

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.5
(852 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
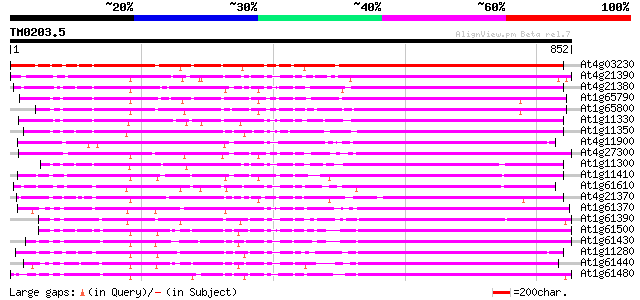
Sequences producing significant alignments: (bits) Value
At4g03230 putative receptor kinase 758 0.0
At4g21390 serine/threonine kinase - like protein 570 e-162
At4g21380 receptor-like serine/threonine protein kinase ARK3 559 e-159
At1g65790 receptor kinase, putative 556 e-158
At1g65800 receptor kinase, putative 545 e-155
At1g11330 receptor-like protein kinase, putative 545 e-155
At1g11350 putative receptor-like protein kinase gi|4008010; simi... 537 e-153
At4g11900 KI domain interacting kinase 1 -like protein 531 e-151
At4g27300 putative receptor protein kinase 530 e-150
At1g11300 putative s-locus protein kinase (PPC:1.7.2) 530 e-150
At1g11410 putative brassinosteroid insensitive protein; similar ... 522 e-148
At1g61610 516 e-146
At4g21370 receptor kinase - like protein 507 e-144
At1g61370 S-like receptor protein kinase; member of gene cluster... 489 e-138
At1g61390 S-like receptor protein kinase; member of gene cluster... 488 e-138
At1g61500 487 e-137
At1g61430 S-like receptor protein kinase; member of gene cluster... 484 e-136
At1g11280 serine/threonine kinase like protein 480 e-135
At1g61440 S-like receptor protein kinase; member of gene cluster... 479 e-135
At1g61480 S-like receptor protein kinase; member of gene cluster... 478 e-135
>At4g03230 putative receptor kinase
Length = 852
Score = 758 bits (1956), Expect = 0.0
Identities = 423/869 (48%), Positives = 565/869 (64%), Gaps = 65/869 (7%)
Query: 1 MFFTPFTNMITIFLFHMHCWLLCF-SQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFF 59
M + F M + + + C++ S+ F G TL +G LVSA ++FELGFF
Sbjct: 1 MILSVFFYMFLLHIRRLDCFVAVQDSKTLFKGSTLINDS----HGETLVSAGQRFELGFF 56
Query: 60 SPDLNVTGGKGRYLGIWYYREEGSGLSP--VVWVANRDNPVADDSIGVFRIADDGNLVVL 117
+P N + + RYLGIW+Y L P VVWVANR++PV D S +F I+ DGNL V+
Sbjct: 57 TP--NGSSDERRYLGIWFYN-----LHPLTVVWVANRESPVLDRSC-IFTISKDGNLEVI 108
Query: 118 DTSGIRYWSASNLTNSSSATNRSVKLMDSGNLVLL-DEHVGMKLWESFEHPTDTFLPGMK 176
D+ G YW + + SS + R VKLMD+GNLVL+ D + +W+SF++PTDTFLPGM+
Sbjct: 109 DSKGRVYWD-TGVKPSSVSAERMVKLMDNGNLVLISDGNEANVVWQSFQNPTDTFLPGMR 167
Query: 177 MDKTLELTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPD 236
MD+ + L+ W+S +DP GNFTF+MD++ + +F I + YW+S G+ D
Sbjct: 168 MDENMTLSSWRSFNDPSHGNFTFQMDQEEDKQFIIWKRSMRYWKS-----GISGKFIGSD 222
Query: 237 DISNDVYNLLTNFKE---LKNKTV-----SSYDNTRLLLNSTGVIKVLYRVNFQSDIVW- 287
++ + L+NF E + N +V S Y NTR ++S+G + + W
Sbjct: 223 EMPYAISYFLSNFTETVTVHNASVPPLFTSLYTNTRFTMSSSGQAQYF---RLDGERFWA 279
Query: 288 --WYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKS 345
W +PR C YN CGNF SCN N+++C CLPGF R + V GD S C+R+S
Sbjct: 280 QIWAEPRDECSVYNACGNFGSCNSKNEEMCKCLPGF--RPNFLEKWVKGDFSGG-CSRES 336
Query: 346 TSCGANT----NTFLNLTMMKIGSPDIKVSAQDENECKFRCISMCSQTQCQACSYVPIPV 401
CG + + FLNL+++++GSPD + A +E EC+ C++ C QCQA SY + +
Sbjct: 337 RICGKDGVVVGDMFLNLSVVEVGSPDSQFDAHNEKECRAECLNNC---QCQAYSYEEVDI 393
Query: 402 QQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPT-----RKGNPKSTLS 456
Q + CWIW ++L LKE YLG R +F+RVA DI R G K+ +
Sbjct: 394 LQSN---TKCWIWLEDLNNLKEGYLGS--RNVFIRVAVPDIGSHVERGRGRYGEAKTPVV 448
Query: 457 LILGIALPGVVILACICILA---YVCRRKIALKLKQESESILRQRGRFYDSERHVKDLID 513
LI+ + IL + A ++ RRK+ +L + DSERH+K+LI+
Sbjct: 449 LIIVVTFTSAAILVVLSSTASYVFLQRRKVNKELGSIPRGV-----HLCDSERHIKELIE 503
Query: 514 KEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLS 573
++ D++GI+VP F+ E+IL AT FS+ANKLG+GG+GPVYKG G +EIAVKRLS
Sbjct: 504 SGRFKQDDSQGIDVPSFELETILYATSNFSNANKLGQGGFGPVYKGMFPGDQEIAVKRLS 563
Query: 574 SVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKS 633
S QG++EFKNEVVLIAKLQHRNLVRL GYC+ G+EK+L+YEYMP+KSLD F+FD
Sbjct: 564 RCSGQGLEEFKNEVVLIAKLQHRNLVRLLGYCVAGEEKLLLYEYMPHKSLDFFIFDRKLC 623
Query: 634 ALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFG 693
LDW+MR +I+LGIARGLLYLHQDSRLR+IHRDLKTSNILLD EM PKISDFGLARIFG
Sbjct: 624 QRLDWKMRCNIILGIARGLLYLHQDSRLRIIHRDLKTSNILLDEEMNPKISDFGLARIFG 683
Query: 694 GKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTL 753
G ET ANT RVVGTYGYMSPEYAL+G FS KSD+FSFGVV++E ISGK+NTGF++ + +L
Sbjct: 684 GSETSANTNRVVGTYGYMSPEYALEGLFSFKSDVFSFGVVVIETISGKRNTGFHEPEKSL 743
Query: 754 SLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML 813
SLLG+AW LW + ++L+D +L E+ F++C +VGLLCVQ++P+DRP MSNVV ML
Sbjct: 744 SLLGHAWDLWKAERGIELLDQALQESCETEGFLKCLNVGLLCVQEDPNDRPTMSNVVFML 803
Query: 814 -DSETATLPTPKQPTFFTRKDLSSTASSS 841
SE ATLPTPKQP F R+ SS+ +SS
Sbjct: 804 GSSEAATLPTPKQPAFVLRRCPSSSKASS 832
>At4g21390 serine/threonine kinase - like protein
Length = 849
Score = 570 bits (1468), Expect = e-162
Identities = 359/902 (39%), Positives = 517/902 (56%), Gaps = 106/902 (11%)
Query: 2 FFTPFTNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEITG--NGTVLVSAAKKFELGFF 59
FF + +++FL+ + A +T+ G+ + N LVS K FELGFF
Sbjct: 3 FFRKTSLYLSLFLYFF------LYESSMAANTIRRGESLRDGINHKPLVSPQKTFELGFF 56
Query: 60 SPDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDT 119
SP + R+LGIWY E VVWVANR P++D S GV I++DGNLV+LD
Sbjct: 57 SPGSSTH----RFLGIWYGNIEDKA---VVWVANRATPISDQS-GVLMISNDGNLVLLDG 108
Query: 120 SGIRYWSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDK 179
I WS++ +++++ NR V + D+GN VL + +WESF HPTDTFLP M++
Sbjct: 109 KNITVWSSNIESSTTNNNNRVVSIHDTGNFVLSETDTDRPIWESFNHPTDTFLPQMRVRV 168
Query: 180 TLE------LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLY-WQSEEQGDGVMNPE 232
+ W+S +DP GN++ +D + + W+S + +
Sbjct: 169 NPQTGDNHAFVSWRSETDPSPGNYSLGVDPSGAPEIVLWEGNKTRKWRSGQWNSAIFTGI 228
Query: 233 SNPDDISNDVYNLLTNFKELKNKTVSSY-----DNTRLLLNSTGVIKVLYRVNFQSDIVW 287
N ++N +Y ++T S Y + +LL KVLY + ++ W
Sbjct: 229 PNMSLLTNYLYGF--KLSSPPDETGSVYFTYVPSDPSVLLR----FKVLYN-GTEEELRW 281
Query: 288 ------WY----QPRTTCLTYNVCGNFSSCN-DDNDKLCTCLPGFGRRSPLNDYTVGGDT 336
W +P + C YN CG F C+ ++ +C+C+ G+ + S + +++ G
Sbjct: 282 NETLKKWTKFQSEPDSECDQYNRCGKFGICDMKGSNGICSCIHGYEQVS-VGNWSRGCRR 340
Query: 337 SSLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVSAQ---DENECKFRCISMCSQTQCQA 393
+ L ++ S G + FL L +K+ PD ++ D +C+ RC+ CS C A
Sbjct: 341 RTPLKCERNISVGEDE--FLTLKSVKL--PDFEIPEHNLVDPEDCRERCLRNCS---CNA 393
Query: 394 CSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKS 453
S V G+ C IW Q+L L++ GG L +R+A S++ E N K+
Sbjct: 394 YSLVG------GIG---CMIWNQDLVDLQQFEAGGSS--LHIRLADSEVGE-----NRKT 437
Query: 454 TLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLID 513
+++I+ + L GV+++ +L + +RK K S + G+ D+ V DL
Sbjct: 438 KIAVIVAV-LVGVILIGIFALLLWRFKRK-----KDVSGAYC---GKNTDTSVVVADLTK 488
Query: 514 KEG------------LEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKL 561
+ +E K E+P F +I +AT+ F N+LGRGG+GPVYKG L
Sbjct: 489 SKETTSAFSGSVDIMIEGKAVNTSELPVFSLNAIAIATNDFCKENELGRGGFGPVYKGVL 548
Query: 562 QGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNK 621
+ GREIAVKRLS S QG+ EFKNE++LIAKLQHRNLVRL G C +G+EK+L+YEYMPNK
Sbjct: 549 EDGREIAVKRLSGKSGQGVDEFKNEIILIAKLQHRNLVRLLGCCFEGEEKMLVYEYMPNK 608
Query: 622 SLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQP 681
SLD F+FD TK AL+DW++RF I+ GIARGLLYLH+DSRLR+IHRDLK SN+LLD EM P
Sbjct: 609 SLDFFLFDETKQALIDWKLRFSIIEGIARGLLYLHRDSRLRIIHRDLKVSNVLLDAEMNP 668
Query: 682 KISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGK 741
KISDFG+ARIFGG + EANT RVVGTYGYMSPEYA++G FS KSD++SFGV+LLEI+SGK
Sbjct: 669 KISDFGMARIFGGNQNEANTVRVVGTYGYMSPEYAMEGLFSVKSDVYSFGVLLLEIVSGK 728
Query: 742 KNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPD 801
+NT + SL+GYAW L+T + +L+D + + + +RC HV +LCVQD
Sbjct: 729 RNTSLRSSEHG-SLIGYAWYLYTHGRSEELVDPKIRVTCSKREALRCIHVAMLCVQDSAA 787
Query: 802 DRPNMSNVVIMLDSETATLPTPKQPTFFTRK----DLSSTASSSLQF-------DSSIVE 850
+RPNM++V++ML+S+TATL P+QPTF + + D++ SS Q+ S++V
Sbjct: 788 ERPNMASVLLMLESDTATLAAPRQPTFTSTRRNSIDVNFALDSSQQYIVSSNEITSTVVL 847
Query: 851 GR 852
GR
Sbjct: 848 GR 849
>At4g21380 receptor-like serine/threonine protein kinase ARK3
Length = 850
Score = 559 bits (1441), Expect = e-159
Identities = 356/873 (40%), Positives = 501/873 (56%), Gaps = 88/873 (10%)
