

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
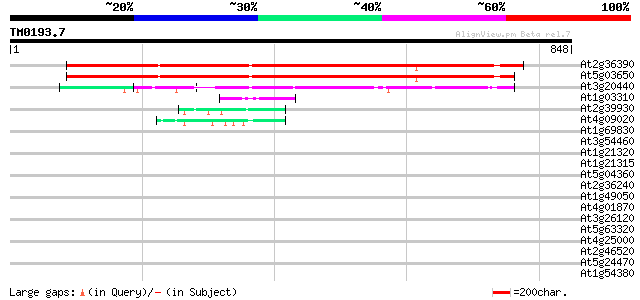
Query= TM0193.7
(848 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g36390 starch branching enzyme class II (sbe2-1) 820 0.0
At5g03650 1,4-alpha-glucan branching enzyme protein soform SBE2.... 818 0.0
At3g20440 1,4-alpha-glucan branching enzyme- like protein 418 e-117
At1g03310 putative isoamylase 55 1e-07
At2g39930 putative isoamylase 55 2e-07
At4g09020 isoamylase-like protein 53 8e-07
At1g69830 alpha-amylase like protein 40 0.007
At3g54460 RING finger -like protein 33 0.67
At1g21320 hypothetical protein 32 1.5
At1g21315 putative protein 32 1.5
At5g04360 pullulanase-like protein (starch debranching enzyme) 31 2.6
At2g36240 putative salt-inducible protein 31 3.3
At1g49050 unknown protein 31 3.3
At4g01870 unknown protein 30 4.4
At3g26120 RNA-binding protein, putative 30 4.4
At5g63320 putative protein 30 5.7
At4g25000 alpha-amylase like protein 30 5.7
At2g46520 putative cellular apoptosis susceptibility protein 30 5.7
At5g24470 pseudo-response regulator 5 (APRR5) 30 7.4
At1g54380 unknown protein 30 7.4
>At2g36390 starch branching enzyme class II (sbe2-1)
Length = 858
Score = 820 bits (2118), Expect = 0.0
Identities = 394/702 (56%), Positives = 502/702 (71%), Gaps = 26/702 (3%)
Query: 87 ILHIDPAIKPFKDHFKCRLKRYIDQKKLIEEYEGGLEEFAQGYLKFGFNREEGGIVYREW 146
I IDP + ++H R +Y ++ I++ EGGLE F++GY FGF R GI YREW
Sbjct: 160 IYDIDPMLNSHRNHLDYRYGQYRKLREEIDKNEGGLEAFSRGYEIFGFTRSATGITYREW 219
Query: 147 APAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPDVA-GNPAIPHNSRVKFRFRHGGV 205
AP A+ A +IGDFN WN + M +N FGVW I +P+ A G+PAIPH SRVK R
Sbjct: 220 APGAKAASLIGDFNNWNAKSDVMARNDFGVWEIFLPNNADGSPAIPHGSRVKIRMDTPSG 279
Query: 206 WADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSS 265
D IPAWIKY+ P + PY+GVY+DPP ++Y FK+PRP KP + RIYE+HVGMSS
Sbjct: 280 IKDSIPAWIKYSVQPPGEI--PYNGVYYDPPEEDKYAFKHPRPKKPTSLRIYESHVGMSS 337
Query: 266 SEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPED 325
+EP+IN+Y F DD+LPRI+ YN VQ+MA+ EH+YYASFGYHVTNFFA SSR GTP+D
Sbjct: 338 TEPKINTYANFRDDVLPRIKKLGYNAVQIMAIQEHAYYASFGYHVTNFFAPSSRFGTPDD 397
Query: 326 LKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDRGYHKLWDSR 385
LK LIDKAH LGL VLMD+VHSHAS N DGL+ FD + YFH+G RGYH +WDSR
Sbjct: 398 LKSLIDKAHELGLVVLMDIVHSHASKNTLDGLDMFDG---TDGQYFHSGSRGYHWMWDSR 454
Query: 386 LFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGVNIAFSGDYNEYFSEAT 445
