

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0191.15
(398 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
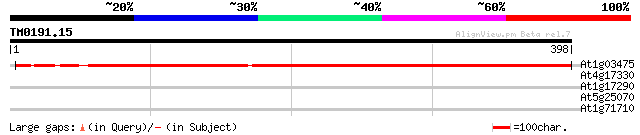
At1g03475 coproporphyrinogen III oxidase (LIN2) 642 0.0
At4g17330 G2484-1 protein 31 1.4
At1g17290 alanine aminotransferase (ALAAT1) 30 2.3
At5g25070 unknown protein 28 6.8
At1g71710 RIBOSOMAL PROTEIN, putative (At1g71710) 28 8.9
>At1g03475 coproporphyrinogen III oxidase (LIN2)
Length = 386
Score = 642 bits (1655), Expect = 0.0
Identities = 314/394 (79%), Positives = 340/394 (85%), Gaps = 13/394 (3%)
Query: 5 ATIVSPPSYAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVSIEKE 64
+T++S P++A F+S SP L R++ I+ K +L R VSIEKE
Sbjct: 6 STLLSSPTFAP--FSSHRLHYSPNPSTL---RFSRPIRNKPNLAL------RCSVSIEKE 54
Query: 65 TPEAERPETFLRGGDTAEALSSSSSVRARFEKMIREAQDSVCDAIEAADGGAKFKEDVWS 124
PE ERP TFLR D SSSSSVRARFE MIR AQDSVCDAIEA +GG KFKEDVWS
Sbjct: 55 VPETERPFTFLRDSDDVTPSSSSSSVRARFETMIRAAQDSVCDAIEAIEGGPKFKEDVWS 114
Query: 125 RPGGGGGISRVLQDGAVWEKAGVNVSVVYGVMPPDAYRAAKGAAASPDEKPGPVPFFAAG 184
RPGGGGGISRVLQDG V+EKAGVNVSVVYGVMPP+AYRAAKG+A+ D+KPGPVPFFAAG
Sbjct: 115 RPGGGGGISRVLQDGNVFEKAGVNVSVVYGVMPPEAYRAAKGSAS--DQKPGPVPFFAAG 172
Query: 185 ISSVLHPKNPFAPTMHFNYRYFETDAPKDAPGAPKQWWFGGGTDLTPAYIFEEDVKHFHS 244
+SSVLHPKNPFAPT+HFNYRYFETDAPKD PGAP+QWWFGGGTD TPAYIFEEDVKHFHS
Sbjct: 173 VSSVLHPKNPFAPTLHFNYRYFETDAPKDVPGAPRQWWFGGGTDFTPAYIFEEDVKHFHS 232
Query: 245 IQKQACDKFDPTFYPRFKKWCDDYFYIKHRGERRGLGGIFFDDLNDYDQEMLLSFATECA 304
IQKQACDKFDP+FYPRFKKWCDDYFYIKHR ERRGLGGIFFDDLNDYDQEMLLSFATECA
Sbjct: 233 IQKQACDKFDPSFYPRFKKWCDDYFYIKHRDERRGLGGIFFDDLNDYDQEMLLSFATECA 292
Query: 305 NSVVPSYIPIIEKRKDTPFTDHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIESILV 364
NSVVP+YIPI+EKRKD FT+ KAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIESILV
Sbjct: 293 NSVVPAYIPIVEKRKDMEFTEQHKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIESILV 352
Query: 365 SLPLTSRWEYDHKPEEGSEEWKLLDACINPKEWI 398
SLPL++RWEYDHKPEEG+EEWKLLDACINPKEWI
Sbjct: 353 SLPLSARWEYDHKPEEGTEEWKLLDACINPKEWI 386
>At4g17330 G2484-1 protein
Length = 2037
Score = 30.8 bits (68), Expect = 1.4
Identities = 28/87 (32%), Positives = 41/87 (46%), Gaps = 10/87 (11%)
Query: 3 SCATIVSPPSYAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVSIE 62
S AT+VSP P +S T++ ++K S+R+ +T S+G K R G S++
Sbjct: 827 SQATLVSPIVVGSPSTSSLDKTAAKSSKAKSERK-----PRRTSKSVG-KETSRKGTSVK 880
Query: 63 KETPEAERPETFLRGGDTAEALSSSSS 89
TP E F GG T S +S
Sbjct: 881 GATP----IEQFQSGGKTNAVNQSLAS 903
>At1g17290 alanine aminotransferase (ALAAT1)
Length = 488
Score = 30.0 bits (66), Expect = 2.3
Identities = 15/50 (30%), Positives = 25/50 (50%)
Query: 98 IREAQDSVCDAIEAADGGAKFKEDVWSRPGGGGGISRVLQDGAVWEKAGV 147
I+ +D++ D IEA DG D++ G G+ ++Q EK G+
Sbjct: 124 IKGLRDAIADGIEARDGFPADPNDIFMTDGASPGVHMMMQLLITSEKDGI 173
>At5g25070 unknown protein
Length = 736
Score = 28.5 bits (62), Expect = 6.8
Identities = 21/71 (29%), Positives = 34/71 (47%), Gaps = 1/71 (1%)
Query: 41 IQTKTMTSLGSKGAVRAGVSIEKETPEAERPETFLRGGDTAEALSSSSSVRARFEKMIRE 100
I+ K S + A +EK+ EA E F +E+L++ R ++R+
Sbjct: 218 IRKKRQASENLRLASTTHEELEKQLEEAIETEDFDAAERISESLAAKERDRLALLALLRQ 277
Query: 101 AQDSVCDAIEA 111
A +S CDAIE+
Sbjct: 278 A-ESDCDAIES 287
>At1g71710 RIBOSOMAL PROTEIN, putative (At1g71710)
Length = 664
Score = 28.1 bits (61), Expect = 8.9
Identities = 22/89 (24%), Positives = 35/89 (38%), Gaps = 8/89 (8%)
Query: 33 SKRRWNPLIQTKTMTSLGSKGAVRAGVSIEKETPEAERPETFLRGGDTAEALSSSSSVRA 92
S+R W L+ K + G A E E +A D++ S R
Sbjct: 18 SQRLWAKLVMRKWLNISGRDPEYGADTDNESENEDAREDND-----DSSSDEEGGSGSRG 72
Query: 93 RFEKMIREAQDSVCDA---IEAADGGAKF 118
R K+ A+D++ A ++AA A+F
Sbjct: 73 RESKVYENAEDAIAAASAVVDAAAAAAEF 101
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.135 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,268,815
Number of Sequences: 26719
Number of extensions: 478907
Number of successful extensions: 1137
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1134
Number of HSP's gapped (non-prelim): 6
length of query: 398
length of database: 11,318,596
effective HSP length: 101
effective length of query: 297
effective length of database: 8,619,977
effective search space: 2560133169
effective search space used: 2560133169
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0191.15


