

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.5
(770 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
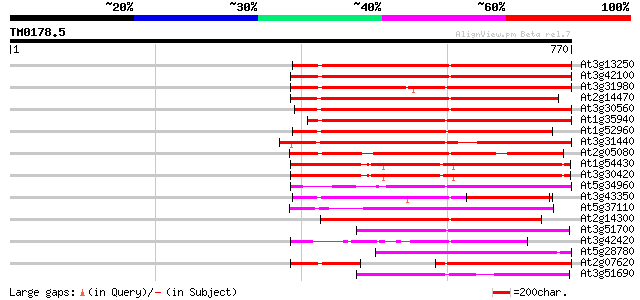
Score E
Sequences producing significant alignments: (bits) Value
At3g13250 hypothetical protein 392 e-109
At3g42100 putative protein 386 e-107
At3g31980 hypothetical protein 378 e-105
At2g14470 pseudogene 371 e-103
At3g30560 hypothetical protein 360 1e-99
At1g35940 hypothetical protein 358 5e-99
At1g52960 hypothetical protein 355 4e-98
At3g31440 hypothetical protein 342 6e-94
At2g05080 putative helicase 317 1e-86
At1g54430 hypothetical protein 316 3e-86
At3g30420 hypothetical protein 307 1e-83
At5g34960 putative protein 282 4e-76
At3g43350 putative protein 279 4e-75
At5g37110 putative helicase 272 5e-73
At2g14300 pseudogene; similar to MURA transposase of maize Muta... 263 4e-70
At3g51700 unknown protein 206 3e-53
At3g42420 putative protein 196 4e-50
At5g28780 putative protein 191 2e-48
At2g07620 putative helicase 189 7e-48
At3g51690 putative protein 171 2e-42
>At3g13250 hypothetical protein
Length = 1419
Score = 392 bits (1007), Expect = e-109
Identities = 198/385 (51%), Positives = 268/385 (69%), Gaps = 8/385 (2%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
K+ASLIIWDE PM++K CFEALD+S +DIIK FGG V+V GGDFRQ+LPVI
Sbjct: 1039 KEASLIIWDEAPMMSKFCFEALDKSFSDIIKRVDNK----VFGGKVMVFGGDFRQVLPVI 1094
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVGDGT 507
+ R+EIV S++N+SYLW HCKV++LT NM +L N S EI+EF+DWLL VGDG
Sbjct: 1095 NGAGRAEIVMSSLNASYLWDHCKVLRLTKNMRLLNNDLSVDEAKEIQEFSDWLLAVGDGR 1154
Query: 508 VKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAPTLE 565
V ++ E +I+IP LLI++ P+ + Y P H + FFQ RAILAP E
Sbjct: 1155 VNEPNDGEVIIDIPEELLIQEADNPIEAISREIYGDPTKLHEISDPKFFQRRAILAPKNE 1214
Query: 566 SVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKL 625
V IN +ML + +E YLS D++ SD DS N T +FLN K S +P+H+++L
Sbjct: 1215 DVNTINQYMLEHLDSEERIYLSADSIDPSDSDSLKNPV-ITPDFLNSIKVSGMPHHSLRL 1273
Query: 626 KVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNS 685
KVG P+ML+RN+D GLCN TR+ + L +I+ A ++ G+++G+ +IP +++TPS++
Sbjct: 1274 KVGAPVMLLRNLDPKGGLCNGTRLQITQLCSHIVEAKVITGDRIGQIVYIPLINITPSDT 1333
Query: 686 DIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLK 745
+PFK +RRQFP+++ F MTINKSQGQSL VGLYLP+PVF+HGQLYVALSRV S+ GLK
Sbjct: 1334 KLPFKMRRRQFPLSVAFVMTINKSQGQSLEQVGLYLPKPVFSHGQLYVALSRVTSKTGLK 1393
Query: 746 MLIIDDEGVVSNTTRNVMYQEVFDN 770
+LI+D EG + T NV+++EVF N
Sbjct: 1394 ILILDKEGKIQKQTTNVVFKEVFQN 1418
>At3g42100 putative protein
Length = 1752
Score = 386 bits (991), Expect = e-107
