL---QEDDAKAIDIPHFHLESILDATNN 513
R+Q + I+ + + + +L+++ R +E+ +++P +++ ATNN
Sbjct: 463 RKQKRSITIQTPIVDLVRSQDSLMNELVKASRSYTSKENKTDYLELPLMEWKALAMATNN 522
Query: 514 FAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRL 573
F+ NKLGQGGFG VYKG G+EIAVKRLS S QG +EF NEV LIA+LQH NLVRL
Sbjct: 523 FSTDNKLGQGGFGIVYKGMLLDGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRL 582
Query: 574 LGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRL 625
LG CV+ EKML+YEY+ N SLD+ +FD W RF II GIARGLLYLH+DSR
Sbjct: 583 LGCCVDKGEKMLIYEYLENLSLDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRC 642
Query: 626 RIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHF 685
RIIHRDLKASN+LLD+ PKISDFG+ARIFG ++T NT RVVGTYGYMSPEYA+DG F
Sbjct: 643 RIIHRDLKASNVLLDKNMTPKISDFGMARIFGREETEANTRRVVGTYGYMSPEYAMDGIF 702
Query: 686 SVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFID----QTLS 741
S+KSDVFSFGV++LEIISGKRN GFY +L+LLG+ W WK G+ L+ +D LS
Sbjct: 703 SMKSDVFSFGVLLLEIISGKRNKGFYNSNRDLNLLGFVWRHWKEGKELEIVDPINIDALS 762
Query: 742 QTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSR 801
E ++C+ +GLLC+QE +RP MS+V+ MLGSE+ +P P+ P F + R
Sbjct: 763 SEFPTHEILRCIQIGLLCVQERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCVGRSSLEV 822
Query: 802 ASTSS--KMETFSRNDLTVTLENGR 824
S+SS + + + N +T+++ + R
Sbjct: 823 DSSSSTQRDDECTVNQVTLSVIDAR 847
>At1g11300 putative s-locus protein kinase (PPC:1.7.2)
Length = 820
Score = 572 bits (1473), Expect = e-163
Identities = 338/858 (39%), Positives = 489/858 (56%), Gaps = 83/858 (9%)
Query: 2 VPIFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYV 61
V I +L F + +L A++ + L D +T++S+ F GFF+P S++ RY
Sbjct: 11 VCILVLSCFFLSVSL--AQERAFFSGKLNDS--ETIVSSFRTFRFGFFSPVNSTS--RYA 64
Query: 62 GIRYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKN 121
GI Y+ ++ QTV+WVAN+D P+ DS G S+++DGNL V D + W TN+ +S+ +
Sbjct: 65 GIWYNSVSVQTVIWVANKDKPINDSSGVISVSQDGNLVVTDGQRRVLWSTNVSTQASANS 124
Query: 122 MTVKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKMDDS-------ITLTSWRSHDD 174
+LLDSGNL++ + LW+SF PTD++LP M + + +T+TSW+S D
Sbjct: 125 TVAELLDSGNLVLKEASSDAYLWESFKYPTDSWLPNMLVGTNARIGGGNVTITSWKSPSD 184
Query: 175 PAPGNFS-----------FEQDQGENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLL 223
P+PG+++ F + N +W+ W + F G ++ + + L
Sbjct: 185 PSPGSYTAALVLAAYPELFIMNNNNNNSTVWRSGP--WNGQM---FNGLPDVYAGV--FL 237
Query: 224 SNFTLRISPNNTIPFLTSSLYSNTRLVMTYWGQLQYLKMD---SMKMWLMVWVEPRDRCS 280
F + N ++ ++ + M Y G + ++ D + + W + P C
Sbjct: 238 YRFIVNDDTNGSVTMSYANDSTLRYFYMDYRGSV--IRRDWSETRRNWTVGLQVPATECD 295
Query: 281 VFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSE----DAKSD 336
+ CG F +CN + + +C C+ GFRP ++ W+ G++SGGC+R+ + E + +D
Sbjct: 296 NYRRCGEFATCNPRKNPLCSCIRGFRPRNLIEWNNGNWSGGCTRRVPLQCERQNNNGSAD 355
Query: 337 TFLSLRMMKVGNPD-AQFNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICW 395
FL LR MK+ PD A+ + +E EC CL C C A A G C
Sbjct: 356 GFLRLRRMKL--PDFARRSEASEPECLRTCLQTCSCIA--------AAHGLGYG----CM 401
Query: 396 IWSQDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSS 455
IW+ L + +E G DL++R+A S+I+ + P +I +F++ +
Sbjct: 402 IWNGSLVDSQELSASGLDLYIRLAHSEIKTKDKR-------PILIGTILAGGIFVV---A 451
Query: 456 TCIYLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILDATNNFA 515
C+ L RR K R K + + +E+ + K ++P F + + ATNNF+
Sbjct: 452 ACVLLARRIVMKKRAKKKGRDAEQIFERVEA-LAGGNKGKLKELPLFEFQVLAAATNNFS 510
Query: 516 IANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLG 575
+ NKLGQGGFGPVYKGK GQEIAVKRLS SGQGLEE NEVV+I++LQHRNLV+LLG
Sbjct: 511 LRNKLGQGGFGPVYKGKLQEGQEIAVKRLSRASGQGLEELVNEVVVISKLQHRNLVKLLG 570
Query: 576 YCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRI 627
C+ G+E+MLVYE+MP +SLD ++FD W RF II GI RGLLYLH DSRLRI
Sbjct: 571 CCIAGEERMLVYEFMPKKSLDYYLFDSRRAKLLDWKTRFNIINGICRGLLYLHRDSRLRI 630
Query: 628 IHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSV 687
IHRDLKASNILLDE PKISDFGLARIF G + NT RVVGTYGYM+PEYA+ G FS
Sbjct: 631 IHRDLKASNILLDENLIPKISDFGLARIFPGNEDEANTRRVVGTYGYMAPEYAMGGLFSE 690
Query: 688 KSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAE 747
KSDVFS GV++LEIISG+RN+ +LL Y W +W G +D + +
Sbjct: 691 KSDVFSLGVILLEIISGRRNS-------NSTLLAYVWSIWNEGEINSLVDPEIFDLLFEK 743
Query: 748 ECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIR-RCPSSRASTSS 806
E KC+++GLLC+QE +RP++S V ML SE +P P++PAF+ R P + +S +S
Sbjct: 744 EIHKCIHIGLLCVQEAANDRPSVSTVCSMLSSEIADIPEPKQPAFISRNNVPEAESSENS 803
Query: 807 KMETFSRNDLTVTLENGR 824
++ S N++T+T GR
Sbjct: 804 DLKD-SINNVTITDVTGR 820
>At4g27300 putative receptor protein kinase
Length = 815
Score = 571 bits (1471), Expect = e-163
Identities = 349/855 (40%), Positives = 496/855 (57%), Gaps = 80/855 (9%)
Query: 4 IFLLCSFIFTFNLCSAKD--TITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGR-RY 60
+F L F+ + +L A D IT L+DG DTL S F+LGFF+ + + R+
