

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0201.10
(137 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
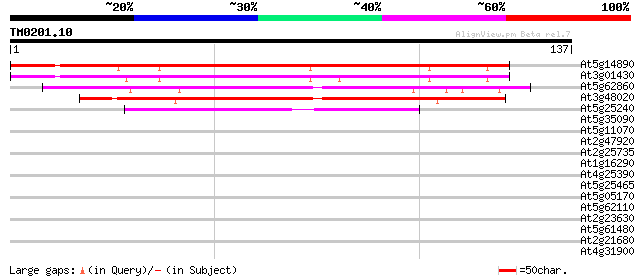
At5g14890 putative protein 126 3e-30
At3g01430 unknown protein 120 3e-28
At5g62860 unknown protein 99 7e-22
At3g48020 unknown protein 96 6e-21
At5g25240 unknown protein 51 2e-07
At5g35090 unknown protein 36 0.005
At5g11070 unknown protein 34 0.021
At2g47920 unknown protein (At2g47920) 28 1.1
At2g25735 unknown protein 28 1.5
At1g16290 hypothetical protein 28 1.9
At4g25390 receptor kinase-like protein 27 3.3
At5g25465 unknown protein 27 4.3
At5g05170 cellulose synthase catalytic subunit (Ath-B) 27 4.3
At5g62110 putative protein 26 5.6
At2g23630 putative pectinesterase 26 5.6
At5g61480 leucine-rich receptor-like protein kinase - like protein 26 7.4
At2g21680 hypothetical protein 26 7.4
At4g31900 putative protein 25 9.6
>At5g14890 putative protein
Length = 733
Score = 126 bits (317), Expect = 3e-30
Identities = 68/140 (48%), Positives = 87/140 (61%), Gaps = 19/140 (13%)
Query: 1 MQEVLFAKQACICYLIPCSSSSETS----LSWWERVRSP---ENKEWWWAHGWNKVREWS 53
M E +FAK+ C C+++PC SS+ S WW+R+R+ E E WW GW K+REWS
Sbjct: 579 MHEAMFAKRGC-CFILPCLGSSQPSGPNGSVWWQRIRTVDKLEPDERWWVSGWMKMREWS 637
Query: 54 EIIVGPKWKTFIRRFNRNN----NRAGAASYDKKGSFHYDSLSYALNFDDGE-----EDV 104
EI+ GPKWKTFIRRF RN+ G + + SF YDS SY+LNFDDG+ ED
Sbjct: 638 EIVAGPKWKTFIRRFGRNHCCNGGIDGGCNRPEHVSFRYDSWSYSLNFDDGKQTGHFEDE 697
Query: 105 YSYGGFSTRFA--SVPASTK 122
+ Y +S RFA S+P STK
Sbjct: 698 FPYRDYSMRFAAPSLPVSTK 717
>At3g01430 unknown protein
Length = 180
Score = 120 bits (300), Expect = 3e-28
Identities = 66/153 (43%), Positives = 88/153 (57%), Gaps = 32/153 (20%)
Query: 1 MQEVLFAKQACICYLIPCSSSSETSLS----WWERVRSP---ENKEWWWAHGWNKVREWS 53
M E LFAK+ C C+L+PC +SS+ S WW+R+ + E E WW GW ++REWS
Sbjct: 17 MHEALFAKRGC-CFLMPCLASSQPSTRGGSVWWQRITTVDKLEPDERWWIRGWRRMREWS 75
Query: 54 EIIVGPKWKTFIRRFNRNN------NRAGAAS-----------YDKKGSFHYDSLSYALN 96
E++ GP+WKT+IRRF R+N R G +S +G F YD LSY+LN
Sbjct: 76 ELVAGPRWKTYIRRFGRSNCCGGGGGRVGNSSGGCGGGAMPNRSSDQGKFRYDQLSYSLN 135
Query: 97 FDDGE-----EDVYSYGGFSTRFA--SVPASTK 122
FDDG +D + Y +S RFA S+P STK
Sbjct: 136 FDDGNQTGHFDDEFPYRDYSMRFAAPSLPVSTK 168
>At5g62860 unknown protein
Length = 377
Score = 99.0 bits (245), Expect = 7e-22
Identities = 60/137 (43%), Positives = 76/137 (54%), Gaps = 20/137 (14%)
Query: 9 QACICYLIPCSSSSETSLSW--WERVRSPENKEW---------WWAHGWNKVREWSEIIV 57
Q C C+ S S T++ + W R+R+ ++ WW K+REWSEI+
Sbjct: 228 QRCCCFPSFRRSRSSTAVGYSSWGRIRTVDDSNHSGDHGDEPRWWIRASLKIREWSEIVA 287
Query: 58 GPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNF-DDGEEDVY-SYGG---FST 112
GP+WKTFIRRFNR+ R +D F YD LSY+LNF DD EED Y GG FST
Sbjct: 288 GPRWKTFIRRFNRDPRR--GRDWDASEKFQYDPLSYSLNFDDDDEEDEYVGLGGLRSFST 345
Query: 113 RFASVP--ASTKPTFTP 127
RFASVP + P +P
Sbjct: 346 RFASVPVYSGKAPAISP 362
>At3g48020 unknown protein
Length = 135
Score = 95.9 bits (237), Expect = 6e-21
Identities = 48/111 (43%), Positives = 68/111 (61%), Gaps = 10/111 (9%)
Query: 18 CSSSSETSLSWWERVRSPENKE-WWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAG 76
