

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0190.3
(329 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
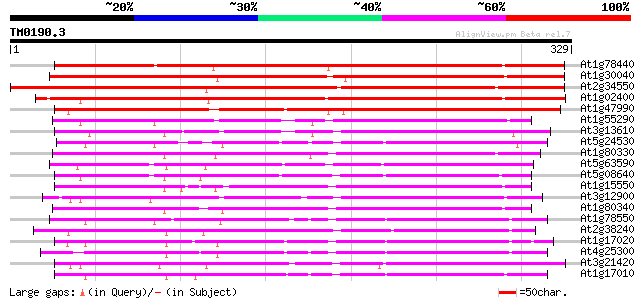
Score E
Sequences producing significant alignments: (bits) Value
At1g78440 putative dioxygenase 373 e-104
At1g30040 unknown protein 370 e-103
At2g34550 putative gibberellin 2-oxidase 342 2e-94
At1g02400 unknown protein 275 2e-74
At1g47990 dioxygenase, putative 273 1e-73
At1g55290 leucoanthocyanidin dioxygenase 2, putative 157 6e-39
At3g13610 unknown protein 156 1e-38
At5g24530 flavanone 3-hydroxylase-like protein 149 2e-36
At1g80330 putative gibberellin 3 beta-hydroxylase 149 2e-36
At5g63590 flavonol synthase 146 1e-35
At5g08640 flavonol synthase (FLS) (sp|Q96330) 145 2e-35
At1g15550 Gibberillin 3 beta-hydroxylase (AtGA3ox1) 145 4e-35
At3g12900 hypothetical protein 140 7e-34
At1g80340 gibberellin 3 beta-hydroxylase 140 9e-34
At1g78550 unknown protein 140 1e-33
At2g38240 putative anthocyanidin synthase 139 2e-33
At1g17020 SRG1-like protein 138 4e-33
At4g25300 SRG1-like protein 138 5e-33
At3g21420 unknown protein 136 2e-32
At1g17010 SRG1-like protein 136 2e-32
>At1g78440 putative dioxygenase
Length = 329
Score = 373 bits (957), Expect = e-104
Identities = 184/306 (60%), Positives = 236/306 (76%), Gaps = 9/306 (2%)
Query: 27 IPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGL 86
IPV+D+S P++K +VKACE+FGFFKVINHGVS E +S LE E ++FFS+P +EK +
Sbjct: 18 IPVIDMSDPESKHALVKACEDFGFFKVINHGVSAELVSVLEHETVDFFSLPKSEKTQVA- 76
Query: 87 ANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN----VPVHSQNPEKFRCVLDDYLCAVRN 142
PFGYGN +IG NGDVGWVEYLL+ N + P ++P FR L++Y +VR
Sbjct: 77 GYPFGYGNSKIGRNGDVGWVEYLLMNANHDSGSGPLFPSLLKSPGTFRNALEEYTTSVRK 136
Query: 143 MGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACP---EMGLDGENLIGFGE 199
M ++L+ + +GL I+P+N LSKL+ D+ +DS+ R+NHYP CP + G+N+IGFGE
Sbjct: 137 MTFDVLEKITDGLGIKPRNTLSKLVSDQNTDSILRLNHYPPCPLSNKKTNGGKNVIGFGE 196
Query: 200 HTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRV 259
HTDPQIIS+LRSNNTSGLQI+L DGSWISVPPD++SFF NVGDSLQVMTNGRF+SV+HRV
Sbjct: 197 HTDPQIISVLRSNNTSGLQINLNDGSWISVPPDHTSFFFNVGDSLQVMTNGRFKSVRHRV 256
Query: 260 LANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLADNR 319
LAN +SR+SMIYF GP L+++IAPL L+ ++E LY+EFTW EYKNS+Y SRL+DNR
Sbjct: 257 LANCKKSRVSMIYFAGPSLTQRIAPLTCLI-DNEDERLYEEFTWSEYKNSTYNSRLSDNR 315
Query: 320 LGHFER 325
L FER
Sbjct: 316 LQQFER 321
>At1g30040 unknown protein
Length = 341
Score = 370 bits (950), Expect = e-103
Identities = 181/308 (58%), Positives = 244/308 (78%), Gaps = 10/308 (3%)
Query: 24 SFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEK 83
S +IPVV+L+ P+AK+ IVKACEEFGFFKV+NHGV E +++LE EA+ FF +P + K +
Sbjct: 28 SHSIPVVNLADPEAKTRIVKACEEFGFFKVVNHGVRPELMTRLEQEAIGFFGLPQSLKNR 87
Query: 84 TGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP----VHSQNPEKFRCVLDDYLCA 139
G P+GYGNKRIG NGDVGW+EYLLL N +++ P V Q P+ FR +++Y+
Sbjct: 88 AGPPEPYGYGNKRIGPNGDVGWIEYLLLNANPQLSSPKTSAVFRQTPQIFRESVEEYMKE 147
Query: 140 VRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLI--GF 197
++ + ++L+++AE L I+P++ LSK+L D++SDS R+NHYPA E + E ++ GF
Sbjct: 148 IKEVSYKVLEMVAEELGIEPRDTLSKMLRDEKSDSCLRLNHYPAAEE---EAEKMVKVGF 204
Query: 198 GEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKH 257