Sbjct: 7 LFSLSLFLISSSLSVALDYNVITPKEFLKDG--DTLSSPDQVFQLGFFSLDQEEQPQHRF 64
Query: 61 VGIRYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSK 120
+G+ Y + P VVWVANR+NPL + G +++ G+L++ D K+ W ++ T +SK
Sbjct: 65 LGLWY--MEPFAVVWVANRNNPLYGTSGFLNLSSLGDLQLFDGEHKALWSSSSSSTKASK 122
Query: 121 ---NMTVKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKMDDSI------TLTSWRS 171
N +K+ SGNLI SD + + +LWQSF P +T L GMK+ + +L+SW++
Sbjct: 123 TANNPLLKISCSGNLISSDGE-EAVLWQSFDYPMNTILAGMKLGKNFKTQMEWSLSSWKT 181
Query: 172 HDDPAPGNFSFEQD-QGENQFVIWKR---SMKYWKSSVSG-KFVGTGEMSSAISYLLSNF 226
DP+PG+F+ D +G Q ++ K S Y S +G F G M S F
Sbjct: 182 LKDPSPGDFTLSLDTRGLPQLILRKNGDSSYSYRLGSWNGLSFTGAPAMGRENSLFDYKF 241
Query: 227 TLRISPNNTIPFLTSSLYSNTRLVMTYWGQLQYLKMDSMKMWLMVWVEPRDRCSVFNACG 286
T N + S RLV+ G+L W++ P D C ++ CG
Sbjct: 242 TSSAQEVNYSWTPRHRIVS--RLVLNNTGKLHRFIQSKQNQWILANTAPEDECDYYSICG 299
Query: 287 NFGSC--NSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMM 344
+ C NSK C CL GF+P S W+ + GC + E K D F+ +
Sbjct: 300 AYAVCGINSKNTPSCSCLQGFKPKSGRKWNISRGAYGCVHEIPTNCE--KKDAFVKFPGL 357
Query: 345 KVGNPDAQ---FNAKNE---EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWS 398
K+ PD ++AKNE E+CK++C +NC C AY+ + + +G C +W
Sbjct: 358 KL--PDTSWSWYDAKNEMTLEDCKIKCSSNCSCTAYANTDIREGGKG--------CLLWF 407
Query: 399 QDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCI 458
DL ++ E G D+++R+ F+ IE R + + V AVV +++ +
Sbjct: 408 GDLVDMREYSSFGQDVYIRMGFAKIEFKG----REVVGMVVGSVVAIAVVLVVVFAC--- 460
Query: 459 YLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILDATNNFAIAN 518
R++ + RG + G +ED +D+P F ++I AT++F+ N
Sbjct: 461 -FRKKIMKRYRG-----------ENFRKGIEEED----LDLPIFDRKTISIATDDFSYVN 504
Query: 519 KLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCV 578
LG+GGFGPVYKGK GQEIAVKRLS+ SGQG+EEFKNEV LIA+LQHRNLVRLLG C+
Sbjct: 505 FLGRGGFGPVYKGKLEDGQEIAVKRLSANSGQGVEEFKNEVKLIAKLQHRNLVRLLGCCI 564
Query: 579 EGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHR 630
+G+E ML+YEYMPN+SLD FIFD W R II G+ARG+LYLH+DSRLRIIHR
Sbjct: 565 QGEECMLIYEYMPNKSLDFFIFDERRSTELDWKKRMNIINGVARGILYLHQDSRLRIIHR 624
Query: 631 DLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSD 690
DLKA N+LLD + NPKISDFGLA+ FGG + +T+RVVGTYGYM PEYA+DGHFSVKSD
Sbjct: 625 DLKAGNVLLDNDMNPKISDFGLAKSFGGDQSESSTNRVVGTYGYMPPEYAIDGHFSVKSD 684
Query: 691 VFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQT-LSQTCKAEEC 749
VFSFGV+VLEII+GK N GF +H+L+LLG+ W +W R ++ ++ L +T E
Sbjct: 685 VFSFGVLVLEIITGKTNRGFRHADHDLNLLGHVWKMWVEDREIEVPEEEWLEETSVIPEV 744
Query: 750 MKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKME 809
++C++V LLC+Q+ P +RPTM++V+ M GS+S +LP P +P F R + SS +
Sbjct: 745 LRCIHVALLCVQQKPEDRPTMASVVLMFGSDS-SLPHPTQPGFFTNR---NVPDISSSLS 800
Query: 810 TFSRNDLTVTLENGR 824
S+N++++T+ GR
Sbjct: 801 LRSQNEVSITMLQGR 815
>At4g11900 KI domain interacting kinase 1 -like protein
Length = 849
Score = 558 bits (1438), Expect = e-159
Identities = 340/836 (40%), Positives = 486/836 (57%), Gaps = 89/836 (10%)
Query: 13 TFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRR--YVGIRYHKLAP 70
+F + S+ DTI+ N L G +T++S+G FELG FTP + R Y+G+ Y ++P
Sbjct: 20 SFQVSSSTDTISTNQPLS--GFETIVSSGDIFELGLFTPTPDTYDHRNYYIGMWYRHVSP 77
Query: 71 QTVVWVANRDNPLPDSGGAFSITE-DGNLRVLDRTG------------------------ 105
QT+VWVANR++PL + + DGNL + D
Sbjct: 78 QTIVWVANRESPLGGDASTYLLKILDGNLILHDNISATRKSHTEGTSRRSPQKISEGNLL 137
Query: 106 --KSFWGTNLERTSSSKNMTVKLLDSGNLIVSD--DKVKKILWQSFANPTDTFLPGMKMD 161
++ W T + +S SK++ L DSGNL++ D + +LWQSF +P+DT+LPG K+
Sbjct: 138 FHETVWSTGVN-SSMSKDVQAVLFDSGNLVLRDGPNSSAAVLWQSFDHPSDTWLPGGKIR 196
Query: 162 -DSITLTSWRSHDDPAPGNFSFEQDQGENQFV-IWKRSMKYWKSSVSGKFVGTGEMSSAI 219
S TSW S DP+PG +S E D + V +W RS YW S ++ + + +
Sbjct: 197 LGSQLFTSWESLIDPSPGRYSLEFDPKLHSLVTVWNRSKSYWSSGPLYDWLQSFKGFPEL 256
Query: 220 SYLLSNFTLRISPNNTIPFLTSSL--YSNTRLVMTYWGQ--LQYLKMDSMKMWLMVWVEP 275
+FTL + + ++T S+ S RLVM GQ LQ +D ++ W ++ +P
Sbjct: 257 QGTKLSFTLNMDES----YITFSVDPQSRYRLVMGVSGQFMLQVWHVD-LQSWRVILSQP 311
Query: 276 RDRCSVFNACGNFGSCNSKYDSM-CKCLPGF-RPNSVENWSAGDFSGGCSRKTNV-CSED 332
+RC V+N+CG+FG CN + C+C+PGF R S + + D+SGGC R+T + C +
Sbjct: 312 DNRCDVYNSCGSFGICNENREPPPCRCVPGFKREFSQGSDDSNDYSGGCKRETYLHCYK- 370
Query: 333 AKSDTFLSLRMMKVGNPDAQFNAKNE---EECKLECLNNCQCYAYSYEEYEKARQGDSVD 389
++D FL + MK+ + C C+ +C C AY A G+
Sbjct: 371 -RNDEFLPIENMKLATDPTTASVLTSGTFRTCASRCVADCSCQAY-------ANDGNK-- 420
Query: 390 PNAICWIWSQDLNNLEE-EYEGGCDLHVRVAFSDIE-GDSYQTERNLSSPTI---ILVTF 444
C +W++D NL++ + G +R+A S+I ++ +TE + + +L +
Sbjct: 421 ----CLVWTKDAFNLQQLDANKGHTFFLRLASSNISTANNRKTEHSKGKSIVLPLVLASL 476
Query: 445 TAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHL 504
A + CI R RR+ K R E++ R+L+E G + DDA ++ + +L
Sbjct: 477 VATAACFVGLYCCISSRIRRKKKQR------DEKHSRELLEGGLI--DDAGE-NMCYLNL 527
