

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0193.4
(1140 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
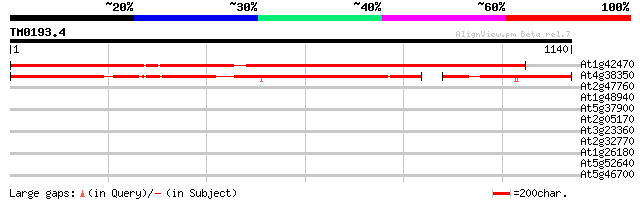
Score E
Sequences producing significant alignments: (bits) Value
At1g42470 Niemann-Pick C disease protein-like protein 1538 0.0
At4g38350 putative protein 979 0.0
At2g47760 Not56-like protein 34 0.54
At1g48940 similar to early nodulins gb|AAD23007.1 33 0.71
At5g37900 putative protein 33 1.2
At2g05170 unknown protein 31 4.6
At3g23360 hypothetical protein 30 6.0
At2g32770 purple acid phosphatase precursor (PAP13) like protein 30 6.0
At1g26180 unknown protein (At1g26180) 30 6.0
At5g52640 heat-shock protein 30 7.9
At5g46700 senescence-associated protein 5-like protein 30 7.9
>At1g42470 Niemann-Pick C disease protein-like protein
Length = 1248
Score = 1538 bits (3983), Expect = 0.0
Identities = 759/1048 (72%), Positives = 877/1048 (83%), Gaps = 27/1048 (2%)
Query: 1 MYDICGTRSDGKVLNCPFGSPAVKPDDLLSSKIQSMCPTITGNVCCTKAQFDTLQTQVQQ 60
MYDICG RSDGKVLNCPF P+VKPDDLLSSKIQS+CPTITGNVCCT+ QFDTL++QVQQ
Sbjct: 21 MYDICGARSDGKVLNCPFNIPSVKPDDLLSSKIQSLCPTITGNVCCTETQFDTLRSQVQQ 80
Query: 61 AIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFVSDAFGEG 120
AIPF+VGCPACLRNFLNLFCELTCSP+QSLFINVTS K NSTV GI Y+++D FG G
Sbjct: 81 AIPFIVGCPACLRNFLNLFCELTCSPDQSLFINVTSTTKVKNNSTVDGIQYYITDDFGAG 140
Query: 121 LYESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSS 180
+YESCK+VKFGS NSRA+ F+GAGA+NFKEWF FIG+KA N PGSPY I F P SS
Sbjct: 141 MYESCKNVKFGSSNSRALDFLGAGAKNFKEWFTFIGQKAGVNLPGSPYGIAFLPTPPVSS 200
Query: 181 GMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFIL 240
GM+PMNVS YSC D SLGCSCGDCPS++ CS+ A K +SCSIK+GSL VKCVDFIL
Sbjct: 201 GMRPMNVSIYSCGDESLGCSCGDCPSAATCSSKAEVPTQKKHSCSIKIGSLEVKCVDFIL 260
Query: 241 AVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMIEDV 300
A+LYI+L+ +FLG L H +R +K T + +S + G + NQ+K + + QM+++
Sbjct: 261 AILYIVLVSLFLGGGLLHPVRGKKKTSQMGTLSE--ASGERNSVNQQKPDTIQSQMLQNT 318
Query: 301 PQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEK 360
PQ RN +LS VQG+++NFY KYG VARHP VL L++++VLLLC+GLIRFKVETRP+K
Sbjct: 319 PQ-RNWGQLSTVQGHLANFYGKYGIWVARHPTLVLCLSVSVVLLLCVGLIRFKVETRPDK 377
Query: 361 LWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKK 420
LWVG GS+AA+EKQFFD+HLAPFYRIEQLI+ATV + +P I++ DNI+ LF++QKK
Sbjct: 378 LWVGSGSRAAEEKQFFDTHLAPFYRIEQLIIATVQTSSHEKAPEILTDDNIKLLFDIQKK 437
Query: 421 VDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEHLNYCFQQYS 480
VD +RAN+SG MVSL DICMKPL +DCATQSVLQ+F +
Sbjct: 438 VDGLRANHSGSMVSLTDICMKPLGEDCATQSVLQHF-----------------------T 474