Query: 6 FTNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLN 64
F + T F F + L+ F + +TL+ + +T + +VS FELGFF P L+
Sbjct: 7 FYHSYTFFFFFL---LILFPAYSISANTLSASESLTISSNNTIVSPGNVFELGFFKPGLD 63
Query: 65 VTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRY 124
YLGIWY + VWVANRD P++ SIG +I+D NLVVLD S
Sbjct: 64 SRW----YLGIWY---KAISKRTYVWVANRDTPLSS-SIGTLKISDS-NLVVLDQSDTPV 114
Query: 125 WSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMK---LWESFEHPTDTFLPGMKMDKTL 181
WS +NLT + +L+D+GN VL D LW+SF+ PTDT LP MK+
Sbjct: 115 WS-TNLTGGDVRSPLVAELLDNGNFVLRDSKNSAPDGVLWQSFDFPTDTLLPEMKLGWDA 173
Query: 182 E------LTCWKSLSDPGRGNFTFKMDKKWENRFAILN-QGQLYWQSEEQG---DGVMNP 231
+ + WKS DP G+F+FK++ + + N + ++Y G GV P
Sbjct: 174 KTGFNRFIRSWKSPDDPSSGDFSFKLETEGFPEIFLWNRESRMYRSGPWNGIRFSGV--P 231
Query: 232 ESNPDDISNDVYNLLTNFKELKN--KTVSSYDNTRLLLNSTGVI-KVLYRVNFQSDIVWW 288
E P + V+N T+ +E+ + S +RL ++S+G++ + + Q+ +W
Sbjct: 232 EMQPFEYM--VFNFTTSKEEVTYSFRITKSDVYSRLSISSSGLLQRFTWIETAQNWNQFW 289
Query: 289 YQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRK 344
Y P+ C Y CG + C+ + +C C+ GF R+P L D + G C RK
Sbjct: 290 YAPKDQCDEYKECGVYGYCDSNTSPVCNCIKGFKPRNPQVWGLRDGSDG-------CVRK 342
Query: 345 ST-SCGANTNTFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPI 399
+ SCG F+ L MK+ PD ++ D EC+ +C+ C+ C A + I
Sbjct: 343 TLLSCGGGDG-FVRLKKMKL--PDTTTASVDRGIGVKECEQKCLRDCN---CTAFANTDI 396
Query: 400 PVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLIL 459
RG S C WT L ++ GG D L+VR+A +D+E+ + + I+
Sbjct: 397 ----RGSG-SGCVTWTGELFDIRNYAKGGQD--LYVRLAATDLEDKRNRS------AKII 443
Query: 460 GIALPGVVILACICILAYVCRRKIALKLKQESESI---LRQRGRFYD-----SERHVKDL 511
G ++ V+L I+ ++ +RK + E+ + LR R + S RH+
Sbjct: 444 GSSIGVSVLLLLSFIIFFLWKRKQKRSILIETPIVDHQLRSRDLLMNEVVISSRRHIS-- 501
Query: 512 IDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKR 571
E + + +E+P +FE + +AT+ FS+ANKLG+GG+G VYKGKL G+E+AVKR
Sbjct: 502 ------RENNTDDLELPLMEFEEVAMATNNFSNANKLGQGGFGIVYKGKLLDGQEMAVKR 555
Query: 572 LSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPT 631
LS S QG EFKNEV LIA+LQH NLVRL C+ EK+LIYEY+ N SLD+ +FD +
Sbjct: 556 LSKTSVQGTDEFKNEVKLIARLQHINLVRLLACCVDAGEKMLIYEYLENLSLDSHLFDKS 615
Query: 632 KSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARI 691
+++ L+WQMRFDI+ GIARGLLYLHQDSR R+IHRDLK SNILLD M PKISDFG+ARI
Sbjct: 616 RNSKLNWQMRFDIINGIARGLLYLHQDSRFRIIHRDLKASNILLDKYMTPKISDFGMARI 675
Query: 692 FGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKG 751
FG ETEANT++VVGTYGYMSPEYA+DG FS KSD+FSFGV+LLEIIS K+N GFY
Sbjct: 676 FGRDETEANTRKVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEIISSKRNKGFYNSDR 735
Query: 752 TLSLLGYAWKLWTENKLLDLMDLSLGEA---YNANQFIRCTHVGLLCVQDEPDDRPNMSN 808
L+LLG W+ W E K L+++D + ++ + ++ +RC +GLLCVQ+ +DRP MS
Sbjct: 736 DLNLLGCVWRNWKEGKGLEIIDPIITDSSSTFRQHEILRCIQIGLLCVQERAEDRPTMSL 795
Query: 809 VVIMLDSETATLPTPKQPTFFTRKDLSSTASSS 841
V++ML SE+ T+P PK P + + L T SSS
Sbjct: 796 VILMLGSESTTIPQPKAPGYCLERSLLDTDSSS 828
>At1g65790 receptor kinase, putative
Length = 843
Score = 556 bits (1434), Expect = e-158
Identities = 345/859 (40%), Positives = 499/859 (57%), Gaps = 70/859 (8%)
Query: 15 FHMHCWLLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLNVTGGKGRYL 73
F + L+ F + +TL+ + +T + ++S ++ FELGFF+P YL
Sbjct: 11 FFIFLILILFLAFSVSPNTLSATESLTISSNKTIISPSQIFELGFFNP----ASSSRWYL 66
Query: 74 GIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNS 133
GIWY + + VWVANRDNP++ + G +I+ + NLV+ D S WS +N+T
Sbjct: 67 GIWY---KIIPIRTYVWVANRDNPLSSSN-GTLKISGN-NLVIFDQSDRPVWS-TNITGG 120
Query: 134 SSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLE------LTCWK 187
+ + +L+D+GN +L D + + LW+SF+ PTDT L MK+ + L WK
Sbjct: 121 DVRSPVAAELLDNGNFLLRDSNNRL-LWQSFDFPTDTLLAEMKLGWDQKTGFNRILRSWK 179
Query: 188 SLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLT 247
+ DP G F+ K++ F I ++ + ++S M S P I D ++
Sbjct: 180 TTDDPSSGEFSTKLETSEFPEFYICSKESILYRSGPWNG--MRFSSVPGTIQVDY--MVY 235
Query: 248 NFKELKNKTVSSYD------NTRLLLNSTGVIKVLYRVNFQSDIVW---WYQPRTTCLTY 298
NF K + SY +RL LNS G+++ L F++ W WY P+ C Y
Sbjct: 236 NFTASKEEVTYSYRINKTNLYSRLYLNSAGLLQRL--TWFETTQSWKQLWYSPKDLCDNY 293
Query: 299 NVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKSTSCGANTNTFLNL 358
VCGNF C+ ++ C C+ GF P+N+ S C RK+ + F L
Sbjct: 294 KVCGNFGYCDSNSLPNCYCIKGF---KPVNEQAWDLRDGSAGCMRKTRLSCDGRDGFTRL 350
Query: 359 TMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIW 414
MK+ PD + D CK RC+ C+ T ++ ++ G S C IW
Sbjct: 351 KRMKL--PDTTATIVDREIGLKVCKERCLEDCNCT-----AFANADIRNGG---SGCVIW 400
Query: 415 TQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICI 474
T+ + ++ GG D L+VR+A +++E+ K I+G ++ ++L +
Sbjct: 401 TREILDMRNYAKGGQD--LYVRLAAAELEDKRIKNEK------IIGSSIGVSILLLLSFV 452
Query: 475 LAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLI-DKEGL--EEKDNEGIEVPYFD 531
+ + +RK + ++ ++ + R + + + D++ + G +EK +E +E+P +
Sbjct: 453 IFHFWKRKQKRSITIQTPNVDQVRSQ----DSLINDVVVSRRGYTSKEKKSEYLELPLLE 508
Query: 532 FESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIA 591
E++ AT+ FS+ NKLG+GG+G VYKG+L G+EIAVKRLS +SSQG EF NEV LIA
Sbjct: 509 LEALATATNNFSNDNKLGQGGFGIVYKGRLLDGKEIAVKRLSKMSSQGTDEFMNEVRLIA 568
Query: 592 KLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARG 651
KLQH NLVRL G C+ EK+LIYEY+ N SLD+ +FD T+S+ L+WQ RFDI+ GIARG
Sbjct: 569 KLQHINLVRLLGCCVDKGEKMLIYEYLENLSLDSHLFDQTRSSNLNWQKRFDIINGIARG 628
Query: 652 LLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYM 711
LLYLHQDSR R+IHRDLK SN+LLD M PKISDFG+ARIFG +ETEANT+RVVGTYGYM
Sbjct: 629 LLYLHQDSRCRIIHRDLKASNVLLDKNMTPKISDFGMARIFGREETEANTRRVVGTYGYM 688
Query: 712 SPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDL 771
SPEYA+DG FS KSD+FSFGV+LLEIISGK+N GFY L+LLG+ W+ W E L++
Sbjct: 689 SPEYAMDGIFSMKSDVFSFGVLLLEIISGKRNKGFYNSNRDLNLLGFVWRHWKEGNELEI 748
Query: 772 MDL----SLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPT 827
+D SL + ++ +RC +GLLCVQ+ +DRP MS+V++ML SET +P PK+P
Sbjct: 749 VDPINIDSLSSKFPTHEILRCIQIGLLCVQERAEDRPVMSSVMVMLGSETTAIPQPKRPG 808
Query: 828 F-FTRKDLSSTASSSLQFD 845
F R L + +SSS Q D
Sbjct: 809 FCIGRSPLEADSSSSTQRD 827
>At1g65800 receptor kinase, putative
Length = 847
Score = 545 bits (1404), Expect = e-155
Identities = 343/832 (41%), Positives = 489/832 (58%), Gaps = 64/832 (7%)
Query: 40 ITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVA 99
I+ N T+ +S ++ FELGFF+PD YLGIWY + + VWVANRDNP++
Sbjct: 38 ISSNKTI-ISPSQIFELGFFNPD----SSSRWYLGIWY---KIIPIRTYVWVANRDNPLS 89
Query: 100 DDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMK 159
+ G +I+D+ NLV+ D S WS +N+T + + +L+D GN VL D
Sbjct: 90 SSN-GTLKISDN-NLVIFDQSDRPVWS-TNITGGDVRSPVAAELLDYGNFVLRDSKNNKP 146
Query: 160 ---LWESFEHPTDTFLPGMKMDKTLE-------LTCWKSLSDPGRGNFTFKMDKKWENRF 209
LW+SF+ PTDT L MKM + L WK+ DP G+F+ K+ F
Sbjct: 147 SGFLWQSFDFPTDTLLSDMKMGWDNKSGGFNRILRSWKTTDDPSSGDFSTKLRTSGFPEF 206
Query: 210 AILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVSSYDNTR----- 264
I N+ + ++S G + N S+ + Y + +F E + V SY +
Sbjct: 207 YIYNKESITYRS---GPWLGNRFSSVPGMKPVDY-IDNSFTENNQQVVYSYRVNKTNIYS 262
Query: 265 -LLLNSTGVIKVL-YRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFG 322
L L+STG+++ L + QS WY P+ C Y CGN+ C+ + +C C+ GF
Sbjct: 263 ILSLSSTGLLQRLTWMEAAQSWKQLWYSPKDLCDNYKECGNYGYCDANTSPICNCIKGF- 321
Query: 323 RRSPLNDYTVGGDTSSLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVSAQDEN----EC 378
P+N+ D S+ C RK+ + F+ L M++ PD ++ D+ EC
Sbjct: 322 --EPMNEQAALRD-DSVGCVRKTKLSCDGRDGFVRLKKMRL--PDTTETSVDKGIGLKEC 376
Query: 379 KFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVA 438
+ RC+ C+ T ++ ++ G S C IW+ L ++ GG D L+VRVA
Sbjct: 377 EERCLKGCNCT-----AFANTDIRNGG---SGCVIWSGGLFDIRNYAKGGQD--LYVRVA 426
Query: 439 KSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQR 498
D+E+ K K + +G+++ +++L+ I + ++K ++ ++ ++R +
Sbjct: 427 AGDLEDKRIKS--KKIIGSSIGVSI--LLLLSFIIFHFWKRKQKRSITIQTPIVDLVRSQ 482
Query: 499 GRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYK 558
+ VK E K + +E+P +++++ +AT+ FS NKLG+GG+G VYK