Query: 505 ESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIAR 564
I+ ATN+F+ KLG+GGFGPVYKGK P G E+A+KRLS S QGL EFKNEVVLI +
Sbjct: 528 HDIMVATNSFSRKKKLGEGGFGPVYKGKLPNGMEVAIKRLSKKSSQGLTEFKNEVVLIIK 587
Query: 565 LQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGL 616
LQH+NLVRLLGYCVEGDEK+L+YEYM N+SLD +FD W+ R KI+ G RGL
Sbjct: 588 LQHKNLVRLLGYCVEGDEKLLIYEYMSNKSLDGLLFDSLKSRELDWETRMKIVNGTTRGL 647
Query: 617 LYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMS 676
YLHE SRLRIIHRDLKASNILLD+E NPKISDFG ARIFG K +T R+VGT+GYMS
Sbjct: 648 QYLHEYSRLRIIHRDLKASNILLDDEMNPKISDFGTARIFGCKQIDDSTQRIVGTFGYMS 707
Query: 677 PEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFI 736
PEYAL G S KSD++SFGV++LEIISGK+ T F + + SL+ Y W W + + I
Sbjct: 708 PEYALGGVISEKSDIYSFGVLLLEIISGKKATRFVHNDQKHSLIAYEWESWCETKGVSII 767
Query: 737 DQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAF 792
D+ + + EE M+C+++ LLC+Q+ P +RP +S +++ML S NTLP P++P F
Sbjct: 768 DEPMCCSYSLEEAMRCIHIALLCVQDHPKDRPMISQIVYML-SNDNTLPIPKQPTF 822
>At1g11295 putative s-locus protein kinase (PPC:1.7.2)
Length = 820
Score = 555 bits (1429), Expect = e-158
Identities = 328/855 (38%), Positives = 480/855 (55%), Gaps = 79/855 (9%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ +L F ++ A + + L D +T++S+ F GFF+P S+N RY GI
Sbjct: 11 VHVLSLSCFFLSVSLAHERALFSGTLNDS--ETIVSSFRTFRFGFFSPVNSTN--RYAGI 66
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y+ + QTV+WVAN+D P+ DS G SI+EDGNL V D + W TN+ +S+ +
Sbjct: 67 WYNSIPVQTVIWVANKDTPINDSSGVISISEDGNLVVTDGQRRVLWSTNVSTRASANSTV 126
Query: 124 VKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKMDDS-------ITLTSWRSHDDPA 176
+LL+SGNL++ D LW+SF PTD++LP M + + IT+TSW + DP+
Sbjct: 127 AELLESGNLVLKDANTDAYLWESFKYPTDSWLPNMLVGTNARTGGGNITITSWTNPSDPS 186
Query: 177 PGNFS-----------FEQDQGENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSN 225
PG+++ F + +N +W+ W + F G ++ + L
Sbjct: 187 PGSYTAALVLAPYPELFIFNNNDNNATVWRSGP--WNGLM---FNGLPDVYPGL--FLYR 239
Query: 226 FTLRISPNNTIPFLTSSLYSNTRLVMTYWG-QLQYLKMDSMKMWLMVWVEPRDRCSVFNA 284
F + N + ++ + L + Y G ++ ++ + W + P C +++
Sbjct: 240 FKVNDDTNGSATMSYANDSTLRHLYLDYRGFAIRRDWSEARRNWTLGSQVPATECDIYSR 299
Query: 285 CGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSE----DAKSDTFLS 340
CG + +CN + + C C+ GFRP ++ W+ G++SGGC RK + E +D FL
Sbjct: 300 CGQYTTCNPRKNPHCSCIKGFRPRNLIEWNNGNWSGGCIRKLPLQCERQNNKGSADRFLK 359
Query: 341 LRMMKVGNPD-AQFNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQ 399
L+ MK+ PD A+ + +E EC + CL +C C A+++ C IW++
Sbjct: 360 LQRMKM--PDFARRSEASEPECFMTCLQSCSCIAFAH------------GLGYGCMIWNR 405
Query: 400 DLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIY 459
L + + G DL +R+A S+ + + P +I + +F++ +TC+
Sbjct: 406 SLVDSQVLSASGMDLSIRLAHSEFKTQDRR-------PILIGTSLAGGIFVV---ATCVL 455
Query: 460 LRRRRQAKIRGIKLYGSERYVRDLIES--GRLQEDDAKAIDIPHFHLESILDATNNFAIA 517
L RR K R K + +E+ G +E K ++P F + + AT+NF+++
Sbjct: 456 LARRIVMKKRAKKKGTDAEQIFKRVEALAGGSRE---KLKELPLFEFQVLATATDNFSLS 512
Query: 518 NKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYC 577
NKLGQGGFGPVYKG GQEIAVKRLS SGQGLEE EVV+I++LQHRNLV+L G C
Sbjct: 513 NKLGQGGFGPVYKGMLLEGQEIAVKRLSQASGQGLEELVTEVVVISKLQHRNLVKLFGCC 572
Query: 578 VEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIH 629
+ G+E+MLVYE+MP +SLD +IFD W+ RF+II GI RGLLYLH DSRLRIIH
Sbjct: 573 IAGEERMLVYEFMPKKSLDFYIFDPREAKLLDWNTRFEIINGICRGLLYLHRDSRLRIIH 632
Query: 630 RDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKS 689
RDLKASNILLDE PKISDFGLARIF G + NT RVVGTYGYM+PEYA+ G FS KS
Sbjct: 633 RDLKASNILLDENLIPKISDFGLARIFPGNEDEANTRRVVGTYGYMAPEYAMGGLFSEKS 692
Query: 690 DVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEEC 749
DVFS GV++LEIISG+RN+ +LL + W +W G +D + +E
Sbjct: 693 DVFSLGVILLEIISGRRNS-------HSTLLAHVWSIWNEGEINGMVDPEIFDQLFEKEI 745
Query: 750 MKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKME 809
KCV++ LLC+Q+ +RP++S V ML SE +P P++PAF+ R + S
Sbjct: 746 RKCVHIALLCVQDAANDRPSVSTVCMMLSSEVADIPEPKQPAFMPRNVGLEAEFSESIAL 805
Query: 810 TFSRNDLTVTLENGR 824
S N++T+T +GR
Sbjct: 806 KASINNVTITDVSGR 820
>At1g61380 S-like receptor protein kinase
Length = 805
Score = 551 bits (1421), Expect = e-157
Identities = 329/814 (40%), Positives = 459/814 (55%), Gaps = 67/814 (8%)
Query: 36 TLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWVANRDNPLPDSGGAFSITED 95
TL S GG++ELGFF+PN + N +YVGI + K+ P+ VVWVANRD P+ S +I+ +
Sbjct: 34 TLSSPGGFYELGFFSPNNTQN--QYVGIWFKKIVPRVVVWVANRDTPVTSSAANLTISSN 91
Query: 96 GNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIVSDDKVKKILWQSFANPTDTFL 155
G+L +LD W T TS+ + +LLD+GN +V DD LWQSF + +T L
Sbjct: 92 GSLILLDGKQDVIWSTGKAFTSNKCH--AELLDTGNFVVIDDVSGNKLWQSFEHLGNTML 149
Query: 156 P--GMKMDDSI----TLTSWRSHDDPAPGNFSFE-QDQGENQFVIWKRSMKYWKSSVSGK 208
P + D S LT+W+S+ DP+PG FS E Q Q +I + S+ YW+ K