Query: 481 SADQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAF 540
S + C+SAFK PLDP+T LGGFSG +S ASAF+VTYPV+N +D +GN+T KAVAWEKAF
Sbjct: 475 STESCLSAFKGPLDPTTALGGFSGNSFSEASAFLVTYPVDNFVDNKGNKTEKAVAWEKAF 534
Query: 541 IQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDT 600
IQL KDELLPM Q++NLTL+FSSESSIEEELKRESTAD ITI +SYLVMFAYISLTLGD+
Sbjct: 535 IQLAKDELLPMVQAKNLTLSFSSESSIEEELKRESTADVITIAISYLVMFAYISLTLGDS 594
Query: 601 PHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNM 660
P SFYI+SKVLLGLSGV+LVMLSVLGSV FSA+G+KSTLIIMEVIPFLVLAVGVDNM
Sbjct: 595 PRLKSFYITSKVLLGLSGVLLVMLSVLGSVGFFSAVGMKSTLIIMEVIPFLVLAVGVDNM 654
Query: 661 CILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFA 720
CILVHAVKRQ ELPLE RISNAL+EVGPSITLASL+E+LAFAVG+FI MPA RVFSMFA
Sbjct: 655 CILVHAVKRQEQELPLERRISNALMEVGPSITLASLAEILAFAVGAFIKMPAVRVFSMFA 714
Query: 721 ALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLAR 780
ALAVLLDF+LQ+TAFVALIV D +R EDKRVDCFPCIK +KG+ QRK GLL R
Sbjct: 715 ALAVLLDFLLQITAFVALIVFDFRRTEDKRVDCFPCIKTSKSSISAEKGVGQRKAGLLTR 774
Query: 781 YMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNV 840
YMKEVHAP+LS W VKIVVIA F A+A IALSTRIEPGLEQ+IVLP+DSYLQGYFNN+
Sbjct: 775 YMKEVHAPVLSHWIVKIVVIAFFFGLAMAGIALSTRIEPGLEQQIVLPQDSYLQGYFNNI 834
Query: 841 SEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPA 900
S YLRIGPPLYFV+KNYNYSSES HTNQLCSI++CN +SLLNEI++ASL PE SYIAKPA
Sbjct: 835 STYLRIGPPLYFVLKNYNYSSESRHTNQLCSINKCNPNSLLNEIARASLTPELSYIAKPA 894
Query: 901 ASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSC-VSGACKDCTTCFRHS 959
ASWLDDFLVW+SPEAFGCCRKFTNG++CPPDDQPPCC P SC +S CKDCTTCFRH+
Sbjct: 895 ASWLDDFLVWLSPEAFGCCRKFTNGTFCPPDDQPPCCPPGQTSCGLSEVCKDCTTCFRHA 954
Query: 960 DLRNDRTSTMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHT 1019
DL +DR ST QF++KLPWFL+ALPSADCAKGGHGAY+SSVDL+GY +GIIQASSFRTYHT
Sbjct: 955 DLSSDRPSTTQFKEKLPWFLNALPSADCAKGGHGAYSSSVDLQGYANGIIQASSFRTYHT 1014
Query: 1020 PLNKQVDYVNSMRAAREFSSKVSDSLKV 1047
PLNKQVD+VNSMRAA+EFS+KVS SLK+
Sbjct: 1015 PLNKQVDFVNSMRAAQEFSAKVSRSLKM 1042
>At4g38350 putative protein
Length = 1055
Score = 979 bits (2531), Expect = 0.0
Identities = 512/860 (59%), Positives = 627/860 (72%), Gaps = 85/860 (9%)
Query: 1 MYDICGTRSDGKVLNCPFGSPAVKPDDLLSSKIQSMCPTITGNVCCTKAQFDTLQTQVQQ 60
MYDICG RSDGKVLNCP+ SP+++PD+L S+KIQS+CPTI+GNVCCT+ QFDTL++QVQQ
Sbjct: 1 MYDICGHRSDGKVLNCPYASPSIQPDELFSAKIQSLCPTISGNVCCTETQFDTLRSQVQQ 60
Query: 61 AIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFVSDAFGEG 120
A+PFLVGCPACLRNFLNLFCEL+CSPNQSLFINVTSV + GN TV GIDY ++D FGEG
Sbjct: 61 AVPFLVGCPACLRNFLNLFCELSCSPNQSLFINVTSVAEVSGNLTVDGIDYHITDTFGEG 120
