

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0180b.2
(57 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
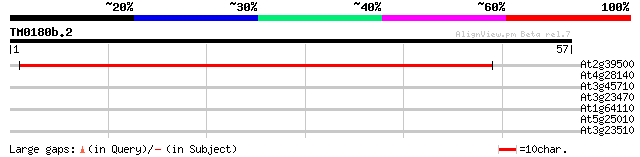
Score E
Sequences producing significant alignments: (bits) Value
At2g39500 unknown protein 82 7e-17
At4g28140 putative DNA-binding protein 26 3.3
At3g45710 putative transporter protein 25 5.6
At3g23470 cyclopropane-fatty-acyl-phospholipid synthase, putative 25 7.3
At1g64110 unknown protein 25 7.3
At5g25010 putative protein 25 9.6
At3g23510 cyclopropane-fatty-acyl-phospholipid synthase, putative 25 9.6
>At2g39500 unknown protein
Length = 55
Score = 81.6 bits (200), Expect = 7e-17
Identities = 32/48 (66%), Positives = 42/48 (86%)
Query: 2 ELKNVVKDKKFWVASFLIAWAAALQGHMMWLQRQDSFKHKFRNLDDDD 49
++K +VKDK+FW+ASF+I WAA LQGHMMWLQ+Q+SFK KF +D+DD
Sbjct: 6 DVKTIVKDKRFWIASFIIVWAAGLQGHMMWLQKQESFKQKFGTIDEDD 53
>At4g28140 putative DNA-binding protein
Length = 292
Score = 26.2 bits (56), Expect = 3.3
Identities = 17/64 (26%), Positives = 28/64 (43%), Gaps = 9/64 (14%)
Query: 2 ELKNVVKDKKFWVASFLIAWAAA---------LQGHMMWLQRQDSFKHKFRNLDDDDRSD 52
E++ + W+ +F A AA L+GH L + F +K L D + SD
Sbjct: 157 EIRKPRSRARLWLGTFDTAEEAAMAYDRQAFKLRGHSATLNFPEHFVNKESELHDSNSSD 216
Query: 53 EPQP 56
+ +P
Sbjct: 217 QKEP 220
>At3g45710 putative transporter protein
Length = 556
Score = 25.4 bits (54), Expect = 5.6
Identities = 9/19 (47%), Positives = 14/19 (73%)
Query: 37 SFKHKFRNLDDDDRSDEPQ 55
S+ +K+RNL DDD +P+
Sbjct: 534 SWFYKYRNLKDDDHEQDPK 552
>At3g23470 cyclopropane-fatty-acyl-phospholipid synthase, putative
Length = 283
Score = 25.0 bits (53), Expect = 7.3
Identities = 13/50 (26%), Positives = 28/50 (56%), Gaps = 1/50 (2%)
Query: 5 NVVKDKKFWVASFLIAWAAALQGHMMWLQRQDSFKHKFRNLDDD-DRSDE 53
N+VK + +W FL A A+++ ++ + +Q++ +N+ D S+E
Sbjct: 40 NLVKKRGWWTPVFLTAGLASVKYYLKHVLKQNTLTQARKNISSHYDLSNE 89
>At1g64110 unknown protein
Length = 824
Score = 25.0 bits (53), Expect = 7.3
Identities = 10/32 (31%), Positives = 18/32 (56%)
Query: 18 LIAWAAALQGHMMWLQRQDSFKHKFRNLDDDD 49
L++W + L+ M +Q QD+ H L ++D
Sbjct: 338 LVSWKSQLERDMNMIQTQDNRNHIMEVLSEND 369
>At5g25010 putative protein
Length = 286
Score = 24.6 bits (52), Expect = 9.6
Identities = 10/18 (55%), Positives = 12/18 (66%)
Query: 36 DSFKHKFRNLDDDDRSDE 53
DS HKF N+DDD +E
Sbjct: 146 DSILHKFINIDDDSFRNE 163
>At3g23510 cyclopropane-fatty-acyl-phospholipid synthase, putative
Length = 456
Score = 24.6 bits (52), Expect = 9.6
Identities = 11/41 (26%), Positives = 21/41 (50%)
Query: 5 NVVKDKKFWVASFLIAWAAALQGHMMWLQRQDSFKHKFRNL 45
N+ K + +W FL A A+ + + + RQ++ RN+
Sbjct: 125 NLTKKRGWWTPMFLTAGLASAKYFLKHVSRQNTLTQARRNI 165
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.133 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,470,886
Number of Sequences: 26719
Number of extensions: 44776
Number of successful extensions: 134
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 127
Number of HSP's gapped (non-prelim): 7
length of query: 57
length of database: 11,318,596
effective HSP length: 33
effective length of query: 24
effective length of database: 10,436,869
effective search space: 250484856
effective search space used: 250484856
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0180b.2


