

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0197.8
(242 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
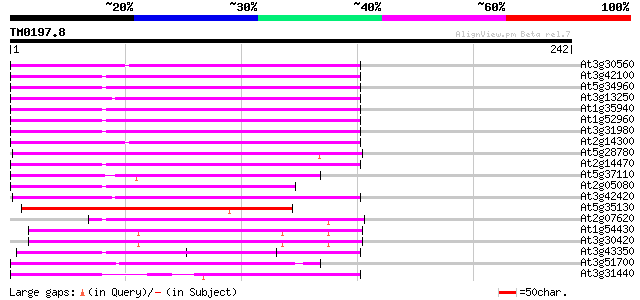
Score E
Sequences producing significant alignments: (bits) Value
At3g30560 hypothetical protein 102 1e-22
At3g42100 putative protein 102 2e-22
At5g34960 putative protein 101 4e-22
At3g13250 hypothetical protein 101 4e-22
At1g35940 hypothetical protein 100 7e-22
At1g52960 hypothetical protein 100 1e-21
At3g31980 hypothetical protein 98 5e-21
At2g14300 pseudogene; similar to MURA transposase of maize Muta... 97 6e-21
At5g28780 putative protein 95 4e-20
At2g14470 pseudogene 93 1e-19
At5g37110 putative helicase 90 9e-19
At2g05080 putative helicase 90 9e-19
At3g42420 putative protein 89 2e-18
At5g35130 putative protein 87 1e-17
At2g07620 putative helicase 85 3e-17
At1g54430 hypothetical protein 85 3e-17
At3g30420 hypothetical protein 82 3e-16
At3g43350 putative protein 80 1e-15
At3g51700 unknown protein 62 2e-10
At3g31440 hypothetical protein 57 7e-09
>At3g30560 hypothetical protein
Length = 1473
Score = 102 bits (255), Expect = 1e-22
Identities = 62/151 (41%), Positives = 79/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF DR IL PT V IN +ML + E Y SS+S S ++ +T FL
Sbjct: 1254 PKFFQDRAILCPTNDDVNSINDHMLSKLTGEEKIYRSSDSIDPSDTRADK-NPVYTPDFL 1312
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N K G+ NH L LKV P+MLLRN+D GL NGTRL +V LG++ + ++TGT
Sbjct: 1313 NKIKISGLPNHLLWLKVGCPVMLLRNLDSHGGLMNGTRLQIVRLGDKLVQGRILTGTRVG 1372
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V I M L PSD + F L A
Sbjct: 1373 KLVIIPRMPLTPSDRRLPFKMKRRQFPLSVA 1403
>At3g42100 putative protein
Length = 1752
Score = 102 bits (253), Expect = 2e-22
Identities = 60/151 (39%), Positives = 82/151 (53%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL PT + V IN YML + + E YLS++S + + + T FL
Sbjct: 1532 PKFFQRRAILAPTNEDVNTINQYMLEHLKSEERIYLSADS-IDPTDSDSLANPVITPDFL 1590
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ + GM +H L LKV P+MLLRN+D GLCNGTRL + +L ++ + A VIT
Sbjct: 1591 NSIQLTGMPHHALRLKVGAPVMLLRNLDPKGGLCNGTRLQITQLAKQVVQAKVITRDRIG 1650
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
D V I +++L PSD F L A
Sbjct: 1651 DIVLIPLINLTPSDTKLPFKMRRRQFPLSVA 1681
>At5g34960 putative protein
Length = 1033
Score = 101 bits (251), Expect = 4e-22
Identities = 60/151 (39%), Positives = 81/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL PT + V IN Y+L + A E YLS++S + + + + T FL
Sbjct: 813 PKFFQGRAILAPTNEDVNTINEYLLEQLHAEERIYLSADS-IDPTDSNSLNNPVITPDFL 871
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K G+ NH L LKV P+MLLRN+D GLCNGTRL + +L + + A VITG
Sbjct: 872 NSIKLAGLPNHSLRLKVSAPVMLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDIIG 931
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V I ++L P+D F L A
Sbjct: 932 HIVLIPTVNLTPTDTKLPFKMRRRQFPLSVA 962
>At3g13250 hypothetical protein
Length = 1419
Score = 101 bits (251), Expect = 4e-22