LFNY +WEVLR+LLSN RWWLEE+KFDGFRFDGVTSM+Y HHG+ + F+G+YNEYF +T
Sbjct: 455 LFNYGSWEVLRYLLSNARWWLEEYKFDGFRFDGVTSMMYTHHGLQVEFTGNYNEYFGYST 514
Query: 446 DVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEVGIGFDYRLAMAIPDKWID 505
DVDAVVYLML N LIH + P+A V+ EDVSGMP P+ + G+GFDYRL MA+ DKWI+
Sbjct: 515 DVDAVVYLMLVNDLIHGLYPEAIVVGEDVSGMPAFCVPVEDGGVGFDYRLHMAVADKWIE 574
Query: 506 YLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGDKTFSFLLMDEEIYSGMSC 565
LK K+D +W + +I+ +LTNRR+ EKCV YAESHDQ++VGDKT +F LMD+++Y M+
Sbjct: 575 LLK-KRDEDWQVGDITFTLTNRRWGEKCVVYAESHDQALVGDKTIAFWLMDKDMYDFMAV 633
Query: 566 LADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGHPEWIDFPR-----------E 614
A+P ++RGIALHKMI ITM LGGEGYLNFMGNEFGHPEWIDFPR
Sbjct: 634 DRQATPRVDRGIALHKMIRLITMGLGGEGYLNFMGNEFGHPEWIDFPRTDQHLPDGRVIA 693
Query: 615 GNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQIVSSTNEEDKVI 674
GN SY+KCRR++ L D ++LRY + FD+AM L++ + F+ S Q +S +E D+VI
Sbjct: 694 GNNGSYDKCRRRFDLGDAEYLRYHGLQEFDRAMQNLEETYGFMTSEHQYISRKDEGDRVI 753
Query: 675 VFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGGHGRVGHNVDHFTAPE 734
VFERG+L+FVFNFH +Y Y++GC +PGKY++ LDSD FGG R+ + + FT+
Sbjct: 754 VFERGNLLFVFNFHWTNSYSDYRIGCSVPGKYKIVLDSDNSLFGGFNRLDDSAEFFTS-- 811
Query: 735 GIPGVPESNFNNRPNSFKILSPPRTCVVYYRVDESQEENSIS 776
+ ++RP SF + +P RT VVY VD+ ++ S
Sbjct: 812 ------DGRHDDRPCSFMVYAPCRTAVVYAAVDDDDDDERSS 847
>At5g03650 1,4-alpha-glucan branching enzyme protein soform SBE2.2
precursor
Length = 805
Score = 818 bits (2114), Expect = 0.0
Identities = 391/689 (56%), Positives = 499/689 (71%), Gaps = 26/689 (3%)
Query: 87 ILHIDPAIKPFKDHFKCRLKRYIDQKKLIEEYEGGLEEFAQGYLKFGFNREEGGIVYREW 146
I IDP ++ + +H R +Y ++ I++YEGGLE F++GY K GF+R + GI YREW
Sbjct: 125 IYEIDPMLRTYNNHLDYRYGQYKRLREEIDKYEGGLEAFSRGYEKLGFSRSDAGITYREW 184
Query: 147 APAAQEAQIIGDFNEWNGSNHPMEKNQFGVWSIKIPD-VAGNPAIPHNSRVKFRFRHGGV 205
AP A+ A +IGDFN WN + M +N+FGVW I +P+ G+PAIPH SRVK R
Sbjct: 185 APGAKAASLIGDFNNWNSNADIMTRNEFGVWEIFLPNNTDGSPAIPHGSRVKIRMDTPSG 244
Query: 206 WADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSS 265
D IPAWIK++ P + P++G+Y+DPP E+Y FK+P+P +PK+ RIYEAHVGMSS
Sbjct: 245 IKDSIPAWIKFSVQAPGEI--PFNGIYYDPPEEEKYVFKHPQPKRPKSLRIYEAHVGMSS 302
Query: 266 SEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPED 325
+EP +N+Y F DD+LPRI+ YN VQ+MA+ EHSYYASFGYHVTNFFA SSR GTPE+
Sbjct: 303 TEPMVNTYANFRDDVLPRIKKLGYNAVQIMAIQEHSYYASFGYHVTNFFAPSSRCGTPEE 362
