

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0195a.11
(377 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
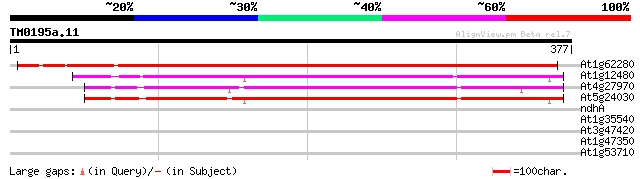
Score E
Sequences producing significant alignments: (bits) Value
At1g62280 unknown protein 424 e-119
At1g12480 unknown protein 256 2e-68
At4g27970 unknown protein 251 6e-67
At5g24030 unknown protein 249 1e-66
ndhA -chloroplast genome- NADH dehydrogenase ND1 30 2.2
At1g35540 auxin response factor, putative 30 2.2
At3g47420 putative sugar transporter protein 30 2.8
At1g47350 hypothetical protein 30 2.8
At1g53710 cell division control protein like protein 29 3.7
>At1g62280 unknown protein
Length = 385
Score = 424 bits (1089), Expect = e-119
Identities = 213/363 (58%), Positives = 265/363 (72%), Gaps = 5/363 (1%)
Query: 6 PKPEIALVIDNTVSTIISREPPSFIVARRLLASLNSVLTKLHAGYFRISLSLGGQALLWK 65
P+ EI + IDN++ + S+E + + + + L S L LHAGYFRISLSL QALLWK
Sbjct: 4 PRQEIHIEIDNSIPS--SKEFKTGLADAKPVV-LMSALRSLHAGYFRISLSLCSQALLWK 60
Query: 66 TLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHH 125
+I P + ++ H+ +PS AF +LW LAL T L LY L+C+F F+ VK EFLH+
Sbjct: 61 IMIAP--ESPSMSHMHSKLPSMAFHLLWYLALVTQVSLCFLYALKCIFFFDKVKEEFLHY 118
Query: 126 VGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRFL 185
+GVNYL+AP ISW L+LQSAP + P + Y L+W+FAVPV+ LD+K+YGQWFT KRFL
Sbjct: 119 IGVNYLYAPSISWLLMLQSAPMMEPNSVLYQTLFWIFAVPVLTLDIKLYGQWFTTEKRFL 178
Query: 186 STAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRLP 245
S ANP SQ+SVI NLV A+ AA MGW E A+C+FSLGMVHYLV+FVTLYQRL GGN P
Sbjct: 179 SMLANPASQVSVIANLVAARGAAEMGWNECALCMFSLGMVHYLVIFVTLYQRLPGGNNFP 238
Query: 246 VLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKRF 305
LRP+FFLF AAP +ASLAW SI G FD ++KMLFFLSLF+FMSLVCRP LF++SMKRF
Sbjct: 239 AKLRPIFFLFVAAPAMASLAWNSICGTFDAVAKMLFFLSLFIFMSLVCRPNLFKKSMKRF 298
Query: 306 NVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLALTVFTFINSKMLL 365
NVAWWAYSFP+T LA+ S YA+EVK + LML+ ++SVL+ L + V T NS LL
Sbjct: 299 NVAWWAYSFPLTFLALDSVQYAQEVKDPVGSGLMLIFSSISVLIFLGMMVLTAANSNRLL 358
Query: 366 PDD 368
D
Sbjct: 359 RHD 361
>At1g12480 unknown protein
Length = 556
Score = 256 bits (653), Expect = 2e-68
Identities = 144/335 (42%), Positives = 197/335 (57%), Gaps = 13/335 (3%)
Query: 43 LTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLAL 102
L + G F I L L QA+LW L KS + +H+ P LV+W +L L
Sbjct: 185 LLRFPIGCFGICLGLSSQAVLWLALA-----KSPATNFLHITPLIN-LVVWLFSLVVLVS 238
Query: 103 LSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYL--VLWW 160
+S Y+L+C+F+F VK E+ H V VN+ FAPW+ L S P + YL +W
Sbjct: 239 VSFTYILKCIFYFEAVKREYFHPVRVNFFFAPWVVCMFLAISVPPMFSPNRKYLHPAIWC 298
Query: 161 VFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLF 220
VF P L++KIYGQW + GKR L ANP+S LSV+GN VGA A+ +GW E A L+