Sbjct: 150 PQSSLMYDTSNGKKRVLTTWKSNSDPSPGEFSLEITPQIPTQGLIRRGSVPYWRCGPWAK 209
Query: 209 FVGTGEMSSAISYLLSNFTLRISPNNTIPFLTSSL--YSNTRLVMTYWGQLQYLKMDSMK 266
+G SY+ ++ + T F S+L Y+ + + +T G+++ L D
Sbjct: 210 TRFSGISGIDASYVSPFSVVQDTAAGTGSFSYSTLRNYNLSYVTLTPEGKMKIL-WDDGN 268
Query: 267 MWLMVWVEPRDRCSVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKT 326
W + P + C ++ CG +G C C+CL GF P S E W G+++ GC R+T
Sbjct: 269 NWKLHLSLPENPCDLYGRCGPYGLCVRSDPPKCECLKGFVPKSDEEWGKGNWTSGCVRRT 328
Query: 327 NVCSEDAKS------DTFLSLRMMKVGNPDAQFNAK--NEEECKLECLNNCQCYAYSYEE 378
+ + S DT + RM V PD A N E+C CL NC C A++Y
Sbjct: 329 KLSCQAKSSMKTQGKDTDIFYRMTDVKTPDLHQFASFLNAEQCYQGCLGNCSCTAFAYIS 388
Query: 379 YEKARQGDSVDPNAICWIWSQDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPT 438
C +W+ +L + + G L +R+A S++ G S +
Sbjct: 389 ------------GIGCLVWNGELADTVQFLSSGEFLFIRLASSELAGSSRRK-------I 429
Query: 439 IILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAID 498
I+ T + +FLI++ + + R R + D ++G + D ++
Sbjct: 430 IVGTTVSLSIFLILVFAAIMLWRYRAKQN--------------DAWKNG-FERQDVSGVN 474
Query: 499 IPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNE 558
F + +I ATNNF+ +NKLGQGGFGPVYKGK G+EI VKRL+S SGQG EEF NE
Sbjct: 475 F--FEMHTIRTATNNFSPSNKLGQGGFGPVYKGKLVDGKEIGVKRLASSSGQGTEEFMNE 532
Query: 559 VVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIIL 610
+ LI++LQHRNLVRLLGYC++G+EK+L+YE+M N+SLD FIFD W RF II
Sbjct: 533 ITLISKLQHRNLVRLLGYCIDGEEKLLIYEFMVNKSLDIFIFDPCLKFELDWPKRFNIIQ 592
Query: 611 GIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVG 670
GIARGLLYLH DSRLR+IHRDLK SNILLD+ NPKISDFGLAR+F G NT RVVG
Sbjct: 593 GIARGLLYLHRDSRLRVIHRDLKVSNILLDDRMNPKISDFGLARMFQGTQYQDNTRRVVG 652
Query: 671 TYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVG 730
T GYMSPEYA G FS KSD++SFGV++LEIISGKR + F + LL Y W W
Sbjct: 653 TLGYMSPEYAWAGLFSEKSDIYSFGVLMLEIISGKRISRFIYGDESKGLLAYTWDSWCET 712
Query: 731 RTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREP 790
+ +D+ L+ TC+A E +CV +GLLC+Q + ++RP VL ML S ++ LP P++P
Sbjct: 713 GGSNLLDRDLTDTCQAFEVARCVQIGLLCVQHEAVDRPNTLQVLSMLTSATD-LPVPKQP 771
Query: 791 AFVIRRCPSSRASTSSKMETFSRNDLTVTLENGR 824
F + ++ + S N++T ++ GR
Sbjct: 772 IFAVHTLNDMPMLQANSQDFLSVNEMTESMIQGR 805
>At1g61500
Length = 804
Score = 549 bits (1415), Expect = e-156
Identities = 340/847 (40%), Positives = 476/847 (56%), Gaps = 76/847 (8%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L F+FT S+ IT + L G TL SA +ELGFF+PN + + +YVGI
Sbjct: 8 LHLFTMFLFTLLSGSSSAVITTESPLSMG--QTLSSANEVYELGFFSPNNTQD--QYVGI 63
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
+ P+ VVWVANR+ P+ DS +I+ G+L +L+ + W + + T SS
Sbjct: 64 WFKDTIPRVVVWVANREKPVTDSTAYLAISSSGSLLLLNGKHGTVWSSGV--TFSSSGCR 121
Query: 124 VKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKMDDSIT------LTSWRSHDDPAP 177
+L DSGNL V D+ ++ LWQSF + DT L + ++ LTSW+S+ DP+P
Sbjct: 122 AELSDSGNLKVIDNVSERALWQSFDHLGDTLLHTSSLTYNLATAEKRVLTSWKSYTDPSP 181
Query: 178 GNFSFE-QDQGENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSNFTLRISPNNTI 236
G+F + Q +Q + + S YW+S K TG SY FTL N +
Sbjct: 182 GDFLGQITPQVPSQGFVMRGSTPYWRSGPWAKTRFTGIPFMDESYT-GPFTLHQDVNGS- 239
Query: 237 PFLT--SSLYSNTRLVMTYWGQLQYLKMDSMKMWLMVWVEPRDRCSVFNACGNFGSCNSK 294
+LT Y +R+ +T G ++ + + M W + + P+ C + ACG FG C
Sbjct: 240 GYLTYFQRDYKLSRITLTSEGSIKMFRDNGMG-WELYYEAPKKLCDFYGACGPFGLCVMS 298
Query: 295 YDSMCKCLPGFRPNSVENWSAGDFSGGCSRKT------NVCSEDAKSDTFLSLRMMKVGN 348
MCKC GF P SVE W G+++GGC R T N EDA D F + +K +
Sbjct: 299 PSPMCKCFRGFVPKSVEEWKRGNWTGGCVRHTELDCLGNSTGEDA--DDFHQIANIKPPD 356
Query: 349 PDAQFNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEY 408
++ N EEC C++NC C A++Y + C +W+QDL + +
Sbjct: 357 FYEFASSVNAEECHQRCVHNCSCLAFAYIK------------GIGCLVWNQDLMDAVQFS 404
Query: 409 EGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKI 468
G L +R+A S+++G+ + I+ + ++ +IL T + R R I
Sbjct: 405 ATGELLSIRLARSELDGNKRKKT--------IVASIVSLTLFMILGFTAFGVWRCRVEHI 456
Query: 469 RGIKLYGSERYVRDLIESGRLQEDDAKAIDIP---HFHLESILDATNNFAIANKLGQGGF 525
I S ++D K D+P F + +I +ATNNF+++NKLGQGGF
Sbjct: 457 AHI--------------SKDAWKNDLKPQDVPGLDFFDMHTIQNATNNFSLSNKLGQGGF 502
Query: 526 GPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKML 585
G VYKGK G+EIAVKRLSS SGQG EEF NE+VLI++LQHRNLVR+LG C+E +EK+L
Sbjct: 503 GSVYKGKLQDGKEIAVKRLSSSSGQGKEEFMNEIVLISKLQHRNLVRVLGCCIEEEEKLL 562
Query: 586 VYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNI 637
+YE+M N+SLD F+FD W RF II GIARGLLYLH DSRLR+IHRDLK SNI
Sbjct: 563 IYEFMVNKSLDTFLFDSRKRLEIDWPKRFDIIQGIARGLLYLHHDSRLRVIHRDLKVSNI 622
Query: 638 LLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVV 697
LLDE+ NPKISDFGLAR++ G + NT RVVGT GYMSPEYA G FS KSD++SFGV+
Sbjct: 623 LLDEKMNPKISDFGLARMYQGTEYQDNTRRVVGTLGYMSPEYAWTGMFSEKSDIYSFGVL 682
Query: 698 VLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGL 757