Query: 326 LKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDRGYHKLWDSR 385
LK LID+AH LGL VLMD+VHSHAS N DGLN FD + YFH+G RGYH +WDSR
Sbjct: 363 LKSLIDRAHELGLVVLMDIVHSHASKNTLDGLNMFDG---TDAHYFHSGPRGYHWMWDSR 419
Query: 386 LFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGVNIAFSGDYNEYFSEAT 445
LFNY +WEVLR+LLSN RWWLEE+KFDGFRFDGVTSM+Y HHG+++ F+G+Y EYF T
Sbjct: 420 LFNYGSWEVLRYLLSNARWWLEEYKFDGFRFDGVTSMMYTHHGLSVGFTGNYTEYFGLET 479
Query: 446 DVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEVGIGFDYRLAMAIPDKWID 505
DVDAV YLML N +IH + P+A + EDVSGMP P+ + G+GFDYRL MAI DKWI+
Sbjct: 480 DVDAVNYLMLVNDMIHGLYPEAITVGEDVSGMPTFCIPVQDGGVGFDYRLHMAIADKWIE 539
Query: 506 YLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGDKTFSFLLMDEEIYSGMSC 565
LK K+D +W M +I +LTNRR+SEKC+SYAESHDQ++VGDKT +F LMD+++Y M+
Sbjct: 540 MLK-KRDEDWQMGDIIYTLTNRRWSEKCISYAESHDQALVGDKTIAFWLMDKDMYDFMAV 598
Query: 566 LADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGHPEWIDFPR-----------E 614
++P I+RGIALHKMI ITM LGGEGYLNFMGNEFGHPEWIDFPR
Sbjct: 599 DRPSTPLIDRGIALHKMIRLITMGLGGEGYLNFMGNEFGHPEWIDFPRGEQRLSDGSVIP 658
Query: 615 GNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQIVSSTNEEDKVI 674
GN +SY+KCRR++ L D D+LRY+ + FD+AM L++ + F+ S Q +S +E D+VI
Sbjct: 659 GNNFSYDKCRRRFDLGDADYLRYRGLQEFDQAMQHLEENYGFMTSEHQFISRKDEADRVI 718
Query: 675 VFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGGHGRVGHNVDHFTAPE 734
VFERGDLVFVFNFH ++Y Y++GC PGKY++ LDSD FGG R+ ++FT
Sbjct: 719 VFERGDLVFVFNFHWTSSYFDYRIGCSKPGKYKIVLDSDDPLFGGFNRLDRKAEYFTY-- 776
Query: 735 GIPGVPESNFNNRPNSFKILSPPRTCVVY 763
+ ++ RP SF + +P RT VVY
Sbjct: 777 ------DGLYDERPCSFMVYAPCRTAVVY 799
>At3g20440 1,4-alpha-glucan branching enzyme- like protein
Length = 869
Score = 418 bits (1075), Expect = e-117
Identities = 222/581 (38%), Positives = 338/581 (57%), Gaps = 58/581 (9%)
Query: 188 PAIPHNSRVKFRFRHGGVWADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPR 247
PA+PH S+ + F +R+PAW Y V P ++W+P Y++KY +
Sbjct: 334 PAVPHGSKYRLYFNTPDGPLERVPAWATY--VQPEDEGKQAYAIHWEPSPEAAYKWKYSK 391
Query: 248 PPKPKAPRIYEAHVGMSSSEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFG 307
P P++ RIYE HVG+S SEP++++++EF
Sbjct: 392 PKVPESLRIYECHVGISGSEPKVSTFEEFTKK---------------------------- 423
Query: 308 YHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQ 367
VTNFFA SSR GTP+D K L+D+AH LGL V +D+VHS+A+ + GL+ FD S