Sbjct: 299 VFMGPYFFLELKIYGQWLSGGKRRLCKVANPSSHLSVVGNFVGAILASKVGWDEVAKFLW 358
Query: 221 SLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKML 280
++G HYLV+FVTLYQRL LP L PV+ +F AAP AS+AW +I G FD S+
Sbjct: 359 AVGFAHYLVVFVTLYQRLPTSEALPKELHPVYSMFIAAPSAASIAWNTIYGQFDGCSRTC 418
Query: 281 FFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLML 340
FF++LFL++SLV R F + +F+VAWW+Y+FP+T ++A+ YAE V G S L L
Sbjct: 419 FFIALFLYISLVARINFF--TGFKFSVAWWSYTFPMTTASVATIKYAEAVPGYPSRALAL 476
Query: 341 VLLALSVLVSLALTVFTFINS---KMLLPDDDPIA 372
L +S + L V T +++ + L P+D IA
Sbjct: 477 TLSFISTAMVCVLFVSTLLHAFVWQTLFPNDLAIA 511
>At4g27970 unknown protein
Length = 519
Score = 251 bits (640), Expect = 6e-67
Identities = 138/327 (42%), Positives = 196/327 (59%), Gaps = 16/327 (4%)
Query: 51 FRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLR 110
+ + L + QA++WKTL T + HV ++ VLW ++L L +S+ YL +
Sbjct: 148 YGMCLGVSSQAIMWKTLA--TTEAEKFLHVTQVINH----VLWWISLLLLLAVSITYLFK 201
Query: 111 CLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAP--FVAPKTATYLVLWWVFAVPVVV 168
+ F V+ EF H + VN+ FAP IS L P ++ +T LW+ P++
Sbjct: 202 TILFFEAVRREFRHPIRVNFFFAPLISILFLALGIPHSIISHLPST---LWYFLMAPILF 258
Query: 169 LDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYL 228
L++KIYGQW + G+R LS ANPT+ LS++GN GA A MG KE + F++G+ +YL
Sbjct: 259 LEMKIYGQWMSGGQRRLSKVANPTNHLSIVGNFAGALLGASMGLKEGPIFFFAIGLAYYL 318
Query: 229 VLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
VLFVTLYQRL LP L PVFFLF AAP VAS+AW I FD S++ +F+SLFL+
Sbjct: 319 VLFVTLYQRLPTNETLPKELHPVFFLFVAAPAVASMAWTKISASFDLGSRLAYFISLFLY 378
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVL---LAL 345
SLVCR LFR +F++AWWAY+FP+T +A A+ Y++EV G + +L +V+ L
Sbjct: 379 FSLVCRINLFRGF--KFSLAWWAYTFPMTAVASATIKYSDEVTGVATKILSVVMSGAATL 436
Query: 346 SVLVSLALTVFTFINSKMLLPDDDPIA 372
+V+ L LTV + L P+D IA
Sbjct: 437 TVIAVLGLTVMHAFVQRDLFPNDVVIA 463
>At5g24030 unknown protein
Length = 635
Score = 249 bits (637), Expect = 1e-66
Identities = 136/327 (41%), Positives = 199/327 (60%), Gaps = 16/327 (4%)
Query: 51 FRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLR 110
F + L + QA++WKTL T + + HV + LW +++ + ++ +YLL+
Sbjct: 263 FGMCLGVSSQAIMWKTLA--TAEPTKFLHVPLWINQG----LWFISVALILTIATIYLLK 316
Query: 111 CLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYL--VLWWVFAVPVVV 168
+ F V+ E+ H + +N+ FAP+IS L P P T L LW++ P +
Sbjct: 317 IILFFEAVRREYYHPIRINFFFAPFISLLFLALGVP---PSIITDLPHFLWYLLMFPFIC 373
Query: 169 LDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYL 228
L++KIYGQW + G+R LS ANPT+ LSV+GN VGA A MG +E + +++GM HYL
Sbjct: 374 LELKIYGQWMSGGQRRLSRVANPTNHLSVVGNFVGALLGASMGLREGPIFFYAVGMAHYL 433
Query: 229 VLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
VLFVTLYQRL LP L PVFFLF AAP VAS+AW + G FD SK+ +F+++FL+