Identities = 196/388 (50%), Positives = 268/388 (68%), Gaps = 8/388 (2%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N K+ASLIIWDE PM++K CFE+LD+S DI+ + FGG VVV GGDFRQ+L
Sbjct: 1368 NLIKEASLIIWDEAPMMSKFCFESLDKSFYDILNNKDNK----VFGGKVVVFGGDFRQVL 1423
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVG 504
PVI+ R EIV S++N+SYLW HCKV+KLT NM +L S+ EI++F+DWLL VG
Sbjct: 1424 PVINGAGRVEIVMSSLNASYLWDHCKVLKLTKNMRLLSGGLSSEEAKEIQQFSDWLLAVG 1483
Query: 505 DGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAP 562
DG + ++ E LI+IP LLI++ P+ + Y P H + FFQ RAILAP
Sbjct: 1484 DGRINEPNDGEALIDIPEELLIKEAGNPIEAISKEIYGDPSELHMINDPKFFQRRAILAP 1543
Query: 563 TLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHA 622
T E V IN +ML + +E YLS D++ +D DS N T +FLN + + +P+HA
Sbjct: 1544 TNEDVNTINQYMLEHLKSEERIYLSADSIDPTDSDSLANPV-ITPDFLNSIQLTGMPHHA 1602
Query: 623 IKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTP 682
++LKVG P+ML+RN+D GLCN TR+ + L K ++ A ++ +++G+ IP ++LTP
Sbjct: 1603 LRLKVGAPVMLLRNLDPKGGLCNGTRLQITQLAKQVVQAKVITRDRIGDIVLIPLINLTP 1662
Query: 683 SNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRK 742
S++ +PFK +RRQFP+++ FAMTINKSQGQSL VGLYLP+PVF+HGQLYVALSRV S+K
Sbjct: 1663 SDTKLPFKMRRRQFPLSVAFAMTINKSQGQSLEQVGLYLPKPVFSHGQLYVALSRVTSKK 1722
Query: 743 GLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
GLK+LI+D +G + T NV+++EVF N
Sbjct: 1723 GLKILILDKDGNMQKQTTNVVFKEVFQN 1750
>At3g31980 hypothetical protein
Length = 1099
Score = 378 bits (971), Expect = e-105
Identities = 196/389 (50%), Positives = 265/389 (67%), Gaps = 10/389 (2%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N K+ASLIIWDE PM++++CFE+LDRSLND+I PFGG VVV GGDFRQ+L
Sbjct: 714 NMLKEASLIIWDEAPMMSRYCFESLDRSLNDVIGNIDGK----PFGGKVVVFGGDFRQVL 769
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVG 504
PVI R+EIV +A+NSSYLW+HCKV+ LT NM ++ N EIKEF++WLL VG
Sbjct: 770 PVIHGAGRAEIVLAALNSSYLWEHCKVLTLTKNMRLMSNDLDKDEAEEIKEFSNWLLAVG 829
Query: 505 DGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNS---FFQERAILA 561
DG V ++ E LI+IP LLI+ +P+ + Y L LQ N+ FFQ+RAIL
Sbjct: 830 DGRVSEPNDGEVLIDIPEELLIKDANDPIEAITKAVYGDL-DLLQPNNDPKFFQQRAILC 888
Query: 562 PTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNH 621
P V IN+ ML + G+ YLS D++ D S +N T +FLN K S +PNH
Sbjct: 889 PRNTDVNTINDIMLDKLNGELVTYLSADSIDPQDAAS-LNNPVLTPDFLNSIKLSGLPNH 947
Query: 622 AIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLT 681
+ LK+G P+ML+RNID GLCN TR+ V + +I+ A ++ G+++G+ I + ++
Sbjct: 948 NLTLKIGTPVMLLRNIDPKGGLCNGTRLQVTQMGNHILEARVITGDRVGDKVIIIKSQIS 1007