Identities = 60/151 (39%), Positives = 80/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL P + V IN YML + + E YLS++S S + + T FL
Sbjct: 1200 PKFFQRRAILAPKNEDVNTINQYMLEHLDSEERIYLSADSIDPS-DSDSLKNPVITPDFL 1258
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K GM +H L LKV P+MLLRN+D GLCNGTRL + +L + A VITG
Sbjct: 1259 NSIKVSGMPHHSLRLKVGAPVMLLRNLDPKGGLCNGTRLQITQLCSHIVEAKVITGDRIG 1318
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V+I ++++ PSD F L A
Sbjct: 1319 QIVYIPLINITPSDTKLPFKMRRRQFPLSVA 1349
>At1g35940 hypothetical protein
Length = 1678
Score = 100 bits (249), Expect = 7e-22
Identities = 59/151 (39%), Positives = 80/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL P + V IN Y+L + A E YLS++S + + + T FL
Sbjct: 1458 PKFFQGRAILAPKNEDVNTINEYLLEQLDAEERIYLSADS-IDPTDSDSLNNPVITPDFL 1516
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K G+ NH L LKV P+MLLRN+D GLCNGTRL + +L + + A VITG
Sbjct: 1517 NSIKLPGLPNHSLCLKVGAPVMLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDRIG 1576
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
+ V I ++L P+D F L A
Sbjct: 1577 NIVLIPTVNLTPTDTKLPFKMRRRQFPLSVA 1607
>At1g52960 hypothetical protein
Length = 924
Score = 99.8 bits (247), Expect = 1e-21
Identities = 60/151 (39%), Positives = 80/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL PT + V IN +M+ M+ E YLSS+S + + S + ++ FL
Sbjct: 729 PKFFQGRAILCPTNEDVNSINEHMMSMLDGEERIYLSSDS-IDPADTSSANNDAYSADFL 787
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ + G+ NH L LKV P+MLLRN+D GLCNGTRL V ++ + I A ITG
Sbjct: 788 NSVRVSGLPNHCLRLKVGCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVG 847
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V I M + PSD F L A
Sbjct: 848 KIVLIPRMLITPSDTRLPFKMRRRQFPLSVA 878
>At3g31980 hypothetical protein
Length = 1099
Score = 97.8 bits (242), Expect = 5e-21
Identities = 59/151 (39%), Positives = 76/151 (50%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL P V IN ML + YLS++S + + + + T FL
Sbjct: 878 PKFFQQRAILCPRNTDVNTINDIMLDKLNGELVTYLSADS-IDPQDAASLNNPVLTPDFL 936
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K G+ NH L LK+ P+MLLRNID GLCNGTRL V ++G + A VITG
Sbjct: 937 NSIKLSGLPNHNLTLKIGTPVMLLRNIDPKGGLCNGTRLQVTQMGNHILEARVITGDRVG 996
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
DKV I + PSD F + A
Sbjct: 997 DKVIIIKSQISPSDTKLPFRMRRRQFPIAVA 1027
>At2g14300 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 1230
Score = 97.4 bits (241), Expect = 6e-21
Identities = 60/151 (39%), Positives = 77/151 (50%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF + IL PT V IN +ML + E Y SSNS S ++ +T FL
Sbjct: 1053 PKFFQHKAILCPTNDDVNSINDHMLSKLTGEERIYRSSNSIDPSDTRADK-NPIYTPDFL 1111
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N K G++NH L LKV P+MLLRN GL NGTRL +V LG++ + ++TGT
Sbjct: 1112 NKIKISGLANHLLRLKVGCPVMLLRNFYPHGGLMNGTRLQIVRLGDKLVQGRILTGTRVG 1171
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V I M L PSD + F L A