Sbjct: 483 DSLMNEL--VKASRSYTSKENK-TDYLELPLMEWKALAMATNNFSTDNKLGQGGFGIVYK 539
Query: 559 GKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYM 618
G L G+EIAVKRLS +SSQG EF NEV LIAKLQH NLVRL G C+ EK+LIYEY+
Sbjct: 540 GMLLDGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRLLGCCVDKGEKMLIYEYL 599
Query: 619 PNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGE 678
N SLD+ +FD T+S+ L+WQ RFDI+ GIARGLLYLHQDSR R+IHRDLK SN+LLD
Sbjct: 600 ENLSLDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRCRIIHRDLKASNVLLDKN 659
Query: 679 MQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEII 738
M PKISDFG+ARIFG +ETEANT+RVVGTYGYMSPEYA+DG FS KSD+FSFGV+LLEII
Sbjct: 660 MTPKISDFGMARIFGREETEANTRRVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEII 719
Query: 739 SGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDL----SLGEAYNANQFIRCTHVGLL 794
SGK+N GFY L+LLG+ W+ W E K L+++D +L + ++ +RC +GLL
Sbjct: 720 SGKRNKGFYNSNRDLNLLGFVWRHWKEGKELEIVDPINIDALSSEFPTHEILRCIQIGLL 779
Query: 795 CVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFT-RKDLSSTASSSLQFD 845
CVQ+ +DRP MS+V++ML SET +P PK+P F R L +SSS Q D
Sbjct: 780 CVQERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCVGRSSLEVDSSSSTQRD 831
>At1g11330 receptor-like protein kinase, putative
Length = 840
Score = 545 bits (1404), Expect = e-155
Identities = 346/859 (40%), Positives = 480/859 (55%), Gaps = 83/859 (9%)
Query: 14 LFHMHCWLLCFSQLCFAGDTLNVGQEITGNGT-VLVSAAKKFELGFFSPDLNVTGGKGRY 72
L + C L +LCF D + I + + L+ + F GFF+P + T + RY
Sbjct: 13 LLLLACTCLLSRRLCFGEDRITFSSPIKDSESETLLCKSGIFRFGFFTPVNSTT--RLRY 70
Query: 73 LGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTN 132
+GIWY E + VVWVAN+D+P+ D S GV I DGNL V D WS +N++
Sbjct: 71 VGIWY---EKIPIQTVVWVANKDSPINDTS-GVISIYQDGNLAVTDGRNRLVWS-TNVSV 125
Query: 133 SSSATNRSVKLMDSGNLVLLDE-HVGMKLWESFEHPTDTFLPGMKMDK------TLELTC 185
+ V+LMDSGNL+L D + G LWESF+HP D+F+P M + L+LT
Sbjct: 126 PVAPNATWVQLMDSGNLMLQDNRNNGEILWESFKHPYDSFMPRMTLGTDGRTGGNLKLTS 185
Query: 186 WKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISN-DVYN 244
W S DP GN+T + I W+S V N D + D +N
Sbjct: 186 WTSHDDPSTGNYTAGIAPFTFPELLIWKNNVPTWRSGPWNGQVFIGLPNMDSLLFLDGFN 245
Query: 245 LLTNFKELKNKTVS-SYDNTRLL----LNSTGVIKVLYRVNFQSDIVWWYQ----PRTTC 295
L ++ T+S SY N + L+ G+I Y+ ++ + + W P T C
Sbjct: 246 LNSD----NQGTISMSYANDSFMYHFNLDPEGII---YQKDWSTSMRTWRIGVKFPYTDC 298
Query: 296 LTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKST-SC------ 348
Y CG F SC+ + C C+ GF P N+ G S C RK+ C
Sbjct: 299 DAYGRCGRFGSCHAGENPPCKCVKGF---VPKNNTEWNGGNWSNGCMRKAPLQCERQRNV 355
Query: 349 -----GANTNTFLNLTMMKIGSPDIKVSAQDENECKFRCISMCSQTQCQACSYVPIPVQQ 403
G + FL L MK+ + S E C C+ CS C A +Y
Sbjct: 356 SNGGGGGKADGFLKLQKMKVPI-SAERSEASEQVCPKVCLDNCS---CTAYAY------D 405
Query: 404 RGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIAL 463
RG+ C +W+ +L ++ G D LF+RVA S+++ S L++++ +
Sbjct: 406 RGIG---CMLWSGDLVDMQSFLGSGID--LFIRVAHSELKT-------HSNLAVMIAAPV 453
Query: 464 PGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNE 523
GV+++A +C+L CR+ K S ++ +R S+ E N+
Sbjct: 454 IGVMLIAAVCVLL-ACRKYKKRPAKDRSAELMFKRMEALTSDN-----------ESASNQ 501
Query: 524 GI--EVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQ 581
E+P F+F+ + +TD FS NKLG+GG+GPVYKGKL G+EIAVKRLS S QG++
Sbjct: 502 IKLKELPLFEFQVLATSTDSFSLRNKLGQGGFGPVYKGKLPEGQEIAVKRLSRKSGQGLE 561
Query: 582 EFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMR 641
E NEVV+I+KLQHRNLV+L G CI+G+E++L+YEYMP KSLDA++FDP K +LDW+ R
Sbjct: 562 ELMNEVVVISKLQHRNLVKLLGCCIEGEERMLVYEYMPKKSLDAYLFDPMKQKILDWKTR 621
Query: 642 FDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANT 701
F+I+ GI RGLLYLH+DSRL++IHRDLK SNILLD + PKISDFGLARIF E EANT
Sbjct: 622 FNIMEGICRGLLYLHRDSRLKIIHRDLKASNILLDENLNPKISDFGLARIFRANEDEANT 681
Query: 702 QRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWK 761
+RVVGTYGYMSPEYA++G FS KSD+FS GV+ LEIISG++N+ ++ + L+LL YAWK
Sbjct: 682 RRVVGTYGYMSPEYAMEGFFSEKSDVFSLGVIFLEIISGRRNSSSHKEENNLNLLAYAWK 741
Query: 762 LWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLP 821
LW + + L D ++ + + +C H+GLLCVQ+ +DRPN+SNV+ ML +E +L
Sbjct: 742 LWNDGEAASLADPAVFDKCFEKEIEKCVHIGLLCVQEVANDRPNVSNVIWMLTTENMSLA 801
Query: 822 TPKQPTFFTRKDLSSTASS 840
PKQP F R+ S SS
Sbjct: 802 DPKQPAFIVRRGASEAESS 820
>At1g11350 putative receptor-like protein kinase gi|4008010; similar
to EST gb|AI999419.1
Length = 830
Score = 537 bits (1384), Expect = e-153
Identities = 340/844 (40%), Positives = 475/844 (55%), Gaps = 68/844 (8%)
Query: 21 LLCFS-QLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYR 79
L+CFS +LC A D + E + TV VS F GFFSP GRY GIW+
Sbjct: 11 LICFSLRLCLATDVITFSSEFRDSETV-VSNHSTFRFGFFSP----VNSTGRYAGIWF-- 63
Query: 80 EEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNR 139
+ VVWVAN ++P+ +DS G+ I+ +GNLVV+D G +WS +N+ +A
Sbjct: 64 -NNIPVQTVVWVANSNSPI-NDSSGMVSISKEGNLVVMDGRGQVHWS-TNVLVPVAANTF 120
Query: 140 SVKLMDSGNLVLLDE-HVGMK-LWESFEHPTDTFLPGM------KMDKTLELTCWKSLSD 191
+L+++GNLVLL + G + LWESFEHP + +LP M K ++L+L WKS D
Sbjct: 121 YARLLNTGNLVLLGTTNTGDEILWESFEHPQNIYLPTMSLATDTKTGRSLKLRSWKSPFD 180
Query: 192 PGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKE 251
P G ++ + + L W+S N D N ++ L +
Sbjct: 181 PSPGRYSAGLIPLPFPELVVWKDDLLMWRSGPWNGQYFIGLPNMDYRIN-LFELTLSSDN 239
Query: 252 LKNKTVSSYDNTRL---LLNSTG-VIKVLYRVNFQSDIVWWYQPRTTCLTYNVCGNFSSC 307
+ ++S NT L LL+S G V + + V Q W P T C TY CG F+SC
Sbjct: 240 RGSVSMSYAGNTLLYHFLLDSEGSVFQRDWNVAIQEWKTWLKVPSTKCDTYATCGQFASC 299
Query: 308 --NDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKST-SCGANTNT--------FL 356
N + C C+ GF +S ++ G T C RK+ C + N F+
Sbjct: 300 RFNPGSTPPCMCIRGFKPQS-YAEWNNGNWTQG--CVRKAPLQCESRDNNDGSRKSDGFV 356
Query: 357 NLTMMKIGSPDIKVSAQDENECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQ 416
+ MK+ + S +E +C C+ CS C A S+ RG+ C +W+
Sbjct: 357 RVQKMKVPHNPQR-SGANEQDCPESCLKNCS---CTAYSF------DRGIG---CLLWSG 403
Query: 417 NLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILA 476
NL ++E G ++R+A S+ ++ T + + L+ G V+LA I
Sbjct: 404 NLMDMQE--FSGTGVVFYIRLADSEFKKRTNRSIVITVTLLVGAFLFAGTVVLALWKIAK 461
Query: 477 YVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESIL 536
+ + K + +L +R S L+++ L+E +P F+F+ +
Sbjct: 462 H--------REKNRNTRLLNERMEALSSNDVGAILVNQYKLKE-------LPLFEFQVLA 506
Query: 537 VATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHR 596
VAT+ FS NKLG+GG+G VYKG+LQ G +IAVKRLS S QG++EF NEVV+I+KLQHR
Sbjct: 507 VATNNFSITNKLGQGGFGAVYKGRLQEGLDIAVKRLSRTSGQGVEEFVNEVVVISKLQHR 566
Query: 597 NLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLH 656
NLVRL G+CI+G+E++L+YE+MP LDA++FDP K LLDW+ RF+I+ GI RGL+YLH
Sbjct: 567 NLVRLLGFCIEGEERMLVYEFMPENCLDAYLFDPVKQRLLDWKTRFNIIDGICRGLMYLH 626
Query: 657 QDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYA 716
+DSRL++IHRDLK SNILLD + PKISDFGLARIF G E E +T RVVGTYGYM+PEYA
Sbjct: 627 RDSRLKIIHRDLKASNILLDENLNPKISDFGLARIFQGNEDEVSTVRVVGTYGYMAPEYA 686
Query: 717 LDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSL 776
+ G FS KSD+FS GV+LLEI+SG++N+ FY +L YAWKLW + + L+D +
Sbjct: 687 MGGLFSEKSDVFSLGVILLEIVSGRRNSSFYNDGQNPNLSAYAWKLWNTGEDIALVDPVI 746
Query: 777 GEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRKDLSS 836
E N+ RC HVGLLCVQD +DRP+++ V+ ML SE + LP PKQP F R+ S
Sbjct: 747 FEECFENEIRRCVHVGLLCVQDHANDRPSVATVIWMLSSENSNLPEPKQPAFIPRRGTSE 806
Query: 837 TASS 840
SS
Sbjct: 807 VESS 810
>At4g11900 KI domain interacting kinase 1 -like protein
Length = 849
Score = 531 bits (1368), Expect = e-151
Identities = 334/858 (38%), Positives = 473/858 (54%), Gaps = 84/858 (9%)
Query: 12 IFLFHMHCWLLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGR 71
+FL + + Q+ + DT++ Q ++G T+ VS+ FELG F+P + +
Sbjct: 8 VFLLYYGVLVFLSFQVSSSTDTISTNQPLSGFETI-VSSGDIFELGLFTPTPDTYDHRNY 66
Query: 72 YLGIWYYREEGSGLSP--VVWVANRDNPVADDSIGVFRIADDGNLVVLD---------TS 120
Y+G+WY +SP +VWVANR++P+ D+ DGNL++ D T
Sbjct: 67 YIGMWYRH-----VSPQTIVWVANRESPLGGDASTYLLKILDGNLILHDNISATRKSHTE 121
Query: 121 GIRYWSASNLT---------------NSSSATNRSVKLMDSGNLVLLD--EHVGMKLWES 163
G S ++ NSS + + L DSGNLVL D LW+S