Query: 682 PSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSR 741
PS++ +PF+ +RRQFP+ + FAMTINKSQGQSL VG+YLP+PVF+HGQLYVALSRV S+
Sbjct: 1008 PSDTKLPFRMRRRQFPIAVAFAMTINKSQGQSLKEVGIYLPKPVFSHGQLYVALSRVTSK 1067
Query: 742 KGLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
KGLK+LI+D EG + T NV+++E+F N
Sbjct: 1068 KGLKVLIVDKEGNTQSQTMNVVFKEIFQN 1096
>At2g14470 pseudogene
Length = 1265
Score = 371 bits (952), Expect = e-103
Identities = 187/371 (50%), Positives = 257/371 (68%), Gaps = 8/371 (2%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N KKASL+IWDE PM+++ CFEALD+S +DIIK + FGG VVV GGDFRQ+
Sbjct: 898 NLVKKASLVIWDEAPMMSRFCFEALDKSFSDIIKNTD----NTVFGGKVVVFGGDFRQVF 953
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVG 504
PVI+ R+EIV S++N+SYLW +CKV+KLT N +L N S + EI+EF+DWLL VG
Sbjct: 954 PVINGAGRAEIVMSSLNASYLWDNCKVLKLTKNTRLLANNLSETEAKEIQEFSDWLLAVG 1013
Query: 505 DGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAP 562
DG + ++ +I+IP +LLI +P+ + N Y PK+ H + FFQ RAILA
Sbjct: 1014 DGRINESNDGVAIIDIPEDLLITNADKPIESITNEIYGDPKILHEITDPKFFQGRAILAS 1073
Query: 563 TLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHA 622
E V IN ++L + +E YLS D++ +D DS N T +FLN K +PNH+
Sbjct: 1074 KNEDVNTINEYLLDQLHAEERIYLSADSIDPTDSDSLSNPV-ITPDFLNSIKLPGLPNHS 1132
Query: 623 IKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTP 682
++LKVG P++L+RN+D GLCN TR+ + L I+ A ++ G+++G IP ++LTP
Sbjct: 1133 LRLKVGAPVLLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDRIGHIILIPTVNLTP 1192
Query: 683 SNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRK 742
+N+ +PFK +RRQFP+++ F MTINKS+GQSL HVGLYLP+PVF+HGQLYVALSRV S+K
Sbjct: 1193 TNTKLPFKMRRRQFPLSVAFVMTINKSEGQSLEHVGLYLPKPVFSHGQLYVALSRVTSKK 1252
Query: 743 GLKMLIIDDEG 753
GLK+LI+D +G
Sbjct: 1253 GLKILILDKDG 1263
>At3g30560 hypothetical protein
Length = 1473
Score = 360 bits (925), Expect = 1e-99
Identities = 183/380 (48%), Positives = 256/380 (67%), Gaps = 5/380 (1%)
Query: 391 ASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVISK 450
A LIIWDE PM++K+CFE+LD+SL DI+ T D+PFGG +++ GGDFRQILPVI
Sbjct: 1098 AKLIIWDEAPMMSKYCFESLDKSLKDILSTPE----DMPFGGKLIIFGGDFRQILPVILA 1153
Query: 451 GSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQVGDGTVKT 510
R IV S++NSS+LW++CKV KLT NM L + EI++F+ W+L VG+G +
Sbjct: 1154 AGRELIVKSSLNSSHLWQYCKVFKLTKNMRLLQDIDINEAREIEDFSKWILAVGEGKLNQ 1213
Query: 511 IDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESVEEI 570
++ T I+I ++LI + P+ ++ Y + FFQ+RAIL PT + V I
Sbjct: 1214 PNDGVTQIQIRDDILIPEGDNPIESIIKAVYGTSFDEERDPKFFQDRAILCPTNDDVNSI 1273
Query: 571 NNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVP 630