C S++ S SWW+R+ ++E WW + K+REWSEI+ GP+WKTFIRRFNR+ R
Sbjct: 15 CCSTTVKS-SWWQRIHRNNHQEPRWWVRAFLKIREWSEIVAGPRWKTFIRRFNRDPRR-- 71
Query: 77 AASYDKKGSFHYDSLSYALNFDDGEED------VYSYGGFSTRFASVPAST 121
+D F YD +SY L+F+D ++D V FS R+ASVP ++
Sbjct: 72 GQDWDDSDKFRYDPVSYTLSFEDEDKDDDDEAGVGGVRSFSMRYASVPVAS 122
>At5g25240 unknown protein
Length = 131
Score = 50.8 bits (120), Expect = 2e-07
Identities = 27/72 (37%), Positives = 36/72 (49%), Gaps = 5/72 (6%)
Query: 29 WERVRSPENKEWWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAGAASYDKKGSFHY 88
W E + W + ++E SE I GPKWK FIR F+ +G + F Y
Sbjct: 46 WSGCLQEERRGNWGSEKLKGLKEISEKIAGPKWKNFIRSFS-----SGRKKMRRDVDFTY 100
Query: 89 DSLSYALNFDDG 100
D +Y+LNFDDG
Sbjct: 101 DLKNYSLNFDDG 112
>At5g35090 unknown protein
Length = 159
Score = 36.2 bits (82), Expect = 0.005
Identities = 19/40 (47%), Positives = 22/40 (54%), Gaps = 4/40 (10%)
Query: 86 FHYDSLSYALNFDDGEE----DVYSYGGFSTRFASVPAST 121
FHYD SYALNFD G+E D + FS R P S+
Sbjct: 105 FHYDPSSYALNFDKGDEDDNIDRFPLRNFSARLPHSPPSS 144
>At5g11070 unknown protein
Length = 152
Score = 34.3 bits (77), Expect = 0.021
Identities = 25/68 (36%), Positives = 32/68 (46%), Gaps = 9/68 (13%)
Query: 62 KTFIRRFNRNNNRA-------GAASYDKKGSFHYDSLSYALNFDDG--EEDVYSYGGFST 112
K+ I+ RNNN A S G F YD LSYALNF+D +D S+ F+
Sbjct: 73 KSRIKVTCRNNNCAYNNCVHHHHHSQSYPGDFSYDPLSYALNFEDNVRADDDGSFPNFTA 132
Query: 113 RFASVPAS 120
R P +
Sbjct: 133 RLPQSPVT 140
>At2g47920 unknown protein (At2g47920)
Length = 225
Score = 28.5 bits (62), Expect = 1.1
Identities = 27/102 (26%), Positives = 42/102 (40%), Gaps = 19/102 (18%)
Query: 32 VRSPENKEWWW--AHGWNKVREWSEII---VGPKWKTFIRRFNRNNNRAG--AASYDKK- 83
VR E WWW +H +K +W + + K K ++ + N + A +Y KK
Sbjct: 2 VREEEKSRWWWFESHKSSKHSQWLQSTLAEIDAKTKAMLKLLDGNADSFAQRAETYYKKR 61
Query: 84 -------GSFHYDSLSYALNFDDGEEDVYSYGGFSTRFASVP 118
F+ S A+NFD + S + +R A VP
Sbjct: 62 PELISFVEDFYRAHRSLAVNFD----HLKSSDHYGSRSAKVP 99
>At2g25735 unknown protein
Length = 119
Score = 28.1 bits (61), Expect = 1.5
Identities = 19/63 (30%), Positives = 26/63 (41%), Gaps = 11/63 (17%)
Query: 59 PKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNFDDG------EEDVYSYGGFST 112
PKWK + + +RA + Y+ Y+LNFD G +E FS
Sbjct: 49 PKWKVLLMKLKLLPSRASSTKV-----VAYEPCDYSLNFDQGPGWHDHDEPENLSRSFSC 103
Query: 113 RFA 115
RFA
Sbjct: 104 RFA 106
>At1g16290 hypothetical protein
Length = 455
Score = 27.7 bits (60), Expect = 1.9
Identities = 13/40 (32%), Positives = 18/40 (44%), Gaps = 5/40 (12%)
Query: 17 PCSSSSETSLSWWERVRSPENKEWWWAHG-----WNKVRE 51
P S S T ++W+ SPE+ E W W K +E
Sbjct: 214 PASPGSNTDFTYWDSRASPEDMEDMWNQSEICKEWTKSKE 253
>At4g25390 receptor kinase-like protein
Length = 651
Score = 26.9 bits (58), Expect = 3.3
Identities = 25/102 (24%), Positives = 41/102 (39%), Gaps = 24/102 (23%)
Query: 18 CSSSSETSLSWWERVRSPENKEWWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAG- 76
CS +S+S W R + WW G + G +W R N +++ +G
Sbjct: 452 CSDDGSSSVSQWRRGSGSGSSIDWWLDG----------LSGERW--LRARGNSHDSVSGE 499
Query: 77 -------AASYDKKGSFHYDSLSYALNFDDGEE---DVYSYG 108
