

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0185.17
(469 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
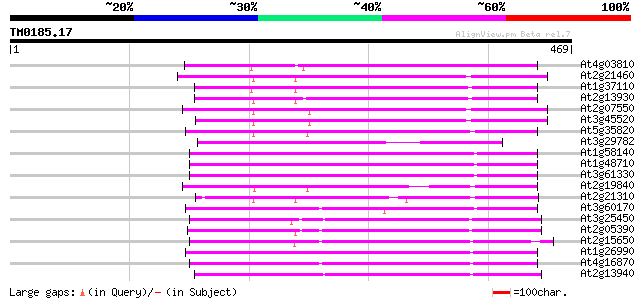
Score E
Sequences producing significant alignments: (bits) Value
At4g03810 putative retrotransposon protein 207 8e-54
At2g21460 putative retroelement pol polyprotein 188 6e-48
At1g37110 181 8e-46
At2g13930 putative retroelement pol polyprotein 179 4e-45
At2g07550 putative retroelement pol polyprotein 178 6e-45
At3g45520 copia-like polyprotein 175 4e-44
At5g35820 copia-like retrotransposable element 174 9e-44
At3g29782 hypothetical protein 172 4e-43
At1g58140 hypothetical protein 169 2e-42
At1g48710 hypothetical protein 169 2e-42
At3g61330 copia-type polyprotein 169 4e-42
At2g19840 copia-like retroelement pol polyprotein 164 1e-40
At2g21310 putative retroelement pol polyprotein 162 4e-40
At3g60170 putative protein 160 2e-39
At3g25450 hypothetical protein 158 7e-39
At2g05390 putative retroelement pol polyprotein 153 2e-37
At2g15650 putative retroelement pol polyprotein 150 1e-36
At1g26990 polyprotein, putative 150 1e-36
At4g16870 retrotransposon like protein 149 4e-36
At2g13940 putative retroelement pol polyprotein 149 4e-36
>At4g03810 putative retrotransposon protein
Length = 964
Score = 207 bits (528), Expect = 8e-54
Identities = 122/307 (39%), Positives = 177/307 (56%), Gaps = 13/307 (4%)
Query: 147 SGKAV-MTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRD--GEI 203
SG AV LYVDD+L+ G D+ ++ K++L + F MKD+GEA ILGI+I RD +I
Sbjct: 628 SGSAVAFLVLYVDDILLLGNDIPLLQSVKTWLGSCFSMKDMGEAAYILGIRIYRDRLNKI 687
Query: 204 IGLSQSHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCP--------VAQLEYSKAIR 255
IGLSQ YI+KVL +F+ D K P L +CP ++++ Y+ AI
Sbjct: 688 IGLSQDTYIDKVLHRFNMHDSKKGFIPMSHGITLSK-TQCPSTHDERERMSKIPYASAIG 746
Query: 256 CLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPS-VL 314
+MYAM TRPD+A + SRY +P + HW VV + KYL T + L+Y G V+
Sbjct: 747 SIMYAMLYTRPDVACALSMTSRYQSDPGESHWIVVRNIFKYLRRTKDKFLVYGGSEELVV 806
Query: 315 EGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV 374
GYT AS+ T D S SG F L GGAVSW S KQ+ +ADST AE+IA + +KE V
Sbjct: 807 SGYTDASFQTDKDDFRSQSGFFFCLNGGAVSWKSTKQSTVADSTTEAEYIAASEAAKEVV 866
Query: 375 *LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVI 434
+R + E+ + P + + ++CD+ +++A + KS+HI R+ ++++I G +
Sbjct: 867 WIRKFITELGVVPSISGPIDLYCDNNGAIAQAKEPKSHQKSKHIQRRYHLIREIIDRGDV 926
Query: 435 SIVFVRT 441
I V T
Sbjct: 927 KISRVST 933
>At2g21460 putative retroelement pol polyprotein
Length = 1333
Score = 188 bits (477), Expect = 6e-48
Identities = 112/319 (35%), Positives = 175/319 (54%), Gaps = 13/319 (4%)
Query: 141 YL**IPSGKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRD 200
Y+ + G V LYVDDML+ + ++ + K LS FDMKDLG A ILG++IIR+
Sbjct: 997 YIKELSDGSRVYLLLYVDDMLVAAKNKEDISQLKEELSQRFDMKDLGAAKRILGMEIIRN 1056
Query: 201 GE--IIGLSQSHYIEKVLKKFDHFDCKSVSTPFDQNTKL-------QPHKRCPVAQLEYS 251
E + LSQ+ Y+ K+L+ ++ + K V TP + K+ Q + + YS
Sbjct: 1057 REENTLWLSQNGYLNKILETYNMAESKHVVTPLGAHLKMRAATVEKQEQDEDYMKSIPYS 1116
Query: 252 KAIRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYP 311
A+ +MYAM TRPD+AY VG +SRY P+++HW V VL+Y+ G++ L Y
Sbjct: 1117 SAVGSIMYAMIGTRPDLAYPVGIISRYMSQPAREHWLGVKWVLRYIKGSLGTKLQYKRSS 1176
