

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.3
(475 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
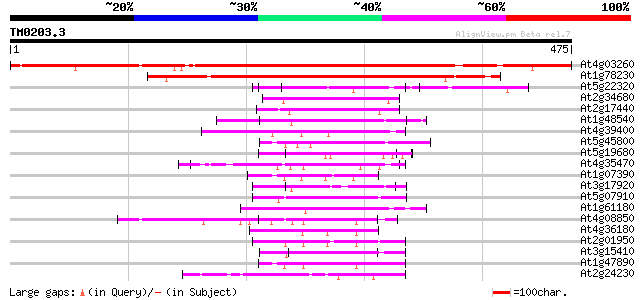
Sequences producing significant alignments: (bits) Value
At4g03260 protein phosphatase regulatory subunit like 468 e-132
At1g78230 unknown protein; similar to ESTs gb|H36831.1, gb|N9684... 242 4e-64
At5g22320 unknown protein 82 8e-16
At2g34680 auxin-induced protein (AIR9) 61 1e-09
At2g17440 unknown protein 55 1e-07
At1g48540 unknown protein 54 2e-07
At4g39400 brassinosteroid insensitive 1 gene (BRI1) 53 3e-07
At5g45800 receptor kinase-like protein 52 7e-07
At5g19680 unknown protein 52 9e-07
At4g35470 unknown protein 52 9e-07
At1g07390 disease resistance protein, putative 50 4e-06
At3g17920 unknown protein 49 5e-06
At5g07910 putative protein 49 8e-06
At1g61180 49 8e-06
At4g08850 receptor protein kinase like protein 48 1e-05
At4g36180 receptor protein kinase like protein 47 2e-05
At2g01950 putative receptor protein kinase 47 2e-05
At3g15410 unknown protein 46 5e-05
At1g47890 disease resistance protein, putative 45 7e-05
At2g24230 putative receptor-like protein kinase 45 9e-05
>At4g03260 protein phosphatase regulatory subunit like
Length = 677
Score = 468 bits (1204), Expect = e-132
Identities = 268/487 (55%), Positives = 352/487 (72%), Gaps = 27/487 (5%)
Query: 1 MKSRSLPNIKASTLSPENHAFKHPSSKSRSSDDLHALGMRKKEGFINESEDQIR--EGQE 58
++S SLPNI A + S ++ FK+ S SRSSDDLHAL R+ + ++E++++++ E Q+
Sbjct: 206 VRSNSLPNI-ADSSSEKSSPFKYSSHHSRSSDDLHALDTRQTDKSVHETDEEVKQEEDQD 264
Query: 59 REDNIGKTEDNHMGEYFDDGFD-SYLLSDLEKDWVMPTTDDISEVKTLQGDNSVDCFGEF 117
R+ ++ + DN+ +DG+D SY S L KDW++P TD++ K L+G+ + + EF
Sbjct: 265 RDYDMHNSGDNNKENLVEDGYDDSYDYSSLAKDWIVPPTDELKLSKFLEGETT-NQQAEF 323
Query: 118 PNKDFKVKRIEDWVVDLQHCG--PPVEEI----NELPESVDPVVDINTINGVTAAGVNHK 171
KD K KRIEDWV DLQH +EI +ELP +PVV +N +A K
Sbjct: 324 SGKDSKFKRIEDWVNDLQHVNLSEEADEITGYDDELPR--EPVV-LNEQATSSAKVDAIK 380
Query: 172 ITPGMEAAKRYISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALP 231
+TPGMEAAK+YISSL+A+A+ AQL +HGLVV+PFLSAFV L+VLNL+GN+IVRITAGALP
Sbjct: 381 LTPGMEAAKKYISSLSASATTAQLVSHGLVVIPFLSAFVGLRVLNLSGNAIVRITAGALP 440
Query: 232 RGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGE 291
RGLH+LNLS+N+IS IEGLRELTRLRVLDLSYNRILR+GHGLASCSSLKELYLAGNKI E
Sbjct: 441 RGLHALNLSKNSISVIEGLRELTRLRVLDLSYNRILRLGHGLASCSSLKELYLAGNKISE 500
Query: 292 VEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQ 351
+EGLHRLLKL++LDLRFNK ST KCLGQLAANY+SLQAI+L+GNP QKNVGDEQL+KYL
Sbjct: 501 IEGLHRLLKLTVLDLRFNKFSTTKCLGQLAANYSSLQAISLEGNPAQKNVGDEQLRKYLL 560
Query: 352 GLLPHLVYYNRQAMKVSTLKDGADRLVRLGTNDRSLRVDRK-TTRKGSHGVAAARRPPST 410
GLLP+LVYYNRQ K + L +L DR LR + K ++RK SHG +++ +P S+
Sbjct: 561 GLLPNLVYYNRQGTKDARLGTSTHQL------DRGLRSELKNSSRKSSHGASSSHKPGSS 614
Query: 411 STHSHRSQTVESPKLSKGKQSHLPPIRTKVS--TQSRHYLDAQSKVLNLTSGHSMRKSRS 468
+ + K SK ++S LPP+ K+S +++ ++ +L + SMR+SRS
Sbjct: 615 TAR----KAPALQKRSKERRSRLPPVGHKLSPAAYENYHVATGDRLSSLRTELSMRRSRS 670
Query: 469 EGTLGAL 475
EGTLG +
Sbjct: 671 EGTLGPI 677
>At1g78230 unknown protein; similar to ESTs gb|H36831.1,
gb|N96841.1, and gb|AA651293.1
Length = 413
Score = 242 bits (617), Expect = 4e-64
Identities = 141/306 (46%), Positives = 195/306 (63%), Gaps = 15/306 (4%)
Query: 117 FPNKDFKVKRIEDWV--VDLQHCGPPVEEINELPE-SVDPVVDINTINGVTAAGVNHKIT 173
F + +KR+++WV +D++ P E+ + L P + + + V +G ++
Sbjct: 90 FSAESSSMKRVDEWVRGLDVETVVPVNEDKDVLAIFPTSPNTERSPLGNVVQSG---NVS 146
Query: 174 PGMEAAKRYISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRG 233
+ A I SL+ ++S A +++ GL +P +S F SLK ++L+ N IV+IT +LP+G
Sbjct: 147 EAIVHANSLIQSLSKSSSVAHISSIGLKAIPSISHFTSLKSIDLSNNFIVQITPASLPKG 206
Query: 234 LHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVE 293
LH+LNLS+N IS IEGLR+LTRLRVLDLSYNRI RIG GL++C+ +KELYLAGNKI VE
Sbjct: 207 LHALNLSKNKISVIEGLRDLTRLRVLDLSYNRISRIGQGLSNCTLIKELYLAGNKISNVE 266
Query: 294 GLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQGL 353
GLHRLLKL +LDL FNKI+T K +GQL ANYNSL A+N+ GNP Q NVG++QL+K + L
Sbjct: 267 GLHRLLKLIVLDLSFNKIATTKAIGQLVANYNSLVALNILGNPIQNNVGEDQLRKTVSSL 326
Query: 354 LPHLVYYNRQAMKV----STLKDGADRLVRLGTNDRSLRVDRKTTRKGSHGVAAARRPPS 409
LP LVY+N+Q +K LKD R G D ++T+ K G A+ PS
Sbjct: 327 LPKLVYHNKQLIKPQRAREVLKDSVAR-AAFGGGDSLHHRRKRTSTKSVVGSAS----PS 381