Sbjct: 424 --VTNFFAASSRYGTPDDFKRLVDEAHGLGLLVFLDIVHSYAAADQMVGLSLFDG---SN 478
Query: 368 ESYFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHH 427
+ YFH G RG+HK W +R+F Y + +VL FL+SNL WW+ E++ DG++F + SM+Y H+
Sbjct: 479 DCYFHYGKRGHHKHWGTRMFKYGDLDVLHFLISNLNWWITEYQVDGYQFHSLASMIYTHN 538
Query: 428 GVNIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEV 487
G +F+ D ++Y ++ D DA++YL+LAN ++H P+ IAED + PGL P+S+
Sbjct: 539 GF-ASFNNDLDDYCNQYVDRDALMYLILANEILHVQHPNIITIAEDATYYPGLCEPVSQG 597
Query: 488 GIGFDYRLAMAIPDKWIDYLKNKKDHEWSM-KEISLSLTNRRYSEKCVSYAESHDQSIVG 546
G+GFDY + ++ + W+ L N D+EWSM K +S + N+ Y++K +SYAE+H+QSI G
Sbjct: 598 GLGFDYYVNLSASEMWVSLLDNVPDNEWSMSKIVSTLVANKEYADKMLSYAENHNQSISG 657
Query: 547 DKTFSFLLMDEEIYSGMSCLADASP----TIERGIALHKMIHFITMSLGGEGYLNFMGNE 602
++F+ E ++ G+ + SP ++RGI+LHKMI IT + GG YLNFMGNE
Sbjct: 658 GRSFA-----EILFGGVD---NGSPGGKELLDRGISLHKMIRLITFTSGGRAYLNFMGNE 709
Query: 603 FGHPEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQ 662
FGHPE ++FP + N +S+ R+W L+++ + F +FDK + LD L+
Sbjct: 710 FGHPERVEFPTQSNNFSFSLANRRWDLLESGVHHHLF--SFDKELMDLDKSKGILSRGLP 767
Query: 663 IVSSTNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGGHGR 722
+ N+ + VI F RG +F+FNFHP +YE Y VG + G+Y + L+SD ++GG G
Sbjct: 768 SIHHVNDANMVISFSRGPFLFIFNFHPSNSYEKYDVGVEEAGEYTMILNSDEVKYGGQGI 827
Query: 723 VGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVY 763
V DH+ + + + N ++ P RT VY
Sbjct: 828 V--TEDHY-----LQRSISKRIDGQRNCLEVFLPSRTAQVY 861
Score = 43.1 bits (100), Expect = 6e-04
Identities = 52/228 (22%), Positives = 86/228 (36%), Gaps = 35/228 (15%)
Query: 76 SATEEDLENIGIL-HIDPAIKPFKDHFKCRLKRYIDQKKLIEEYEGGLEEFAQGYLKFGF 134
S E ++ +G L + A + F + R K D K I + +FA G+ G
Sbjct: 63 SDAEAGVDPVGFLTRLGIADRIFAQFLRERHKALKDLKDEIFKRHFDFRDFASGFELLGM 122
Query: 135 NRE-EGGIVYREWAPAAQEAQIIGDFNEWNGSNHPMEK-----NQFGVWSIKIPDVAGNP 188
+R E + + +W P ++ IIGDFN W+ + + + + +G W I + D
Sbjct: 123 HRHMEHRVDFMDWGPGSRYGAIIGDFNGWSPTENAAREGLFGHDDYGYWFIILEDKLREG 182
Query: 189 AIP-------HNSRVKFRFRHGGVWADRIPAWIKYATVDPTKFAAPYDGVYWDPPLSERY 241
P +N + GV A+ I F D YW+P
Sbjct: 183 EEPDELYFQQYNYVDDYDKGDSGVSAEEI-------------FQKAND-EYWEPGEDRFI 228
Query: 242 QFKYPRPPK-------PKAPRIYEAHVGMSSSEPRINSYKEFADDILP 282
+ ++ P K P +P+ E + +E R +KE D P
Sbjct: 229 KNRFEVPAKLYEQMFGPNSPQTLEELGDIPDAETRYKQWKEEHKDDPP 276
>At1g03310 putative isoamylase
Length = 882
Score = 55.5 bits (132), Expect = 1e-07
Identities = 36/117 (30%), Positives = 62/117 (52%), Gaps = 11/117 (9%)