Query: 312 SV-LEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACS 370
+ GY A C S +G +F LGG +SW S +Q +A ST AE+++L
Sbjct: 1177 DFKVVGYCDADHAACKDRRRSITGLVFTLGGSTISWKSGQQRVVALSTTEAEYMSLTEAV 1236
Query: 371 KETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLIT 430
KE V ++ LL E + + V I CDSQ ++ + + V++ +++HI +R+ +++D+I
Sbjct: 1237 KEAVWMKGLLKE---FGYEQKSVEIFCDSQSAIALSKNNVHHERTKHIDVRYQYIRDIIA 1293
Query: 431 NGVISIVFVRTVKESCRSF 449
NG +V + T K F
Sbjct: 1294 NGDGDVVKIDTEKNPADIF 1312
>At1g37110
Length = 1356
Score = 181 bits (459), Expect = 8e-46
Identities = 104/294 (35%), Positives = 164/294 (55%), Gaps = 10/294 (3%)
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRD--GEIIGLSQSHYI 212
LYVDDMLI G E+ + K LS F+MKD+G A ILGI I RD G ++ LSQ YI
Sbjct: 1036 LYVDDMLIAGASKAEINRVKEQLSTEFEMKDMGGASRILGIDIYRDRKGGVLKLSQEIYI 1095
Query: 213 EKVLKKFDHFDCKSVSTPFDQNTKL----QPHKRCPVAQLEYSKAIRCLMYAMTCTRPDI 268
KVL +F+ K + P + KL + + + YS A+ +MYAM TRPD+
Sbjct: 1096 RKVLDRFNMSGAKMTNAPVGAHFKLAAVREEDECVDTDVVPYSSAVGSIMYAMLGTRPDL 1155
Query: 269 AYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPS-VLEGYTYASWVTCVK 327
AY + +SRY P HW V V++YL G + +L++ + GY +++ +
Sbjct: 1156 AYAICLISRYMSKPGSMHWEAVKWVMRYLKGAQDLNLVFTKEKDFTVTGYCDSNYAADLD 1215
Query: 328 DHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWP 387
S SG +F +GG VSW + Q +A ST AE+IALA +KE + ++ LL ++
Sbjct: 1216 RRRSISGYVFTIGGNTVSWKASLQPVVAMSTTEAEYIALAEAAKEAMWIKGLLQDM---G 1272
Query: 388 KLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
+V I CDSQ + + + VY+ +++HI +R ++++D++ +G + ++ + T
Sbjct: 1273 MQQDKVKIWCDSQSAICLSKNSVYHERTKHIDVRFNYIRDVVESGDVDVLKIHT 1326
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 179 bits (453), Expect = 4e-45
Identities = 107/299 (35%), Positives = 167/299 (55%), Gaps = 17/299 (5%)
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGE--IIGLSQSHYI 212
LYVDDMLI + +EV + K LS F+MKDLG+A ILG++I RD + ++ LSQ Y+
Sbjct: 1013 LYVDDMLIASANKSEVNELKQLLSREFEMKDLGDAKKILGMEISRDRDAGLLTLSQEGYV 1072
Query: 213 EKVLKKFDHFDCKSVSTPFDQNTKL---------QPHKRCPVAQLEYSKAIRCLMYAMTC 263
+KVL+ F + K VSTP + KL + +R + + Y+ I +MY+M
Sbjct: 1073 KKVLRSFQMDNAKPVSTPLGIHFKLKAATDKEYEEQFERMKI--VPYANTIGSIMYSMIG 1130
Query: 264 TRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPS-VLEGYTYASW 322
TRPD+AY +G +SR+ P KDHW V VL+Y+ GT L + +L GY + +
Sbjct: 1131 TRPDLAYSLGVISRFMSKPLKDHWQAVKWVLRYMRGTEKKKLCFRKQEDFLLRGYCDSDY 1190
Query: 323 VTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYE 382
+ S +G +F +GG +SW SK Q +A S+ AE++AL KE + L+ E
Sbjct: 1191 GSNFDTRRSITGYVFTVGGNTISWKSKLQKVVAISSTEAEYMALTEAVKEALWLKGFAAE 1250
Query: 383 ILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
+ V +H DSQ ++ A + V++ +++HI +R ++D+I G+I +V + T
Sbjct: 1251 L---GHSQDYVEVHSDSQSAITLAKNSVHHERTKHIDIRLHFIRDIICAGLIKVVKIAT 1306
>At2g07550 putative retroelement pol polyprotein
Length = 1356
Score = 178 bits (451), Expect = 6e-45
Identities = 102/315 (32%), Positives = 171/315 (53%), Gaps = 13/315 (4%)
Query: 145 IPSGKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGE-- 202
+P G + +YVDD+L+ + + K+ L F+MKDLG A ILG++IIRD
Sbjct: 1024 LPDGSVMYLLIYVDDILVASKNKEAITALKANLGMRFEMKDLGAAKKILGMEIIRDRTLG 1083