Query: 410 TSTHSH 415
H
Sbjct: 382 VHHRGH 387
>At5g22320 unknown protein
Length = 452
Score = 81.6 bits (200), Expect = 8e-16
Identities = 75/244 (30%), Positives = 116/244 (46%), Gaps = 19/244 (7%)
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSI-EGLRELTRLRVLDLSYN 264
+S+ V+L+ L L N I I L + L+SL LSRN IS I + L +L L + LS
Sbjct: 103 ISSLVNLRALILNDNEISSICKLDLLKDLNSLVLSRNPISEIGDSLSKLKNLSKISLSDC 162
Query: 265 RILRIGHGLASCSSLKELYLAGNKI----GEVEGLHRLLKLSILDLRFNKISTAKCLGQL 320
RI IG L SCS LKEL LA N+I E+ RLL L + + ++S + LG L
Sbjct: 163 RIKAIGSSLKSCSDLKELRLANNEIKALPAELAVNKRLLNLDVGNNVITQLSGLEVLGTL 222
Query: 321 AANYNSLQAINLDGNPCQKNVGDEQLKKYLQGLLPHLVYYNRQAMKVSTLKDGADRLVRL 380
+ L+ +N+ GNP N D+ KK LLP + +N Q ++ S+ + +RL
Sbjct: 223 SC----LRNLNIRGNPISDN--DKSAKKVRTLLLPSVNVFNAQPLEKSSRN---AKHIRL 273
Query: 381 GTNDRSLRVDRKTTRKGSHGVAAARRPPSTSTHSHRSQTV-----ESPKLSKGKQSHLPP 435
T+D + + + +R S+ + + V +S K +++
Sbjct: 274 DTDDETFDAYHNKSAEEEQSKEDRKRKKSSKRNKSEEEEVNNEDHKSKKKKSKSNTNVDQ 333
Query: 436 IRTK 439
+ TK
Sbjct: 334 VETK 337
Score = 49.7 bits (117), Expect = 4e-06
Identities = 31/105 (29%), Positives = 54/105 (50%), Gaps = 5/105 (4%)
Query: 231 PRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIG 290
P + LNL ++ + L + L LDL +N + + GL SC +LK L + NK+
Sbjct: 18 PDSVKELNLGHKALTDVSCLSKFKNLEKLDLRFNNLTDL-QGLKSCVNLKWLSVVENKLQ 76
Query: 291 EVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGN 335
+ G+ L KL++L+ NK+ + + L +L+A+ L+ N
Sbjct: 77 SLNGIEALTKLTVLNAGKNKLKSMNEISSLV----NLRALILNDN 117
Score = 47.4 bits (111), Expect = 2e-05
Identities = 38/137 (27%), Positives = 69/137 (49%), Gaps = 6/137 (4%)
Query: 211 SLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIG 270
S+K LNL ++ ++ + + L L+L NN++ ++GL+ L+ L + N++ +
Sbjct: 20 SVKELNLGHKALTDVSCLSKFKNLEKLDLRFNNLTDLQGLKSCVNLKWLSVVENKLQSL- 78
Query: 271 HGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAI 330
+G+ + + L L NK+ + + L+ L L L N+IS+ C L + NSL
Sbjct: 79 NGIEALTKLTVLNAGKNKLKSMNEISSLVNLRALILNDNEISSI-CKLDLLKDLNSLV-- 135
Query: 331 NLDGNPCQKNVGDEQLK 347
L NP + +GD K
Sbjct: 136 -LSRNPISE-IGDSLSK 150
>At2g34680 auxin-induced protein (AIR9)
Length = 1661
Score = 61.2 bits (147), Expect = 1e-09
Identities = 43/124 (34%), Positives = 67/124 (53%), Gaps = 8/124 (6%)
Query: 215 LNLAGNSIVRITAGAL--PRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIG-H 271
L+L G+ I +T+G L L + L N +S++EG+ L R++VLDLS+N G
Sbjct: 273 LDLRGHRIRSLTSGGLHLSPNLEFVYLRDNLLSTLEGIEILNRVKVLDLSFNDFKGPGFE 332
Query: 272 GLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQ-----LAANYNS 326
L +C L++LYLAGN+I + L +L L L + NK+ + Q LAA+ N
Sbjct: 333 PLENCKMLQQLYLAGNQITSLASLPQLPNLEFLSVAQNKLKSLAMASQPRLQVLAASKNK 392
Query: 327 LQAI 330
+ +
Sbjct: 393 ITTL 396
Score = 30.8 bits (68), Expect = 1.7
Identities = 21/77 (27%), Positives = 39/77 (50%), Gaps = 1/77 (1%)
Query: 195 LANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELT 254
LA + + + L +L+ L++A N + + + PR L L S+N I++++ L
Sbjct: 345 LAGNQITSLASLPQLPNLEFLSVAQNKLKSLAMASQPR-LQVLAASKNKITTLKDFPYLP 403
Query: 255 RLRVLDLSYNRILRIGH 271
L L + N +L+I H
Sbjct: 404 VLEHLRVEENPLLKISH 420
>At2g17440 unknown protein
Length = 526
Score = 54.7 bits (130), Expect = 1e-07
Identities = 41/132 (31%), Positives = 74/132 (56%), Gaps = 12/132 (9%)
Query: 210 VSLKVLNLAGNSIVRITAGALPRGLH--SLNLSRNNISSI-EGLRELTRLRVLDLSYNRI 266
++L LNL+GN + + + + R +H L+LS N++S + E + L L+ LD+ N I
Sbjct: 276 LNLVNLNLSGNQLSSLPS-SFNRLIHLEELDLSSNSLSILPESIGSLVSLKKLDVETNNI 334
Query: 267 LRIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKI-------STAKCLG 318
I H ++ CSS++EL N++ + E + +L L IL +R+N I S+ L
Sbjct: 335 EEIPHSISGCSSMEELRADYNRLKALPEAVGKLSTLEILTVRYNNIRQLPTTMSSMANLK 394
Query: 319 QLAANYNSLQAI 330
+L ++N L+++
Sbjct: 395 ELDVSFNELESV 406
Score = 38.5 bits (88), Expect = 0.008
Identities = 34/104 (32%), Positives = 55/104 (52%), Gaps = 11/104 (10%)
Query: 237 LNLSRNNI----SSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV 292
L+LS N I ++I GL LTRL DL NRI ++ + +L L L+GN++ +
Sbjct: 235 LDLSENCIMVLPATIGGLISLTRL---DLHSNRIGQLPESIGDLLNLVNLNLSGNQLSSL 291
Query: 293 -EGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGN 335
+RL+ L LDL N +S L + + SL+ ++++ N
Sbjct: 292 PSSFNRLIHLEELDLSSNSLS---ILPESIGSLVSLKKLDVETN 332