Query: 121 LYESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSS 180
LYESCK+VKFG+MN+RAI F+G GA+NF+EWF FIG+KA PGSPYAI F+ + +SS
Sbjct: 121 LYESCKEVKFGTMNTRAINFVGGGAKNFREWFTFIGQKAPSGFPGSPYAINFKSSIPESS 180
Query: 181 GMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFIL 240
M PMNVS YSC+ CS+ + +SCSI++G L V+C++ +
Sbjct: 181 AMVPMNVSVYSCA----------------CSSPEPLPPHDEDSCSIRIGPLKVRCIELSM 224
Query: 241 AVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMIEDV 300
A++Y++L+ F GWA +R R T+P+ + S +L+ ++ + + I V
Sbjct: 225 ALVYVLLVSCFFGWAGLNRRRNT-----TQPLDS--SKPLLHPVEEDGINSEMKENILGV 277
Query: 301 PQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEK 360
R+ +LS VQ YM+ FYR YGS +AR+P VL +++AIVL LC GL FKVETRPEK
Sbjct: 278 KVQRHA-QLSPVQRYMAKFYRSYGSWIARNPSLVLFMSVAIVLALCSGLYNFKVETRPEK 336
Query: 361 LWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKK 420
LWVGP SKAA+EK+FFD+HL+PFYRIEQLILATVPD + +P IV+ +NI LF++Q+K
Sbjct: 337 LWVGPESKAAEEKKFFDTHLSPFYRIEQLILATVPDPKSGRAPSIVTDENILLLFDIQQK 396
Query: 421 VDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEHLNYCFQQYS 480
YFKMD FDD G VEH YCFQ Y+
Sbjct: 397 ----------------------------------YFKMDSGTFDDYGGVEHAEYCFQHYT 422
Query: 481 SADQCMSAFKAPLDPSTVLGGFSGKDYSG------------------------ASAFIVT 516
S++ C+SAF+AP+DPS VLGGFSG +YS A+AF+VT
Sbjct: 423 SSETCLSAFQAPVDPSAVLGGFSGNNYSEVMVSELGCSVPFDCYSDVKRTLFQATAFVVT 482
Query: 517 YPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKREST 576
YPVNN I + NE A+AVAWEK+FIQL K+ELLPM +S+NL+L+FSSESSIEEELKREST
Sbjct: 483 YPVNNVIGDSSNENARAVAWEKSFIQLAKEELLPMVRSKNLSLSFSSESSIEEELKREST 542
Query: 577 ADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSAL 636
AD ITI SYLVMF YIS+TLGD P +FYISSKVLLGLSGV+LV+LSVLGSV +FSAL
Sbjct: 543 ADVITIAASYLVMFVYISVTLGDAPQFYTFYISSKVLLGLSGVVLVLLSVLGSVGVFSAL 602
Query: 637 GVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASL 696
GVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQP E+ LE RIS+ALVEVGPSITLASL
Sbjct: 603 GVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPREVSLEQRISSALVEVGPSITLASL 662
Query: 697 SEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPC 756
SEVLAFAVG+F+ MPACR+FSMFAALA++LDF LQ+TAFVALIV D +R+ D R+DCFPC
Sbjct: 663 SEVLAFAVGAFVPMPACRIFSMFAALAIMLDFFLQITAFVALIVFDCKRSADNRIDCFPC 722
Query: 757 IKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTR 816
IKV S + +G R+PG L RYMKEVHAP+L +WGVK+VV+A+F AFALASI +S
Sbjct: 723 IKVPSSSRESVEG--GREPGFLERYMKEVHAPVLGLWGVKMVVVAVFFAFALASI-ISRA 779
Query: 817 IEPGLEQEIVLPRDSYLQGY 836
+ I P S+L +
Sbjct: 780 SQASDTSYIAKPAASWLDDF 799
Score = 334 bits (856), Expect = 2e-91
Identities = 175/303 (57%), Positives = 208/303 (67%), Gaps = 62/303 (20%)