+LEIISG++ + F +L+ YAW W R +D +DQ L+ +C E +C+ +GL
Sbjct: 683 MLEIISGEKISRFSYGVEGKTLIAYAWESWSEYRGIDLLDQDLADSCHPLEVGRCIQIGL 742
Query: 758 LCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLT 817
LC+Q P +RP +L ML + S+ LPSP++P F +R S + + N +T
Sbjct: 743 LCVQHQPADRPNTLELLAMLTTTSD-LPSPKQPTFAFH----TRDDESLSNDLITVNGMT 797
Query: 818 VTLENGR 824
++ GR
Sbjct: 798 QSVILGR 804
>At1g61490 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 804
Score = 543 bits (1399), Expect = e-154
Identities = 326/845 (38%), Positives = 483/845 (56%), Gaps = 71/845 (8%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+F C +FT L + IT + L E TL S+ G +ELGFF+PN S N YVGI
Sbjct: 7 VFFACLLLFTVLLRFSYAGITTESPLSV--EQTLSSSNGIYELGFFSPNNSQN--LYVGI 62
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
+ + P+ VVWVANR+ P D+ +I+ +G+L + + W ++ +S
Sbjct: 63 WFKGIIPRVVVWVANRETPTTDTSANLAISSNGSLLLFNGKHGVVW--SIGENFASNGSR 120
Query: 124 VKLLDSGNLIVSDDKVKKILWQSFANPTDTFLP------GMKMDDSITLTSWRSHDDPAP 177
+L D+GNL+V D+ + LW+SF + DT LP + + LTSW++ DP+P
Sbjct: 121 AELTDNGNLVVIDNASGRTLWESFEHFGDTMLPFSSLMYNLATGEKRVLTSWKTDTDPSP 180
Query: 178 GNFSFE-QDQGENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSNFTLRISPNNTI 236
G F + Q +Q +I + S +Y+++ K TG +Y S F+L+ N +
Sbjct: 181 GVFVGQITPQVPSQVLIMRGSTRYYRTGPWAKTRFTGIPLMDDTYA-SPFSLQQDANGS- 238
Query: 237 PFLT--SSLYSNTRLVMTYWGQLQYLKMDSMKMWLMVWVEPRDRCSVFNACGNFGSCNSK 294
F T + +R++++ G ++ + + W + ++ P + C ++ CG FG C
Sbjct: 239 GFFTYFDRSFKLSRIIISSEGSMKRFRHNGTD-WELSYMAPANSCDIYGVCGPFGLCIVS 297
Query: 295 YDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNV-CSEDAKS---DTFLSLRMMKVGNPD 350
CKCL GF P+S E W G+++GGC+R T + C ++ + F + +K+ +
Sbjct: 298 VPLKCKCLKGFVPHSTEEWKRGNWTGGCARLTELHCQGNSTGKDVNIFHPVTNVKLPDFY 357
Query: 351 AQFNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYEG 410
++ + EEC CL+NC C A++Y C IW+Q+L + + G
Sbjct: 358 EYESSVDAEECHQSCLHNCSCLAFAYIH------------GIGCLIWNQNLMDAVQFSAG 405
Query: 411 GCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRG 470
G L +R+A S++ G N + I+ T + +F+I+ S+ + R R + K
Sbjct: 406 GEILSIRLAHSELGG-------NKRNKIIVASTVSLSLFVILTSAAFGFWRYRVKHKAYT 458
Query: 471 IKLYGSERYVRDLIESGRLQEDDAKAIDIP---HFHLESILDATNNFAIANKLGQGGFGP 527
+K +D K+ ++P F + +I ATNNF+++NKLGQGGFG
Sbjct: 459 LK---------------DAWRNDLKSKEVPGLEFFEMNTIQTATNNFSLSNKLGQGGFGS 503
Query: 528 VYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVY 587
VYKGK G+EIAVK+LSS SGQG EEF NE+VLI++LQHRNLVR+LG C+EG+EK+L+Y
Sbjct: 504 VYKGKLQDGKEIAVKQLSSSSGQGKEEFMNEIVLISKLQHRNLVRVLGCCIEGEEKLLIY 563
Query: 588 EYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILL 639
E+M N+SLD F+FD W RF I+ GIARGLLYLH DSRL++IHRDLK SNILL
Sbjct: 564 EFMLNKSLDTFVFDARKKLEVDWPKRFDIVQGIARGLLYLHRDSRLKVIHRDLKVSNILL 623
Query: 640 DEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVL 699
DE+ NPKISDFGLAR++ G T RVVGT GYMSPEYA G FS KSD++SFGV++L
Sbjct: 624 DEKMNPKISDFGLARMYEGTQCQDKTRRVVGTLGYMSPEYAWTGVFSEKSDIYSFGVLLL 683
Query: 700 EIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLC 759
EII G++ + F E +LL YAW W + +D +DQ L+ +C+ E +CV +GLLC
Sbjct: 684 EIIIGEKISRFSYGEEGKTLLAYAWESWGETKGIDLLDQDLADSCRPLEVGRCVQIGLLC 743
Query: 760 LQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVT 819
+Q P +RP +L ML + S+ LPSP++P FV+ S +S + F+ N++T +
Sbjct: 744 VQHQPADRPNTLELLAMLTTTSD-LPSPKQPTFVVH---SRDDESSLSKDLFTVNEMTQS 799
Query: 820 LENGR 824
+ GR
Sbjct: 800 MILGR 804
>At1g61390 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 831
Score = 540 bits (1392), Expect = e-153
Identities = 324/815 (39%), Positives = 465/815 (56%), Gaps = 64/815 (7%)
Query: 36 TLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWVANRDNPLPDSGGAFSITED 95
TL S G +ELGFF+PN S ++YVGI + +APQ VVWVANRD P+ + +I+ +
Sbjct: 55 TLSSPDGVYELGFFSPNNSR--KQYVGIWFKNIAPQVVVWVANRDKPVTKTAANLTISSN 112
Query: 96 GNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIVSDDKVKKILWQSFANPTDTFL 155
G+L +LD T W T TS+ + +LLD+GNL+V DD K LW+SF N +T L
Sbjct: 113 GSLILLDGTQDVIWSTGEAFTSNKCH--AELLDTGNLVVIDDVSGKTLWKSFENLGNTML 170
Query: 156 PGMKMDDSI------TLTSWRSHDDPAPGNFSFE-QDQGENQFVIWKRSMKYWKSSVSGK 208
P + I LTSWRS+ DP+PG F+ E Q Q +I + S YW+S K
Sbjct: 171 PQSSVMYDIPRGKNRVLTSWRSNSDPSPGEFTLEFTPQVPPQGLIRRGSSPYWRSGPWAK 230
Query: 209 FVGTGEMSSAISYLLSNFTLRISPNNTIPFLTSSL--YSNTRLVMTYWGQLQYLKMDSMK 266
+G SY+ L+ T F S L Y + + +T G+++ L D K
Sbjct: 231 TRFSGIPGIDASYVSPFTVLQDVAKGTASFSYSMLRNYKLSYVTLTSEGKMKILWNDG-K 289
Query: 267 MWLMVWVEPRDRCSVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKT 326
W + + P C ++ ACG FG C + C CL GF P S + W G+++ GC R+T
Sbjct: 290 SWKLHFEAPTSSCDLYRACGPFGLCVRSRNPKCICLKGFVPKSDDEWKKGNWTSGCVRRT 349
Query: 327 NVC--------SEDAKSDTFLSLRMMKVGNPDAQFNAKNEEECKLECLNNCQCYAYSYEE 378
+ ++ ++D+F + +K + N E+C +CL NC C A++Y