Sbjct: 122 GTSRRSPQKISEGNLLFHETVWSTGVNSSMSKDVQAVLFDSGNLVLRDGPNSSAAVLWQS 181
Query: 164 FEHPTDTFLPGMKMDKTLEL-TCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSE 222
F+HP+DT+LPG K+ +L T W+SL DP G ++ + D K + + N+ + YW S
Sbjct: 182 FDHPSDTWLPGGKIRLGSQLFTSWESLIDPSPGRYSLEFDPKLHSLVTVWNRSKSYWSSG 241
Query: 223 EQGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVSSYDNTRLLLNSTGVIKV-LYRVNF 281
D + + + P+ + L + +V RL++ +G + ++ V+
Sbjct: 242 PLYDWLQSFKGFPELQGTKLSFTLNMDESYITFSVDPQSRYRLVMGVSGQFMLQVWHVDL 301
Query: 282 QSDIVWWYQPRTTCLTYNVCGNFSSCNDDNDKL-CTCLPGFGRR-----SPLNDYTVGGD 335
QS V QP C YN CG+F CN++ + C C+PGF R NDY+ G
Sbjct: 302 QSWRVILSQPDNRCDVYNSCGSFGICNENREPPPCRCVPGFKREFSQGSDDSNDYSGG-- 359
Query: 336 TSSLLCTRKS-TSCGANTNTFLNLTMMKIGSPDIKVSAQDENE---CKFRCISMCSQTQC 391
C R++ C + FL + MK+ + S C RC++ CS C
Sbjct: 360 -----CKRETYLHCYKRNDEFLPIENMKLATDPTTASVLTSGTFRTCASRCVADCS---C 411
Query: 392 QACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPT-RKGN 450
QA + + + C +WT++ L ++ F+R+A S+I RK
Sbjct: 412 QAYAN----------DGNKCLVWTKDAFNL-QQLDANKGHTFFLRLASSNISTANNRKTE 460
Query: 451 PKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKD 510
S++L + L +V A + Y C I+ +++++ +QR E+H ++
Sbjct: 461 HSKGKSIVLPLVLASLVATAACFVGLYCC---ISSRIRRKK----KQR-----DEKHSRE 508
Query: 511 LIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVK 570
L++ + D+ G + Y + I+VAT+ FS KLG GG+GPVYKGKL G E+A+K
Sbjct: 509 LLEGGLI---DDAGENMCYLNLHDIMVATNSFSRKKKLGEGGFGPVYKGKLPNGMEVAIK 565
Query: 571 RLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDP 630
RLS SSQG+ EFKNEVVLI KLQH+NLVRL GYC++GDEK+LIYEYM NKSLD +FD
Sbjct: 566 RLSKKSSQGLTEFKNEVVLIIKLQHKNLVRLLGYCVEGDEKLLIYEYMSNKSLDGLLFDS 625
Query: 631 TKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLAR 690
KS LDW+ R I+ G RGL YLH+ SRLR+IHRDLK SNILLD EM PKISDFG AR
Sbjct: 626 LKSRELDWETRMKIVNGTTRGLQYLHEYSRLRIIHRDLKASNILLDDEMNPKISDFGTAR 685
Query: 691 IFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYK 750
IFG K+ + +TQR+VGT+GYMSPEYAL G S KSDI+SFGV+LLEIISGKK T F
Sbjct: 686 IFGCKQIDDSTQRIVGTFGYMSPEYALGGVISEKSDIYSFGVLLLEIISGKKATRFVHND 745
Query: 751 GTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVV 810
SL+ Y W+ W E K + ++D + +Y+ + +RC H+ LLCVQD P DRP +S +V
Sbjct: 746 QKHSLIAYEWESWCETKGVSIIDEPMCCSYSLEEAMRCIHIALLCVQDHPKDRPMISQIV 805
Query: 811 IMLDSETATLPTPKQPTF 828
ML ++ TLP PKQPTF
Sbjct: 806 YMLSNDN-TLPIPKQPTF 822
>At4g27300 putative receptor protein kinase
Length = 815
Score = 530 bits (1366), Expect = e-150
Identities = 339/872 (38%), Positives = 492/872 (55%), Gaps = 96/872 (11%)
Query: 14 LFHMHCWLLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYL 73
LF + +L+ S + +E +G L S + F+LGFFS D + R+L
Sbjct: 7 LFSLSLFLISSSLSVALDYNVITPKEFLKDGDTLSSPDQVFQLGFFSLDQEEQP-QHRFL 65
Query: 74 GIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNS 133
G+WY VVWVANR+NP+ S G ++ G+L + D WS+S+ +
Sbjct: 66 GLWYMEPFA-----VVWVANRNNPLYGTS-GFLNLSSLGDLQLFDGEHKALWSSSSSSTK 119
Query: 134 SSAT--NRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLE------LTC 185
+S T N +K+ SGNL+ D + LW+SF++P +T L GMK+ K + L+
Sbjct: 120 ASKTANNPLLKISCSGNLISSDGEEAV-LWQSFDYPMNTILAGMKLGKNFKTQMEWSLSS 178
Query: 186 WKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNL 245
WK+L DP G+FT +D + + + G S G N S + N
Sbjct: 179 WKTLKDPSPGDFTLSLDTRGLPQLILRKNGD---SSYSYRLGSWNGLSFTGAPAMGRENS 235
Query: 246 LTNFKELKNKTVSSYDNT-------RLLLNSTGVIKVLYRVNFQSDIVWWYQPRTTCLTY 298
L ++K + +Y T RL+LN+TG + + I+ P C Y
Sbjct: 236 LFDYKFTSSAQEVNYSWTPRHRIVSRLVLNNTGKLHRFIQSKQNQWILANTAPEDECDYY 295
Query: 299 NVCGNFSSC--NDDNDKLCTCLPGF----GRRSPLNDYTVGGDTSSLLCTRKSTSCGANT 352
++CG ++ C N N C+CL GF GR+ ++ G C + +
Sbjct: 296 SICGAYAVCGINSKNTPSCSCLQGFKPKSGRKWNISRGAYG-------CVHEIPTNCEKK 348
Query: 353 NTFLNLTMMKIGSPDIKVSAQDEN------ECKFRCISMCSQTQCQACSYVPIPVQQRGL 406
+ F+ +K+ PD S D +CK +C S CS T +Y +++ G
Sbjct: 349 DAFVKFPGLKL--PDTSWSWYDAKNEMTLEDCKIKCSSNCSCT-----AYANTDIREGGK 401
Query: 407 NLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGV 466
C +W +L ++E G D +++R+ + IE R+ ++G+ + V
Sbjct: 402 G---CLLWFGDLVDMREYSSFGQD--VYIRMGFAKIEFKGRE---------VVGMVVGSV 447
Query: 467 VILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIE 526
V +A + ++ + C RK +K R RG + ++G+EE+D ++
Sbjct: 448 VAIAVVLVVVFACFRKKIMK---------RYRGENF-----------RKGIEEED---LD 484
Query: 527 VPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNE 586
+P FD ++I +ATD FS N LGRGG+GPVYKGKL+ G+EIAVKRLS+ S QG++EFKNE
Sbjct: 485 LPIFDRKTISIATDDFSYVNFLGRGGFGPVYKGKLEDGQEIAVKRLSANSGQGVEEFKNE 544
Query: 587 VVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILL 646
V LIAKLQHRNLVRL G CI+G+E +LIYEYMPNKSLD F+FD +S LDW+ R +I+
Sbjct: 545 VKLIAKLQHRNLVRLLGCCIQGEECMLIYEYMPNKSLDFFIFDERRSTELDWKKRMNIIN 604
Query: 647 GIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVG 706
G+ARG+LYLHQDSRLR+IHRDLK N+LLD +M PKISDFGLA+ FGG ++E++T RVVG
Sbjct: 605 GVARGILYLHQDSRLRIIHRDLKAGNVLLDNDMNPKISDFGLAKSFGGDQSESSTNRVVG 664
Query: 707 TYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTEN 766
TYGYM PEYA+DG FS KSD+FSFGV++LEII+GK N GF L+LLG+ WK+W E+
Sbjct: 665 TYGYMPPEYAIDGHFSVKSDVFSFGVLVLEIITGKTNRGFRHADHDLNLLGHVWKMWVED 724
Query: 767 KLLDLMDLS-LGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQ 825
+ +++ + L E + +RC HV LLCVQ +P+DRP M++VV+M S+ ++LP P Q
Sbjct: 725 REIEVPEEEWLEETSVIPEVLRCIHVALLCVQQKPEDRPTMASVVLMFGSD-SSLPHPTQ 783
Query: 826 PTFFTRK---DLSSTASSSLQFDSSI--VEGR 852
P FFT + D+SS+ S Q + SI ++GR
Sbjct: 784 PGFFTNRNVPDISSSLSLRSQNEVSITMLQGR 815
>At1g11300 putative s-locus protein kinase (PPC:1.7.2)
Length = 820
Score = 530 bits (1365), Expect = e-150
Identities = 329/818 (40%), Positives = 465/818 (56%), Gaps = 83/818 (10%)
Query: 47 LVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVF 106
+VS+ + F GFFSP RY GIWY + V+WVAN+D P+ +DS GV
Sbjct: 42 IVSSFRTFRFGFFSP----VNSTSRYAGIWY---NSVSVQTVIWVANKDKPI-NDSSGVI 93
Query: 107 RIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEH 166
++ DGNLVV D WS +N++ +SA + +L+DSGNLVL + LWESF++
Sbjct: 94 SVSQDGNLVVTDGQRRVLWS-TNVSTQASANSTVAELLDSGNLVLKEASSDAYLWESFKY 152
Query: 167 PTDTFLPGMKMDKT-------LELTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQ--- 216
PTD++LP M + + +T WKS SDP G++T + I+N
Sbjct: 153 PTDSWLPNMLVGTNARIGGGNVTITSWKSPSDPSPGSYTAALVLAAYPELFIMNNNNNNS 212
Query: 217 LYWQSEEQGDGVMNPESNPDDISND-VYNLLTNFKELKNKTVSSYDNTRLL----LNSTG 271
W+S + N PD + +Y + N + SY N L ++ G
Sbjct: 213 TVWRSGPWNGQMFN--GLPDVYAGVFLYRFIVN-DDTNGSVTMSYANDSTLRYFYMDYRG 269
Query: 272 -VIKVLYRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPL--- 327
VI+ + ++ V P T C Y CG F++CN + LC+C+ GF R+ +
Sbjct: 270 SVIRRDWSETRRNWTVGLQVPATECDNYRRCGEFATCNPRKNPLCSCIRGFRPRNLIEWN 329
Query: 328 NDYTVGGDTSS--LLCTRKSTSCGANTNTFLNLTMMKIGSPDI-KVSAQDENECKFRCIS 384
N GG T L C R++ + A+ FL L MK+ PD + S E EC C+
Sbjct: 330 NGNWSGGCTRRVPLQCERQNNNGSADG--FLRLRRMKL--PDFARRSEASEPECLRTCLQ 385
Query: 385 MCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEE 444
CS C A ++ GL C IW +L +E G D L++R+A S+I+
Sbjct: 386 TCS---CIAAAH--------GLGYG-CMIWNGSLVDSQELSASGLD--LYIRLAHSEIKT 431
Query: 445 PTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDS 504
++ +++G L G + + C+L R+I +K + + +
Sbjct: 432 KDKR-------PILIGTILAGGIFVVAACVLL---ARRIVMKKRAKKKG----------- 470
Query: 505 ERHVKDLIDKEGLEEKDNEGI--EVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQ 562
R + + ++ N+G E+P F+F+ + AT+ FS NKLG+GG+GPVYKGKLQ
Sbjct: 471 -RDAEQIFERVEALAGGNKGKLKELPLFEFQVLAAATNNFSLRNKLGQGGFGPVYKGKLQ 529
Query: 563 GGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKS 622
G+EIAVKRLS S QG++E NEVV+I+KLQHRNLV+L G CI G+E++L+YE+MP KS
Sbjct: 530 EGQEIAVKRLSRASGQGLEELVNEVVVISKLQHRNLVKLLGCCIAGEERMLVYEFMPKKS 589
Query: 623 LDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPK 682
LD ++FD ++ LLDW+ RF+I+ GI RGLLYLH+DSRLR+IHRDLK SNILLD + PK
Sbjct: 590 LDYYLFDSRRAKLLDWKTRFNIINGICRGLLYLHRDSRLRIIHRDLKASNILLDENLIPK 649
Query: 683 ISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKK 742
ISDFGLARIF G E EANT+RVVGTYGYM+PEYA+ G FS KSD+FS GV+LLEIISG++