Query: 203 IIGLSQSHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLE-------YSKAIR 255
++ LSQ Y+ K+L+ ++ + K TP + K Q + + E YS A+
Sbjct: 1084 VLWLSQEGYLNKILETYNMAEAKPAMTPLGAHFKFQAATEQKLIRDEDFMKSVPYSSAVG 1143
Query: 256 CLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-L 314
+MYAM TRPD+AY VG +SR+ P K+HW V VL+Y+ GT+ L Y S +
Sbjct: 1144 SIMYAMLGTRPDLAYPVGIISRFMSQPIKEHWLGVKWVLRYIKGTLKTRLCYKKSSSFSI 1203
Query: 315 EGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV 374
GY A + + S +G +F LGG +SW S Q +A ST +E+++L KE +
Sbjct: 1204 VGYCDADYAADLDKRRSITGLVFTLGGNTISWKSGLQRVVAQSTTESEYMSLTEAVKEAI 1263
Query: 375 *LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVI 434
L+ LL + + + V I CDSQ ++ + + V++ +++HI +++ ++++I++G +
Sbjct: 1264 WLKGLLKD---FGYEQKSVEIFCDSQSAIALSKNNVHHERTKHIDVKYHFIREIISDGTV 1320
Query: 435 SIVFVRTVKESCRSF 449
++ + T K F
Sbjct: 1321 EVLKISTEKNPADIF 1335
>At3g45520 copia-like polyprotein
Length = 1363
Score = 175 bits (444), Expect = 4e-44
Identities = 103/304 (33%), Positives = 166/304 (53%), Gaps = 13/304 (4%)
Query: 156 YVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGE--IIGLSQSHYIE 213
YVDDML+ ++ ++ K LS F+MKDLG A ILGI+II D E ++ LSQ Y+
Sbjct: 1042 YVDDMLVAANNMQAIDALKKELSIKFEMKDLGAAKKILGIEIIIDREAGVLWLSQESYLN 1101
Query: 214 KVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLE-------YSKAIRCLMYAMTCTRP 266
KVLK F+ + K TP + K++ ++ E YS A+ +MYAM TRP
Sbjct: 1102 KVLKTFNMLESKPALTPLGAHLKMKSATEEKLSTEEEYMNSVPYSSAVGSIMYAMIGTRP 1161
Query: 267 DIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTYASWVTC 325
D+AY VG +SR+ P+K+HW V VL+Y+ GT++ L Y + GY A +
Sbjct: 1162 DLAYPVGVVSRFMSQPAKEHWLGVKWVLRYIKGTVDTRLCYKRNSDFSICGYCDADYAAD 1221
Query: 326 VKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILI 385
+ S +G +F LGG +SW S Q +A S+ E+++L KE + L+ LL +
Sbjct: 1222 LDKRRSITGLVFTLGGNTISWKSGLQRVVAQSSTECEYMSLTEAVKEAIWLKGLLKD--- 1278
Query: 386 WPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKES 445
+ + V I CDSQ ++ + + V++ +++HI ++ ++++I +G + + + T K
Sbjct: 1279 FGYEQKNVEIFCDSQSAIALSKNNVHHERTKHIDVKFHFIREIIADGKVEVSKISTEKNP 1338
Query: 446 CRSF 449
F
Sbjct: 1339 ADIF 1342
>At5g35820 copia-like retrotransposable element
Length = 1342
Score = 174 bits (441), Expect = 9e-44
Identities = 106/304 (34%), Positives = 167/304 (54%), Gaps = 13/304 (4%)
Query: 148 GKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGE--IIG 205
G + LYVDDMLI + ++++K K L+ F+MKDLG A ILG++I R+ E I+
Sbjct: 1013 GSYIYLLLYVDDMLIASQNKDQIQKLKESLNREFEMKDLGPARKILGMEITRNREQGILD 1072
Query: 206 LSQSHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQ-------LEYSKAIRCLM 258
LSQS Y+ VL+ F K TP + KL+ +A+ + Y AI +M
Sbjct: 1073 LSQSEYVAGVLRAFGMDQSKVSQTPLGAHFKLRAANEKTLARDAEYMKLVPYPNAIGSIM 1132
Query: 259 YAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGY 317
Y+M +RPD+AY VG +SR+ PSK+HW V V++Y+ GT + L + + GY
Sbjct: 1133 YSMIGSRPDLAYPVGVVSRFMSKPSKEHWQAVKWVMRYMKGTQDTCLRFKKDDKFEIRGY 1192
Query: 318 TYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LR 377
+ + T + S +G +F GG +SW S Q +A ST AE++ALA KE + LR
Sbjct: 1193 CDSDYATDLDRRRSITGFVFTAGGNTISWKSGLQRVVALSTTEAEYMALAEAVKEAIWLR 1252
Query: 378 NLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIV 437
L E+ V + CDSQ ++ + + V++ +++HI +R+ +++ I +G I +V