Sbjct: 350 QLSCHTNSSTKTQGKETDSFYHMTRVKTPDLYQLAGFLNAEQCYQDCLGNCSCTAFAYIS 409
Query: 379 YEKARQGDSVDPNAICWIWSQDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPT 438
C +W+++L + + G L +R+A S++ G + + + T
Sbjct: 410 ------------GIGCLVWNRELVDTVQFLSDGESLSLRLASSELAGSN--RTKIILGTT 455
Query: 439 IILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSE-RYVRDLIESGRLQEDDAKAI 497
+ L F +VF S + R +Q + + ++ S+ + +D+ + D +
Sbjct: 456 VSLSIFVILVFAAYKS----WRYRTKQNEPNPMFIHSSQDAWAKDM------EPQDVSGV 505
Query: 498 DIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKN 557
++ F + +I ATNNF+ +NKLGQGGFGPVYKGK G+EIAVKRLSS SGQG +EF N
Sbjct: 506 NL--FDMHTIRTATNNFSSSNKLGQGGFGPVYKGKLVDGKEIAVKRLSSSSGQGTDEFMN 563
Query: 558 EVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF--------DWDMRFKII 609
E+ LI++LQH+NLVRLLG C++G+EK+L+YEY+ N+SLD F+F DW RF II
Sbjct: 564 EIRLISKLQHKNLVRLLGCCIKGEEKLLIYEYLVNKSLDVFLFDSTLKFEIDWQKRFNII 623
Query: 610 LGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVV 669
G+ARGLLYLH DSRLR+IHRDLK SNILLDE+ PKISDFGLAR+ G NT RVV
Sbjct: 624 QGVARGLLYLHRDSRLRVIHRDLKVSNILLDEKMIPKISDFGLARMSQGTQYQDNTRRVV 683
Query: 670 GTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKV 729
GT GYM+PEYA G FS KSD++SFGV++LEII G++ + F E +LL YAW W
Sbjct: 684 GTLGYMAPEYAWTGVFSEKSDIYSFGVLLLEIIIGEKISRF--SEEGKTLLAYAWESWCE 741
Query: 730 GRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPRE 789
+ +D +DQ L+ + E +CV +GLLC+Q P +RP ++ ML + S LPSP++
Sbjct: 742 TKGVDLLDQALADSSHPAEVGRCVQIGLLCVQHQPADRPNTLELMSMLTTISE-LPSPKQ 800
Query: 790 PAFVIRRCPSSRASTSSKMETFSRNDLTVTLENGR 824
P F + SR S+ + + N++T ++ GR
Sbjct: 801 PTFTVH----SRDDDSTSNDLITVNEITQSVIQGR 831
>At1g61360 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 821
Score = 539 bits (1389), Expect = e-153
Identities = 322/817 (39%), Positives = 454/817 (55%), Gaps = 68/817 (8%)
Query: 36 TLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWVANRDNPLPDSGGAFSITED 95
TL S GG +ELGFF+ N S N +YVGI + K+ P+ +VWVANR+ P+ + +I+ +
Sbjct: 33 TLSSPGGSYELGFFSSNNSGN--QYVGIWFKKVTPRVIVWVANREKPVSSTMANLTISSN 90
Query: 96 GNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIVSDDKVKKILWQSFANPTDTFL 155
G+L +LD W + + TS+ +LLD+GNL+V D+ LWQSF + DT L
Sbjct: 91 GSLILLDSKKDLVWSSGGDPTSNK--CRAELLDTGNLVVVDNVTGNYLWQSFEHLGDTML 148
Query: 156 PGMKMDDSI------TLTSWRSHDDPAPGNFSFE-QDQGENQFVIWKRSMKYWKSS--VS 206
P + I LTSW+S DP+PG F E Q +Q +I K S YW+S
Sbjct: 149 PLTSLMYDIPNNKKRVLTSWKSETDPSPGEFVAEITPQVPSQGLIRKGSSPYWRSGPWAG 208
Query: 207 GKFVGTGEMSSAISYLLSNFTLRISPNNTIPFLTSSLYSNTRLVMTYWGQLQYLKMDSMK 266
+F G EM ++ L ++ F ++ + + +T G L+ + +
Sbjct: 209 TRFTGIPEMDASYVNPLGMVQDEVNGTGVFAFCVLRNFNLSYIKLTPEGSLRITRNNGTD 268
Query: 267 MWLMVWVEPRDRCSVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKT 326
W+ + P C ++ CG FG C MC+CL GF P S E W +G++S GC R+T
Sbjct: 269 -WIKHFEGPLTSCDLYGRCGPFGLCVRSGTPMCQCLKGFEPKSDEEWRSGNWSRGCVRRT 327
Query: 327 NVCSEDAKS--------DTFLSLRMMKVGNPDAQFNAKNEEECKLECLNNCQCYAYSYEE 378
N+ + S D F + +K + + NEE+C CL NC C A+SY
Sbjct: 328 NLSCQGNSSVETQGKDRDVFYHVSNIKPPDSYELASFSNEEQCHQGCLRNCSCTAFSYVS 387
Query: 379 YEKARQGDSVDPNAICWIWSQDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPT 438
C +W+Q+L + + GG L +R+A S++ G + + T
Sbjct: 388 ------------GIGCLVWNQELLDTVKFIGGGETLSLRLAHSELTG-----RKRIKIIT 430
Query: 439 IILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESG---RLQEDDAK 495
+ ++ + + L++++ C R +K GS +D +E LQ D
Sbjct: 431 VATLSLSVCLILVLVACGCWRYR---------VKQNGSSLVSKDNVEGAWKSDLQSQDVS 481
Query: 496 AIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEF 555
++ F + + ATNNF++ NKLGQGGFG VYKGK G+EIAVKRL+S S QG EEF
Sbjct: 482 GLNF--FEIHDLQTATNNFSVLNKLGQGGFGTVYKGKLQDGKEIAVKRLTSSSVQGTEEF 539
Query: 556 KNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFK 607
NE+ LI++LQHRNL+RLLG C++G+EK+LVYEYM N+SLD FIFD W RF
Sbjct: 540 MNEIKLISKLQHRNLLRLLGCCIDGEEKLLVYEYMVNKSLDIFIFDLKKKLEIDWATRFN 599
Query: 608 IILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDR 667
II GIARGLLYLH DS LR++HRDLK SNILLDE+ NPKISDFGLAR+F G +T
Sbjct: 600 IIQGIARGLLYLHRDSFLRVVHRDLKVSNILLDEKMNPKISDFGLARLFHGNQHQDSTGS 659
Query: 668 VVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLW 727
VVGT GYMSPEYA G FS KSD++SFGV++LEII+GK + F + +LL YAW W
Sbjct: 660 VVGTLGYMSPEYAWTGTFSEKSDIYSFGVLMLEIITGKEISSFSYGKDNKNLLSYAWDSW 719
Query: 728 KVGRTLDFIDQTL--SQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLP 785
++ +DQ L S + + E +CV++GLLC+Q I+RP + V+ ML S ++ LP
Sbjct: 720 SENGGVNLLDQDLDDSDSVNSVEAGRCVHIGLLCVQHQAIDRPNIKQVMSMLTSTTD-LP 778
Query: 786 SPREPAFVIRRCPSSRASTSSKMETFSRNDLTVTLEN 822
P +P FV+ + + S+ NDL+ EN
Sbjct: 779 KPTQPMFVLETSDEDSSLSHSQRS----NDLSSVDEN 811
>At1g11280 serine/threonine kinase like protein
Length = 830
Score = 536 bits (1381), Expect = e-152
Identities = 327/829 (39%), Positives = 475/829 (56%), Gaps = 65/829 (7%)