N+ ML+ + G+E Y S D++ SD + N +T +FLN K S +PNH + LKVG P
Sbjct: 1274 NDHMLSKLTGEEKIYRSSDSIDPSDTRADKNPV-YTPDFLNKIKISGLPNHLLWLKVGCP 1332
Query: 631 IMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFK 690
+ML+RN+D GL N TR+ + L ++ IL G ++G+ IPRM LTPS+ +PFK
Sbjct: 1333 VMLLRNLDSHGGLMNGTRLQIVRLGDKLVQGRILTGTRVGKLVIIPRMPLTPSDRRLPFK 1392
Query: 691 FQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIID 750
+RRQFP+++ FAMTINKSQGQSL +VG+YLP+PVF+HGQLYVA+SRVKS+ GLK+LI D
Sbjct: 1393 MKRRQFPLSVAFAMTINKSQGQSLGNVGIYLPKPVFSHGQLYVAMSRVKSKGGLKVLITD 1452
Query: 751 DEGVVSNTTRNVMYQEVFDN 770
+G N T NV+++E+F N
Sbjct: 1453 SKGKQKNETTNVVFKEIFRN 1472
>At1g35940 hypothetical protein
Length = 1678
Score = 358 bits (920), Expect = 5e-99
Identities = 182/365 (49%), Positives = 251/365 (67%), Gaps = 8/365 (2%)
Query: 409 ALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVISKGSRSEIVGSAINSSYLWK 468
+LD+S +DIIK + FGG VVV GGDFRQ+LPVI+ R+EIV S++N+SYLW
Sbjct: 1317 SLDKSFSDIIKNTNNK----VFGGKVVVFGGDFRQVLPVINGAGRAEIVMSSLNASYLWD 1372
Query: 469 HCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVGDGTVKTIDEEETLIEIPPNLLIE 527
HCKV+KLT NM +L N S + EI+EF+DWLL V DG + ++ I+IP +LLI
Sbjct: 1373 HCKVLKLTKNMRLLANNLSATEAKEIQEFSDWLLAVSDGRINEPNDGVATIDIPEDLLIT 1432
Query: 528 QCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAPTLESVEEINNFMLAMIPGDETEY 585
+P+ + N Y PK+ H + FFQ RAILAP E V IN ++L + +E Y
Sbjct: 1433 NADKPIETITNEIYGDPKILHEITDPKFFQGRAILAPKNEDVNTINEYLLEQLDAEERIY 1492
Query: 586 LSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCN 645
LS D++ +D DS +N T +FLN K +PNH++ LKVG P+ML+RN+D GLCN
Sbjct: 1493 LSADSIDPTDSDS-LNNPVITPDFLNSIKLPGLPNHSLCLKVGAPVMLLRNLDPKGGLCN 1551
Query: 646 DTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMT 705
TR+ + L I+ A ++ G+++G IP ++LTP+++ +PFK +RRQFP+++ FAMT
Sbjct: 1552 GTRLQITQLCTQIVEAKVITGDRIGNIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMT 1611
Query: 706 INKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQ 765
INKSQGQSL H+GLYLP+PVF+HGQLYVALSRV S+KGLK+LI+D +G + T NV+++
Sbjct: 1612 INKSQGQSLEHIGLYLPKPVFSHGQLYVALSRVTSKKGLKILILDKDGKLQKQTTNVVFK 1671
Query: 766 EVFDN 770
EVF N
Sbjct: 1672 EVFQN 1676
>At1g52960 hypothetical protein
Length = 924
Score = 355 bits (912), Expect = 4e-98
Identities = 177/357 (49%), Positives = 244/357 (67%), Gaps = 5/357 (1%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
K+A+LIIWDETPM++KHCFE+LDR+L DI+ D PFGG +V GGDFRQ+LPVI
Sbjct: 571 KEANLIIWDETPMMSKHCFESLDRTLRDIMNNPG----DKPFGGKGIVFGGDFRQVLPVI 626
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQVGDGTV 508