+++ +G+ Y + Y N D+ DVYSYG
Sbjct: 500 IAKSCGISSTPSMRGTVCYAAPEYC-NLDNNVSEKCDVYSYG 540
>At5g25465 unknown protein
Length = 280
Score = 26.6 bits (57), Expect = 4.3
Identities = 22/100 (22%), Positives = 36/100 (36%), Gaps = 5/100 (5%)
Query: 17 PCSSSSETSLSWWERVRSPENKEWWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNN--- 73
P +S ET+ + S + W HG ++ ++ PK F++ + N+
Sbjct: 137 PKNSHKETTTASASASASEFSDNWRETHGCADIKNPELYLLNPKNPYFVKTLTKGNDVLY 196
Query: 74 --RAGAASYDKKGSFHYDSLSYALNFDDGEEDVYSYGGFS 111
+ Y K H + Y L D YGG S
Sbjct: 197 VPKTVIKKYGLKFGPHLSPMHYLLPGDKINGSTKIYGGGS 236
>At5g05170 cellulose synthase catalytic subunit (Ath-B)
Length = 1065
Score = 26.6 bits (57), Expect = 4.3
Identities = 15/39 (38%), Positives = 18/39 (45%)
Query: 70 RNNNRAGAASYDKKGSFHYDSLSYALNFDDGEEDVYSYG 108
+ N++A S KK H DS N DD EE V G
Sbjct: 651 KKNSKAKKESDKKKSGRHTDSTVPVFNLDDIEEGVEGAG 689
>At5g62110 putative protein
Length = 691
Score = 26.2 bits (56), Expect = 5.6
Identities = 13/41 (31%), Positives = 21/41 (50%)
Query: 76 GAASYDKKGSFHYDSLSYALNFDDGEEDVYSYGGFSTRFAS 116
G A D G+ H A+N++ DV +Y ++RF+S
Sbjct: 180 GGAIVDSNGNGHLGESLGAMNWNLNNNDVRNYESSTSRFSS 220
>At2g23630 putative pectinesterase
Length = 541
Score = 26.2 bits (56), Expect = 5.6
Identities = 16/49 (32%), Positives = 24/49 (48%), Gaps = 4/49 (8%)
Query: 47 NKVREWSEIIVGPK----WKTFIRRFNRNNNRAGAASYDKKGSFHYDSL 91
+K+R + +GP W R R N A AA + +GSFHY ++
Sbjct: 298 SKIRPSHPLPIGPTYHIHWSMKQARTIRLNLTANAARPNPQGSFHYGTI 346
>At5g61480 leucine-rich receptor-like protein kinase - like protein
Length = 1041
Score = 25.8 bits (55), Expect = 7.4
Identities = 14/43 (32%), Positives = 20/43 (45%), Gaps = 1/43 (2%)
Query: 84 GSFHYDSLSYALNFD-DGEEDVYSYGGFSTRFASVPASTKPTF 125
GS+ Y + YA D + D+YSYG + S +P F
Sbjct: 888 GSYGYIAPEYAYTLQVDKKSDIYSYGVILLEIITGKRSVEPEF 930
>At2g21680 hypothetical protein
Length = 414
Score = 25.8 bits (55), Expect = 7.4
Identities = 10/31 (32%), Positives = 17/31 (54%)
Query: 15 LIPCSSSSETSLSWWERVRSPENKEWWWAHG 45
L+ C ++S L W ++ PE K W++ G
Sbjct: 299 LLFCINTSVYFLGWPIKIYDPEKKTWFYLQG 329
>At4g31900 putative protein
Length = 1067
Score = 25.4 bits (54), Expect = 9.6
Identities = 17/68 (25%), Positives = 28/68 (41%), Gaps = 5/68 (7%)
Query: 55 IIVGPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNFDDGEEDVYSYGGFSTRF 114
I + P+WKT R G K Y LSY ++ + E D+ + RF
Sbjct: 135 IAIRPEWKTVDRIIACREGDDGEEYLVK-----YKELSYRNSYWESESDISDFQNEIQRF 189
Query: 115 ASVPASTK 122
+ +S++
Sbjct: 190 KDINSSSR 197
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.133 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,544,583
Number of Sequences: 26719
Number of extensions: 153386
Number of successful extensions: 418
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 394
Number of HSP's gapped (non-prelim): 20
length of query: 137
length of database: 11,318,596
effective HSP length: 89
effective length of query: 48
effective length of database: 8,940,605
effective search space: 429149040
effective search space used: 429149040
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0201.10