GEHTDPQIIS+LRSNNT+GLQI ++DGSW++VPPD+SSFFINVGD+LQVMTNGRF+SVKH
Sbjct: 205 GEHTDPQIISVLRSNNTAGLQICVKDGSWVAVPPDHSSFFINVGDALQVMTNGRFKSVKH 264
Query: 258 RVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLAD 317
RVLA+ +SR+SMIYFGGPPLS+KIAPLP L+ +++ LYKEFTW +YK+S+Y S+L D
Sbjct: 265 RVLADTRRSRISMIYFGGPPLSQKIAPLPCLVP-EQDDWLYKEFTWSQYKSSAYKSKLGD 323
Query: 318 NRLGHFER 325
RLG FE+
Sbjct: 324 YRLGLFEK 331
>At2g34550 putative gibberellin 2-oxidase
Length = 335
Score = 342 bits (876), Expect = 2e-94
Identities = 175/329 (53%), Positives = 233/329 (70%), Gaps = 7/329 (2%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV++ + A+ + N P IPV+DL+ DAK+ IVKACEEFGFFKVINHGV
Sbjct: 1 MVIVLQPASFDSNLYVNPKCKPRPVLIPVIDLTDSDAKTQIVKACEEFGFFKVINHGVRP 60
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTN----QE 116
+ ++QLE EA+NFF++ + K+K G +PFGYG KRIG NGD+GW+EY+LL N
Sbjct: 61 DLLTQLEQEAINFFALHHSLKDKAGPPDPFGYGTKRIGPNGDLGWLEYILLNANLCLESH 120
Query: 117 VNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF 176
+ P FR +++Y+ ++ M + L+++ E LKI+PK LS+L+ K+SDS
Sbjct: 121 KTTAIFRHTPAIFREAVEEYIKEMKRMSSKFLEMVEEELKIEPKEKLSRLVKVKESDSCL 180
Query: 177 RVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSF 236
R+NHYP E + E IGFGEHTDPQ+ISLLRSN+T GLQI ++DG+W+ V PD+SSF
Sbjct: 181 RMNHYPEKEETPVKEE--IGFGEHTDPQLISLLRSNDTEGLQICVKDGTWVDVTPDHSSF 238
Query: 237 FINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES 296
F+ VGD+LQVMTNGRF+SVKHRV+ N +SR+SMIYF GPPLSEKIAPL S L +++
Sbjct: 239 FVLVGDTLQVMTNGRFKSVKHRVVTNTKRSRISMIYFAGPPLSEKIAPL-SCLVPKQDDC 297
Query: 297 LYKEFTWFEYKNSSYGSRLADNRLGHFER 325
LY EFTW +YK S+Y ++L D RLG FE+
Sbjct: 298 LYNEFTWSQYKLSAYKTKLGDYRLGLFEK 326
>At1g02400 unknown protein
Length = 329
Score = 275 bits (704), Expect = 2e-74
Identities = 148/319 (46%), Positives = 201/319 (62%), Gaps = 11/319 (3%)
Query: 16 KNCMATPCSFTIPVVDLSKPDAKSI---IVKACEEFGFFKVINHGVSMEAISQLESEALN 72
K +++P + PV+D S D + IVKACE GFFKVINHGV E I + E E
Sbjct: 14 KKTISSP-EYNFPVIDFSLNDRSKLSEKIVKACEVNGFFKVINHGVKPEIIKRFEHEGEE 72
Query: 73 FFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQ----EVNVPVHSQNPEK 128
FF+ P ++K + G A+PFGYG K IG NGD+G +EYLLL N + + + +P K
Sbjct: 73 FFNKPESDKLRAGPASPFGYGCKNIGFNGDLGELEYLLLHANPTAVADKSETISHDDPFK 132
Query: 129 FRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMG 188
F +DY+ VR++ CEI+DL E L Q + +S+L+ D +SDS+ R+NHYP P
Sbjct: 133 FSSATNDYIRTVRDLACEIIDLTIENLWGQKSSEVSELIRDVRSDSILRLNHYPPAP-YA 191
Query: 189 LDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMT 248
L G IGFGEH+DPQI+++LRSN+ GL+I RDG WI +P D + FF+ VGD LQ +T
Sbjct: 192 LSGVGQIGFGEHSDPQILTVLRSNDVDGLEICSRDGLWIPIPSDPTCFFVLVGDCLQALT 251
Query: 249 NGRFRSVKHRVLAN-GFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYK 307
NGRF SV+HRVLAN + R+S +YF PPL KI+PLP ++ + Y FTW +YK
Sbjct: 252 NGRFTSVRHRVLANTAKKPRMSAMYFAAPPLEAKISPLPKMV-SPENPRRYNSFTWGDYK 310
Query: 308 NSSYGSRLADNRLGHFERI 326
++Y RL RL F+ +
Sbjct: 311 KATYSLRLDVPRLEFFKTL 329
>At1g47990 dioxygenase, putative
Length = 321
Score = 273 bits (697), Expect = 1e-73
Identities = 151/308 (49%), Positives = 193/308 (62%), Gaps = 17/308 (5%)
Query: 27 IPVVDLS--KPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKT 84
IP++D+S + IVKACE GFFKVINHGV IS++E E++NFF+ P EK+