Score = 35.0 bits (79), Expect = 0.090
Identities = 33/107 (30%), Positives = 55/107 (50%), Gaps = 10/107 (9%)
Query: 212 LKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIE----GLRELTRLRVLDLSYNRIL 267
L ++ LA S++ ++A + LNL + +E L +L+ L LDLS N I+
Sbjct: 189 LSLIKLA--SLIEVSA---KKATQELNLQHRLMDQLEWLPDSLGKLSSLVRLDLSENCIM 243
Query: 268 RIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKIST 313
+ + SL L L N+IG++ E + LL L L+L N++S+
Sbjct: 244 VLPATIGGLISLTRLDLHSNRIGQLPESIGDLLNLVNLNLSGNQLSS 290
Score = 33.5 bits (75), Expect = 0.26
Identities = 41/180 (22%), Positives = 82/180 (44%), Gaps = 18/180 (10%)
Query: 131 VVDLQHCGPPVEEINELPESVDPVVDINTINGVT--AAGVNHKIT--PGMEAAKRYISSL 186
++ L+ ++ LPES+ +V + ++ T + H I+ ME + + L
Sbjct: 298 LIHLEELDLSSNSLSILPESIGSLVSLKKLDVETNNIEEIPHSISGCSSMEELRADYNRL 357
Query: 187 TANASAA-QLANHGLVVVPF---------LSAFVSLKVLNLAGNSIVRITAG-ALPRGLH 235
A A +L+ ++ V + +S+ +LK L+++ N + + + L
Sbjct: 358 KALPEAVGKLSTLEILTVRYNNIRQLPTTMSSMANLKELDVSFNELESVPESLCYAKTLV 417
Query: 236 SLNLSRN--NISSIEGL-RELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV 292
LN+ N N+ S+ GL L +L LD+S N+I + + + S+L+ L N + E+
Sbjct: 418 KLNIGNNFANLRSLPGLIGNLEKLEELDMSNNQIRFLPYSFKTLSNLRVLQTEQNPLEEL 477
>At1g48540 unknown protein
Length = 1063
Score = 53.5 bits (127), Expect = 2e-07
Identities = 38/126 (30%), Positives = 63/126 (49%), Gaps = 2/126 (1%)
Query: 212 LKVLNLAGNSIVRITAGA-LPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIG 270
L V++ A N +V + L SL+LSRN ++ LR T+L+ LDL +N + +
Sbjct: 170 LAVISCACNRLVLMDESLQLLPAAESLDLSRNKFVKVDNLRRCTKLKHLDLGFNHLRTVS 229
Query: 271 HGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAI 330
+ L +L L N + + G+ L L LD+ +N IS L + + + L+ +
Sbjct: 230 YLSQVSCHLVKLVLRNNALTTLRGIENLKSLQGLDVSYNIISNFSEL-EFLWSLSQLKEL 288
Query: 331 NLDGNP 336
L+GNP
Sbjct: 289 WLEGNP 294
Score = 49.3 bits (116), Expect = 5e-06
Identities = 54/192 (28%), Positives = 82/192 (42%), Gaps = 18/192 (9%)
Query: 176 MEAAKRYISSLTANASAAQLANHGLVVVPF-LSAFVSLKVLNLAGNSIVRITAGALPRGL 234
+E +R + LT+ + L + P L F SLKVL L + A L
Sbjct: 76 LEQLRRILRILTSLKVVSTLPSPARDPTPLSLFPFGSLKVLELRRCDLSTSPAKGLIELR 135
Query: 235 HSL-------------NLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKE 281
H+L ++ + I+ I E +L V+ + NR++ + L + +
Sbjct: 136 HTLEKIICHNSTDALRHVFASRIAEITNSPEWNKLAVISCACNRLVLMDESLQLLPAAES 195
Query: 282 LYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNV 341
L L+ NK +V+ L R KL LDL FN + T L Q++ + L + L N
Sbjct: 196 LDLSRNKFVKVDNLRRCTKLKHLDLGFNHLRTVSYLSQVSCH---LVKLVLRNNALTTLR 252
Query: 342 GDEQLKKYLQGL 353
G E LK LQGL
Sbjct: 253 GIENLKS-LQGL 263
Score = 41.2 bits (95), Expect = 0.001
Identities = 46/181 (25%), Positives = 79/181 (43%), Gaps = 22/181 (12%)
Query: 131 VVDLQHCG---PPVEEINELPESVDPVVDINTINGVTAAGVNH--KITPGMEAAKRYISS 185
V++L+ C P + + EL +++ ++ N+ + + + +IT E K + S
Sbjct: 115 VLELRRCDLSTSPAKGLIELRHTLEKIICHNSTDALRHVFASRIAEITNSPEWNKLAVIS 174
Query: 186 LTAN--------------ASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRIT-AGAL 230
N A + L+ + V V L LK L+L N + ++ +
Sbjct: 175 CACNRLVLMDESLQLLPAAESLDLSRNKFVKVDNLRRCTKLKHLDLGFNHLRTVSYLSQV 234
Query: 231 PRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGH--GLASCSSLKELYLAGNK 288
L L L N ++++ G+ L L+ LD+SYN I L S S LKEL+L GN
Sbjct: 235 SCHLVKLVLRNNALTTLRGIENLKSLQGLDVSYNIISNFSELEFLWSLSQLKELWLEGNP 294
Query: 289 I 289
+
Sbjct: 295 V 295
>At4g39400 brassinosteroid insensitive 1 gene (BRI1)
Length = 1196
Score = 53.1 bits (126), Expect = 3e-07
Identities = 58/186 (31%), Positives = 84/186 (44%), Gaps = 17/186 (9%)
Query: 163 VTAAGVNHK-ITPGMEAAKRYISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNS 221
VT+ ++ K + G A + SLT S +H V SL L+L+ NS
Sbjct: 74 VTSIDLSSKPLNVGFSAVSSSLLSLTGLESLFLSNSHINGSVSGFKCSASLTSLDLSRNS 133
Query: 222 ----IVRITAGALPRGLHSLNLSRNNIS---SIEGLRELTRLRVLDLSYNRILR---IGH 271
+ +T+ GL LN+S N + + G +L L VLDLS N I +G
Sbjct: 134 LSGPVTTLTSLGSCSGLKFLNVSSNTLDFPGKVSGGLKLNSLEVLDLSANSISGANVVGW 193
Query: 272 GLAS-CSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTA-KCLGQLAANYNSLQA 329
L+ C LK L ++GNKI + R + L LD+ N ST LG +A LQ
Sbjct: 194 VLSDGCGELKHLAISGNKISGDVDVSRCVNLEFLDVSSNNFSTGIPFLGDCSA----LQH 249
Query: 330 INLDGN 335
+++ GN
Sbjct: 250 LDISGN 255
Score = 37.4 bits (85), Expect = 0.018