Sbjct: 1172 KLVIIPRMSLTPSDRRLPFKMKRRHFPLSVA 1202
>At5g28780 putative protein
Length = 337
Score = 94.7 bits (234), Expect = 4e-20
Identities = 58/153 (37%), Positives = 84/153 (53%), Gaps = 2/153 (1%)
Query: 2 QFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLN 61
Q+ +R ILTP + V+ IN YML + EYLSS S ++ + +LN
Sbjct: 113 QYLTERGILTPHNEYVDEINAYMLSQVGGDSKEYLSSYSIGKADTIGADYEALYHVKYLN 172
Query: 62 NTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAND 121
+ + + HK+ LK +PIM +RN +Q GLCNGTRLIV LGE+ I A ++TGT+A
Sbjct: 173 SLEFPSLPKHKISLKKGVPIMQMRNFNQKEGLCNGTRLIVTNLGEQVIEAQIVTGTHAGK 232
Query: 122 KVHISIMDLVP--SDPNFQLNSEEDSFQLQYAL 152
V I L P S+ F L ++ ++ YA+
Sbjct: 233 MVSIPRFILSPPQSEHPFTLRRQQFPMRVCYAM 265
>At2g14470 pseudogene
Length = 1265
Score = 93.2 bits (230), Expect = 1e-19
Identities = 55/151 (36%), Positives = 78/151 (51%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL + V IN Y+L + A E YLS++S + + + T FL
Sbjct: 1062 PKFFQGRAILASKNEDVNTINEYLLDQLHAEERIYLSADS-IDPTDSDSLSNPVITPDFL 1120
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K G+ NH L LKV P++LLRN+D GLCNGTRL + +L + + A VITG
Sbjct: 1121 NSIKLPGLPNHSLRLKVGAPVLLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDRIG 1180
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
+ I ++L P++ F L A
Sbjct: 1181 HIILIPTVNLTPTNTKLPFKMRRRQFPLSVA 1211
>At5g37110 putative helicase
Length = 1307
Score = 90.1 bits (222), Expect = 9e-19
Identities = 56/137 (40%), Positives = 77/137 (55%), Gaps = 7/137 (5%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GE---WFTT 57
P FF ++ IL PT + V IN ML + E +LSS+S + ++I G+ T
Sbjct: 1098 PIFFQEKAILCPTNEDVNQINETMLDNLQGEEFTFLSSDSL----DPADIGGKNNPALTP 1153
Query: 58 *FLNNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGT 117
FLN+ K + NHKL LK+ P+MLLRNID GL NGTRL + ++G + A ++TG
Sbjct: 1154 DFLNSVKVSRLPNHKLRLKIGCPVMLLRNIDPIGGLMNGTRLRITQMGPFILQAMILTGD 1213
Query: 118 NANDKVHISIMDLVPSD 134
A V I + L PSD
Sbjct: 1214 RAGHLVLIPRLKLAPSD 1230
>At2g05080 putative helicase
Length = 1219
Score = 90.1 bits (222), Expect = 9e-19
Identities = 53/123 (43%), Positives = 72/123 (58%), Gaps = 1/123 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P FF +R IL PT + V IN ML + E +LSS+S L + + T FL
Sbjct: 993 PVFFQERAILCPTNEDVNQINETMLDNLQGEELTFLSSDS-LDTADIGSRNNPVLTPEFL 1051
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
NN K G+SNHKL LK+ P+MLLRNID GL NGTRL ++++ + A ++TG A+
Sbjct: 1052 NNVKVLGLSNHKLRLKIGSPVMLLRNIDPIGGLMNGTRLQIMQMSPFILQAMILTGDRAD 1111
Query: 121 DKV 123
K+
Sbjct: 1112 TKL 1114
>At3g42420 putative protein
Length = 1018
Score = 89.4 bits (220), Expect = 2e-18
Identities = 54/150 (36%), Positives = 81/150 (54%), Gaps = 1/150 (0%)
Query: 2 QFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLN 61
+FF RVIL+PT V IN YML + E +LS++S S + + T LN
Sbjct: 861 KFFLKRVILSPTNDDVHTINQYMLNNMEGEERIFLSADSIDPS-DSDSLKNPVITPDLLN 919
Query: 62 NTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAND 121