Sbjct: 1253 GLAAEMGF---EQDAVEVMCDSQSAIALSKNSVHHERTKHIDVRYHFIREKIADGEIQVV 1309
Query: 438 FVRT 441
+ T
Sbjct: 1310 KIST 1313
>At3g29782 hypothetical protein
Length = 1130
Score = 172 bits (436), Expect = 4e-43
Identities = 99/255 (38%), Positives = 140/255 (54%), Gaps = 27/255 (10%)
Query: 158 DDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSHYIEKVLK 217
+DMLI G++ + + ++K+ L F+MK++G ADVILGIKIIR E I LSQ+HY EK+L
Sbjct: 873 NDMLILGSNTDIINQSKNMLKRYFEMKNMGLADVILGIKIIRTDEGIILSQTHYDEKILD 932
Query: 218 KFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPDIAYVVGRLSR 277
+F H+ + P D L + PV Q++Y++ + LMY TRPD+ + V LSR
Sbjct: 933 RFKHYSNGTAKNPVDPQLHLTKNSGEPVQQVKYARVVGSLMYLTNSTRPDLVHSVNVLSR 992
Query: 278 YTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSVLEGYTYASWVTCVKDHASTSG*IF 337
YT N +H + RV+ Y+ T ++ L Y P+VLE
Sbjct: 993 YTSNQGYNHCKAITRVMNYVCYTKDHGLHYGKEPAVLE---------------------- 1030
Query: 338 NLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMRQVSIHC 397
AVSW S KQT A STM +EFIAL + E LRN L +I +W + + +HC
Sbjct: 1031 -----AVSWKSSKQTVAAKSTMESEFIALDTTAAEAEWLRNFLEDIPMWGIPVPAIRVHC 1085
Query: 398 DSQCTLSKAYSQVYN 412
DSQ + +A S +YN
Sbjct: 1086 DSQSVIGRAQSTLYN 1100
>At1g58140 hypothetical protein
Length = 1320
Score = 169 bits (429), Expect = 2e-42
Identities = 94/292 (32%), Positives = 162/292 (55%), Gaps = 3/292 (1%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSH 210
++ CLYVDD++ G + + E+ K ++ F+M D+G LGI++ ++ I ++Q
Sbjct: 994 LIACLYVDDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEG 1053
Query: 211 YIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPDIAY 270
Y ++VLKKF D V TP + KL + K++ + +TCTRPDI Y
Sbjct: 1054 YAKEVLKKFKMDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILY 1113
Query: 271 VVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTYASWVTCVKDH 329
VG +SRY +P+ H+ R+L+Y+ GT+N+ L Y+ L GY+ + W V D
Sbjct: 1114 AVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDR 1173
Query: 330 ASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKL 389
STSG +F +G A +W SKKQ + ST AE++A +C + LRNLL E+ + +
Sbjct: 1174 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE- 1232
Query: 390 MRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
I D++ ++ A + V++ +S+HI R+ ++++ ++ + + +V+T
Sbjct: 1233 -EPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKT 1283
>At1g48710 hypothetical protein
Length = 1352
Score = 169 bits (429), Expect = 2e-42
Identities = 94/292 (32%), Positives = 162/292 (55%), Gaps = 3/292 (1%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSH 210
++ CLYVDD++ G + + E+ K ++ F+M D+G LGI++ ++ I ++Q
Sbjct: 1026 LIACLYVDDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEG 1085
Query: 211 YIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPDIAY 270
Y ++VLKKF D V TP + KL + K++ + +TCTRPDI Y
Sbjct: 1086 YAKEVLKKFKMDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILY 1145
Query: 271 VVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTYASWVTCVKDH 329
VG +SRY +P+ H+ R+L+Y+ GT+N+ L Y+ L GY+ + W V D
Sbjct: 1146 AVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDR 1205
Query: 330 ASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKL 389
STSG +F +G A +W SKKQ + ST AE++A +C + LRNLL E+ + +