Identities = 29/84 (34%), Positives = 41/84 (48%), Gaps = 5/84 (5%)
Query: 211 SLKVLNLAGNSI--VRITAGALPRG---LHSLNLSRNNISSIEGLRELTRLRVLDLSYNR 265
SL+VL+L+ NSI + L G L L +S N IS + L LD+S N
Sbjct: 174 SLEVLDLSANSISGANVVGWVLSDGCGELKHLAISGNKISGDVDVSRCVNLEFLDVSSNN 233
Query: 266 ILRIGHGLASCSSLKELYLAGNKI 289
L CS+L+ L ++GNK+
Sbjct: 234 FSTGIPFLGDCSALQHLDISGNKL 257
>At5g45800 receptor kinase-like protein
Length = 666
Score = 52.0 bits (123), Expect = 7e-07
Identities = 55/179 (30%), Positives = 83/179 (45%), Gaps = 40/179 (22%)
Query: 212 LKVLNLAGNSIVRITAGALPR------GLHSLNLSRN-----------NISSIEGLREL- 253
L+VL+L+ NS+ G+LP GL S+NLSRN N S + ++EL
Sbjct: 82 LRVLDLSNNSL----DGSLPTWLWSMPGLVSVNLSRNRFGGSIRVIPVNGSVLSAVKELN 137
Query: 254 ---------------TRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKI-GEVEGLHR 297
T L LDLS+N + + GL S S L+ L ++ KI G V+ +
Sbjct: 138 LSFNRFKHAVNFTGFTNLTTLDLSHNSLGVLPLGLGSLSGLRHLDISRCKINGSVKPISG 197
Query: 298 LLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQGLLPH 356
L L LDL N ++ + + N N LQ +NL N +VG ++ +K+ + H
Sbjct: 198 LKSLDYLDLSENSMNGSFPVD--FPNLNHLQFLNLSANRFSGSVGFDKYRKFGKSAFLH 254
>At5g19680 unknown protein
Length = 328
Score = 51.6 bits (122), Expect = 9e-07
Identities = 46/173 (26%), Positives = 70/173 (39%), Gaps = 43/173 (24%)
Query: 211 SLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIG 270
+LK L ++ N + +I L L L N + +E L T+L L L NRI +
Sbjct: 133 TLKELYVSKNEVNKIMEIEHLHNLQILELGSNRLRVMENLENFTKLEELWLGRNRIKVVN 192
Query: 271 --------------------HGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNK 310
G C +L+ELYL+ N I ++EGL L+ L +LD+ NK
Sbjct: 193 LCGLKCIKKISLQSNRLTSMKGFEECVALEELYLSHNGISKMEGLSALVNLRVLDVSNNK 252
Query: 311 ISTAKCLGQLAA---------NYNSLQAIN--------------LDGNPCQKN 340
+++ + L SL+AI L+ NPC K+
Sbjct: 253 LTSVDDIQNLTKLEDLWLNDNQIESLEAITEAVTGSKEKLTTIYLENNPCAKS 305
Score = 51.2 bits (121), Expect = 1e-06
Identities = 32/114 (28%), Positives = 58/114 (50%), Gaps = 6/114 (5%)
Query: 234 LHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVE 293
L L L N ++ + + T+L V D+S+N I + + S+LKELY++ N++ ++
Sbjct: 89 LEELVLRDNKLAKVPDVSIFTKLLVYDISFNEITSLEGISKASSTLKELYVSKNEVNKIM 148
Query: 294 GLHRLLKLSILDLRFNKISTAK------CLGQLAANYNSLQAINLDGNPCQKNV 341
+ L L IL+L N++ + L +L N ++ +NL G C K +
Sbjct: 149 EIEHLHNLQILELGSNRLRVMENLENFTKLEELWLGRNRIKVVNLCGLKCIKKI 202
Score = 42.7 bits (99), Expect = 4e-04
Identities = 34/123 (27%), Positives = 61/123 (48%), Gaps = 7/123 (5%)
Query: 211 SLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEG-LRELTRLRVLDLSYNRI--- 266
S VL+L + + LP L L+L+ N +S ++ + +L+ L+ L L N I
Sbjct: 16 SNNVLDLTSYQLHSLDTVELPPNLIELDLTANRLSGLDSRIAQLSTLKKLSLRQNLIDDS 75
Query: 267 --LRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANY 324
+ H A S L+EL L NK+ +V + KL + D+ FN+I++ + + + ++
Sbjct: 76 AVEPLSHWDA-LSDLEELVLRDNKLAKVPDVSIFTKLLVYDISFNEITSLEGISKASSTL 134
Query: 325 NSL 327
L
Sbjct: 135 KEL 137
Score = 31.6 bits (70), Expect = 1.00
Identities = 18/57 (31%), Positives = 35/57 (60%)
Query: 210 VSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRI 266
V+L+ L L+ N I ++ + L L++S N ++S++ ++ LT+L L L+ N+I
Sbjct: 219 VALEELYLSHNGISKMEGLSALVNLRVLDVSNNKLTSVDDIQNLTKLEDLWLNDNQI 275
>At4g35470 unknown protein
Length = 549
Score = 51.6 bits (122), Expect = 9e-07
Identities = 51/215 (23%), Positives = 89/215 (40%), Gaps = 43/215 (20%)
Query: 154 VVDINTINGVTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGLVVVPFLSAFVSLK 213
+++++ + +K+T +E + L++ S NH +V+ + SL
Sbjct: 213 LIEVSAKKATQEINLQNKLTEQLEWLPDSLGKLSSLTSLDLSENHIVVLPNTIGGLSSLT 272
Query: 214 VLNLAGNSIVRITAGALPRGLHS------LNLSRNNISSI-------------------- 247
L+L N I G LP + LNL N +SS+
Sbjct: 273 KLDLHSNRI-----GQLPESIGELLNLVYLNLGSNQLSSLPSAFSRLVRLEELDLSCNNL 327
Query: 248 ----EGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLS 302
E + L L+ LD+ N I I + + CSSL EL NK+ + E + ++ L
Sbjct: 328 PILPESIGSLVSLKKLDVETNDIEEIPYSIGGCSSLIELRADYNKLKALPEAIGKITTLE 387
Query: 303 ILDLRFNKI-------STAKCLGQLAANYNSLQAI 330
IL +R+N I S+ L +L ++N L+++
Sbjct: 388 ILSVRYNNIRQLPTTMSSLASLKELDVSFNELESV 422
Score = 42.0 bits (97), Expect = 7e-04
Identities = 51/200 (25%), Positives = 99/200 (49%), Gaps = 23/200 (11%)
Query: 144 INELPESVDPVVDINTINGVTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGLVVV 203