Sbjct: 15 IPIIDMSQERSQVSMQIVKACESLGFFKVINHGVDQTTISRMEQESINFFAKPAHEKKSV 74
Query: 85 GLAN-PFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVRNM 143
N PF YG + IG NGD G VEYLL TN ++ F ++ Y+ AV+ +
Sbjct: 75 RPVNQPFRYGFRDIGLNGDSGEVEYLLFHTNDPA-----FRSQLSFSSAVNCYIEAVKQL 129
Query: 144 GCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACP----EMGLDGENL----I 195
EILDL AEGL + P + S+L+ SDSV RVNHYP E L +++ +
Sbjct: 130 AREILDLTAEGLHVPPHS-FSRLISSVDSDSVLRVNHYPPSDQFFGEANLSDQSVSLTRV 188
Query: 196 GFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSV 255
GFGEHTDPQI+++LRSN GLQ+S DG W+SV PD S+F +NVGD LQVMTNGRF SV
Sbjct: 189 GFGEHTDPQILTVLRSNGVGGLQVSNSDGMWVSVSPDPSAFCVNVGDLLQVMTNGRFISV 248
Query: 256 KHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRL 315
+HR L G +SRLS YF GPPL KI PL +++ + LY+ FTW EYK +Y RL
Sbjct: 249 RHRALTYGEESRLSTAYFAGPPLQAKIGPLSAMVMTMNQPRLYQTFTWGEYKKRAYSLRL 308
Query: 316 ADNRLGHF 323
D+RL F
Sbjct: 309 EDSRLDMF 316
>At1g55290 leucoanthocyanidin dioxygenase 2, putative
Length = 361
Score = 157 bits (398), Expect = 6e-39
Identities = 99/296 (33%), Positives = 157/296 (52%), Gaps = 30/296 (10%)
Query: 26 TIPVVDLSKPDAKSI---IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKE 82
+IPV+D+S D KS+ + A EE+GFF+VINHGVSME + +++ FF +PV EK
Sbjct: 61 SIPVIDISNLDEKSVSKAVCDAAEEWGFFQVINHGVSMEVLENMKTATHRFFGLPVEEKR 120
Query: 83 K----TGLANPFGYGNKRIGH-NGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYL 137
K L+ +G H + W +YL L E P+ R +Y+
Sbjct: 121 KFSREKSLSTNVRFGTSFSPHAEKALEWKDYLSLFFVSEAEAS--QLWPDSCRSETLEYM 178
Query: 138 CAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF------RVNHYPACPEMGLDG 191
+ + ++L + E L ++ +DK +S F +N+YP CP +
Sbjct: 179 NETKPLVKKLLRFLGENLNVKE--------LDKTKESFFMGSTRINLNYYPICP----NP 226
Query: 192 ENLIGFGEHTDPQIISLLRSNNTSGLQI-SLRDGSWISVPPDYSSFFINVGDSLQVMTNG 250
E +G G H+D +++L + GL + SL G W+ VPP S IN+GD++Q+M+NG
Sbjct: 227 ELTVGVGRHSDVSSLTILLQDEIGGLHVRSLTTGRWVHVPPISGSLVINIGDAMQIMSNG 286
Query: 251 RFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
R++SV+HRVLANG +R+S+ F P I PL +++ G E+ +YK+ + +Y
Sbjct: 287 RYKSVEHRVLANGSYNRISVPIFVSPKPESVIGPLLEVIENG-EKPVYKDILYTDY 341
>At3g13610 unknown protein
Length = 361
Score = 156 bits (395), Expect = 1e-38
Identities = 96/311 (30%), Positives = 160/311 (50%), Gaps = 35/311 (11%)
Query: 27 IPVVDLSKPDAKSIIVKAC---EEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEK 83
IPV+D+S PD + C E++GFF+VINHGV +E + +++ FF++PV EK K
Sbjct: 62 IPVIDMSNPDEDRVAEAVCDAAEKWGFFQVINHGVPLEVLDDVKAATHKFFNLPVEEKRK 121
Query: 84 TGLANP------FGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYL 137
N FG + + W +YL L E P+ R +Y+
Sbjct: 122 FTKENSLSTTVRFGTSFSPLAEQA-LEWKDYLSLFFVSEAEAEQFW--PDICRNETLEYI 178
Query: 138 CAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF------RVNHYPACPEMGLDG 191
+ M +L+ + + L ++ +D+ +S+F +N+YP CP L
Sbjct: 179 NKSKKMVRRLLEYLGKNLNVKE--------LDETKESLFMGSIRVNLNYYPICPNPDLT- 229
Query: 192 ENLIGFGEHTDPQIISLLRSNNTSGLQI-SLRDGSWISVPPDYSSFFINVGDSLQVMTNG 250
+G G H+D +++L + GL + SL G+W+ VPP SF IN+GD++Q+M+NG
Sbjct: 230 ---VGVGRHSDVSSLTILLQDQIGGLHVRSLASGNWVHVPPVAGSFVINIGDAMQIMSNG 286
Query: 251 RFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKE----ESLYKEFTWFEY 306