Sbjct: 434 VLFVTLYQRLPTNETLPKDLHPVFFLFVAAPSVASMAWAKVTGSFDYGSKVCYFIAIFLY 493
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVL 348
SL R FR +F+++WWAY+FP+T A+A+ YA VK T++ ++ +VL A++ L
Sbjct: 494 FSLAVRINFFRGI--KFSLSWWAYTFPMTGAAIATIRYATVVKSTMTQIMCVVLCAIATL 551
Query: 349 VSLALTVFTFINS---KMLLPDDDPIA 372
V AL V T I++ + L P+D IA
Sbjct: 552 VVFALLVTTIIHAFVLRDLFPNDLAIA 578
>ndhA -chloroplast genome- NADH dehydrogenase ND1
Length = 360
Score = 30.0 bits (66), Expect = 2.2
Identities = 18/53 (33%), Positives = 29/53 (53%), Gaps = 4/53 (7%)
Query: 266 WGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTV 318
+G+ +G F TL+K LFLF+S+ R TL R M + W + P+++
Sbjct: 298 FGTTIGIFITLAKTY----LFLFVSIATRWTLPRLRMDQLLNLGWKFLLPISL 346
>At1g35540 auxin response factor, putative
Length = 661
Score = 30.0 bits (66), Expect = 2.2
Identities = 16/43 (37%), Positives = 23/43 (53%)
Query: 63 LWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSL 105
LWK GP D L V+ P ++ SL+L++ +LLSL
Sbjct: 28 LWKLCAGPLCDIPKLGEKVYYFPQGHIELVSSLSLYSFSLLSL 70
>At3g47420 putative sugar transporter protein
Length = 523
Score = 29.6 bits (65), Expect = 2.8
Identities = 23/70 (32%), Positives = 34/70 (47%), Gaps = 9/70 (12%)
Query: 171 VKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHM---GWKES----AVCLFSLG 223
V + G WF K KR L + + +GN+ G+ AA M GW S V + +G
Sbjct: 182 VAVVGNWFNKKKRGLIMGI--WNAHTSVGNITGSLIAAAMLRYGWGWSFVVPGVIIVVIG 239
Query: 224 MVHYLVLFVT 233
+V+Y L V+
Sbjct: 240 LVNYAFLPVS 249
>At1g47350 hypothetical protein
Length = 528
Score = 29.6 bits (65), Expect = 2.8
Identities = 19/76 (25%), Positives = 32/76 (42%), Gaps = 2/76 (2%)
Query: 220 FSLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKM 279
FS G ++ F + + + +PVL P+ + P +A G+D + K
Sbjct: 177 FSCGYASGMMYFYGMRIKEDDYDGVPVLCNPITGHYATLPAIARFRQAFSFFGYDPIDKQ 236
Query: 280 LFFLSLFLFMSLVCRP 295
F LF+ M+ C P
Sbjct: 237 --FKVLFMIMAYPCSP 250
>At1g53710 cell division control protein like protein
Length = 528
Score = 29.3 bits (64), Expect = 3.7
Identities = 11/23 (47%), Positives = 18/23 (77%)
Query: 84 VPSSAFLVLWSLALFTLALLSLL 106
+PS F+ +W L+LF ++LL+LL
Sbjct: 352 LPSQLFIYMWYLSLFVMSLLALL 374
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.330 0.141 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,771,645
Number of Sequences: 26719
Number of extensions: 297633
Number of successful extensions: 935
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 919
Number of HSP's gapped (non-prelim): 10
length of query: 377
length of database: 11,318,596
effective HSP length: 101
effective length of query: 276
effective length of database: 8,619,977
effective search space: 2379113652
effective search space used: 2379113652
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0195a.11