I +LPES+ ++++ +N G N +++ A R + L+ + L ++
Sbjct: 281 IGQLPESIGELLNLVYLN----LGSN-QLSSLPSAFSRLV-----RLEELDLSCNNLPIL 330
Query: 204 P-FLSAFVSLKVLNLAGNSIVRI--TAGALPRGLHSLNLSRNNISSI-EGLRELTRLRVL 259
P + + VSLK L++ N I I + G L L N + ++ E + ++T L +L
Sbjct: 331 PESIGSLVSLKKLDVETNDIEEIPYSIGGCS-SLIELRADYNKLKALPEAIGKITTLEIL 389
Query: 260 DLSYNRILRIGHGLASCSSLKELYLAGNKIGEV-EGL---HRLLKLSILDLRFNKISTAK 315
+ YN I ++ ++S +SLKEL ++ N++ V E L L+KL+I + + +S +
Sbjct: 390 SVRYNNIRQLPTTMSSLASLKELDVSFNELESVPESLCFATTLVKLNIGNNFADMVSLPR 449
Query: 316 CLGQLAANYNSLQAINLDGN 335
+G N L+ +++ N
Sbjct: 450 SIG----NLEMLEELDISNN 465
Score = 40.8 bits (94), Expect = 0.002
Identities = 33/107 (30%), Positives = 57/107 (52%), Gaps = 10/107 (9%)
Query: 212 LKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIE----GLRELTRLRVLDLSYNRIL 267
L ++ LA S++ ++A + +NL +E L +L+ L LDLS N I+
Sbjct: 205 LSLIKLA--SLIEVSA---KKATQEINLQNKLTEQLEWLPDSLGKLSSLTSLDLSENHIV 259
Query: 268 RIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKIST 313
+ + + SSL +L L N+IG++ E + LL L L+L N++S+
Sbjct: 260 VLPNTIGGLSSLTKLDLHSNRIGQLPESIGELLNLVYLNLGSNQLSS 306
Score = 28.5 bits (62), Expect = 8.4
Identities = 31/94 (32%), Positives = 45/94 (46%), Gaps = 10/94 (10%)
Query: 293 EGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQG 352
+ L +L L+ LDL N I L +SL ++L N +G QL + +
Sbjct: 240 DSLGKLSSLTSLDLSENHIVV---LPNTIGGLSSLTKLDLHSN----RIG--QLPESIGE 290
Query: 353 LLPHLVYYNRQAMKVSTLKDGADRLVRLGTNDRS 386
LL +LVY N + ++S+L RLVRL D S
Sbjct: 291 LL-NLVYLNLGSNQLSSLPSAFSRLVRLEELDLS 323
>At1g07390 disease resistance protein, putative
Length = 976
Score = 49.7 bits (117), Expect = 4e-06
Identities = 44/125 (35%), Positives = 63/125 (50%), Gaps = 19/125 (15%)
Query: 202 VVPFLSAFVSLKVLNLAGNSIVRITAGALP-------RGLHSLNLSRNNIS--SIEGLRE 252
+VPFL+A S++ L+L N + G P L LNL N+ S S +GL +
Sbjct: 127 IVPFLNAATSIRSLHLESNYM----EGVFPPQELSNMTNLRVLNLKDNSFSFLSSQGLTD 182
Query: 253 LTRLRVLDLSYNRI--LRIGHGLASCSSLKELYLAGNKI---GEVEGLHRLLKLSILDLR 307
L VLDLS+N + H L S + LK L L N + +++GL L +L +L LR
Sbjct: 183 FRDLEVLDLSFNGVNDSEASHSL-STAKLKTLDLNFNPLSDFSQLKGLESLQELQVLKLR 241
Query: 308 FNKIS 312
NK +
Sbjct: 242 GNKFN 246
Score = 40.4 bits (93), Expect = 0.002
Identities = 36/109 (33%), Positives = 59/109 (54%), Gaps = 10/109 (9%)
Query: 212 LKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRE-----LTRLRVLDLSYNRI 266
L+ + L+GNS+ ++ L GL L++S N I + ++E LRVL LS N++
Sbjct: 482 LQTILLSGNSLTKLQLPILVHGLQVLDISSNMI--YDSIQEDIGMVFPNLRVLKLSNNQL 539
Query: 267 L-RIGHGLASCSSLKELYLAGNKI-GEV-EGLHRLLKLSILDLRFNKIS 312
+I A+ + L L+L GN G + EGL + L++LD+ N+ S
Sbjct: 540 QGKIFSKHANLTGLVGLFLDGNNFTGSLEEGLLKSKNLTLLDISDNRFS 588
Score = 32.3 bits (72), Expect = 0.59
Identities = 23/67 (34%), Positives = 38/67 (56%), Gaps = 6/67 (8%)
Query: 249 GLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGE-----VEGLHRLLK-LS 302
G+ L +LR LDLS N + + + L + + L+ L L+ N++ V GL +L+ LS
Sbjct: 327 GICRLMKLRELDLSSNALTSLPYCLGNLTHLRTLDLSNNQLNGNLSSFVSGLPSVLEYLS 386
Query: 303 ILDLRFN 309
+LD F+
Sbjct: 387 LLDNNFD 393
>At3g17920 unknown protein
Length = 962
Score = 49.3 bits (116), Expect = 5e-06
Identities = 32/103 (31%), Positives = 54/103 (52%), Gaps = 7/103 (6%)
Query: 234 LHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVE 293
+ SL+LSRN + ++ LR +L+ LDL +N++ +I H L+ + N + +
Sbjct: 193 VESLDLSRNKFAKVDNLRRCNKLKHLDLGFNQLRKISH-LSEVGKFRN-----NALTTLR 246
Query: 294 GLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNP 336
G+ L L LD+ FN IS L + + + L + L+GNP
Sbjct: 247 GIENLKSLEGLDVSFNLISDFSEL-EFLGSLSFLTDLWLEGNP 288
Score = 42.0 bits (97), Expect = 7e-04
Identities = 34/134 (25%), Positives = 60/134 (44%), Gaps = 13/134 (9%)
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGLHSL-------------NLSRNNISSIEGLRE 252
L F LKVL L G + +A L H+L ++ + I+ I+ +
Sbjct: 107 LLPFSKLKVLELRGCDLSTSSAKGLLELRHTLEKLICHNSTDALRHVFASRIAEIKDSPQ 166
Query: 253 LTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKIS 312
+L + + NR++ + L +++ L L+ NK +V+ L R KL LDL FN++
Sbjct: 167 WNKLAFISCACNRLVLMDESLQLLPAVESLDLSRNKFAKVDNLRRCNKLKHLDLGFNQLR 226
Query: 313 TAKCLGQLAANYNS 326
L ++ N+
Sbjct: 227 KISHLSEVGKFRNN 240
Score = 32.0 bits (71), Expect = 0.76
Identities = 27/81 (33%), Positives = 41/81 (50%), Gaps = 9/81 (11%)