Query: 23 ITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWVANRDNP 82
ITI++ L G TL S GG++ELGFF+PN S N +YVGI + K+ P+ VVWVANR+ P
Sbjct: 40 ITISSPLTLG--QTLSSPGGFYELGFFSPNNSQN--QYVGIWFKKITPRVVVWVANREKP 95
Query: 83 LPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIVSDDKVKKI 142
+ +I+ +G+L +LD + W T R S S KLLD+GNL++ DD + +
Sbjct: 96 ITTPVANLTISRNGSLILLDSSKNVVWSTR--RPSISNKCHAKLLDTGNLVIVDDVSENL 153
Query: 143 LWQSFANPTDTFLP------GMKMDDSITLTSWRSHDDPAPGNFSFE-QDQGENQFVIWK 195
LWQSF NP DT LP + + L+SW+SH DP+PG+F Q Q V +
Sbjct: 154 LWQSFENPGDTMLPYSSLMYNLATGEKRVLSSWKSHTDPSPGDFVVRLTPQVPAQIVTMR 213
Query: 196 RSMKYWKSSVSGKFVGTGEMSSAISYLLSNFTLRISPNNTIPFLTSSLYSN--TRLVMTY 253
S Y +S K TG SY S F+L N + S+ TR+++T
Sbjct: 214 GSSVYKRSGPWAKTGFTGVPLMDESYT-SPFSLSQDVGNGTGLFSYLQRSSELTRVIITS 272
Query: 254 WGQLQYLKMDSMKMWLMVWVEPRDRCSVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENW 313
G L+ + + W++ ++ P + C ++ ACG FG C + + CKC+ GF P E W
Sbjct: 273 EGYLKTFRYNGTG-WVLDFITPANLCDLYGACGPFGLCVTSNPTKCKCMKGFVPKYKEEW 331
Query: 314 SAGDFSGGCSRKTNV-CSEDAKSDTF-----LSLRMMKVGNPDAQFNAK--NEEECKLEC 365
G+ + GC R+T + C + + T + R+ V PD A + ++C C
Sbjct: 332 KRGNMTSGCMRRTELSCQANLSTKTQGKGVDVFYRLANVKPPDLYEYASFVDADQCHQGC 391
Query: 366 LNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYEGGCDLHVRVAFSDIEG 425
L+NC C A++Y C +W+ +L + GG L +R+A S++ G
Sbjct: 392 LSNCSCSAFAYIT------------GIGCLLWNHELIDTIRYSVGGEFLSIRLASSELAG 439
Query: 426 DSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIE 485
S +T+ II+ + + +F+I+ + Y R R + + + + +D +
Sbjct: 440 -SRRTK-------IIVGSISLSIFVILAFGSYKYWRYRAKQNVGPTWAFFNNS--QDSWK 489
Query: 486 SGRLQEDDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLS 545
+G L+ + + F + +I ATNNF ++NKLGQGGFGPVYKG ++IAVKRLS
Sbjct: 490 NG-LEPQEISGLTF--FEMNTIRAATNNFNVSNKLGQGGFGPVYKGTLSDKKDIAVKRLS 546
Query: 546 SCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD---- 601
S SGQG EEF NE+ LI++LQHRNLVRLLG C++G+EK+L+YE++ N+SLD F+FD
Sbjct: 547 SSSGQGTEEFMNEIKLISKLQHRNLVRLLGCCIDGEEKLLIYEFLVNKSLDTFLFDLTLK 606
Query: 602 ----WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFG 657
W RF II G++RGLLYLH DS +R+IHRDLK SNILLD++ NPKISDFGLAR+F
Sbjct: 607 LQIDWPKRFNIIQGVSRGLLYLHRDSCMRVIHRDLKVSNILLDDKMNPKISDFGLARMFQ 666
Query: 658 GKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHEL 717
G NT +VVGT GYMSPEYA G FS KSD+++FGV++LEIISGK+ + F E
Sbjct: 667 GTQHQDNTRKVVGTLGYMSPEYAWTGMFSEKSDIYAFGVLLLEIISGKKISSFCCGEEGK 726
Query: 718 SLLGYAWHLWKVGRTLDFIDQTLSQTCK--AEECMKCVNVGLLCLQEDPIERPTMSNVLF 775
+LLG+AW W +D +D+ +S +C E +CV +GLLC+Q+ ++RP ++ V+
Sbjct: 727 TLLGHAWECWLETGGVDLLDEDISSSCSPVEVEVARCVQIGLLCIQQQAVDRPNIAQVVT 786
Query: 776 MLGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVTLENGR 824
M+ S ++ LP P++P F ++ + SK S N +T T GR
Sbjct: 787 MMTSATD-LPRPKQPLFALQIQDQESVVSVSK----SVNHVTQTEIYGR 830
>At4g21370 receptor kinase - like protein
Length = 844
Score = 533 bits (1374), Expect = e-151
Identities = 326/844 (38%), Positives = 471/844 (55%), Gaps = 77/844 (9%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L + N SA +++TI++N T++S GG FELGFF G S Y+GI
Sbjct: 22 LILFPDLSISVNTLSATESLTISSN------KTIVSPGGVFELGFFRILGDS---WYLGI 72
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y K++ +T VWVANRD PL + G I+ + NL +LD + W TNL S ++
Sbjct: 73 WYKKISQRTYVWVANRDTPLSNPIGILKIS-NANLVILDNSDTHVWSTNLTGAVRS-SVV 130
Query: 124 VKLLDSGNLIVSDDKVKK---ILWQSFANPTDTFLPGMKMDDSIT------LTSWRSHDD 174
+LLD+GN ++ K+ + LWQSF PTDT LP MK+ +TSW+S D
Sbjct: 131 AELLDNGNFVLRGSKINESDEFLWQSFDFPTDTLLPQMKLGRDHKRGLNRFVTSWKSSFD 190
Query: 175 PAPGNFSFEQDQ-GENQFVIWKRSMKYWKSSVSG--KFVGTGEMSSAISYLLSNFTLRIS 231
P+ G+F F+ + G +F + ++ ++S +F G EM ++ NFT
Sbjct: 191 PSSGSFMFKLETLGLPEFFGFTSFLEVYRSGPWDGLRFSGILEMQQWDD-IIYNFTEN-R 248
Query: 232 PNNTIPFLTSSLYSNTRLVMTYWGQLQYLKMD-SMKMWLMVWVEPRDRCSVFNACGNFGS 290
F + S +RL + G+L+ + + + W M W P+D C ++ CG +
Sbjct: 249 EEVAYTFRVTDHNSYSRLTINTVGRLEGFTWEPTQQEWNMFWFMPKDTCDLYGICGPYAY 308
Query: 291 CNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNV-CSEDAKSDTFLSLRMMKVGNP 349
C+ C C+ GF+P S ++W++GD +G C RKT + C ED F L MK+
Sbjct: 309 CDMSTSPTCNCIKGFQPLSPQDWASGDVTGRCRRKTQLTCGEDR----FFRLMNMKIPAT 364
Query: 350 DAQFNAKNE--EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEE 407
A K +EC+ +C +C C AY+ + G C IW + ++
Sbjct: 365 TAAIVDKRIGLKECEEKCKTHCNCTAYANSDIRNGGSG--------CIIWIGEFRDIRNY 416
Query: 408 YEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAK 467
G DL VR+A ++ + R + I L+ +++ ++ C + +++++A+
Sbjct: 417 AADGQDLFVRLAAAE-----FGERRTIRGKIIGLIIGISLMLVLSFIIYCFWKKKQKRAR 471
Query: 468 IRGIKLYGSERYVRDLI-------ESGRLQEDDAKAIDIPHFHLESILDATNNFAIANKL 520
+ +R +++LI SGR + + +++P E+++ AT NF+ +N L
Sbjct: 472 ATAAPIGYRDR-IQELIITNGVVMSSGRRLLGEEEDLELPLTEFETVVMATENFSDSNIL 530