Query: 879 SLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCA 938
+L + IS+AS +TSYIAKPAASWLDDFLVW+SPEAFGCCRKFTNGSYCPPDDQ
Sbjct: 771 ALASIISRASQASDTSYIAKPAASWLDDFLVWLSPEAFGCCRKFTNGSYCPPDDQ----- 825
Query: 939 PEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGGHGAYTSS 998
CFRHSDL DR ST QFR+KLPWFL+ALPSADCAKGGHGAYT+S
Sbjct: 826 ----------------CFRHSDLVQDRPSTAQFREKLPWFLNALPSADCAKGGHGAYTNS 869
Query: 999 VDLKGYDSGIIQASSFRTYHTPLNKQVD-------------YVN---------------- 1029
VDLKGY+SG+IQAS FRTYHTPLN Q+D Y+N
Sbjct: 870 VDLKGYESGVIQASEFRTYHTPLNTQIDIFPYSVFYIFFEQYLNIWTVALTNLAIAIVGI 929
Query: 1030 ------------SMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLV 1077
S+ A EF +S + ++SGD++ R +EAL TMGASVFSGITLTKLV
Sbjct: 930 QLNAVSVVNLIMSIGIAVEFCVHISHAFLMSSGDREHRAREALETMGASVFSGITLTKLV 989
Query: 1078 GVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPPSRCTIIEQEENRSS 1137
GVIVL F+R+E+FV+YYFQMYL+LV++GFLHGLVFLPV+LS+ GPP IEQ++ +
Sbjct: 990 GVIVLCFARSEIFVVYYFQMYLALVIIGFLHGLVFLPVILSLAGPPQLNLDIEQQQTDEA 1049
Query: 1138 TSS 1140
+SS
Sbjct: 1050 SSS 1052
>At2g47760 Not56-like protein
Length = 438
Score = 33.9 bits (76), Expect = 0.54
Identities = 19/74 (25%), Positives = 38/74 (50%)
Query: 579 AITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGV 638
A+T+L + + +F Y LG + + VLL ++L++L + + + SAL
Sbjct: 165 AMTLLHASMALFLYRKWHLGMLVFSGAVSVKMNVLLYAPTLLLLLLKAMNIIGVVSALAG 224
Query: 639 KSTLIIMEVIPFLV 652
+ + I+ +PFL+
Sbjct: 225 AALVQILVGLPFLI 238
>At1g48940 similar to early nodulins gb|AAD23007.1
Length = 177
Score = 33.5 bits (75), Expect = 0.71
Identities = 17/45 (37%), Positives = 25/45 (54%)
Query: 794 GVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFN 838
G KIV+++IFV F + S+ T E G E ++P+ S FN
Sbjct: 3 GQKIVLLSIFVCFYVFSLVSCTEFEAGGENGWIIPQSSNQSDIFN 47
>At5g37900 putative protein
Length = 241
Score = 32.7 bits (73), Expect = 1.2
Identities = 22/88 (25%), Positives = 37/88 (42%), Gaps = 3/88 (3%)
Query: 893 TSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQ---PPCCAPEDDSCVSGAC 949
T I +P + L+ LV + FGC F G +++ C P D SG
Sbjct: 43 TCPIFQPMENILESILVTCPNDMFGCTESFLYGKKSTHEEECIFSLCSCPSLDCEYSGRY 102
Query: 950 KDCTTCFRHSDLRNDRTSTMQFRDKLPW 977
+D ++ + + N +T FR +P+
Sbjct: 103 EDLYDHYKLTHISNSYWTTNCFRSSIPY 130
>At2g05170 unknown protein
Length = 932
Score = 30.8 bits (68), Expect = 4.6
Identities = 13/30 (43%), Positives = 20/30 (66%)
Query: 633 FSALGVKSTLIIMEVIPFLVLAVGVDNMCI 662
F + S L++ EV P L++A+G+DN CI
Sbjct: 126 FPEAKITSFLVLEEVPPILLIAIGLDNGCI 155
>At3g23360 hypothetical protein
Length = 256
Score = 30.4 bits (67), Expect = 6.0