Sbjct: 650 ISDFGLARIFPGNEDEANTRRVVGTYGYMAPEYAMGGLFSEKSDVFSLGVILLEIISGRR 709
Query: 743 NTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDD 802
N+ +LL Y W +W E ++ L+D + + + +C H+GLLCVQ+ +D
Sbjct: 710 NS-------NSTLLAYVWSIWNEGEINSLVDPEIFDLLFEKEIHKCIHIGLLCVQEAAND 762
Query: 803 RPNMSNVVIMLDSETATLPTPKQPTFFTRKDLSSTASS 840
RP++S V ML SE A +P PKQP F +R ++ SS
Sbjct: 763 RPSVSTVCSMLSSEIADIPEPKQPAFISRNNVPEAESS 800
>At1g11410 putative brassinosteroid insensitive protein; similar to
EST gb|W43102
Length = 840
Score = 522 bits (1345), Expect = e-148
Identities = 337/868 (38%), Positives = 480/868 (54%), Gaps = 92/868 (10%)
Query: 13 FLFHMHCWLLCFS-QLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGR 71
F F +L F Q C++ +T+ Q + +G V+ S K+F GFFS + K R
Sbjct: 3 FFFIFFIFLFSFLIQSCYSDNTILRSQSLK-DGDVIYSEGKRFAFGFFS----LGNSKLR 57
Query: 72 YLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDT-SGIRYWSASNL 130
Y+GIWY + +VWVANRD+P+ D S G+ + + GNL V + +G ++++
Sbjct: 58 YVGIWYAQVSEQ---TIVWVANRDHPINDTS-GLIKFSTRGNLCVYASGNGTEPIWSTDV 113
Query: 131 TNSSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLE------LT 184
+ KL D GNLVLLD G WESF HPT+T LP MK T + +T
Sbjct: 114 IDMIQEPALVAKLSDLGNLVLLDPVTGKSFWESFNHPTNTLLPFMKFGFTRQSGVDRIMT 173
Query: 185 CWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSE----EQGDGVMNPESNPDDISN 240
W+S DPG GN T++++++ + + L+W++ ++ GV P+ +
Sbjct: 174 SWRSPGDPGSGNITYRIERRGFPQMMMYKGLTLWWRTGSWTGQRWSGV------PEMTNK 227
Query: 241 DVYNL-LTNFKELKNKTVSSYD---NTRLLLNSTGVIKVLYRVNFQSD--IVWWYQPRTT 294
++N+ N + + T D TR++LN TG ++ +R N + I +W P
Sbjct: 228 FIFNISFVNNPDEVSITYGVLDASVTTRMVLNETGTLQ-RFRWNGRDKKWIGFWSAPEDK 286
Query: 295 CLTYNVCGNFSSCNDDNDKL--CTCLPGFGRRSP----LNDYTVGGDTSSLLCTR-KSTS 347
C YN CG C+ + + C+CLPG+ ++P L D + G CTR K+ S
Sbjct: 287 CDIYNHCGFNGYCDSTSTEKFECSCLPGYEPKTPRDWFLRDASDG-------CTRIKADS 339
Query: 348 CGANTNTFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQ 403
F L +KI P+ D N EC+ RC+ CS A +Y
Sbjct: 340 ICNGKEGFAKLKRVKI--PNTSAVNVDMNITLKECEQRCLKNCSCV-AYASAYHESQDGA 396
Query: 404 RGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSD-IEEPTRKGNPKSTLSLILGIA 462
+G C W N+ + G D ++RV KSD +E R G ++
Sbjct: 397 KG-----CLTWHGNMLDTRTYLSSGQD--FYLRVDKSDAVEWKWRIGEEET--------- 440
Query: 463 LPGVVILACICILAYVCRRKIA-------LKLKQESESILRQRGRFYDSERHVKDLIDKE 515
C + C R + K + + ++ + F S ++D E
Sbjct: 441 --------CFNSHQFDCSRDVTTDQFSLLFKEAKTTNTLRKAPSSFAPSSFDLEDSFILE 492
Query: 516 GLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSV 575
LE+K E+P F+ +I AT+ F+ NKLG GG+GPVYKG LQ G EIAVKRLS
Sbjct: 493 ELEDKSRSR-ELPLFELSTIATATNNFAFQNKLGAGGFGPVYKGVLQNGMEIAVKRLSKS 551
Query: 576 SSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSAL 635
S QG++EFKNEV LI+KLQHRNLVR+ G C++ +EK+L+YEY+PNKSLD F+F + A
Sbjct: 552 SGQGMEEFKNEVKLISKLQHRNLVRILGCCVEFEEKMLVYEYLPNKSLDYFIFHEEQRAE 611
Query: 636 LDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGK 695
LDW R I+ GI RG+LYLHQDSRLR+IHRDLK SN+LLD EM PKI+DFGLARIFGG
Sbjct: 612 LDWPKRMGIIRGIGRGILYLHQDSRLRIIHRDLKASNVLLDNEMIPKIADFGLARIFGGN 671
Query: 696 ETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSL 755
+ E +T RVVGTYGYMSPEYA+DGQFS KSD++SFGV++LEII+GK+N+ FY+ +L+L
Sbjct: 672 QIEGSTNRVVGTYGYMSPEYAMDGQFSIKSDVYSFGVLILEIITGKRNSAFYE--ESLNL 729
Query: 756 LGYAWKLWTENKLLDLMDLSLG-EAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLD 814
+ + W W + ++++D +G E Y+ + ++C H+GLLCVQ+ DRP+MS+VV ML
Sbjct: 730 VKHIWDRWENGEAIEIIDKLMGEETYDEGEVMKCLHIGLLCVQENSSDRPDMSSVVFMLG 789
Query: 815 SETATLPTPKQPTFFT-RKDLSSTASSS 841
LP+PK P F R+ + T SS
Sbjct: 790 HNAIDLPSPKHPAFTAGRRRNTKTGGSS 817
>At1g61610
Length = 842
Score = 516 bits (1328), Expect = e-146
Identities = 332/861 (38%), Positives = 475/861 (54%), Gaps = 98/861 (11%)
Query: 7 TNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVT 66
T + T+ +FH LC + C ++ I G L+S + FELGFF+P +
Sbjct: 9 TLVTTLLIFHQ----LCSNVSCSTSNSFTRNHTIR-EGDSLISEDESFELGFFTPKNSTL 63
Query: 67 GGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWS 126
RY+GIWY E VVWVANR+ P+ D G +IADDGNLV+++ WS
Sbjct: 64 ----RYVGIWYKNIEPQ---TVVWVANREKPLLDHK-GALKIADDGNLVIVNGQNETIWS 115
Query: 127 ASNLTNSSSATNRSVK-LMDSGNLVLLDEHVGMK-LWESFEHPTDTFLPGMK------MD 178
TN +N +V L +G+LVL + K WESF +PTDTFLPGM+ +
Sbjct: 116 ----TNVEPESNNTVAVLFKTGDLVLCSDSDRRKWYWESFNNPTDTFLPGMRVRVNPSLG 171
Query: 179 KTLELTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDI 238
+ WKS SDP G ++ +D I + W+S + +
Sbjct: 172 ENRAFIPWKSESDPSPGKYSMGIDPVGALEIVIWEGEKRKWRSGPWNSAIFTGIPDMLRF 231
Query: 239 SNDVYNLLTNFKELKNKTV-------SSYDNTRLLLNSTGVIKVLYRVNFQSDIVWW--- 288
+N +Y + ++ +V S D R + GV + + + DI W
Sbjct: 232 TNYIYGFKLSSPPDRDGSVYFTYVASDSSDFLRFWIRPDGVEE---QFRWNKDIRNWNLL 288
Query: 289 -YQPRTTCLTYNVCGNFSSCNDDND---KLCTCLPGFGRRSPLN-------DYTVGGDTS 337
++P T C YN CGN+S C+D + C+C+ GF P++ D++ G
Sbjct: 289 QWKPSTECEKYNRCGNYSVCDDSKEFDSGKCSCIDGF---EPVHQDQWNNRDFSGGCQRR 345
Query: 338 SLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVSAQDENECKFRCISMCSQTQCQACSYV 397
L +S G F L +K+ V + CK C CS C+A + V
Sbjct: 346 VPLNCNQSLVAGQEDG-FTVLKGIKVPDFGSVVLHNNSETCKDVCARDCS---CKAYALV 401
Query: 398 PIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSL 457
G+ C IWT++L ++ GG+ + +R+A S + G STL +
Sbjct: 402 V------GIG---CMIWTRDLIDMEHFERGGNS--INIRLAGSKLGG----GKENSTLWI 446
Query: 458 ILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGL 517
I+ + ++ CI IL K K+ ++ L ++ + V D+I+
Sbjct: 447 IVFSVIGAFLLGLCIWIL---------WKFKKSLKAFLWKK-----KDITVSDIIENRDY 492
Query: 518 EEKDNEGI--------EVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAV 569
+ + ++P F F+S+ AT F++ NKLG+GG+G VYKG GREIAV
Sbjct: 493 SSSPIKVLVGDQVDTPDLPIFSFDSVASATGDFAEENKLGQGGFGTVYKGNFSEGREIAV 552
Query: 570 KRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFD 629
KRLS S QG++EFKNE++LIAKLQHRNLVRL G CI+ +EK+L+YEYMPNKSLD F+FD
Sbjct: 553 KRLSGKSKQGLEEFKNEILLIAKLQHRNLVRLLGCCIEDNEKMLLYEYMPNKSLDRFLFD 612
Query: 630 PTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLA 689
+K LDW+ R++++ GIARGLLYLH+DSRL++IHRDLK SNILLD EM PKISDFG+A
Sbjct: 613 ESKQGSLDWRKRWEVIGGIARGLLYLHRDSRLKIIHRDLKASNILLDTEMNPKISDFGMA 672
Query: 690 RIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQY 749
RIF ++ ANT RVVGTYGYM+PEYA++G FS KSD++SFGV++LEI+SG+KN F
Sbjct: 673 RIFNYRQDHANTIRVVGTYGYMAPEYAMEGIFSEKSDVYSFGVLILEIVSGRKNVSF--- 729
Query: 750 KGT--LSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMS 807
+GT SL+GYAW LW++ K +++D + + + + +RC HVG+LC QD RPNM
Sbjct: 730 RGTDHGSLIGYAWHLWSQGKTKEMIDPIVKDTRDVTEAMRCIHVGMLCTQDSVIHRPNMG 789
Query: 808 NVVIMLDSETATLPTPKQPTF 828
+V++ML+S+T+ LP P+QPTF
Sbjct: 790 SVLLMLESQTSQLPPPRQPTF 810
>At4g21370 receptor kinase - like protein
Length = 844
Score = 507 bits (1306), Expect = e-144
Identities = 336/866 (38%), Positives = 481/866 (54%), Gaps = 91/866 (10%)
Query: 11 TIFLFHMHCWLLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLNVTGGK 69
T F+F + +L+ F L + +TL+ + +T + +VS FELGFF G
Sbjct: 13 TFFVF-LFFFLILFPDLSISVNTLSATESLTISSNKTIVSPGGVFELGFFR-----ILGD 66
Query: 70 GRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASN 129
YLGIWY + VWVANRD P+++ IG+ +I++ NLV+LD S WS
Sbjct: 67 SWYLGIWYKK---ISQRTYVWVANRDTPLSNP-IGILKISN-ANLVILDNSDTHVWS--- 118
Query: 130 LTNSSSATNRSV--KLMDSGNLVLLDEHVGMK---LWESFEHPTDTFLPGMKMDKTLE-- 182
TN + A SV +L+D+GN VL + LW+SF+ PTDT LP MK+ + +
Sbjct: 119 -TNLTGAVRSSVVAELLDNGNFVLRGSKINESDEFLWQSFDFPTDTLLPQMKLGRDHKRG 177
Query: 183 ----LTCWKSLSDPGRGNFTFKMDKKW-ENRFAILNQGQLYWQSEEQG---DGVMNPESN 234
+T WKS DP G+F FK++ F + ++Y G G++ +
Sbjct: 178 LNRFVTSWKSSFDPSSGSFMFKLETLGLPEFFGFTSFLEVYRSGPWDGLRFSGILEMQQW 237
Query: 235 PDDISNDVYNLLTNFKELKN--KTVSSYDNTRLLLNSTGVIK-VLYRVNFQSDIVWWYQP 291
DDI +YN N +E+ + +RL +N+ G ++ + Q ++W+ P
Sbjct: 238 -DDI---IYNFTENREEVAYTFRVTDHNSYSRLTINTVGRLEGFTWEPTQQEWNMFWFMP 293
Query: 292 RTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKST-SCGA 350
+ TC Y +CG ++ C+ C C+ GF SP D+ G T C RK+ +CG
Sbjct: 294 KDTCDLYGICGPYAYCDMSTSPTCNCIKGFQPLSP-QDWASGDVTGR--CRRKTQLTCGE 350
Query: 351 NTNTFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGL 406