Query: 521 GQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEG 580
G+GGFG VYK IAVKRLS S QG EFKNEV LIARLQH NLVRLL C+
Sbjct: 531 GRGGFGIVYK--------IAVKRLSEMSSQGTNEFKNEVRLIARLQHINLVRLLSCCIYA 582
Query: 581 DEKMLVYEYMPNRSLDAFIFD---------WDMRFKIILGIARGLLYLHEDSRLRIIHRD 631
DEK+L+YEY+ N SLD+ +F+ W RF II GIARGLLYLH+DSR +IIHRD
Sbjct: 583 DEKILIYEYLENGSLDSHLFETTQSSNKLNWQTRFSIINGIARGLLYLHQDSRFKIIHRD 642
Query: 632 LKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDV 691
LKASN+LLD+ PKISDFG+ARIF +T NT +VVGTYGYMSPEYA++G FSVKSDV
Sbjct: 643 LKASNVLLDKNMTPKISDFGMARIFERDETEANTRKVVGTYGYMSPEYAMEGIFSVKSDV 702
Query: 692 FSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKA----- 746
FSFGV+VLEI+SGKRN GF+ + +LLGY W WK G+ L+ +D + + +
Sbjct: 703 FSFGVLVLEIVSGKRNRGFHNSGQDNNLLGYTWENWKEGKGLEIVDSIIVDSSSSMSLFQ 762
Query: 747 -EECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTS 805
E ++C+ +GLLC+QE +RP MS+V+ MLGSE SPR P + +RR +S
Sbjct: 763 PHEVLRCIQIGLLCVQERAEDRPKMSSVVLMLGSEKGEYFSPRRPGYCVRRSSLDTDDSS 822
Query: 806 SKME 809
S E
Sbjct: 823 SSTE 826
>At1g61550
Length = 802
Score = 533 bits (1373), Expect = e-151
Identities = 337/845 (39%), Positives = 470/845 (54%), Gaps = 76/845 (8%)
Query: 8 CSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHK 67
C T L + IT + L G TL S G FELGFF+PN S N YVGI +
Sbjct: 6 CFLFSTLLLSFSYAAITPTSPLSIG--QTLSSPNGIFELGFFSPNNSRN--LYVGIWFKG 61
Query: 68 LAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLL 127
+ P+TVVWVANR+N + D+ +I+ +G+L + D + W T T +S + +L
Sbjct: 62 IIPRTVVWVANRENSVTDATADLAISSNGSLLLFDGKHSTVWSTG--ETFASNGSSAELS 119
Query: 128 DSGNLIVSDDKVKKILWQSFANPTDTFLPGMKM------DDSITLTSWRSHDDPAPGNF- 180
DSGNL+V D LWQSF + DT LP + + L+SW+S+ DP PG F
Sbjct: 120 DSGNLLVIDKVSGITLWQSFEHLGDTMLPYSSLMYNPGTGEKRVLSSWKSYTDPLPGEFV 179
Query: 181 SFEQDQGENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSNFTLRISPNNTIPFLT 240
+ Q Q I + S YW+S K TG + SY F+++ N ++ F
Sbjct: 180 GYITTQVPPQGFIMRGSKPYWRSGPWAKTRFTGVPLTDESYT-HPFSVQQDANGSVYF-- 236
Query: 241 SSLYSNTR---LVMTYWGQLQYLKMDSMKMWLMVWVEPRDRCSVFNACGNFGSCNSKYDS 297
S L N + LV+T G L+ + W++ P + C + CG FG C
Sbjct: 237 SHLQRNFKRSLLVLTSEGSLKVTHHNGTD-WVLNIDVPANTCDFYGVCGPFGLCVMSIPP 295
Query: 298 MCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSL--RMMKVGNPDAQ--F 353
CKC GF P E W G+++GGC R+T + + + +++ + + PD
Sbjct: 296 KCKCFKGFVPQFSEEWKRGNWTGGCVRRTELLCQGNSTGRHVNVFHPVANIKPPDFYEFV 355
Query: 354 NAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAI-CWIWSQDLNNLEEEYEGGC 412
++ + EEC CL+NC C A++Y N I C IW+Q+L ++ + GG
Sbjct: 356 SSGSAEECYQSCLHNCSCLAFAYI-------------NGIGCLIWNQELMDVMQFSVGGE 402
Query: 413 DLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRR--RQAKIRG 470
L +R+A S++ G N TII + +F+ + S+ + R R A +
Sbjct: 403 LLSIRLASSEMGG-------NQRKKTIIASIVSISLFVTLASAAFGFWRYRLKHNAIVSK 455
Query: 471 IKLYGSERYVRDLIESGRLQEDDAKAIDIP---HFHLESILDATNNFAIANKLGQGGFGP 527
+ L G+ R +D K+ D+ F +++I ATNNF++ NKLGQGGFGP
Sbjct: 456 VSLQGAWR-------------NDLKSEDVSGLYFFEMKTIEIATNNFSLVNKLGQGGFGP 502
Query: 528 VYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVY 587
VYKGK G+EIAVKRLSS SGQG EEF NE++LI++LQH NLVR+LG C+EG+E++LVY
Sbjct: 503 VYKGKLQDGKEIAVKRLSSSSGQGKEEFMNEILLISKLQHINLVRILGCCIEGEERLLVY 562
Query: 588 EYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILL 639
E+M N+SLD FIFD W RF II GIARGLLYLH DSRLRIIHRD+K SNILL
Sbjct: 563 EFMVNKSLDTFIFDSRKRVEIDWPKRFSIIQGIARGLLYLHRDSRLRIIHRDVKVSNILL 622
Query: 640 DEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVL 699
D++ NPKISDFGLAR++ G NT R+VGT GYMSPEYA G FS KSD +SFGV++L
Sbjct: 623 DDKMNPKISDFGLARMYEGTKYQDNTRRIVGTLGYMSPEYAWTGVFSEKSDTYSFGVLLL 682
Query: 700 EIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLC 759
E+ISG++ + F + +LL YAW W + F+D+ + +C E +CV +GLLC
Sbjct: 683 EVISGEKISRFSYDKERKNLLAYAWESWCENGGVGFLDKDATDSCHPSEVGRCVQIGLLC 742
Query: 760 LQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVT 819
+Q P +RP +L ML + S+ LP P+EP F + S S +S + T N++T +
Sbjct: 743 VQHQPADRPNTLELLSMLTTTSD-LPLPKEPTFAVH--TSDDGSRTSDLITV--NEVTQS 797
Query: 820 LENGR 824
+ GR
Sbjct: 798 VVLGR 802
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,183,121
Number of Sequences: 26719
Number of extensions: 940904
Number of successful extensions: 5733
Number of sequences better than 10.0: 1022
Number of HSP's better than 10.0 without gapping: 870
Number of HSP's successfully gapped in prelim test: 152
Number of HSP's that attempted gapping in prelim test: 2045
Number of HSP's gapped (non-prelim): 1236
length of query: 824
length of database: 11,318,596
effective HSP length: 108
effective length of query: 716
effective length of database: 8,432,944
effective search space: 6037987904
effective search space used: 6037987904
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0203.7