++SV+HRVLANG+ +R+S+ F P I PLP ++ G+E + LY ++ + +
Sbjct: 287 LYKSVEHRVLANGYNNRISVPIFVNPKPESVIGPLPEVIANGEEPIYRDVLYSDYVKYFF 346
Query: 307 KNSSYGSRLAD 317
+ + G + D
Sbjct: 347 RKAHDGKKTVD 357
>At5g24530 flavanone 3-hydroxylase-like protein
Length = 341
Score = 149 bits (376), Expect = 2e-36
Identities = 105/309 (33%), Positives = 159/309 (50%), Gaps = 39/309 (12%)
Query: 28 PVVDLSKPDAKSIIVK---ACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEK- 83
P++DLS D +I + AC FGFF+VINHGV+ + I ++ S A FFSM + EK K
Sbjct: 39 PLIDLSSTDRSFLIQQIHQACARFGFFQVINHGVNKQIIDEMVSVAREFFSMSMEEKMKL 98
Query: 84 --------TGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHS------QNPEKF 129
T L+ F + + + W +YL L P+H NP F
Sbjct: 99 YSDDPTKTTRLSTSFNVKKEEVNN-----WRDYLRLHC-----YPIHKYVNEWPSNPPSF 148
Query: 130 RCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGL 189
+ ++ Y VR +G +I +L++E L ++ K+ + K+L ++ VN+YP CPE L
Sbjct: 149 KEIVSKYSREVREVGFKIEELISESLGLE-KDYMKKVLGEQGQHMA--VNYYPPCPEPEL 205
Query: 190 DGENLIGFGEHTDPQIISLLRSNNT-SGLQISLRDGSWISVPPDYSSFFINVGDSLQVMT 248
G HTDP +++L + T GLQI L DG W +V P +F IN+GD LQ ++
Sbjct: 206 T----YGLPAHTDPNALTILLQDTTVCGLQI-LIDGQWFAVNPHPDAFVINIGDQLQALS 260
Query: 249 NGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES--LYKEFTWFEY 306
NG ++SV HR + N RLS+ F P ++P L + +E+ +YK+FT+ EY
Sbjct: 261 NGVYKSVWHRAVTNTENPRLSVASFLCPADCAVMSPAKPLWEAEDDETKPVYKDFTYAEY 320
Query: 307 KNSSYGSRL 315
+ L
Sbjct: 321 YKKFWSRNL 329
>At1g80330 putative gibberellin 3 beta-hydroxylase
Length = 355
Score = 149 bits (376), Expect = 2e-36
Identities = 91/296 (30%), Positives = 150/296 (49%), Gaps = 15/296 (5%)
Query: 26 TIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTG 85
+IPV+DLS PD ++I A + +G F++ NHG+S + + +ES + F MP K +
Sbjct: 48 SIPVIDLSNPDVTTLIGDASKTWGAFQIANHGISQKLLDDIESLSKTLFDMPSERKLEAA 107
Query: 86 LANP--FGYGNKRIGHNGDVG-WVEYLLLKTNQEVNV--PVHSQNPEKFRCVLDDYLCAV 140
++ GYG RI + W E + + N + + K+ ++ +Y+ +
Sbjct: 108 SSDKGVSGYGEPRISPFFEKKMWSEGFTIADDSYRNHFNTLWPHDHTKYCGIIQEYVDEM 167
Query: 141 RNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV----FRVNHYPACPEMGLDGENLIG 196
+ +L + L + +++ ++K V R+NHYP CPE E +G
Sbjct: 168 EKLASRLLYCILGSLGVTVEDIEWAHKLEKSGSKVGRGAIRLNHYPVCPEP----ERAMG 223
Query: 197 FGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVK 256
HTD I+++L +NT GLQ+ + W++V P +N+GD +++NG+ SV
Sbjct: 224 LAAHTDSTILTILHQSNTGGLQVFREESGWVTVEPAPGVLVVNIGDLFHILSNGKIPSVV 283
Query: 257 HRVLANGFQSRLSMIY-FGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSY 311
HR N +SR+S+ Y +GGP +IAP+ S L G E SLY+ TW EY Y
Sbjct: 284 HRAKVNHTRSRISIAYLWGGPAGDVQIAPI-SKLTGPAEPSLYRSITWKEYLQIKY 338
>At5g63590 flavonol synthase
Length = 308
Score = 146 bits (369), Expect = 1e-35
Identities = 94/296 (31%), Positives = 158/296 (52%), Gaps = 20/296 (6%)
Query: 24 SFTIPVVDLSKPDAK---SIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTE 80
S IPV+DLS PD + S +VKA +E+G F+V+NHG+ E I +L + FF +P TE
Sbjct: 11 SLDIPVIDLSNPDEELVASAVVKASQEWGIFQVVNHGIPTELILRLLQVGMEFFELPETE 70
Query: 81 KEKTGLANPF------GYGNK-RIGHNGDVGWVEYLLLKT--NQEVNVPVHSQNPEKFRC 131
KE +A P GY K + G WV++L + VN +NP ++
Sbjct: 71 KE--AVAKPEDSLDIEGYRTKYQKDLEGRNAWVDHLFHRIWPPSRVNHKFWPKNPPEYIE 128
Query: 132 VLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDG 191