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVVLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE- 1264
Query: 390 MRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
I D++ ++ A + V++ +S+HI R+ ++++ ++ + + +V+T
Sbjct: 1265 -EPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKT 1315
>At3g61330 copia-type polyprotein
Length = 1352
Score = 169 bits (427), Expect = 4e-42
Identities = 94/292 (32%), Positives = 162/292 (55%), Gaps = 3/292 (1%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSH 210
++ CLYVDD++ G + + E+ K ++ F+M D+G LGI++ ++ I ++Q
Sbjct: 1026 LIACLYVDDLIFTGNNPSIFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEG 1085
Query: 211 YIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPDIAY 270
Y ++VLKKF D V TP + KL + K++ + +TCTRPDI Y
Sbjct: 1086 YAKEVLKKFKIDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILY 1145
Query: 271 VVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTYASWVTCVKDH 329
VG +SRY +P+ H+ R+L+Y+ GT+N+ L Y+ L GY+ + W V D
Sbjct: 1146 AVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDR 1205
Query: 330 ASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKL 389
STSG +F +G A +W SKKQ + ST AE++A +C + LRNLL E+ + +
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE- 1264
Query: 390 MRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
I D++ ++ A + V++ +S+HI R+ ++++ ++ + + +V+T
Sbjct: 1265 -EPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKT 1315
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 164 bits (414), Expect = 1e-40
Identities = 102/307 (33%), Positives = 167/307 (54%), Gaps = 29/307 (9%)
Query: 145 IPSGKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEI- 203
+ SG+ + LYVDD+LI D V K+ L++ F+MKDLG+A ILG++I+RD +
Sbjct: 821 LQSGEYIYLLLYVDDILIASRDKRTVCDLKALLNSEFEMKDLGDAKKILGMEIVRDRKAG 880
Query: 204 -IGLSQSHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQ-------LEYSKAIR 255
+ +SQ Y+ KVL F K V TP + KL+P V + + Y A+
Sbjct: 881 TMSISQEGYLLKVLGNFGMDQAKPVFTPMGAHFKLKPATDEEVMRQSEVMRAVPYQSAVG 940
Query: 256 CLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIY-NGYPSVL 314
LMY+M TRPD+A+ VG + R+ P K+HW V +L+Y+ G+I+ L Y N +L
Sbjct: 941 SLMYSMIGTRPDLAHSVGLVCRFMSKPLKEHWQAVKWILRYIRGSIDRKLCYKNEGELIL 1000
Query: 315 EGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV 374
EGY + + + STSG +A S+ AE++AL +KE +
Sbjct: 1001 EGYCDSDYAADKEGRRSTSG----------------VKVVALSSTEAEYMALTDGAKEAI 1044
Query: 375 *LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVI 434
L+ + E+ + + V+IHCDSQ ++ A + VY+ +++HI +++ ++DL+ NG +
Sbjct: 1045 WLKGHVSELGF---VQKTVNIHCDSQSAIALAKNAVYHERTKHIDVKYHFIRDLVNNGEV 1101
Query: 435 SIVFVRT 441
++ + T
Sbjct: 1102 QVLKIDT 1108
>At2g21310 putative retroelement pol polyprotein
Length = 838
Score = 162 bits (410), Expect = 4e-40
Identities = 106/304 (34%), Positives = 167/304 (54%), Gaps = 29/304 (9%)
Query: 156 YVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGE--IIGLSQSHYIE 213
++DD+ + V++ K LS+ F+MKDLG A ILG++I R+ E I+ LSQ Y++
Sbjct: 519 FIDDLT--RKNKETVKELKERLSSEFEMKDLGPARKILGMEITRNREENILELSQRSYLQ 576
Query: 214 KVLKKFDHFDCKSVSTPFDQNTKL-------QPHKRCPVAQLEYSKAIRCLMYAMTCTRP 266
KVLK F +CK V TP + K + + + Y+ A+ +MY+M +RP