+ F L MKI P + D+ EC+ +C +T C +Y ++ G
Sbjct: 351 DR--FFRLMNMKI--PATTAAIVDKRIGLKECEEKC-----KTHCNCTAYANSDIRNGG- 400
Query: 407 NLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGV 466
S C IW ++ G D LFVR+A ++ E R+ + LI+GI+L V
Sbjct: 401 --SGCIIWIGEFRDIRNYAADGQD--LFVRLAAAEFGE--RRTIRGKIIGLIIGISLMLV 454
Query: 467 VILACICILAYVCRRKIA----LKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDN 522
+ C +R A + + + ++ G S R + +
Sbjct: 455 LSFIIYCFWKKKQKRARATAAPIGYRDRIQELIITNGVVMSSGRRLLG----------EE 504
Query: 523 EGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQE 582
E +E+P +FE++++AT+ FSD+N LGRGG+G VYK IAVKRLS +SSQG E
Sbjct: 505 EDLELPLTEFETVVMATENFSDSNILGRGGFGIVYK--------IAVKRLSEMSSQGTNE 556
Query: 583 FKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSA-LLDWQMR 641
FKNEV LIA+LQH NLVRL CI DEKILIYEY+ N SLD+ +F+ T+S+ L+WQ R
Sbjct: 557 FKNEVRLIARLQHINLVRLLSCCIYADEKILIYEYLENGSLDSHLFETTQSSNKLNWQTR 616
Query: 642 FDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANT 701
F I+ GIARGLLYLHQDSR ++IHRDLK SN+LLD M PKISDFG+ARIF ETEANT
Sbjct: 617 FSIINGIARGLLYLHQDSRFKIIHRDLKASNVLLDKNMTPKISDFGMARIFERDETEANT 676
Query: 702 QRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWK 761
++VVGTYGYMSPEYA++G FS KSD+FSFGV++LEI+SGK+N GF+ +LLGY W+
Sbjct: 677 RKVVGTYGYMSPEYAMEGIFSVKSDVFSFGVLVLEIVSGKRNRGFHNSGQDNNLLGYTWE 736
Query: 762 LWTENKLLDLMDLSLGEA------YNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDS 815
W E K L+++D + ++ + ++ +RC +GLLCVQ+ +DRP MS+VV+ML S
Sbjct: 737 NWKEGKGLEIVDSIIVDSSSSMSLFQPHEVLRCIQIGLLCVQERAEDRPKMSSVVLMLGS 796
Query: 816 ETATLPTPKQPTFFTRKDLSSTASSS 841
E +P++P + R+ T SS
Sbjct: 797 EKGEYFSPRRPGYCVRRSSLDTDDSS 822
>At1g61370 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 814
Score = 489 bits (1258), Expect = e-138
Identities = 323/869 (37%), Positives = 482/869 (55%), Gaps = 97/869 (11%)
Query: 12 IFLFHMHCWLLCFSQLCFAGDT----LNVGQEITG-NGTVLVSAAKKFELGFFSPDLNVT 66
+F + L+ F FA T L++GQ ++ NGT +ELGFFSP+
Sbjct: 7 VFFASLLFLLIIFPSCAFAAITRASPLSIGQTLSSPNGT--------YELGFFSPN---- 54
Query: 67 GGKGRYLGIWYYREEGSGLSP--VVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRY 124
+ +Y+GIW+ ++P VVWVANRD PV +++ + I +G+L++++
Sbjct: 55 NSRNQYVGIWF-----KNITPRVVVWVANRDKPVTNNAANL-TINSNGSLILVEREQNVV 108
Query: 125 WSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKM------D 178
WS + + S+ +L+++GNLVL+D LWESFEH DT L + +
Sbjct: 109 WS---IGETFSSNELRAELLENGNLVLIDGVSERNLWESFEHLGDTMLLESSVMYDVPNN 165
Query: 179 KTLELTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQ----SEEQGDGVMNPESN 234
K L+ WK+ +DP G F ++ + + I+ + YW+ + + G+ + +
Sbjct: 166 KKRVLSSWKNPTDPSPGEFVAELTTQVPPQGFIMRGSRPYWRGGPWARVRFTGIPEMDGS 225
Query: 235 ---PDDISNDVY---NLLTNFKELKNKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDIVWW 288
DIS DV LT E +N +S L S G +K+++ N +
Sbjct: 226 HVSKFDISQDVAAGTGSLTYSLERRNSNLSY-----TTLTSAGSLKIIWN-NGSGWVTDL 279
Query: 289 YQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGG-DTSSLLCTR 343
P ++C YN CG F C N C CL GF +S ++T G ++L C
Sbjct: 280 EAPVSSCDVYNTCGPFGLCIRSNPPKCECLKGFVPKSDEEWNKRNWTGGCMRRTNLSCDV 339
Query: 344 KSTSCGANTNTFLNLTMMKIGSPDIK--VSAQDENECKFRCISMCSQTQCQACSYVPIPV 401
S++ N + + + PD +S +E +C+ RC+ CS C A SY+
Sbjct: 340 NSSATAQANNGDIFDIVANVKPPDFYEYLSLINEEDCQQRCLGNCS---CTAFSYI---- 392
Query: 402 QQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGI 461
+Q G C +W + L + + GG+ L +R+A S++ R K ++ I+ I
Sbjct: 393 EQIG-----CLVWNRELVDVMQFVAGGET--LSIRLASSELAGSNRV---KIIVASIVSI 442
Query: 462 ALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKD 521
++ +++ A Y ++ + + E+ D+ R E L+ +D
Sbjct: 443 SVFMILVFASYWYWRYKAKQNDSNPIPLETSQ---------DAWR--------EQLKPQD 485
Query: 522 NEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQ 581
V +FD ++IL T+ FS NKLG+GG+GPVYKG LQ G+EIA+KRLSS S QG++
Sbjct: 486 -----VNFFDMQTILTITNNFSMENKLGQGGFGPVYKGNLQDGKEIAIKRLSSTSGQGLE 540
Query: 582 EFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMR 641
EF NE++LI+KLQHRNLVRL G CI+G+EK+LIYE+M NKSL+ F+FD TK LDW R
Sbjct: 541 EFMNEIILISKLQHRNLVRLLGCCIEGEEKLLIYEFMANKSLNTFIFDSTKKLELDWPKR 600
Query: 642 FDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANT 701
F+I+ GIA GLLYLH+DS LRV+HRD+K SNILLD EM PKISDFGLAR+F G + +ANT
Sbjct: 601 FEIIQGIACGLLYLHRDSCLRVVHRDMKVSNILLDEEMNPKISDFGLARMFQGTQHQANT 660
Query: 702 QRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWK 761
+RVVGT GYMSPEYA G FS KSDI++FGV+LLEII+GK+ + F + +LL +AW
Sbjct: 661 RRVVGTLGYMSPEYAWTGMFSEKSDIYAFGVLLLEIITGKRISSFTIGEEGKTLLEFAWD 720
Query: 762 LWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLP 821
W E+ DL+D + + + ++ RC +GLLC+Q + DRPN++ V+ ML + T LP
Sbjct: 721 SWCESGGSDLLDQDISSSGSESEVARCVQIGLLCIQQQAGDRPNIAQVMSML-TTTMDLP 779
Query: 822 TPKQPTFFTRKDLSSTASSSLQFDSSIVE 850
PKQP F + S + S ++ ++I +
Sbjct: 780 KPKQPVFAMQVQESDSESKTMYSVNNITQ 808
>At1g61390 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 831
Score = 488 bits (1256), Expect = e-138
Identities = 327/839 (38%), Positives = 463/839 (54%), Gaps = 90/839 (10%)
Query: 44 GTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSP--VVWVANRDNPVADD 101
G L S +ELGFFSP+ + +Y+GIW+ ++P VVWVANRD PV
Sbjct: 53 GQTLSSPDGVYELGFFSPN----NSRKQYVGIWF-----KNIAPQVVVWVANRDKPVTKT 103
Query: 102 SIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLW 161
+ + I+ +G+L++LD + WS S+ +L+D+GNLV++D+ G LW
Sbjct: 104 AANL-TISSNGSLILLDGTQDVIWSTGEAFTSNKC---HAELLDTGNLVVIDDVSGKTLW 159
Query: 162 ESFEHPTDTFLPGMKM------DKTLELTCWKSLSDPGRGNFTFKMDKKWENRFAILNQG 215
+SFE+ +T LP + K LT W+S SDP G FT + + + I
Sbjct: 160 KSFENLGNTMLPQSSVMYDIPRGKNRVLTSWRSNSDPSPGEFTLEFTPQVPPQGLIRRGS 219
Query: 216 QLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVS-------SYDNTRLLLN 268
YW+S G S I + T +++ T S +Y + + L
Sbjct: 220 SPYWRS---GPWAKTRFSGIPGIDASYVSPFTVLQDVAKGTASFSYSMLRNYKLSYVTLT 276
Query: 269 STGVIKVLYRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLN 328
S G +K+L+ + +S + + P ++C Y CG F C + C CL GF +S +
Sbjct: 277 SEGKMKILWN-DGKSWKLHFEAPTSSCDLYRACGPFGLCVRSRNPKCICLKGFVPKS--D 333
Query: 329 DYTVGGDTSSLLCTRKSTSC---------GANTNTFLNLTMMKIGSPDIKVSAQDEN--E 377
D G+ +S R SC G T++F ++T +K +PD+ A N +
Sbjct: 334 DEWKKGNWTSGCVRRTQLSCHTNSSTKTQGKETDSFYHMTRVK--TPDLYQLAGFLNAEQ 391
Query: 378 CKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRV 437
C C+ CS C A +Y+ G+ C +W + L + G+ L +R+
Sbjct: 392 CYQDCLGNCS---CTAFAYIS------GIG---CLVWNRELVDTVQFLSDGES--LSLRL 437
Query: 438 AKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQ 497
A S++ R +ILG V L+ IL + + + KQ + +
Sbjct: 438 ASSELAGSNRT-------KIILGTT----VSLSIFVILVFAAYKSWRYRTKQNEPNPM-- 484
Query: 498 RGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVY 557
F S + D K+ +E +D G+ + FD +I AT+ FS +NKLG+GG+GPVY
Sbjct: 485 ---FIHSSQ---DAWAKD-MEPQDVSGVNL--FDMHTIRTATNNFSSSNKLGQGGFGPVY 535
Query: 558 KGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEY 617
KGKL G+EIAVKRLSS S QG EF NE+ LI+KLQH+NLVRL G CIKG+EK+LIYEY
Sbjct: 536 KGKLVDGKEIAVKRLSSSSGQGTDEFMNEIRLISKLQHKNLVRLLGCCIKGEEKLLIYEY 595
Query: 618 MPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDG 677
+ NKSLD F+FD T +DWQ RF+I+ G+ARGLLYLH+DSRLRVIHRDLK SNILLD
Sbjct: 596 LVNKSLDVFLFDSTLKFEIDWQKRFNIIQGVARGLLYLHRDSRLRVIHRDLKVSNILLDE 655
Query: 678 EMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEI 737
+M PKISDFGLAR+ G + + NT+RVVGT GYM+PEYA G FS KSDI+SFGV+LLEI
Sbjct: 656 KMIPKISDFGLARMSQGTQYQDNTRRVVGTLGYMAPEYAWTGVFSEKSDIYSFGVLLLEI 715
Query: 738 ISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQ 797
I G+K + F + T LL YAW+ W E K +DL+D +L ++ + + RC +GLLCVQ
Sbjct: 716 IIGEKISRFSEEGKT--LLAYAWESWCETKGVDLLDQALADSSHPAEVGRCVQIGLLCVQ 773
Query: 798 DEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRKDLSSTASSSL----QFDSSIVEGR 852
+P DRPN ++ ML + + LP+PKQPTF + S+ L + S+++GR
Sbjct: 774 HQPADRPNTLELMSML-TTISELPSPKQPTFTVHSRDDDSTSNDLITVNEITQSVIQGR 831
>At1g61500
Length = 804
Score = 487 bits (1254), Expect = e-137
Identities = 329/838 (39%), Positives = 462/838 (54%), Gaps = 98/838 (11%)