V ++Y ++ + +I++ ++EGL ++ + L + L + + + ++N+YP CP D
Sbjct: 129 VNEEYASHIKKLSEKIMEWLSEGLGLRHE-ALKEGLGGETIEYLMKINYYPPCP----DP 183
Query: 192 ENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGR 251
E ++G +HTD I+LL +N GLQ + +D WI S + +GD M+NG+
Sbjct: 184 ELVVGAPDHTDVNGITLLVANEALGLQ-AFKDNQWIDAEYTTSGIIVIIGDQFLRMSNGK 242
Query: 252 FRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYK 307
++SV+HR + ++R+S F L + PLP L+ G + +K + + +YK
Sbjct: 243 YKSVEHRAKMDKEKTRISWPVFVESSLDQVFGPLPELITGDENVPKFKPYVYKDYK 298
>At5g08640 flavonol synthase (FLS) (sp|Q96330)
Length = 336
Score = 145 bits (367), Expect = 2e-35
Identities = 94/292 (32%), Positives = 157/292 (53%), Gaps = 21/292 (7%)
Query: 27 IPVVDLSKPDAKSI---IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEK 83
IPVVDLS PD +S+ +VKA EE+G F+V+NHG+ E I +L+ FF +P +EKE
Sbjct: 43 IPVVDLSDPDEESVRRAVVKASEEWGLFQVVNHGIPTELIRRLQDVGRKFFELPSSEKE- 101
Query: 84 TGLANP------FGYGNK-RIGHNGDVGWVEYLL--LKTNQEVNVPVHSQNPEKFRCVLD 134
+A P GYG K + G WV++L + VN +NP ++R V +
Sbjct: 102 -SVAKPEDSKDIEGYGTKLQKDPEGKKAWVDHLFHRIWPPSCVNYRFWPKNPPEYREVNE 160
Query: 135 DYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENL 194
+Y V+ + +L ++++GL ++ ++ L + L + ++ + ++N+YP CP L
Sbjct: 161 EYAVHVKKLSETLLGILSDGLGLK-RDALKEGLGGEMAEYMMKINYYPPCPRPDL----A 215
Query: 195 IGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRS 254
+G HTD I+LL N GLQ+ +D W S+ +++GD + ++NGR+++
Sbjct: 216 LGVPAHTDLSGITLLVPNEVPGLQV-FKDDHWFDAEYIPSAVIVHIGDQILRLSNGRYKN 274
Query: 255 VKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
V HR + ++R+S F PP + + PLP L G +K F + +Y
Sbjct: 275 VLHRTTVDKEKTRMSWPVFLEPPREKIVGPLPE-LTGDDNPPKFKPFAFKDY 325
>At1g15550 Gibberillin 3 beta-hydroxylase (AtGA3ox1)
Length = 358
Score = 145 bits (365), Expect = 4e-35
Identities = 93/291 (31%), Positives = 152/291 (51%), Gaps = 21/291 (7%)
Query: 27 IPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGL 86
IP++DL PDA + I AC +G F++ NHGV + + +E + F +PV K K+
Sbjct: 57 IPLIDLDHPDATNQIGHACRTWGAFQISNHGVPLGLLQDIEFLTGSLFGLPVQRKLKSAR 116
Query: 87 ANP--FGYGNKRIGH--NGDVGWVEYLLLKTNQEVNV-----PVHSQNPEKFRCVLDDYL 137
+ GYG RI N + W E + T +N P H N + ++++Y
Sbjct: 117 SETGVSGYGVARIASFFNKQM-WSEGFTI-TGSPLNDFRKLWPQHHLN---YCDIVEEYE 171
Query: 138 CAVRNMGCEILDLMAEGLKIQPKNV-LSKLLMDKQ-SDSVFRVNHYPACPEMGLDGENLI 195
++ + +++ L L + +++ + L D + + ++NHYP CPE + +
Sbjct: 172 EHMKKLASKLMWLALNSLGVSEEDIEWASLSSDLNWAQAALQLNHYPVCPEP----DRAM 227
Query: 196 GFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSV 255
G HTD ++++L NNT+GLQ+ D W++VPP S +NVGD +++NG F+SV
Sbjct: 228 GLAAHTDSTLLTILYQNNTAGLQVFRDDLGWVTVPPFPGSLVVNVGDLFHILSNGLFKSV 287
Query: 256 KHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
HR N ++RLS+ + GP KI+P+P L+ E LY+ TW EY
Sbjct: 288 LHRARVNQTRARLSVAFLWGPQSDIKISPVPKLV-SPVESPLYQSVTWKEY 337
>At3g12900 hypothetical protein
Length = 357
Score = 140 bits (354), Expect = 7e-34
Identities = 91/304 (29%), Positives = 157/304 (50%), Gaps = 22/304 (7%)
Query: 20 ATPCSFTIPVVDLSK---PDAKSI---IVKACEEFGFFKVINHGVSMEAISQLESEALNF 73
A C T P+ DLS P K + IV+A E GFF+V+NHGVS+E + L+S A F
Sbjct: 50 ALTCEATQPI-DLSNLDGPQHKEVAKQIVEAAETLGFFQVVNHGVSVELLELLKSSAHEF 108
Query: 74 FSMPVTEK----EKTGLANPFGYGNKRIGHNGD-VGWVEYLLLKTNQEVNVPVHSQNPEK 128