+ R EIV +A+NSSY+W+HCKV++LT NM L S +I+ F+ W+L VGDG +
Sbjct: 627 NGAGREEIVFAALNSSYIWEHCKVLELTKNMRLLANISEHEKRDIEYFSKWILDVGDGKI 686
Query: 509 KTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESVE 568
++ LI+IP LI +P+ ++ Y + FFQ RAIL PT E V
Sbjct: 687 SQPNDGIALIDIPEEFLINGDNDPVESIIEAVYGNTFMEEKDPKFFQGRAILCPTNEDVN 746
Query: 569 EINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVG 628
IN M++M+ G+E YLS D++ +D S N + ++++FLN + S +PNH ++LKVG
Sbjct: 747 SINEHMMSMLDGEERIYLSSDSIDPADTSSA-NNDAYSADFLNSVRVSGLPNHCLRLKVG 805
Query: 629 VPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIP 688
P+ML+RN+D GLCN TR+ V + +I A + GN++G+ IPRM +TPS++ +P
Sbjct: 806 CPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTRLP 865
Query: 689 FKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLK 745
FK +RRQFP+++ FAMTINKSQGQ+L VGLYLPRPVF+HGQLYVA+SRV S+ G K
Sbjct: 866 FKMRRRQFPLSVAFAMTINKSQGQTLESVGLYLPRPVFSHGQLYVAISRVTSKTGTK 922
>At3g31440 hypothetical protein
Length = 536
Score = 342 bits (876), Expect = 6e-94
Identities = 187/408 (45%), Positives = 256/408 (61%), Gaps = 37/408 (9%)
Query: 371 MRYQPAIFVTVPQKP-----NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHG 425
+R P F KP N K+ASL+I D+ PM+++ CFEALD+S +DIIK
Sbjct: 156 IRLNPDEFSVCKIKPKSDLANLVKEASLVICDKAPMMSRFCFEALDKSFSDIIKNT---- 211
Query: 426 YDIPFGGIVVVLGGDFRQILPVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNA 484
Y+ FGG VVV GDFRQ+LPVI+ R+EIV S++N+SYLW HCKV+KLT NM +L N
Sbjct: 212 YNKVFGGKVVVFSGDFRQVLPVINGAGRAEIVMSSLNASYLWDHCKVLKLTKNMRLLANN 271
Query: 485 TSTSSPAEIKEFADWLLQVGDGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAY--P 542
S + EI EF+DWLL VGDG + +++ +I+IP +LLI +P+ + N Y P
Sbjct: 272 LSETEAKEIHEFSDWLLAVGDGRINEPNDDVAIIDIPKDLLITNADKPIEWITNEIYGDP 331
Query: 543 KLAHNLQKNSFFQERAILAPTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNA 602
K+ H + FFQ RAILAP E V IN ++L + +E YLS D++ +D DS +N
Sbjct: 332 KILHEITDPKFFQGRAILAPKNEDVNTINEYLLEQLHAEERIYLSADSIDPTDSDS-LNN 390
Query: 603 EWFTSEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVAT 662
T +FLN K GLCN R+ + L I+ A
Sbjct: 391 PVITPDFLNSIKLP------------------------GGLCNGARLQITQLFTEIVEAK 426
Query: 663 ILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLP 722
++ G+++G IP ++LTP+++ +PFK +RRQFP+++ FAMTINKSQGQSL HVGLYLP
Sbjct: 427 VITGDRIGHIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTINKSQGQSLEHVGLYLP 486
Query: 723 RPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
+PVF+HGQLYVALSRV S+KGLK+LI+D G + T N++++EVF N
Sbjct: 487 KPVFSHGQLYVALSRVTSKKGLKILILDKNGKLQKQTTNIVFKEVFQN 534