Identities = 31/110 (28%), Positives = 52/110 (47%), Gaps = 9/110 (8%)
Query: 789 ILSIWGVKIVVIAIFVAFALASIALSTR-----IEPGLEQEIVLPRDSYL---QGYFNNV 840
I SI ++VV A + ST+ I PG E++ PR+S L N+
Sbjct: 139 IASIGDHRVVVCKDGEAHQIRDRKASTKHWSQFIFPGEEEDESDPRNSELVVITEKINSD 198
Query: 841 SEYLRIGPP-LYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASL 889
+E++ IG P ++ V+K+ + H ++C + LN ISK+S+
Sbjct: 199 TEFIIIGSPGIWEVMKSQEAINLIRHIEDPKEAAKCLAKEALNRISKSSI 248
>At2g32770 purple acid phosphatase precursor (PAP13) like protein
Length = 516
Score = 30.4 bits (67), Expect = 6.0
Identities = 18/46 (39%), Positives = 24/46 (52%), Gaps = 1/46 (2%)
Query: 442 PLDKDCATQSVLQYFKMDPRNFDDSGAVEHLNYCFQQYSSADQCMS 487
PLD +C QS++QY + D R A H QQYSS + M+
Sbjct: 101 PLDPNCV-QSIVQYREFDVRRTRKQNATGHSIVYNQQYSSENGFMN 145
>At1g26180 unknown protein (At1g26180)
Length = 289
Score = 30.4 bits (67), Expect = 6.0
Identities = 24/73 (32%), Positives = 40/73 (53%), Gaps = 2/73 (2%)
Query: 571 LKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFY-ISSKVLLGLSGVILVMLSVLGS 629
L RE+ ITI + + + + ++ L P S Y I VLL SG +++L +L S
Sbjct: 208 LIRENILAVITINIHASLSYVFTAMGLNIMPSMSLIYMIFGTVLLLNSGFFVLLLHLLYS 267
Query: 630 VAIFSALGVKSTL 642
+ + + LG+KS+L
Sbjct: 268 IFL-TRLGMKSSL 279
>At5g52640 heat-shock protein
Length = 705
Score = 30.0 bits (66), Expect = 7.9
Identities = 25/81 (30%), Positives = 39/81 (47%), Gaps = 5/81 (6%)
Query: 542 QLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYIS--LTLG- 598
Q ++D + S T+ + ++ I EEL++ + AD V LVM Y + LT G
Sbjct: 595 QALRDSSMSGYMSSKKTMEINPDNGIMEELRKRAEADKNDKSVKDLVMLLYETALLTSGF 654
Query: 599 --DTPHPSSFYISSKVLLGLS 617
D P+ + I + LGLS
Sbjct: 655 SLDEPNTFAARIHRMLKLGLS 675
>At5g46700 senescence-associated protein 5-like protein
Length = 269
Score = 30.0 bits (66), Expect = 7.9
Identities = 19/56 (33%), Positives = 29/56 (50%), Gaps = 1/56 (1%)
Query: 1071 ITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP-PSR 1125
I L L G I ++ T + V+Y M + +VLLG L G +++ + P PSR
Sbjct: 52 ILLVGLAGFIGGFWRITWLLVVYLIAMLILIVLLGCLVGFIYMVTIRGSGHPEPSR 107
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,787,639
Number of Sequences: 26719
Number of extensions: 1042860
Number of successful extensions: 2542
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 2523
Number of HSP's gapped (non-prelim): 15
length of query: 1140
length of database: 11,318,596
effective HSP length: 110
effective length of query: 1030
effective length of database: 8,379,506
effective search space: 8630891180
effective search space used: 8630891180
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0193.4