Query: 44 GTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSI 103
G L SA + +ELGFFSP+ + +Y+GIW+ + + VVWVANR+ PV D S
Sbjct: 36 GQTLSSANEVYELGFFSPN----NTQDQYVGIWF---KDTIPRVVVWVANREKPVTD-ST 87
Query: 104 GVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLWES 163
I+ G+L++L+ WS S +T SSS +L DSGNL ++D LW+S
Sbjct: 88 AYLAISSSGSLLLLNGKHGTVWS-SGVTFSSSGCR--AELSDSGNLKVIDNVSERALWQS 144
Query: 164 FEHPTDTFLPGMKMDKTLE------LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQL 217
F+H DT L + L LT WKS +DP G+F ++ + ++ ++
Sbjct: 145 FDHLGDTLLHTSSLTYNLATAEKRVLTSWKSYTDPSPGDFLGQITPQVPSQGFVMRGSTP 204
Query: 218 YWQSEE-----------QGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVSSYDNTRLL 266
YW+S + P + D++ Y LT F+ Y +R+
Sbjct: 205 YWRSGPWAKTRFTGIPFMDESYTGPFTLHQDVNGSGY--LTYFQR-------DYKLSRIT 255
Query: 267 LNSTGVIKVLYRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSP 326
L S G IK ++R N +++ P+ C Y CG F C +C C GF +S
Sbjct: 256 LTSEGSIK-MFRDNGMGWELYYEAPKKLCDFYGACGPFGLCVMSPSPMCKCFRGFVPKS- 313
Query: 327 LNDYTVGGDTSSLLCTRKS------TSCGANTNTFLNLTMMKIGSPDIKVSAQDEN--EC 378
+ ++ G T C R + S G + + F + +K PD A N EC
Sbjct: 314 VEEWKRGNWTGG--CVRHTELDCLGNSTGEDADDFHQIANIK--PPDFYEFASSVNAEEC 369
Query: 379 KFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVA 438
RC+ CS C A +Y+ +G+ C +W Q+L + G+ L +R+A
Sbjct: 370 HQRCVHNCS---CLAFAYI------KGIG---CLVWNQDLMDAVQFSATGE--LLSIRLA 415
Query: 439 KSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQR 498
+S+++ RK K+ ++ I+ + L +IL + CR
Sbjct: 416 RSELDGNKRK---KTIVASIVSLTL--FMILGFTAFGVWRCR------------------ 452
Query: 499 GRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYK 558
+ H+ K L+ +D G++ +FD +I AT+ FS +NKLG+GG+G VYK
Sbjct: 453 ---VEHIAHISKDAWKNDLKPQDVPGLD--FFDMHTIQNATNNFSLSNKLGQGGFGSVYK 507
Query: 559 GKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYM 618
GKLQ G+EIAVKRLSS S QG +EF NE+VLI+KLQHRNLVR+ G CI+ +EK+LIYE+M
Sbjct: 508 GKLQDGKEIAVKRLSSSSGQGKEEFMNEIVLISKLQHRNLVRVLGCCIEEEEKLLIYEFM 567
Query: 619 PNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGE 678
NKSLD F+FD K +DW RFDI+ GIARGLLYLH DSRLRVIHRDLK SNILLD +
Sbjct: 568 VNKSLDTFLFDSRKRLEIDWPKRFDIIQGIARGLLYLHHDSRLRVIHRDLKVSNILLDEK 627
Query: 679 MQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEII 738
M PKISDFGLAR++ G E + NT+RVVGT GYMSPEYA G FS KSDI+SFGV++LEII
Sbjct: 628 MNPKISDFGLARMYQGTEYQDNTRRVVGTLGYMSPEYAWTGMFSEKSDIYSFGVLMLEII 687
Query: 739 SGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQD 798
SG+K + F +L+ YAW+ W+E + +DL+D L ++ + + RC +GLLCVQ
Sbjct: 688 SGEKISRFSYGVEGKTLIAYAWESWSEYRGIDLLDQDLADSCHPLEVGRCIQIGLLCVQH 747
Query: 799 EPDDRPNMSNVVIMLDSETATLPTPKQPT--FFTRKD--LSSTASSSLQFDSSIVEGR 852
+P DRPN ++ ML + T+ LP+PKQPT F TR D LS+ + S++ GR
Sbjct: 748 QPADRPNTLELLAML-TTTSDLPSPKQPTFAFHTRDDESLSNDLITVNGMTQSVILGR 804
>At1g61430 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 806
Score = 484 bits (1245), Expect = e-136
Identities = 334/865 (38%), Positives = 462/865 (52%), Gaps = 112/865 (12%)
Query: 24 FSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGS 83
F FAG T I G L S+ +ELGFFS + + +YLGIW+
Sbjct: 18 FMSFSFAGITKESPFSI---GQTLSSSNGVYELGFFS----LNNSQNQYLGIWF-----K 65
Query: 84 GLSP--VVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSV 141
+ P VVWVANR+ PV D + + I+ +G+L++ + WS ++ S+ +
Sbjct: 66 SIIPQVVVWVANREKPVTDSAANL-GISSNGSLLLSNGKHGVVWSTGDIFASNGSR---A 121
Query: 142 KLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLE------LTCWKSLSDPGRG 195
+L D GNLV +D+ G LW+SFEH +T LP M L LT WKS +DP G
Sbjct: 122 ELTDHGNLVFIDKVSGRTLWQSFEHLGNTLLPTSIMMYNLVAGEKRGLTAWKSYTDPSPG 181
Query: 196 NFTFKMDKKWENRFAILNQGQLYWQ-----------SEEQGDGVMNPESNPDDISNDVYN 244
F + + ++ I+ Y++ S + + +P D++ Y
Sbjct: 182 EFVALITPQVPSQGIIMRGSTRYYRTGPWAKTRFTGSPQMDESYTSPFILTQDVNGSGYF 241
Query: 245 LLTNFKELKNKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDIVWWYQPRTTCLTYNVCGNF 304
V +R++L S G +KVL + + P +C Y VCG F
Sbjct: 242 SF----------VERGKPSRMILTSEGTMKVLVHNGMDWESTY-EGPANSCDIYGVCGPF 290
Query: 305 SSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKS------TSCGANTNTFLNL 358
C C C GF + ++ G TS C R++ S G + N F
Sbjct: 291 GLCVVSIPPKCKCFKGFVPKFA-KEWKKGNWTSG--CVRRTELHCQGNSSGKDANVFY-- 345
Query: 359 TMMKIGSPDIKVSAQDEN--ECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQ 416
T+ I PD A +N EC C+ CS C A SY+P G+ C +W++
Sbjct: 346 TVPNIKPPDFYEYANSQNAEECHQNCLHNCS---CLAFSYIP------GIG---CLMWSK 393
Query: 417 NLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGN-PKSTLSLILGIALPGVVILACICIL 475
+L ++ G+ L +R+A+S+++ RK ST+SL L VI
Sbjct: 394 DLMDTRQFSAAGE--LLSIRLARSELDVNKRKMTIVASTVSLTL------FVIFGFAAFG 445
Query: 476 AYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESI 535
+ CR + H+ + + L+ +D G+E +F+ +I
Sbjct: 446 FWRCR---------------------VEHNAHISNDAWRNFLQSQDVPGLE--FFEMNAI 482
Query: 536 LVATDYFSDANKLGRGGYGPVYK---GKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAK 592
AT+ FS +NKLG GG+G VYK GKLQ GREIAVKRLSS S QG QEF NE+VLI+K
Sbjct: 483 QTATNNFSLSNKLGPGGFGSVYKARNGKLQDGREIAVKRLSSSSGQGKQEFMNEIVLISK 542
Query: 593 LQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGL 652
LQHRNLVR+ G C++G EK+LIY ++ NKSLD FVFD K LDW RF+I+ GIARGL
Sbjct: 543 LQHRNLVRVLGCCVEGTEKLLIYGFLKNKSLDTFVFDARKKLELDWPKRFEIIEGIARGL 602
Query: 653 LYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMS 712
LYLH+DSRLRVIHRDLK SNILLD +M PKISDFGLAR+F G + + T+RVVGT GYMS
Sbjct: 603 LYLHRDSRLRVIHRDLKVSNILLDEKMNPKISDFGLARMFQGTQYQEKTRRVVGTLGYMS 662
Query: 713 PEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLM 772
PEYA G FS KSDI+SFGV+LLEIISGKK + F + +LL YAW+ W E + ++ +
Sbjct: 663 PEYAWTGVFSEKSDIYSFGVLLLEIISGKKISSFSYGEEGKALLAYAWECWCETREVNFL 722
Query: 773 DLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTF--FT 830
D +L ++ + ++ RC +GLLCVQ EP DRPN ++ ML + T+ LP PK+PTF T
Sbjct: 723 DQALADSSHPSEVGRCVQIGLLCVQHEPADRPNTLELLSML-TTTSDLPLPKKPTFVVHT 781
Query: 831 RKDLSSTASSSL---QFDSSIVEGR 852
RKD S + S + + S+++GR
Sbjct: 782 RKDESPSNDSMITVNEMTESVIQGR 806
>At1g11280 serine/threonine kinase like protein
Length = 830
Score = 480 bits (1235), Expect = e-135
Identities = 327/862 (37%), Positives = 471/862 (53%), Gaps = 93/862 (10%)
Query: 10 ITIFLFHMHCWLLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGK 69
I I LF WL F +A T++ + G L S +ELGFFSP+ +
Sbjct: 18 IGIVLFPWFLWLSLFLSCGYAAITISSPLTL---GQTLSSPGGFYELGFFSPN----NSQ 70
Query: 70 GRYLGIWYYREEGSGLSP--VVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSA 127
+Y+GIW+ + ++P VVWVANR+ P+ + I+ +G+L++LD+S WS
Sbjct: 71 NQYVGIWFKK-----ITPRVVVWVANREKPITTP-VANLTISRNGSLILLDSSKNVVWS- 123
Query: 128 SNLTNSSSATNRS-VKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLE---- 182
T S +N+ KL+D+GNLV++D+ LW+SFE+P DT LP + L
Sbjct: 124 ---TRRPSISNKCHAKLLDTGNLVIVDDVSENLLWQSFENPGDTMLPYSSLMYNLATGEK 180
Query: 183 --LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEE------QGDGVMNPE-S 233
L+ WKS +DP G+F ++ + + + +Y +S G +M+ +
Sbjct: 181 RVLSSWKSHTDPSPGDFVVRLTPQVPAQIVTMRGSSVYKRSGPWAKTGFTGVPLMDESYT 240
Query: 234 NPDDISNDVYNLLTNFKELKNKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDIVWWYQPRT 293
+P +S DV N F L+ S + TR+++ S G +K +R N ++ + P
Sbjct: 241 SPFSLSQDVGNGTGLFSYLQR----SSELTRVIITSEGYLKT-FRYNGTGWVLDFITPAN 295
Query: 294 TCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKSTSCGANTN 353
C Y CG F C N C C+ GF + ++ G TS + R SC AN +
Sbjct: 296 LCDLYGACGPFGLCVTSNPTKCKCMKGFVPKYK-EEWKRGNMTSGCM-RRTELSCQANLS 353
Query: 354 T---------FLNLTMMKIGSPDIKVSAQ--DENECKFRCISMCSQTQCQACSYVPIPVQ 402
T F L +K PD+ A D ++C C+S CS C A +Y+
Sbjct: 354 TKTQGKGVDVFYRLANVK--PPDLYEYASFVDADQCHQGCLSNCS---CSAFAYIT---- 404
Query: 403 QRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIA 462
G+ C +W L +GG+ L +R+A S++ G+ ++ +
Sbjct: 405 --GIG---CLLWNHELIDTIRYSVGGEF--LSIRLASSELA-----GSRRTKI------- 445
Query: 463 LPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDN 522
+ G + L+ ILA+ + + KQ DS K GLE ++
Sbjct: 446 IVGSISLSIFVILAFGSYKYWRYRAKQNVGPTWAFFNNSQDSW--------KNGLEPQEI 497
Query: 523 EGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQE 582