Query: 317 SSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDR 376
+S +K ++ K HS G+ VL++VV +H +++ L G D SY++ +
Sbjct: 439 NSLESAVNSMKVMVKKLHSEGIEVLLEVVFTHTADS--GALRGID-----DSSYYY---K 488
Query: 377 GYHKLWDSRLFNYANWEVLRFL-LSNLRWWLEEFKFDGFRFDGVTSMLYHHHGVNIA 432
G DS+ + N+ V++ L L +LR+W+ EF DGF F +S+L HG ++
Sbjct: 489 GRANDLDSKSYLNCNYPVVQQLVLESLRYWVTEFHVDGFCFINASSLLRGVHGEQLS 545
>At2g39930 putative isoamylase
Length = 783
Score = 55.1 bits (131), Expect = 2e-07
Identities = 53/193 (27%), Positives = 78/193 (39%), Gaps = 33/193 (17%)
Query: 256 IYEAHVG-----MSSSEPRINSYKEFADDILPRIRANNYNTVQLMAVME------HSYYA 304
IYE HV SS +Y+ A+ L ++ N ++LM E +SY
Sbjct: 220 IYEMHVRGFTRHESSKIEFPGTYQGVAEK-LDHLKELGINCIELMPCHEFNELEYYSYNT 278
Query: 305 SFGYHVTNFFAVSS------------------RSGTPEDLKYLIDKAHSLGLHVLMDVVH 346
G H NF+ S+ + K L+ +AH G+ V+MDVV
Sbjct: 279 ILGDHRVNFWGYSTIGFFSPMIRYASASSNNFAGRAINEFKILVKEAHKRGIEVIMDVVL 338
Query: 347 SHASNNVTDGLNGFDVGQVSQESYFHTGDRG--YHKLWDSRLFNYANWEVLRFLLSNLRW 404
+H + G F V Y+ +G Y+ FN + V +F+L LR+
Sbjct: 339 NHTAEGNEKGPI-FSFRGVDNSVYYMLAPKGEFYNYSGCGNTFNCNHPVVRQFILDCLRY 397
Query: 405 WLEEFKFDGFRFD 417
W+ E DGFRFD
Sbjct: 398 WVTEMHVDGFRFD 410
>At4g09020 isoamylase-like protein
Length = 702
Score = 52.8 bits (125), Expect = 8e-07
Identities = 64/240 (26%), Positives = 94/240 (38%), Gaps = 55/240 (22%)
Query: 223 KFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGM------SSSEPRIN-SYKE 275
+F YD + P +K+P P+ K IYE +V S +P I SY
Sbjct: 137 QFYGTYD--FESSPFDWGDDYKFPNIPE-KDLVIYEMNVRAFTADESSGMDPAIGGSYLG 193
Query: 276 FADDILPRIRANNYNTVQLMAVMEHSYYA-------------SFGYHVTNFFAVSSRSGT 322
F + I P ++ N V+L+ V E ++GY NFFA SR +
Sbjct: 194 FIEKI-PHLQDLGINAVELLPVFEFDELELQRRSNPRDHMVNTWGYSTVNFFAPMSRYAS 252
Query: 323 PE--------DLKYLIDKAHSLG---------LHVLMDVVHSHASNN-----VTDGLNGF 360
E + K ++ HS G L V++DVV++H + T G
Sbjct: 253 GEGDPIKASKEFKEMVKALHSAGIEKYSYKFSLQVILDVVYNHTNEADDKYPYTTSFRGI 312
Query: 361 DVGQVSQESYFHTGDRGYHKLWDSRLFNYANWE---VLRFLLSNLRWWLEEFKFDGFRFD 417
D ++ D L S N N V+ +L +LR W+ E+ DGFRFD
Sbjct: 313 D------NKVYYMLDPNNQLLNFSGCGNTLNCNHPVVMELILDSLRHWVTEYHVDGFRFD 366
>At1g69830 alpha-amylase like protein
Length = 887
Score = 39.7 bits (91), Expect = 0.007
Identities = 35/127 (27%), Positives = 57/127 (44%), Gaps = 16/127 (12%)
Query: 307 GYHVTNFFAVSSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHAS----------NNVTDG 356