+ K G+ +H+L LK+ PI+LLRN+D GLCNGTRL + ++ + + A VI G + D
Sbjct: 920 SIKLSGLPHHELRLKIGAPIILLRNLDPKGGLCNGTRLQITQMTIQVLQAKVIIGDRSGD 979
Query: 122 KVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V I ++++ PS+ F + A
Sbjct: 980 IVLIPLINITPSNMKLPFRMRRRQFPVSLA 1009
>At5g35130 putative protein
Length = 414
Score = 86.7 bits (213), Expect = 1e-17
Identities = 48/119 (40%), Positives = 72/119 (60%), Gaps = 2/119 (1%)
Query: 6 DRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLNNTKS 65
+R ILTP + V+ I YML + ++Y SS++ ++ S +T +LN+ +
Sbjct: 271 ERTILTPRNETVDEITNYMLTQLPRTSNKYFSSDNIGKADTISTDYESLYTVEYLNSLEF 330
Query: 66 *GMSNHKLLLKVDIPIMLLRNIDQAAGL--CNGTRLIVVELGERYIGATVITGTNANDK 122
G+ HKL LKV PIMLLRN++Q L CNGTRLI+ LG+R I ++TGT+A ++
Sbjct: 331 RGLPKHKLTLKVGAPIMLLRNLNQKEDLCKCNGTRLIITRLGKRLIEGEIVTGTHAGER 389
>At2g07620 putative helicase
Length = 1241
Score = 85.1 bits (209), Expect = 3e-17
Identities = 49/121 (40%), Positives = 70/121 (57%), Gaps = 3/121 (2%)
Query: 35 YLSSNSALRSYEHSEI*GEWFTT*FLNNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLC 94
YLS++S + + + + FT FLN+ K G+SNH L LK+ P+MLL+NID GLC
Sbjct: 1054 YLSADS-IDPQDPASLNNPVFTPYFLNSIKLSGLSNHNLTLKIGTPVMLLKNIDPKGGLC 1112
Query: 95 NGTRLIVVELGERYIGATVITGTNANDKVHISIMDLVPSDPN--FQLNSEEDSFQLQYAL 152
NGTRL V ++G + A VITG DKV I + PSD F++ + + +A+
Sbjct: 1113 NGTRLQVTQMGNHILEARVITGDRVRDKVIIIKAQISPSDTKLPFRMRRRQFPIAVAFAM 1172
Query: 153 R 153
R
Sbjct: 1173 R 1173
>At1g54430 hypothetical protein
Length = 1639
Score = 85.1 bits (209), Expect = 3e-17
Identities = 55/148 (37%), Positives = 83/148 (55%), Gaps = 4/148 (2%)
Query: 9 ILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEW-FTT*FLNNTKS*G 67
+LTP + V+ IN Y+L + EY S++S R +E E + +LN+ + G
Sbjct: 1418 VLTPRNETVDEINDYLLSKVPGLAKEYFSADSIDRDEALTEEGFEMSYPMEYLNSLEFPG 1477
Query: 68 MSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITG-TNANDKVHIS 126
+ H+L LKV +PIMLLRN++Q GLCNGTRLIV LG++ + A +++ T KV I
Sbjct: 1478 LPAHRLCLKVGVPIMLLRNLNQKEGLCNGTRLIVTHLGDKVLKAEILSDTTKERKKVLIP 1537
Query: 127 IMDLVPSDPN--FQLNSEEDSFQLQYAL 152
+ L P D F L + ++ YA+
Sbjct: 1538 RIILSPQDSKHPFTLRRRQFPVRMCYAM 1565
>At3g30420 hypothetical protein
Length = 837
Score = 81.6 bits (200), Expect = 3e-16
Identities = 53/148 (35%), Positives = 82/148 (54%), Gaps = 4/148 (2%)
Query: 9 ILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEW-FTT*FLNNTKS*G 67
+LTP + V+ IN Y+L + EY S++S + +E E + +LN+ + G
Sbjct: 616 VLTPRNETVDEINDYLLSKVPGLAKEYFSADSIDQDEALTEEGFEMSYPMEYLNSLEFPG 675
Query: 68 MSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITG-TNANDKVHIS 126
+ H+L LKV +PIMLLRN++Q GLCNGTRL V LG++ + A +++ T KV I
Sbjct: 676 LPAHRLCLKVGVPIMLLRNLNQKEGLCNGTRLTVTHLGDKVLKAEILSDTTKKRKKVLIP 735
Query: 127 IMDLVPSDPN--FQLNSEEDSFQLQYAL 152
+ L P D F L + ++ YA+
Sbjct: 736 RIILSPQDSKHPFTLRRRQFPVRMCYAM 763
>At3g43350 putative protein
Length = 830
Score = 79.7 bits (195), Expect = 1e-15
Identities = 48/112 (42%), Positives = 64/112 (56%), Gaps = 1/112 (0%)