Query: 212 LKVLNLAGNSIVRITAGALPRGLHSLNLSRNN-ISSIEGLRELTRLRVLDLSYNRILRIG 270
LK L+L N + +I+ L + RNN ++++ G+ L L LD+S+N I
Sbjct: 215 LKHLDLGFNQLRKISH------LSEVGKFRNNALTTLRGIENLKSLEGLDVSFNLISDFS 268
Query: 271 --HGLASCSSLKELYLAGNKI 289
L S S L +L+L GN I
Sbjct: 269 ELEFLGSLSFLTDLWLEGNPI 289
>At5g07910 putative protein
Length = 268
Score = 48.5 bits (114), Expect = 8e-06
Identities = 44/135 (32%), Positives = 71/135 (52%), Gaps = 7/135 (5%)
Query: 206 LSAFVSLKVLNLAGNSIVRIT--AGALPRGLHSLNLSRNNISSI-EGLRELTRLRVLDLS 262
L SLKVL L GN I + G L R L L++SRN + + + + L L +L++S
Sbjct: 87 LGKLQSLKVLMLDGNRISCLPDELGQLVR-LEQLSISRNMLIYLPDTIGSLRNLLLLNVS 145
Query: 263 YNRILRIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKISTAKCLGQLA 321
NR+ + + SC+SL+E+ N + E+ L L++L L L N+++ + L
Sbjct: 146 NNRLKSLPESVGSCASLEEVQANDNVVEELPASLCNLIQLKSLSLDNNQVN--QIPDGLL 203
Query: 322 ANYNSLQAINLDGNP 336
+ SLQ ++L NP
Sbjct: 204 IHCKSLQNLSLHNNP 218
Score = 32.7 bits (73), Expect = 0.45
Identities = 30/121 (24%), Positives = 54/121 (43%), Gaps = 4/121 (3%)
Query: 216 NLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLAS 275
N AG S + G+ L S+ E + +R LDL++N+I + ++
Sbjct: 7 NTAGGSKANRISRWRSTGIVGLRDSKLKTFPDEVIEMERAVRTLDLTHNKIADVPGEISK 66
Query: 276 CSSLKELYLAGNKIGEVEG-LHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDG 334
+++ L +A N + + G L +L L +L L N+IS CL L+ +++
Sbjct: 67 LINMQRLLIADNLVERLPGNLGKLQSLKVLMLDGNRIS---CLPDELGQLVRLEQLSISR 123
Query: 335 N 335
N
Sbjct: 124 N 124
>At1g61180
Length = 889
Score = 48.5 bits (114), Expect = 8e-06
Identities = 49/163 (30%), Positives = 84/163 (51%), Gaps = 12/163 (7%)
Query: 196 ANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEG--LREL 253
A GL +P + + +++ ++L N I IT + L +L L N + ++ G +R +
Sbjct: 497 AGVGLHEIPKVKDWGAVRKMSLMDNDIEEITCESKCSELTTLFLQSNKLKNLPGAFIRYM 556
Query: 254 TRLRVLDLSYNR-ILRIGHGLASCSSLKELYLAGNKIGEVE-GLHRLLKLSILDLRF-NK 310
+L VLDLSYNR ++ ++ SL+ L L+ I + GL L KL+ LDL + ++
Sbjct: 557 QKLVVLDLSYNRDFNKLPEQISGLVSLQFLDLSNTSIEHMPIGLKELKKLTFLDLTYTDR 616
Query: 311 ISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQGL 353
+ + + +L SL+ + L G+ K GD + K LQ L
Sbjct: 617 LCSISGISRLL----SLRLLRLLGS---KVHGDASVLKELQQL 652
Score = 35.8 bits (81), Expect = 0.053
Identities = 32/100 (32%), Positives = 50/100 (50%), Gaps = 16/100 (16%)
Query: 234 LHSLNLSRNNISSIE-GLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKI--- 289
L L+LS +I + GL+EL +L LDL+Y L G++ SL+ L L G+K+
Sbjct: 583 LQFLDLSNTSIEHMPIGLKELKKLTFLDLTYTDRLCSISGISRLLSLRLLRLLGSKVHGD 642
Query: 290 ----GEVEGLHRLLKLSI--------LDLRFNKISTAKCL 317
E++ L L +L+I LD R K+ + C+
Sbjct: 643 ASVLKELQQLQNLQELAITVSAELISLDQRLAKLISNLCI 682
>At4g08850 receptor protein kinase like protein
Length = 1045
Score = 48.1 bits (113), Expect = 1e-05
Identities = 65/259 (25%), Positives = 117/259 (45%), Gaps = 32/259 (12%)
Query: 92 VMPTTDDISEVKTLQGDNSVDCFGEFPNKDFKVKRIEDWVVDLQHC-GPPVEEINELPES 150
+ P + +E+ LQ D + + G P+ + ++E+ +D H GP + + +
Sbjct: 374 IPPGIANSTELTVLQVDTN-NFTGFLPDTICRGGKLENLTLDDNHFEGPVPKSLRDCKSL 432
Query: 151 VDPVVDINTING--VTAAGVNHKITPGMEAAKRYISSLTANASAAQ------LANHGLV- 201
+ N+ +G A GV + + + L+AN +Q L+N+ +
Sbjct: 433 IRVRFKGNSFSGDISEAFGVYPTLNFIDLSNNNFHGQLSANWEQSQKLVAFILSNNSITG 492
Query: 202 -VVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLN------LSRNNISSI--EGLRE 252
+ P + L L+L+ N I G LP + ++N L+ N +S G+R
Sbjct: 493 AIPPEIWNMTQLSQLDLSSNRIT----GELPESISNINRISKLQLNGNRLSGKIPSGIRL 548
Query: 253 LTRLRVLDLSYNRI-LRIGHGLASCSSLKELYLAGNKIGEV--EGLHRLLKLSILDLRFN 309
LT L LDLS NR I L + L + L+ N + + EGL +L +L +LDL +N
Sbjct: 549 LTNLEYLDLSSNRFSSEIPPTLNNLPRLYYMNLSRNDLDQTIPEGLTKLSQLQMLDLSYN 608
Query: 310 KISTAKCLGQLAANYNSLQ 328
++ G++++ + SLQ
Sbjct: 609 QLD-----GEISSQFRSLQ 622
Score = 43.9 bits (102), Expect = 2e-04
Identities = 37/109 (33%), Positives = 64/109 (57%), Gaps = 9/109 (8%)
Query: 211 SLKVLNLAGN---SIVRITAGALPRGLHSLNLSRNNISSI--EGLRELTRLRVLDLSYNR 265
+L+ L+L+ N S + T LPR L+ +NLSRN++ EGL +L++L++LDLSYN+
Sbjct: 551 NLEYLDLSSNRFSSEIPPTLNNLPR-LYYMNLSRNDLDQTIPEGLTKLSQLQMLDLSYNQ 609
Query: 266 I-LRIGHGLASCSSLKELYLAGNKI-GEV-EGLHRLLKLSILDLRFNKI 311
+ I S +L+ L L+ N + G++ +L L+ +D+ N +
Sbjct: 610 LDGEISSQFRSLQNLERLDLSHNNLSGQIPPSFKDMLALTHVDVSHNNL 658
>At4g36180 receptor protein kinase like protein