GY + + ++SR GT ++LK + K H +G+ VL D V +H N
Sbjct: 544 GYMPKDLYNLNSRYGTIDELKDTVKKFHKVGIKVLGDAVLNHRCAHFKNQNGVWNLFGGR 603
Query: 357 LNGFDVGQVSQESYFHTGDRGYHKLWD----SRLFNYANWEVLRFLLSNLRWWLEEFKFD 412
LN D V+ + +F RG D + +++ V + + L W +EE +D
Sbjct: 604 LNWDDRAVVADDPHFQ--GRGNKSSGDNFHAAPNIDHSQDFVRKDIKEWLCWMMEEVGYD 661
Query: 413 GFRFDGV 419
G+R D V
Sbjct: 662 GWRLDFV 668
>At3g54460 RING finger -like protein
Length = 1378
Score = 33.1 bits (74), Expect = 0.67
Identities = 28/101 (27%), Positives = 44/101 (42%), Gaps = 4/101 (3%)
Query: 24 DLAKQNSVELVLGYRNPKGC-NRFSFGSRRSIHERVSTGFKGVAVITDNKSAMSATEEDL 82
D NS+ V +++P C N+ SF + R + E S FK ++ + + + +
Sbjct: 443 DQFTSNSMSAVKRFQSPSSCRNQVSFEAFRPLLESKSLPFKQARLMDPDDQTLESKNSNF 502
Query: 83 ENIGILHIDPAIKPFKDHFKCRLKRYIDQKKLIEEYEGGLE 123
EN HI PA K +CR +K L+ Y G E
Sbjct: 503 ENEFETHI-PASLDLK--AQCRKSLGNVRKNLLPAYNGASE 540
>At1g21320 hypothetical protein
Length = 381
Score = 32.0 bits (71), Expect = 1.5
Identities = 36/156 (23%), Positives = 61/156 (39%), Gaps = 25/156 (16%)
Query: 221 PTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPR-INSYKEFADD 279
P K + PPL+ P P+P+ P+ ++ S P +N Y
Sbjct: 7 PLKVRGDSHKIIKKPPLA-------PPHPQPQPPQTHQQEPSQSRPPPGPVNIYT----- 54
Query: 280 ILPRI---RANNYNTVQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSL 336
+ PRI NN+ T+ + S T + SS + P+D ++D +H L
Sbjct: 55 VTPRIIHTHPNNFMTLVQRLTGQTSTS-------TTSSSSSSSTSEPKDTSTMVDTSHGL 107
Query: 337 GLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFH 372
V A N+++ L F G+ + + Y+H
Sbjct: 108 ISPAARFAVTEKA--NISNELGTFVGGEGTMDQYYH 141
>At1g21315 putative protein
Length = 239
Score = 32.0 bits (71), Expect = 1.5
Identities = 37/164 (22%), Positives = 62/164 (37%), Gaps = 25/164 (15%)
Query: 221 PTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPRINSYKEFADDI 280
P K + PPL+ P P+P+ P+ ++ S P +
Sbjct: 18 PLKVRGDSHKIIKKPPLA-------PPHPQPQPPQTHQQEPSQSRPPPG----PVIIYTV 66
Query: 281 LPRI---RANNYNT-VQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDKAHSL 336
PRI NN+ T VQ + + S Y SS + P+D ++D +H L
Sbjct: 67 SPRIIHTHPNNFMTLVQRLTGKTSTSTTSSSY--------SSSTSAPKDASTMVDTSHGL 118
Query: 337 GLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDRGYHK 380
V A N+++ L F G+ + + Y+H +H+
Sbjct: 119 ISPAARFAVTEKA--NISNELGTFVGGEGTMDQYYHYHHHHHHQ 160
>At5g04360 pullulanase-like protein (starch debranching enzyme)
Length = 965
Score = 31.2 bits (69), Expect = 2.6