Query: 4 FHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLNNT 63
+ R IL PT + V IN +M+ M+ E YLSS+S + + S + FLNN
Sbjct: 441 YQGRAILCPTNEDVNSINEHMMRMLDGEERIYLSSDS-IDPADISSANNAAYLADFLNNV 499
Query: 64 KS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVIT 115
+ G+ NH L LKV P+MLLRN+D GLCNGTRL V ++ + I A IT
Sbjct: 500 RVYGLPNHCLRLKVGCPVMLLRNMDPNKGLCNGTRLQVTQMTDTIIQARFIT 551
Score = 55.5 bits (132), Expect = 3e-08
Identities = 30/75 (40%), Positives = 37/75 (49%)
Query: 77 VDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDKVHISIMDLVPSDPN 136
+ P+MLLRN+D GLCNGTRL V ++ + I A ITG V I M + PSD
Sbjct: 594 IGCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTR 653
Query: 137 FQLNSEEDSFQLQYA 151
F L A
Sbjct: 654 LPFKMRRKQFALSVA 668
Score = 53.5 bits (127), Expect = 1e-07
Identities = 30/75 (40%), Positives = 36/75 (48%)
Query: 77 VDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDKVHISIMDLVPSDPN 136
V P+MLLRN+D GLCNGTRL V ++ + I A ITG V I M + P D
Sbjct: 710 VGCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPLDTR 769
Query: 137 FQLNSEEDSFQLQYA 151
F L A
Sbjct: 770 LPFKMRRKQFALSVA 784
>At3g51700 unknown protein
Length = 344
Score = 62.4 bits (150), Expect = 2e-10
Identities = 43/134 (32%), Positives = 67/134 (49%), Gaps = 4/134 (2%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P F+ +R IL T + IN YML + E + +++ ++ S + FL
Sbjct: 115 PDFYQERAILCHTNDVADEINDYMLSQLQGEETKCYGADTIYPTHA-SPNDKMLYPLEFL 173
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K G + KL LKV P+MLLR++ L GTRL + + + A +ITG N
Sbjct: 174 NSIKIPGFPDFKLRLKVGAPVMLLRDLAPYGWLRKGTRLQITRVETFVLEAMIITGNNHG 233
Query: 121 DKVHISIMDLVPSD 134
+KV ++ +PSD
Sbjct: 234 EKV---LIPRIPSD 244
>At3g31440 hypothetical protein
Length = 536
Score = 57.4 bits (137), Expect = 7e-09
Identities = 45/154 (29%), Positives = 64/154 (41%), Gaps = 31/154 (20%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL P + V IN Y+L + A E YLS++S +
Sbjct: 340 PKFFQGRAILAPKNEDVNTINEYLLEQLHAEERIYLSADS-------------------I 380
Query: 61 NNTKS*GMSNHKLLLKVDIPIM---LLRNIDQAAGLCNGTRLIVVELGERYIGATVITGT 117
+ T S ++N P++ L +I GLCNG RL + +L + A VITG
Sbjct: 381 DPTDSDSLNN---------PVITPDFLNSIKLPGGLCNGARLQITQLFTEIVEAKVITGD 431
Query: 118 NANDKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V I ++L P+D F L A
Sbjct: 432 RIGHIVLIPTVNLTPTDTKLPFKMRRRQFPLSVA 465
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.341 0.150 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,776,311
Number of Sequences: 26719
Number of extensions: 177209
Number of successful extensions: 502
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 449
Number of HSP's gapped (non-prelim): 36
length of query: 242
length of database: 11,318,596
effective HSP length: 96
effective length of query: 146
effective length of database: 8,753,572
effective search space: 1278021512
effective search space used: 1278021512
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0197.8