Length = 1136
Score = 47.4 bits (111), Expect = 2e-05
Identities = 41/138 (29%), Positives = 60/138 (42%), Gaps = 29/138 (21%)
Query: 204 PFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISS--IEGLRELTRLRVLDL 261
P + SL+V N+AGN + LP L L++S N S GL LT+L++L+L
Sbjct: 134 PAMRNLTSLEVFNVAGNRLSGEIPVGLPSSLQFLDISSNTFSGQIPSGLANLTQLQLLNL 193
Query: 262 SYNRIL-------------------------RIGHGLASCSSLKELYLAGNKIGEV--EG 294
SYN++ + +++CSSL L + N+IG V
Sbjct: 194 SYNQLTGEIPASLGNLQSLQYLWLDFNLLQGTLPSAISNCSSLVHLSASENEIGGVIPAA 253
Query: 295 LHRLLKLSILDLRFNKIS 312
L KL +L L N S
Sbjct: 254 YGALPKLEVLSLSNNNFS 271
Score = 38.9 bits (89), Expect = 0.006
Identities = 47/152 (30%), Positives = 69/152 (44%), Gaps = 35/152 (23%)
Query: 192 AAQLANHGLVVVPFLSAFVSLKVLNLAGNSI---VRITAGALPRGLHSLNLSRNNISSIE 248
A Q N VV S+ VSL+ +NL+ NS + T G L R L SL+LS N+IS
Sbjct: 530 ALQGNNFSGVVPEGFSSLVSLRYVNLSSNSFSGEIPQTFGFL-RLLVSLSLSDNHISG-- 586
Query: 249 GLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKI-GEVEG-LHRLLKLSILDL 306
I + +CS+L+ L L N++ G + L RL +L +LDL
Sbjct: 587 -------------------SIPPEIGNCSALEVLELRSNRLMGHIPADLSRLPRLKVLDL 627
Query: 307 RFN--------KISTAKCLGQLAANYNSLQAI 330
N +IS + L L+ ++N L +
Sbjct: 628 GQNNLSGEIPPEISQSSSLNSLSLDHNHLSGV 659
>At2g01950 putative receptor protein kinase
Length = 1143
Score = 47.0 bits (110), Expect = 2e-05
Identities = 50/141 (35%), Positives = 67/141 (47%), Gaps = 15/141 (10%)
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPR--GLHSLNLSRNNISSI--EGLRELTRLRVLDL 261
LS+ VS+ L+ +GNSI + +L L SLNLS NN + EL L+ LDL
Sbjct: 200 LSSCVSMTYLDFSGNSISGYISDSLINCTNLKSLNLSYNNFDGQIPKSFGELKLLQSLDL 259
Query: 262 SYNRIL-----RIGHGLASCSSLKELYLAGNKIGEV--EGLHRLLKLSILDLRFNKISTA 314
S+NR+ IG +C SL+ L L+ N V E L L LDL N IS
Sbjct: 260 SHNRLTGWIPPEIGD---TCRSLQNLRLSYNNFTGVIPESLSSCSWLQSLDLSNNNIS-G 315
Query: 315 KCLGQLAANYNSLQAINLDGN 335
+ ++ SLQ + L N
Sbjct: 316 PFPNTILRSFGSLQILLLSNN 336
Score = 32.3 bits (72), Expect = 0.59
Identities = 40/145 (27%), Positives = 63/145 (42%), Gaps = 28/145 (19%)
Query: 210 VSLKVLNLAGNSIVRITAGALPRG-------LHSLNLSRNNISSI---EGLRELTRLRVL 259
++L L L+ + ++ G LP L S+ LS NN + + +L+ L
Sbjct: 127 LTLTHLELSSSGLI----GTLPENFFSKYSNLISITLSYNNFTGKLPNDLFLSSKKLQTL 182
Query: 260 DLSYNRILRIGHG----LASCSSLKELYLAGNKIGEV--EGLHRLLKLSILDLRFNKIST 313
DLSYN I G L+SC S+ L +GN I + L L L+L +N
Sbjct: 183 DLSYNNITGPISGLTIPLSSCVSMTYLDFSGNSISGYISDSLINCTNLKSLNLSYNNFD- 241
Query: 314 AKCLGQLAANYNS---LQAINLDGN 335
GQ+ ++ LQ+++L N
Sbjct: 242 ----GQIPKSFGELKLLQSLDLSHN 262
>At3g15410 unknown protein
Length = 562
Score = 45.8 bits (107), Expect = 5e-05
Identities = 32/101 (31%), Positives = 57/101 (55%), Gaps = 5/101 (4%)
Query: 237 LNLSRNNISSIEG-LRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV-EG 294
LN+S N +S + + ELT ++ LD+S+N I + + S SL +L + N++ E+ +
Sbjct: 73 LNVSHNKLSQLPAAIGELTAMKSLDVSFNSISELPEQIGSAISLVKLDCSSNRLKELPDS 132
Query: 295 LHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGN 335
+ R L LS L N+IS+ L + N + L ++++GN
Sbjct: 133 IGRCLDLSDLKATNNQISS---LPEDMVNCSKLSKLDVEGN 170
Score = 43.1 bits (100), Expect = 3e-04
Identities = 31/104 (29%), Positives = 52/104 (49%), Gaps = 4/104 (3%)
Query: 212 LKVLNLAGNSIVRITAGALPRG--LHSLNLSRNNISSI-EGLRELTRLRVLDLSYNRILR 268
L L++ GN + ++ + L LN +N + + + + L+RL LDL N+I
Sbjct: 162 LSKLDVEGNKLTALSENHIASWTMLAELNACKNMLGVLPQNIGSLSRLIRLDLHQNKISS 221
Query: 269 IGHGLASCSSLKELYLAGNKIGEVEG-LHRLLKLSILDLRFNKI 311
+ + CSSL E YL N + + + L +L LDLR N++
Sbjct: 222 VPPSIGGCSSLVEFYLGINSLSTLPAEIGDLSRLGTLDLRSNQL 265
Score = 38.5 bits (88), Expect = 0.008
Identities = 35/127 (27%), Positives = 61/127 (47%), Gaps = 7/127 (5%)
Query: 189 NASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLH-----SLNLSRNN 243
N +L N+ L +P L F + L + S+ ++ P+ H L LSR
Sbjct: 429 NLMCLKLDNNPLNQIP-LDGFQVVSGLQILDLSVNAVSFREHPKFCHLPQLRELYLSRIQ 487
Query: 244 ISSI-EGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLS 302
+S + E + L+ L +LDL+ N + I G+ + +SLK L ++ N I + LL+ +
Sbjct: 488 LSEVPEDILNLSNLIILDLNQNSLQSIPKGIKNMTSLKHLDISNNNISSLPPELGLLEPT 547
Query: 303 ILDLRFN 309
+ LR +
Sbjct: 548 LEVLRLD 554
Score = 31.6 bits (70), Expect = 1.00
Identities = 47/187 (25%), Positives = 79/187 (42%), Gaps = 21/187 (11%)
Query: 123 KVKRIEDWV---VDLQHCGPPVEEINELPESVDPVVDINTI----NGVTAAGVNH----K 171