Sbjct: 577 KVLKTFRMDECKPVKTPLAPHMKFVAATETEAEEQADQMKSIPYANAVGSIMYSMIGSRP 636
Query: 267 DIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPS-VLEGYTYASWVTC 325
D+A+ VG +SR+ P +HW V VL+Y+ G+I+ L Y V+ GY C
Sbjct: 637 DLAHPVGVISRFMSKPLMNHWLGVKWVLRYIKGSIDSRLQYKREGDFVVTGY-------C 689
Query: 326 VKDHA-------STSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRN 378
DH+ ST+G F +GG VSW S Q +A S+ AE+IAL +KE L++
Sbjct: 690 DSDHSGDRDRSRSTTGYTFTVGGNIVSWRSCLQPVVALSSTEAEYIALTEAAKEAYWLKD 749
Query: 379 LLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVF 438
L+ E+ V IHCDSQ ++ A + V++ +++HI R +++D IT+G + ++
Sbjct: 750 LMNELGF---KQGAVDIHCDSQSAIALAKNAVHHERTKHIQRRFHYIRDAITDGEVKVLK 806
Query: 439 VRTV 442
+ TV
Sbjct: 807 ISTV 810
>At3g60170 putative protein
Length = 1339
Score = 160 bits (404), Expect = 2e-39
Identities = 96/298 (32%), Positives = 156/298 (52%), Gaps = 7/298 (2%)
Query: 148 GKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLS 207
G ++ LYVDD++ G+D ++ K + F+M DLG+ LGI++ + I +
Sbjct: 1012 GNILIVSLYVDDLIFTGSDKAMCDEFKKSMMLEFEMSDLGKMKHFLGIEVKQSDGGIFIC 1071
Query: 208 QSHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCP-VAQLEYSKAIRCLMYAMTCTRP 266
Q Y +VL +F + +V P TKL + V + + + + LMY +T TRP
Sbjct: 1072 QRRYAREVLARFGMDESNAVKNPIVPGTKLTKDENGEKVDETMFKQLVGSLMY-LTVTRP 1130
Query: 267 DIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPS---VLEGYTYASWV 323
D+ Y V +SR+ NP HW R+L+YL GT+ + Y + L +T + +
Sbjct: 1131 DLMYGVCLISRFMSNPRMSHWLAAKRILRYLKGTVELGIFYRRRKNRSLKLMAFTDSDYA 1190
Query: 324 TCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEI 383
+ D STSG +F + GA+ W SKKQ +A ST AE+IA A C+ + V LR +L ++
Sbjct: 1191 GDLNDRRSTSGFVFLMASGAICWASKKQPVVALSTTEAEYIAAAFCACQCVWLRKVLEKL 1250
Query: 384 LIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
K I+CD+ T+ + V +GKS+HI +R +++DL+ V+ + + T
Sbjct: 1251 GAEEK--SATVINCDNSSTIQLSKHPVLHGKSKHIEVRFHYLRDLVNGDVVKLEYCPT 1306
>At3g25450 hypothetical protein
Length = 1343
Score = 158 bits (399), Expect = 7e-39
Identities = 94/298 (31%), Positives = 169/298 (56%), Gaps = 9/298 (3%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSH 210
++ +YVDD+L+ G++L+ + K + F+M DLG+ LGI++++ + I L Q
Sbjct: 1008 LVVAVYVDDLLVTGSNLDIILNFKKGMVGKFEMSDLGKLTYYLGIEVLQSKDGITLKQER 1067
Query: 211 YIEKVLKKFDHFDCKSVSTPFDQN---TKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPD 267
Y +K+L++ C +V+TP + +K Q KR + + +Y + I CL Y + TRPD
Sbjct: 1068 YAKKILEEAGMSKCNTVNTPMIASLELSKAQDEKR--IDETDYRRNIGCLRYLLH-TRPD 1124
Query: 268 IAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIY-NGYPSVLEGYTYASWVTCV 326
++Y VG LSRY P + H + ++L+YL GT ++ L + G + L GY+ +S +
Sbjct: 1125 LSYNVGILSRYLQEPRESHGAALKQILRYLQGTTSHGLYFKKGENAGLIGYSDSSHNVDL 1184
Query: 327 KDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIW 386
D ST G IF L ++W S+KQ + S+ AEF+A +K+ + L+ LL E++
Sbjct: 1185 DDGKSTGGHIFYLNDCPITWCSQKQQVVTLSSCEAEFMAATEAAKQAIWLQELLAEVI-- 1242
Query: 387 PKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKE 444
+V+I D++ ++ + V++G+S+HI R+ +++ + NG I + V V++
Sbjct: 1243 GTECEKVTIRVDNKSAIALTKNPVFHGRSKHIHRRYHFIRECVENGQIEVEHVPGVRQ 1300
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 153 bits (386), Expect = 2e-37
Identities = 91/299 (30%), Positives = 167/299 (55%), Gaps = 7/299 (2%)