Identities = 27/103 (26%), Positives = 47/103 (45%), Gaps = 20/103 (19%)
Query: 325 DLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDRGYHKLWDS 384
+ + ++ + GL+V++DVV++H + G +ES GY+ +S
Sbjct: 460 EFRKMVQALNCTGLNVVLDVVYNHLHAS----------GPHDKESVLDKIVPGYYLRRNS 509
Query: 385 RLF--------NYAN--WEVLRFLLSNLRWWLEEFKFDGFRFD 417
F N A+ + V R + +L W+ +K DGFRFD
Sbjct: 510 DGFIENSTCVNNTASEHYMVDRLIRDDLLNWVVNYKVDGFRFD 552
>At2g36240 putative salt-inducible protein
Length = 497
Score = 30.8 bits (68), Expect = 3.3
Identities = 24/84 (28%), Positives = 37/84 (43%), Gaps = 6/84 (7%)
Query: 235 PPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPRINSYKEFADDILPRIRANNYNTV-- 292
PPL E Y + P PP PK P I S P+ +++ F ++ LP + T+
Sbjct: 24 PPLPEIY--RIPNPP-PKLPEISIPPTLTLSPSPKHSNFVNFLENNLPHHQTLTPQTLLG 80
Query: 293 -QLMAVMEHSYYASFGYHVTNFFA 315
+ H YA + + V N+ A
Sbjct: 81 FLRSKIRNHPLYAHYDFAVFNWAA 104
>At1g49050 unknown protein
Length = 583
Score = 30.8 bits (68), Expect = 3.3
Identities = 28/103 (27%), Positives = 46/103 (44%), Gaps = 8/103 (7%)
Query: 505 DYLKNKKDHEWSMKEIS-----LSLTNRRYSEKCVSYAESHDQSIVGDKTFSFLLMDEEI 559
DY DH +SM ++ L L N +E + + +DQ G + L D +
Sbjct: 282 DYEIEYADHSYSMGVLTKDKFHLKLHNGSLAESDIVFGCGYDQQ--GLLLNTLLKTDGIL 339
Query: 560 YSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNE 602
+ ++ S RGI + + H + L GEGY+ FMG++
Sbjct: 340 GLSRAKISLPSQLASRGIISNVVGHCLASDLNGEGYI-FMGSD 381
>At4g01870 unknown protein
Length = 652
Score = 30.4 bits (67), Expect = 4.4
Identities = 14/33 (42%), Positives = 18/33 (54%)
Query: 716 EFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRP 748
E G R+ + PE IPG PES F++RP
Sbjct: 60 ERNGSARIYKTRSGISKPEQIPGAPESYFHDRP 92
>At3g26120 RNA-binding protein, putative
Length = 615
Score = 30.4 bits (67), Expect = 4.4
Identities = 12/38 (31%), Positives = 21/38 (54%)
Query: 236 PLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPRINSY 273
P+S + F Y PP P + + G+S +EPR+ ++
Sbjct: 25 PISSGFHFPYTPPPPQLPPPLPPSSYGLSPTEPRVFTF 62
>At5g63320 putative protein
Length = 569
Score = 30.0 bits (66), Expect = 5.7
Identities = 16/44 (36%), Positives = 24/44 (54%), Gaps = 2/44 (4%)
Query: 764 YRVDESQEENSISNLVGVQETSTAADIVANIPDGSSASKEREVS 807
+ VD E + + + VG E ST D V +PD +A ER++S
Sbjct: 18 FSVDSLNELDQLEHTVG--EKSTTMDAVVLVPDEETAPPERQIS 59
>At4g25000 alpha-amylase like protein
Length = 423
Score = 30.0 bits (66), Expect = 5.7
Identities = 30/128 (23%), Positives = 44/128 (33%), Gaps = 33/128 (25%)