F+ EK ++ + YG + + W +Y+ + + H P+
Sbjct: 109 FAQAPEEKSMYLKEVSPSKLVKYGTSFVPDKEKAIEWKDYVSMLYTNDSEALQHW--PQP 166
Query: 129 FRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMG 188
R V ++L + M +++++ E + + + LM + + +N+YP CP
Sbjct: 167 CREVALEFLNSSMEMVKNVVNILMENVGVTLEEEKMNGLMGTK---MVNMNYYPTCPSPE 223
Query: 189 LDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMT 248
L +G G H+D ++++L + GL + L +G W +PP + + IN+GD+LQ+++
Sbjct: 224 LT----VGVGRHSDMGMLTVLLQDGIGGLYVKLDNGEWAEIPPVHGALVINIGDTLQILS 279
Query: 249 NGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKN 308
NG+++S +HRV SR+S+ F P S+K+ PLP ++K + YKEF + +Y N
Sbjct: 280 NGKYKSAEHRVRTTNIGSRVSVPIFTAPNPSQKVGPLPEVVK-RDGVARYKEFLFQDYMN 338
Query: 309 SSYG 312
+ +G
Sbjct: 339 NFFG 342
>At1g80340 gibberellin 3 beta-hydroxylase
Length = 347
Score = 140 bits (353), Expect = 9e-34
Identities = 90/292 (30%), Positives = 147/292 (49%), Gaps = 21/292 (7%)
Query: 26 TIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTG 85
T+P++DLS +++ AC +G F++ NHGV + +E + F +PV K K
Sbjct: 49 TLPLIDLSDIHVATLVGHACTTWGAFQITNHGVPSRLLDDIEFLTGSLFRLPVQRKLKAA 108
Query: 86 LANP--FGYGNKRIGHNGDVG-WVEYLLLKTNQEVNVPVHS------QNPEKFRCVLDDY 136
+ GYG RI + W E + + P+H + K+ ++++Y
Sbjct: 109 RSENGVSGYGVARIASFFNKKMWSEGFTV-----IGSPLHDFRKLWPSHHLKYCEIIEEY 163
Query: 137 LCAVRNMGCEILDLMAEGLKIQPKNVL-SKLLMDKQ-SDSVFRVNHYPACPEMGLDGENL 194
++ + +++ L ++ K++ + D Q + +V ++NHYP CPE +
Sbjct: 164 EEHMQKLAAKLMWFALGSLGVEEKDIQWAGPNSDFQGTQAVIQLNHYPKCPEP----DRA 219
Query: 195 IGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRS 254
+G HTD ++++L NNT+GLQ+ D W++ PP S +NVGD L ++TNG F S
Sbjct: 220 MGLAAHTDSTLMTILYQNNTAGLQVFRDDVGWVTAPPVPGSLVVNVGDLLHILTNGIFPS 279
Query: 255 VKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
V HR N +SR SM Y GPP I+PLP L+ + LY TW +Y
Sbjct: 280 VLHRARVNHVRSRFSMAYLWGPPSDIMISPLPKLV-DPLQSPLYPSLTWKQY 330
>At1g78550 unknown protein
Length = 356
Score = 140 bits (352), Expect = 1e-33
Identities = 94/306 (30%), Positives = 165/306 (53%), Gaps = 24/306 (7%)
Query: 24 SFTIPVVDLSKPDAKSIIVK-------ACEEFGFFKVINHGVSMEAISQLESEALNFFSM 76
S IPV+D+++ + S + AC+++GFF+++NHG+ + +LE+E FF++
Sbjct: 50 SSEIPVIDMTRLCSVSAMDSELKKLDFACQDWGFFQLVNHGIDSSFLEKLETEVQEFFNL 109
Query: 77 PVTEKEK----TGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVH--SQNPEKFR 130
P+ EK+K +G FG N + N + W + +L T + H S+ P FR
Sbjct: 110 PMKEKQKLWQRSGEFEGFGQVNI-VSENQKLDWGDMFILTTEPIRSRKSHLFSKLPPPFR 168
Query: 131 CVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLD 190
L+ Y V+++ + MA L+I+ + + L D S+ ++N+YP CP+
Sbjct: 169 ETLETYSSEVKSIAKILFAKMASVLEIKHEEMED--LFDDVWQSI-KINYYPPCPQP--- 222
Query: 191 GENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTN 249
+ ++G +H+D ++ LL+ N GLQI +DG W+ V P + +NVG+ L+++TN
Sbjct: 223 -DQVMGLTQHSDAAGLTILLQVNQVEGLQIK-KDGKWVVVKPLRDALVVNVGEILEIITN 280
Query: 250 GRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNS 309
GR+RS++HR + N + RLS+ F P I P SL+ K + L+K + EY ++
Sbjct: 281 GRYRSIEHRAVVNSEKERLSVAMFHSPGKETIIRPAKSLVDRQK-QCLFKSMSTQEYFDA 339
Query: 310 SYGSRL 315
+ +L
Sbjct: 340 FFTQKL 345
>At2g38240 putative anthocyanidin synthase
Length = 353
Score = 139 bits (350), Expect = 2e-33
Identities = 85/304 (27%), Positives = 156/304 (50%), Gaps = 16/304 (5%)