Query: 149 KAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQ 208
K ++ +YVDD+L+ G+ L+ + K ++ F+M DLG+ LGI+++ I L Q
Sbjct: 967 KLLIVAIYVDDLLVTGSSLDLILCFKKDMAGKFEMSDLGQLTYYLGIEVLHRKNGIILRQ 1026
Query: 209 SHYIEKVLKKFDHFDCKSVSTPFDQNTKL--QPHKRCPVAQLEYSKAIRCLMYAMTCTRP 266
Y K++++ +C V P +L ++C + + +Y + I CL Y + TRP
Sbjct: 1027 ERYAMKIIEEAGMSNCNPVLIPMAAGLELCKAQEEKC-ITERDYRRMIGCLRY-IVHTRP 1084
Query: 267 DIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSL-IYNGYPSVLEGYTYASWVTC 325
D++Y VG LSRY P + H + + +VL+YL GT+++ L + G+ S L GY+ +S
Sbjct: 1085 DLSYCVGVLSRYLQQPRESHGNALKQVLRYLKGTMSHGLYLKRGFKSGLVGYSDSSHSAD 1144
Query: 326 VKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILI 385
+ D ST+G IF L ++W S+KQ +A S+ AEF+A +K+ + L++L E+
Sbjct: 1145 LDDGKSTAGHIFYLHQCPITWCSQKQQVVALSSCEAEFMAATEAAKQAIWLQDLFAEVC- 1203
Query: 386 WPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKE 444
+V I D++ ++ + V++G+S+HI R+ +++ + N ++ + V V++
Sbjct: 1204 -GTTSEKVMIRVDNKSAIALTKNLVFHGRSKHIHRRYHFIRECVENNLVEVDHVPGVEQ 1261
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 150 bits (379), Expect = 1e-36
Identities = 91/308 (29%), Positives = 163/308 (52%), Gaps = 14/308 (4%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSH 210
++ LYVDD++I G + + + K + + F+M DLG + LG+++ +D I LSQ
Sbjct: 1025 LIVSLYVDDLIITGNNTHLINTFKKNMKDEFEMTDLGLLNYFLGMEVNQDDSGIFLSQEK 1084
Query: 211 YIEKVLKKFDHFDCKSVSTPFDQNTK---LQPHKRCPVAQLEYSKAIRCLMYAMTCTRPD 267
Y K++ KF + KSVSTP K ++ + +Y + + L+Y + +RPD
Sbjct: 1085 YANKLIDKFGMKESKSVSTPLTPQGKRKGVEGDDKEFADPTKYRRIVGGLLY-LCASRPD 1143
Query: 268 IAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTYASWVTCV 326
+ Y LSRY +PS H+ RVL+Y+ GT N+ +++ + L GY+ + W +
Sbjct: 1144 VMYASSYLSRYMSSPSIQHYQEAKRVLRYVKGTSNFGVLFTSKETPRLVGYSDSDWGGSL 1203
Query: 327 KDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIW 386
+D ST+G +F LG W S KQ +A ST AE+IA+ A + + + L+ L + +
Sbjct: 1204 EDKKSTTGYVFTLGLAMFCWQSCKQQTVAQSTAEAEYIAVCAATNQAIWLQRLFEDFGL- 1262
Query: 387 PKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKESC 446
K + I CD++ ++ + V + +++HI +++ V++ G+I + E C
Sbjct: 1263 -KFKEGIPILCDNKSAIAIGRNPVQHRRTKHIEIKYHFVREAEHKGLIQL-------EYC 1314
Query: 447 RSFDERSD 454
+ D+ +D
Sbjct: 1315 KGEDQLAD 1322
>At1g26990 polyprotein, putative
Length = 1436
Score = 150 bits (379), Expect = 1e-36
Identities = 91/295 (30%), Positives = 151/295 (50%), Gaps = 3/295 (1%)
Query: 148 GKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLS 207
G ++ +YVDD+LI T + S LS+ F ++DLGE LGI+I R+ + I L
Sbjct: 1110 GFFLVVLVYVDDILIASTTEAASAELTSQLSSFFQLRDLGEPKFFLGIEIARNADGISLC 1169
Query: 208 QSHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPD 267
Q Y+ +L D DCK S P + N KL + + + I + + TRPD
Sbjct: 1170 QRKYVLDLLASSDFSDCKPSSIPMEPNQKLSKDTGTLLEDGKQYRRILGKLQYLCLTRPD 1229
Query: 268 IAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTYASWVTCV 326
I + V +L++Y+ P+ H ++++L+YL GTI L Y + L G++ + W TC
Sbjct: 1230 INFAVSKLAQYSSAPTDIHLQALHKILRYLKGTIGQGLFYGADTNFDLRGFSDSDWQTCP 1289
Query: 327 KDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIW 386
+G +G VSW SKKQ ++ S+ AE+ A++ +KE + L +L I
Sbjct: 1290 DTRRCVTGFAIFVGNSLVSWRSKKQDVVSMSSAEAEYRAMSVATKELIWLGYILTAFKI- 1348