G+ +F+ +I AT+ F+ +NKLG+GG+GPVYKG L ++IAVKRLSS S QG +E
Sbjct: 498 SGLT--FFEMNTIRAATNNFNVSNKLGQGGFGPVYKGTLSDKKDIAVKRLSSSSGQGTEE 555
Query: 583 FKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRF 642
F NE+ LI+KLQHRNLVRL G CI G+EK+LIYE++ NKSLD F+FD T +DW RF
Sbjct: 556 FMNEIKLISKLQHRNLVRLLGCCIDGEEKLLIYEFLVNKSLDTFLFDLTLKLQIDWPKRF 615
Query: 643 DILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQ 702
+I+ G++RGLLYLH+DS +RVIHRDLK SNILLD +M PKISDFGLAR+F G + + NT+
Sbjct: 616 NIIQGVSRGLLYLHRDSCMRVIHRDLKVSNILLDDKMNPKISDFGLARMFQGTQHQDNTR 675
Query: 703 RVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKL 762
+VVGT GYMSPEYA G FS KSDI++FGV+LLEIISGKK + F + +LLG+AW+
Sbjct: 676 KVVGTLGYMSPEYAWTGMFSEKSDIYAFGVLLLEIISGKKISSFCCGEEGKTLLGHAWEC 735
Query: 763 WTENKLLDLMDLSLGEAYN--ANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATL 820
W E +DL+D + + + + RC +GLLC+Q + DRPN++ VV M+ S T L
Sbjct: 736 WLETGGVDLLDEDISSSCSPVEVEVARCVQIGLLCIQQQAVDRPNIAQVVTMMTSAT-DL 794
Query: 821 PTPKQPTFFTR-KDLSSTASSS 841
P PKQP F + +D S S S
Sbjct: 795 PRPKQPLFALQIQDQESVVSVS 816
>At1g61440 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 792
Score = 479 bits (1234), Expect = e-135
Identities = 330/851 (38%), Positives = 457/851 (52%), Gaps = 126/851 (14%)
Query: 21 LLCFSQLCFAGDT----LNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIW 76
LL F +A T L++GQ ++ + V +ELGFFS + +Y+GIW
Sbjct: 8 LLLFISFSYAEITKESPLSIGQTLSSSNGV-------YELGFFS----FNNSQNQYVGIW 56
Query: 77 YYREEGSGLSP--VVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSS 134
+ G+ P VVWVANR+ PV D + + I+ G+L++++ WS ++ S
Sbjct: 57 F-----KGIIPRVVVWVANREKPVTDSAANLV-ISSSGSLLLINGKHDVVWSTGEISASK 110
Query: 135 SATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLE------LTCWKS 188
+ +L D GNL++ D G LWESFEH +T LP M L L+ WKS
Sbjct: 111 GS---HAELSDYGNLMVKDNVTGRTLWESFEHLGNTLLPLSTMMYNLVTGEKRGLSSWKS 167
Query: 189 LSDPGRGNFTFKMDKKWENRFAILNQGQLYWQS-----------EEQGDGVMNPESNPDD 237
+DP G+F ++ + ++ ++ Y+++ + + +P S D
Sbjct: 168 YTDPSPGDFWVQITPQVPSQGFVMRGSTPYYRTGPWAKTRYTGIPQMDESYTSPFSLHQD 227
Query: 238 ISNDVYNLLTNFKELKNKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDIVWWYQPRTTCLT 297
++ Y + F+ Y +R++L S G +KVL R N + P +C
Sbjct: 228 VNGSGY--FSYFER-------DYKLSRIMLTSEGSMKVL-RYNGLDWKSSYEGPANSCDI 277
Query: 298 YNVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKS------TSCGAN 351
Y VCG F C + C C GF +S + ++ G TS C R++ S G +
Sbjct: 278 YGVCGPFGFCVISDPPKCKCFKGFVPKS-IEEWKRGNWTSG--CARRTELHCQGNSTGKD 334
Query: 352 TNTFLNLTMMKIGSPDIKVSAQ--DENECKFRCISMCSQTQCQACSYVPIPVQQRGLNLS 409
N F T+ I PD A D C C+ CS C A +Y+P G+
Sbjct: 335 ANVFH--TVPNIKPPDFYEYANSVDAEGCYQSCLHNCS---CLAFAYIP------GIG-- 381
Query: 410 PCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRK-----GNPKSTLSLILGIALP 464
C +W+++L + GG+ L +R+A S+++ RK TL +ILG A
Sbjct: 382 -CLMWSKDLMDTMQFSAGGEI--LSIRLAHSELDVHKRKMTIVASTVSLTLFVILGFATF 438
Query: 465 GVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEG 524
G R R + +D+ R+ L+ +D G
Sbjct: 439 G----------------------------FWRNRVKHHDAWRN--------DLQSQDVPG 462
Query: 525 IEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFK 584
+E +F+ +I AT FS +NKLG GG+G VYKGKLQ GREIAVKRLSS S QG QEF
Sbjct: 463 LE--FFEMNTIQTATSNFSLSNKLGHGGFGSVYKGKLQDGREIAVKRLSSSSEQGKQEFM 520
Query: 585 NEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDI 644
NE+VLI+KLQHRNLVR+ G C++G EK+LIYE+M NKSLD FVF K LDW RFDI
Sbjct: 521 NEIVLISKLQHRNLVRVLGCCVEGKEKLLIYEFMKNKSLDTFVFGSRKRLELDWPKRFDI 580
Query: 645 LLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRV 704
+ GI RGLLYLH+DSRLRVIHRDLK SNILLD +M PKISDFGLAR+F G + + T+RV
Sbjct: 581 IQGIVRGLLYLHRDSRLRVIHRDLKVSNILLDEKMNPKISDFGLARLFQGSQYQDKTRRV 640
Query: 705 VGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWT 764
VGT GYMSPEYA G FS KSDI+SFGV+LLEIISG+K + F + +LL Y W+ W
Sbjct: 641 VGTLGYMSPEYAWTGVFSEKSDIYSFGVLLLEIISGEKISRFSYGEEGKALLAYVWECWC 700
Query: 765 ENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPK 824
E + ++L+D +L ++ + + RC +GLLCVQ +P DRPN ++ ML + T+ LP PK
Sbjct: 701 ETRGVNLLDQALDDSSHPAEVGRCVQIGLLCVQHQPADRPNTLELLSML-TTTSDLPLPK 759
Query: 825 QPTF--FTRKD 833
QPTF TR D
Sbjct: 760 QPTFAVHTRND 770
>At1g61480 S-like receptor protein kinase; member of gene cluster;
similar to EST gb|T41816
Length = 809
Score = 478 bits (1230), Expect = e-135
Identities = 324/880 (36%), Positives = 471/880 (52%), Gaps = 105/880 (11%)
Query: 1 MFFTPFTNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFS 60
MFF +ITIFL +AG T I G L S+ +ELGFFS
Sbjct: 7 MFFASLL-LITIFL-----------SFSYAGITRESPLSI---GKTLSSSNGVYELGFFS 51
Query: 61 PDLNVTGGKGRYLGIWYYREEGSGLSP--VVWVANRDNPVADDSIGVFRIADDGNLVVLD 118
+ +Y+GIW+ G+ P VVWVANR+ PV D + + I+ +G+L++ +
Sbjct: 52 ----FNNSQNQYVGIWF-----KGIIPRVVVWVANREKPVTDSAANL-TISSNGSLLLFN 101
Query: 119 TSGIRYWSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMD 178
+ WS S+ + +L D+GNLV++D + G LWESFEH DT LP +
Sbjct: 102 ENHSVVWSIGETFASNGSR---AELTDNGNLVVIDNNSGRTLWESFEHFGDTMLPFSNLM 158
Query: 179 KTLE------LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPE 232
L LT WKS +DP G+FT ++ + ++ + + YW+S
Sbjct: 159 YNLATGEKRVLTSWKSHTDPSPGDFTVQITPQVPSQACTMRGSKTYWRSGPWAKTRFTGI 218
Query: 233 SNPDDISNDVYNLL--TNFKELKNKTVSSYDNTRLLLNSTGVIKVL------YRVNFQSD 284
DD ++L TN ++ + +++ S G +K+ + +NF++
Sbjct: 219 PVMDDTYTSPFSLQQDTNGSGSFTYFERNFKLSYIMITSEGSLKIFQHNGMDWELNFEA- 277
Query: 285 IVWWYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRK 344
P +C Y CG F C C C GF +S + ++ G T C R
Sbjct: 278 ------PENSCDIYGFCGPFGICVMSVPPKCKCFKGFVPKS-IEEWKRGNWTDG--CVRH 328
Query: 345 ST-SCGANTNT-----FLNLTMMKIGSPDIKVSAQ--DENECKFRCISMCSQTQCQACSY 396
+ C NTN F ++ +K PD A D C C+ CS C A +Y
Sbjct: 329 TELHCQGNTNGKTVNGFYHVANIK--PPDFYEFASFVDAEGCYQICLHNCS---CLAFAY 383
Query: 397 VPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLS 456
+ N C +W Q+L + GG+ L +R+A S++ R K ++
Sbjct: 384 I---------NGIGCLMWNQDLMDAVQFSAGGEI--LSIRLASSELGGNKRN---KIIVA 429
Query: 457 LILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEG 516
I+ ++L ++ A C L Y + ++ K+ + + K+ + +
Sbjct: 430 SIVSLSLFVILAFAAFCFLRYKVKHTVSAKISKIAS----------------KEAWNND- 472
Query: 517 LEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVS 576
LE +D G++ +F+ +I ATD FS +NKLG+GG+G VYKGKLQ G+EIAVKRLSS S
Sbjct: 473 LEPQDVSGLK--FFEMNTIQTATDNFSLSNKLGQGGFGSVYKGKLQDGKEIAVKRLSSSS 530
Query: 577 SQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALL 636
QG +EF NE+VLI+KLQH+NLVR+ G CI+G+E++L+YE++ NKSLD F+FD K +
Sbjct: 531 GQGKEEFMNEIVLISKLQHKNLVRILGCCIEGEERLLVYEFLLNKSLDTFLFDSRKRLEI 590
Query: 637 DWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKE 696
DW RF+I+ GIARGL YLH+DS LRVIHRDLK SNILLD +M PKISDFGLAR++ G E
Sbjct: 591 DWPKRFNIIEGIARGLHYLHRDSCLRVIHRDLKVSNILLDEKMNPKISDFGLARMYQGTE 650
Query: 697 TEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLL 756
+ NT+RV GT GYM+PEYA G FS KSDI+SFGV+LLEII+G+K + F + +LL
Sbjct: 651 YQDNTRRVAGTLGYMAPEYAWTGMFSEKSDIYSFGVILLEIITGEKISRFSYGRQGKTLL 710
Query: 757 GYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSE 816
YAW+ W E+ +DL+D + ++ + + RC +GLLCVQ +P DRPN ++ ML +
Sbjct: 711 AYAWESWCESGGIDLLDKDVADSCHPLEVERCVQIGLLCVQHQPADRPNTMELLSML-TT 769
Query: 817 TATLPTPKQPTFFTRKDLSSTASSSL----QFDSSIVEGR 852
T+ L +PKQPTF + S L + S++ GR
Sbjct: 770 TSDLTSPKQPTFVVHTRDEESLSQGLITVNEMTQSVILGR 809
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,676,848
Number of Sequences: 26719
Number of extensions: 969171
Number of successful extensions: 5690
Number of sequences better than 10.0: 1010
Number of HSP's better than 10.0 without gapping: 869
Number of HSP's successfully gapped in prelim test: 141
Number of HSP's that attempted gapping in prelim test: 2201
Number of HSP's gapped (non-prelim): 1243
length of query: 852
length of database: 11,318,596
effective HSP length: 108
effective length of query: 744
effective length of database: 8,432,944
effective search space: 6274110336
effective search space used: 6274110336
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0203.5