Query: 15 VKNCMATPCSFTIPVVDLS----KPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEA 70
V N + IPV+D++ KP+ ++ ACEE+GFF+++NHGV+ + ++
Sbjct: 36 VFNTTQSDAGIEIPVLDMNDVWGKPEGLRLVRSACEEWGFFQMVNHGVTHSLMERVRGAW 95
Query: 71 LNFFSMPVTEKEKTGLANPF--GYGNKR-IGHNGDVGWVEYLLLK--TNQEVNVPVHSQN 125
FF +P+ EK K + GYG++ + + + W +Y L + N
Sbjct: 96 REFFELPLEEKRKYANSPDTYEGYGSRLGVVKDAKLDWSDYFFLNYLPSSIRNPSKWPSQ 155
Query: 126 PEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACP 185
P K R +++ Y VR + + + ++E L ++P ++ L + + R N YP CP
Sbjct: 156 PPKIRELIEKYGEEVRKLCERLTETLSESLGLKPNKLMQALGGGDKVGASLRTNFYPKCP 215
Query: 186 EMGLDGENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSL 244
+ L +G H+DP I+ LL +GLQ+ DG W+++ ++ +N+GD L
Sbjct: 216 QPQLT----LGLSSHSDPGGITILLPDEKVAGLQVRRGDG-WVTIKSVPNALIVNIGDQL 270
Query: 245 QVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWF 304
Q+++NG ++SV+H+V+ N R+S+ +F P + P+ L+ + +LYK +
Sbjct: 271 QILSNGIYKSVEHQVIVNSGMERVSLAFFYNPRSDIPVGPIEELVTANR-PALYKPIRFD 329
Query: 305 EYKN 308
EY++
Sbjct: 330 EYRS 333
>At1g17020 SRG1-like protein
Length = 358
Score = 138 bits (348), Expect = 4e-33
Identities = 96/307 (31%), Positives = 167/307 (54%), Gaps = 23/307 (7%)
Query: 27 IPVVDL----SKPDAKSIIVK---ACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVT 79
IP++D+ S S + K AC+E+GFF+++NHG+ + +++SE +FF++P+
Sbjct: 53 IPIIDMKRLCSSTTMDSEVEKLDFACKEWGFFQLVNHGIDSSFLDKVKSEIQDFFNLPME 112
Query: 80 EKEKTGLANPF--GYGNKRI-GHNGDVGWVEYLLLKTNQEVNVP---VHSQNPEKFRCVL 133
EK+K G+G + + + W + L T Q V + + + P FR L
Sbjct: 113 EKKKFWQRPDEIEGFGQAFVVSEDQKLDWAD-LFFHTVQPVELRKPHLFPKLPLPFRDTL 171
Query: 134 DDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGEN 193
+ Y V+++ ++ MA L+I+P+ L KL D S R+N+YP CP+ +
Sbjct: 172 EMYSSEVQSVAKILIAKMARALEIKPEE-LEKLFDDVDSVQSMRMNYYPPCPQP----DQ 226
Query: 194 LIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRF 252
+IG H+D ++ L++ N+ GLQI +DG W+ V P ++F +N+GD L+++TNG +
Sbjct: 227 VIGLTPHSDSVGLTVLMQVNDVEGLQIK-KDGKWVPVKPLPNAFIVNIGDVLEIITNGTY 285
Query: 253 RSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYG 312
RS++HR + N + RLS+ F + +++ P SL++ K + +K T EY N
Sbjct: 286 RSIEHRGVVNSEKERLSIATFHNVGMYKEVGPAKSLVERQK-VARFKRLTMKEY-NDGLF 343
Query: 313 SRLADNR 319
SR D +
Sbjct: 344 SRTLDGK 350
>At4g25300 SRG1-like protein
Length = 356
Score = 138 bits (347), Expect = 5e-33
Identities = 96/304 (31%), Positives = 165/304 (53%), Gaps = 22/304 (7%)
Query: 19 MATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPV 78
M+ CS T ++ K D+ AC+E+GFF+++NHG+ +++++SE +FF++P+
Sbjct: 57 MSLLCSSTSMDSEIDKLDS------ACKEWGFFQLVNHGMESSFLNKVKSEVQDFFNLPM 110
Query: 79 TEKEKTGLANPF--GYGNKRI-GHNGDVGWVEYLLLKTNQEVNVP---VHSQNPEKFRCV 132
EK+ G+G + + W + L T Q V + + + P FR
Sbjct: 111 EEKKNLWQQPDEIEGFGQVFVVSEEQKLDWADMFFL-TMQPVRLRKPHLFPKLPLPFRDT 169
Query: 133 LDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGE 192
LD Y V+++ +L +A LKI+P+ + KL D+ + R+N+YP CPE +
Sbjct: 170 LDMYSAEVKSIAKILLGKIAVALKIKPEE-MDKLFDDELGQRI-RLNYYPRCPEP----D 223
Query: 193 NLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGR 251
+IG H+D ++ LL++N GLQI ++ W+SV P ++ +NVGD L+++TNG
Sbjct: 224 KVIGLTPHSDSTGLTILLQANEVEGLQIK-KNAKWVSVKPLPNALVVNVGDILEIITNGT 282
Query: 252 FRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSY 311