Query: 387 PKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
++CD++ L A + V++ +++HI V++ I G++ +FVRT
Sbjct: 1349 -PFTHPAYLYCDNEAALHIANNSVFHERTKHIENDCHKVRECIEAGILKTIFVRT 1402
>At4g16870 retrotransposon like protein
Length = 1474
Score = 149 bits (375), Expect = 4e-36
Identities = 93/293 (31%), Positives = 153/293 (51%), Gaps = 5/293 (1%)
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSH 210
V +YVDD+++ G+D + ++ + L+ F +KD + LGI+ R + + L Q
Sbjct: 1153 VYVLVYVDDIIVTGSDKSSIDAVLTSLAERFSIKDPTDLHYFLGIEATRTKQGLHLMQRK 1212
Query: 211 YIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQL-EYSKAIRCLMYAMTCTRPDIA 269
YI+ +L K + D K V TP + KL H + EY + L Y + TRPDIA
Sbjct: 1213 YIKDLLAKHNMADAKPVLTPLPTSPKLTLHGGTKLNDASEYRSVVGSLQY-LAFTRPDIA 1271
Query: 270 YVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYN-GYPSVLEGYTYASWVTCVKD 328
Y V RLS+ P++DHW RVL+YL GT + + + P L ++ A W D
Sbjct: 1272 YAVNRLSQLMPQPTEDHWQAAKRVLRYLAGTSTHGIFLDTTSPLNLHAFSDADWAGDSDD 1331
Query: 329 HASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPK 388
+ ST+ + LG +SW SKKQ +A S+ +E+ A+A + E L +LL ++ I +
Sbjct: 1332 YVSTNAYVIYLGKNPISWSSKKQRGVARSSTESEYRAVANAASEVKWLCSLLSKLHI--R 1389
Query: 389 LMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
L + SI CD+ + V++ + +HI + + V+++I +G + + V T
Sbjct: 1390 LPIRPSIFCDNIGATYLCANPVFHSRMKHIAIDYHFVRNMIQSGALRVSHVST 1442
>At2g13940 putative retroelement pol polyprotein
Length = 1501
Score = 149 bits (375), Expect = 4e-36
Identities = 94/292 (32%), Positives = 157/292 (53%), Gaps = 5/292 (1%)
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSHYIEK 214
+YVDD+LI G D ++K K +LS F MKDLG+ LGI++ R E I LSQ Y
Sbjct: 1183 IYVDDLLICGNDGYMLQKFKDYLSRCFSMKDLGKLKYFLGIEVSRGPEGIFLSQRKYALD 1242
Query: 215 VLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLE-YSKAIRCLMYAMTCTRPDIAYVVG 273
V+ + + TP +QN L ++ + Y + + L+Y + TRP+++Y V
Sbjct: 1243 VIADSGNLGSRPAHTPLEQNHHLASDDGPLLSDPKPYRRLVGRLLYLLH-TRPELSYSVH 1301
Query: 274 RLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTYASWVTCVKDHAST 332
L+++ NP + H+ RV++YL G+ ++ N P + LE Y + W +C S
Sbjct: 1302 VLAQFMQNPREAHFDAALRVVRYLKGSPGQGILLNADPDLTLEVYCDSDWQSCPLTRRSI 1361
Query: 333 SG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMRQ 392
S + LGG +SW +KKQ ++ S+ AE+ A++ KE LR LL E+ I +
Sbjct: 1362 SAYVVLLGGSPISWKTKKQDTVSHSSAEAEYRAMSYALKEIKWLRKLLKELGI--EQSTP 1419
Query: 393 VSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKE 444
++CDS+ + A + V++ +++HI V+D + +G+I+ VRT ++
Sbjct: 1420 ARLYCDSKAAIHIAANPVFHERTKHIESDCHSVRDAVRDGIITTQHVRTTEQ 1471
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.351 0.154 0.546
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,550,105
Number of Sequences: 26719
Number of extensions: 368115
Number of successful extensions: 1803
Number of sequences better than 10.0: 94
Number of HSP's better than 10.0 without gapping: 90
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1501
Number of HSP's gapped (non-prelim): 121
length of query: 469
length of database: 11,318,596
effective HSP length: 103
effective length of query: 366
effective length of database: 8,566,539
effective search space: 3135353274
effective search space used: 3135353274
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0185.17