++K + D + +DL +I+ LPE + ++ + N +TA NH
Sbjct: 125 RLKELPDSIGRCLDLSDLKATNNQISSLPEDMVNCSKLSKLDVEGNKLTALSENHIASWT 184
Query: 172 ITPGMEAAKRYISSLTAN-ASAAQLANHGL------VVVPFLSAFVSLKVLNLAGNSIVR 224
+ + A K + L N S ++L L V P + SL L NS+
Sbjct: 185 MLAELNACKNMLGVLPQNIGSLSRLIRLDLHQNKISSVPPSIGGCSSLVEFYLGINSLST 244
Query: 225 ITA--GALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKEL 282
+ A G L R L +L+L N + +L LDLS N + + L + ++L++L
Sbjct: 245 LPAEIGDLSR-LGTLDLRSNQLKEYPVGACKLKLSYLDLSNNSLTGLHPELGNMTTLRKL 303
Query: 283 YLAGNKI 289
L GN +
Sbjct: 304 VLVGNPL 310
Score = 30.8 bits (68), Expect = 1.7
Identities = 64/247 (25%), Positives = 100/247 (39%), Gaps = 43/247 (17%)
Query: 206 LSAFVSLKVLNLAGNSIVRITA--GALPRGLHSLNLSRNNISSI---------------- 247
L L VLN++ N + ++ A G L + SL++S N+IS +
Sbjct: 64 LKNLACLVVLNVSHNKLSQLPAAIGELT-AMKSLDVSFNSISELPEQIGSAISLVKLDCS 122
Query: 248 -EGLRELTRL--RVLDLS-----YNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLL 299
L+EL R LDLS N+I + + +CS L +L + GNK+ + H
Sbjct: 123 SNRLKELPDSIGRCLDLSDLKATNNQISSLPEDMVNCSKLSKLDVEGNKLTALSENHIAS 182
Query: 300 KLSILDLRFNKISTAKCLGQLAANYNSL-QAINLDGNPCQKNVGDEQLKKYLQGLLPHLV 358
+ +L K LG L N SL + I LD + + + + G LV
Sbjct: 183 WTMLAELNACK----NMLGVLPQNIGSLSRLIRLDLHQNKISSVPPSI-----GGCSSLV 233
Query: 359 YYNRQAMKVSTLKDGADRLVRLGTNDRSLRVDRKTTRKGSHGVAAARRPPSTSTHSHRSQ 418
+ +STL L RLGT +D ++ + + V A + S S+ S
Sbjct: 234 EFYLGINSLSTLPAEIGDLSRLGT------LDLRSNQLKEYPVGACKLKLSYLDLSNNSL 287
Query: 419 TVESPKL 425
T P+L
Sbjct: 288 TGLHPEL 294
>At1g47890 disease resistance protein, putative
Length = 1019
Score = 45.4 bits (106), Expect = 7e-05
Identities = 43/138 (31%), Positives = 71/138 (51%), Gaps = 17/138 (12%)
Query: 211 SLKVLNLAGNSIVRITAGALP-------RGLHSLNLSRNNISSI--EGLRELTRLRVLDL 261
SL++L+L+ N++ G+LP L L+L N++S E T+LR LD+
Sbjct: 636 SLEILDLSNNNL----NGSLPWCLETLMSSLSDLDLRNNSLSGSLPEIFMNATKLRSLDV 691
Query: 262 SYNRIL-RIGHGLASCSSLKELYLAGNKIGEV--EGLHRLLKLSILDLRFNKI-STAKCL 317
S+NR+ ++ L CSSL+ L + N+I ++ L+ L KL +L L NK T +
Sbjct: 692 SHNRMEGKLPGSLTGCSSLEVLNVGSNRINDMFPFELNSLQKLQVLVLHSNKFHGTLHNV 751
Query: 318 GQLAANYNSLQAINLDGN 335
+ + LQ I++ N
Sbjct: 752 DGVWFGFPQLQIIDVSHN 769
>At2g24230 putative receptor-like protein kinase
Length = 853
Score = 45.1 bits (105), Expect = 9e-05
Identities = 63/199 (31%), Positives = 94/199 (46%), Gaps = 19/199 (9%)
Query: 147 LPESVDPVVDINTINGVTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGLVVVPFL 206
+PE+VD +V + + + G I G+ + +S + S+ QL G + F
Sbjct: 155 IPEAVDSLVSLRVLK-LDHNGFQMSIPRGLLGCQSLVS---IDLSSNQL--EGSLPDGFG 208
Query: 207 SAFVSLKVLNLAGNSI-VRITAGALPRGLHSLNLSRNNI-SSIEGLRELTRLRVLDLSYN 264
SAF L+ L+LAGN I R T A + + LN+S N S+ G+ + T L V DLS N
Sbjct: 209 SAFPKLETLSLAGNKIHGRDTDFADMKSISFLNISGNQFDGSVTGVFKET-LEVADLSKN 267
Query: 265 RILRIGHGLASCS----SLKELYLAGNKI-GEVEGLHRLLKLSILDL---RFNKISTAKC 316
R GH + SL L L+ N++ G ++ L L KL L+L RFN+ +
Sbjct: 268 RFQ--GHISSQVDSNWFSLVYLDLSENELSGVIKNLTLLKKLKHLNLAWNRFNRGMFPRI 325
Query: 317 LGQLAANYNSLQAINLDGN 335
Y +L NL G+
Sbjct: 326 EMLSGLEYLNLSNTNLSGH 344
Score = 38.9 bits (89), Expect = 0.006
Identities = 34/117 (29%), Positives = 55/117 (46%), Gaps = 15/117 (12%)
Query: 252 ELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGE--VEGLHRLLKLSILDLRFN 309
+L++L+ LDLS N+I + S ++LK L L+ NKI + +L +LD+ +N
Sbjct: 90 KLSKLQSLDLSNNKISALPSDFWSLNTLKNLNLSFNKISGSFSSNVGNFGQLELLDISYN 149
Query: 310 KISTAKCLGQLAANYNSLQAINLDGN-----------PCQKNVGDEQLKKYLQGLLP 355
S A + + + SL+ + LD N CQ V + L+G LP
Sbjct: 150 NFSGA--IPEAVDSLVSLRVLKLDHNGFQMSIPRGLLGCQSLVSIDLSSNQLEGSLP 204
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.132 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,370,959
Number of Sequences: 26719
Number of extensions: 450814
Number of successful extensions: 1613
Number of sequences better than 10.0: 198
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 165
Number of HSP's that attempted gapping in prelim test: 1360
Number of HSP's gapped (non-prelim): 342
length of query: 475
length of database: 11,318,596
effective HSP length: 103
effective length of query: 372
effective length of database: 8,566,539
effective search space: 3186752508
effective search space used: 3186752508
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0203.3