Query: 317 SSRSGTPEDLKYLIDKAHSLGLHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDR 376
SS+ G+ +LK LI + G+ L D+V +H + D G+ YF G
Sbjct: 86 SSKYGSEAELKSLIKALNQKGIKALADIVINHRTAERKDDKCGY--------CYFEGGTS 137
Query: 377 GYHKLWDSRL-------------------------FNYANWEVLRFLLSNLRWWLEEFKF 411
WD ++ N V + L + W E F
Sbjct: 138 DDRLDWDPSFVCRNDPKFPGTGNLDTGGDFDGAPDIDHLNPRVQKELSEWMNWLKTEIGF 197
Query: 412 DGFRFDGV 419
G+RFD V
Sbjct: 198 HGWRFDYV 205
>At2g46520 putative cellular apoptosis susceptibility protein
Length = 972
Score = 30.0 bits (66), Expect = 5.7
Identities = 25/90 (27%), Positives = 44/90 (48%), Gaps = 12/90 (13%)
Query: 262 GMSSSEPRINSYKEFADDILPRIRANNYNTVQLMAVMEHSYYASFGYHVTNFFAVSSRSG 321
G S S I+ FA+ ILP +++ + N+ ++ + F H+ FA+
Sbjct: 451 GASVSTDLIDVQNFFANIILPELQSRDVNSFPMLKAGSLKFLTMFRSHIPKPFAMQL--- 507
Query: 322 TPEDLKYLIDKAHSLGLHVLMDVVHSHASN 351
PE +++L KA S +VVHS+A++
Sbjct: 508 FPELVRFL--KAES-------NVVHSYAAS 528
>At5g24470 pseudo-response regulator 5 (APRR5)
Length = 667
Score = 29.6 bits (65), Expect = 7.4
Identities = 18/79 (22%), Positives = 33/79 (40%), Gaps = 12/79 (15%)
Query: 15 TAHNSRNKQDLAKQNSVELVLGYRNPKGCNRFSFGSRRSIHERVSTGF------------ 62
T +R + ++ +EL L R P S G R S+H ++ F
Sbjct: 385 TGRRNREESVAQYESRIELDLSLRRPNASENQSSGDRPSLHPSSASAFTRYVHRPLQTQC 444
Query: 63 KGVAVITDNKSAMSATEED 81
V+TD + ++A+++D
Sbjct: 445 SASPVVTDQRKNVAASQDD 463
>At1g54380 unknown protein
Length = 515
Score = 29.6 bits (65), Expect = 7.4
Identities = 25/101 (24%), Positives = 44/101 (42%), Gaps = 3/101 (2%)
Query: 234 DPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPRINS-YKEFAD-DILPRIRANNYNT 291
D L E F+ + + +A R+ EA +E + S +++ D D LP+I N
Sbjct: 80 DGALEESLNFE-EKEQESEAQRLLEAEKRRLLAEIELGSIFRKSVDVDTLPKIEETMDND 138
Query: 292 VQLMAVMEHSYYASFGYHVTNFFAVSSRSGTPEDLKYLIDK 332
V + +++H+ +H + TP LK + DK
Sbjct: 139 VDKIELVDHTALVDVVHHPKRPGTAQNEKDTPRKLKKIGDK 179
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,890,860
Number of Sequences: 26719
Number of extensions: 990514
Number of successful extensions: 2113
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 2069
Number of HSP's gapped (non-prelim): 30
length of query: 848
length of database: 11,318,596
effective HSP length: 108
effective length of query: 740
effective length of database: 8,432,944
effective search space: 6240378560
effective search space used: 6240378560
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0193.7