+RS++HR + N + RLS+ F L ++I P+ SL++ K + +K T EY N +
Sbjct: 283 YRSIEHRGVVNSEKERLSVAAFHNIGLGKEIGPMRSLVERHK-AAFFKSVTTEEYFNGLF 341
Query: 312 GSRL 315
L
Sbjct: 342 SREL 345
>At3g21420 unknown protein
Length = 364
Score = 136 bits (342), Expect = 2e-32
Identities = 91/316 (28%), Positives = 159/316 (49%), Gaps = 24/316 (7%)
Query: 27 IPVVDLSK---PDAKSI------IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMP 77
IPV+DLSK PD + +ACE++GFF+VINHG+ +E + +E A FF MP
Sbjct: 55 IPVIDLSKLSKPDNDDFFFEILKLSQACEDWGFFQVINHGIEVEVVEDIEEVASEFFDMP 114
Query: 78 VTEKEKTGL--ANPFGYGNKRI-GHNGDVGWVEYLLLKTN--QEVNVPVHSQNPEKFRCV 132
+ EK+K + GYG I + + W L + Q N + P +F
Sbjct: 115 LEEKKKYPMEPGTVQGYGQAFIFSEDQKLDWCNMFALGVHPPQIRNPKLWPSKPARFSES 174
Query: 133 LDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGE 192
L+ Y +R + +L +A L ++ + M ++ R+N+YP C L
Sbjct: 175 LEGYSKEIRELCKRLLKYIAISLGLKEERFEE---MFGEAVQAVRMNYYPPCSSPDL--- 228
Query: 193 NLIGFGEHTDPQIISLLRSNNTS--GLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNG 250
++G H+D +++L+ + S GLQI L+D +W+ V P ++ IN+GD+++V++NG
Sbjct: 229 -VLGLSPHSDGSALTVLQQSKNSCVGLQI-LKDNTWVPVKPLPNALVINIGDTIEVLSNG 286
Query: 251 RFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSS 310
+++SV+HR + N + RL+++ F P +I P+ L+ Y+ + +Y
Sbjct: 287 KYKSVEHRAVTNREKERLTIVTFYAPNYEVEIEPMSELVDDETNPCKYRSYNHGDYSYHY 346
Query: 311 YGSRLADNRLGHFERI 326
++L + F +I
Sbjct: 347 VSNKLQGKKSLDFAKI 362
>At1g17010 SRG1-like protein
Length = 361
Score = 136 bits (342), Expect = 2e-32
Identities = 88/302 (29%), Positives = 166/302 (54%), Gaps = 21/302 (6%)
Query: 27 IPVVDLSKPDAKSIIVK-------ACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVT 79
IP++D+++ + + + AC+E+GFF+++NHG+ + +++SE +FF++P+
Sbjct: 54 IPIIDMNRLCSSTAVDSEVEKLDFACKEYGFFQLVNHGIDPSFLDKIKSEIQDFFNLPME 113
Query: 80 EKEKTGLANPF--GYGNKRI-GHNGDVGWVE--YLLLKTNQEVNVPVHSQNPEKFRCVLD 134
EK+K G+G + + + W + +L+++ Q + + P FR LD
Sbjct: 114 EKKKLWQTPAVMEGFGQAFVVSEDQKLDWADLFFLIMQPVQLRKRHLFPKLPLPFRDTLD 173
Query: 135 DYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENL 194
Y V+++ +L MA+ L+I+P+ V ++ D S+ R+N+YP CP+ L +
Sbjct: 174 MYSTRVKSIAKILLAKMAKALQIKPEEV-EEIFGDDMMQSM-RMNYYPPCPQPNL----V 227
Query: 195 IGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFR 253
G H+D ++ LL+ N GLQI ++G W V P ++F +NVGD L+++TNG +R
Sbjct: 228 TGLIPHSDAVGLTILLQVNEVDGLQIK-KNGKWFFVKPLQNAFIVNVGDVLEIITNGTYR 286
Query: 254 SVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGS 313
S++HR + N + RLS+ F + ++I P SL++ +E + ++ +Y N +
Sbjct: 287 SIEHRAMVNLEKERLSIATFHNTGMDKEIGPARSLVQ-RQEAAKFRSLKTKDYLNGLFSR 345
Query: 314 RL 315
L
Sbjct: 346 EL 347
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,731,906
Number of Sequences: 26719
Number of extensions: 338403
Number of successful extensions: 1068
Number of sequences better than 10.0: 119
Number of HSP's better than 10.0 without gapping: 111
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 740
Number of HSP's gapped (non-prelim): 135
length of query: 329
length of database: 11,318,596
effective HSP length: 100
effective length of query: 229
effective length of database: 8,646,696
effective search space: 1980093384
effective search space used: 1980093384
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0190.3