>At2g05080 putative helicase
Length = 1219
Score = 317 bits (813), Expect = 1e-86
Identities = 173/379 (45%), Positives = 237/379 (61%), Gaps = 34/379 (8%)
Query: 384 KPNY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQ 443
+ N K+ASLIIWDE PM+++HCFE+LDRSL+DI PFGG VVV GGDFRQ
Sbjct: 841 RANLVKEASLIIWDEAPMMSRHCFESLDRSLSDICGNCDNK----PFGGKVVVFGGDFRQ 896
Query: 444 ILPVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQV 503
+LPVI ++IV +A+NSSYLW HCKV+ LT NM L + +W+L V
Sbjct: 897 VLPVIPGADTADIVMAALNSSYLWSHCKVLTLTKNMCLFSE-------------EWILAV 943
Query: 504 GDGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLA--HNLQKNSFFQERAILA 561
GDG + ++ E LI+IP LI + K+P+ + Y + H + FFQERAIL
Sbjct: 944 GDGRIGEPNDGEALIDIPSEFLITKAKDPIQAICTEIYGDITKIHEQKDPVFFQERAILC 1003
Query: 562 PTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNH 621
PT E V +IN ML + G+E +LS D+L +D S N T EFLN+ K + NH
Sbjct: 1004 PTNEDVNQINETMLDNLQGEELTFLSSDSLDTADIGSRNNPV-LTPEFLNNVKVLGLSNH 1062
Query: 622 AIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLT 681
++LK+G P+ML+RNID GL N TR+ + ++ +I+ A IL G++
Sbjct: 1063 KLRLKIGSPVMLLRNIDPIGGLMNGTRLQIMQMSPFILQAMILTGDR------------- 1109
Query: 682 PSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSR 741
+++ +PF+ +R Q P+ +CFAMTINKSQGQSL VG++LPRP F+H QLYVA+SRV S+
Sbjct: 1110 -ADTKLPFRMRRTQLPLAVCFAMTINKSQGQSLKRVGIFLPRPCFSHSQLYVAISRVTSK 1168
Query: 742 KGLKMLIIDDEGVVSNTTR 760
GLK+LI++DEG T+
Sbjct: 1169 SGLKILIVNDEGKPQKQTK 1187
>At1g54430 hypothetical protein
Length = 1639
Score = 316 bits (810), Expect = 3e-86
Identities = 184/395 (46%), Positives = 244/395 (61%), Gaps = 23/395 (5%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N K LIIWDE PM ++H FEA+DR+L DI+ FGG V+LGGDFRQIL
Sbjct: 1247 NVLSKVDLIIWDEAPMAHRHTFEAVDRTLRDILSVGDEKALTKTFGGKTVLLGGDFRQIL 1306
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQVGD 505
PVI +G+R E V +AIN SYLW+ C L+ NM +Q P EIK FA+W+LQVGD
Sbjct: 1307 PVIPQGTRQETVSAAINRSYLWESCHKYLLSQNMRVQ-------PEEIK-FAEWILQVGD 1358
Query: 506 GTV--KTI----DEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAI 559
G KT D+EE I I NLL+ + + PL L +P + Q + A+
Sbjct: 1359 GEAPRKTHGIDDDQEEDNIIIDKNLLLPETENPLEVLCRSVFPDFTNTFQDLENLKGTAV 1418
Query: 560 LAPTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTS----EFLNDFKC 615
L P E+V+EIN+++L+ +PG EY S D++ D D + E F E+LN +
Sbjct: 1419 LTPRNETVDEINDYLLSKVPGLAKEYFSADSI---DRDEALTEEGFEMSYPMEYLNSLEF 1475
Query: 616 SEIPNHAIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGN-KMGETTF 674
+P H + LKVGVPIML+RN++Q GLCN TR+IV L ++ A IL+ K +
Sbjct: 1476 PGLPAHRLCLKVGVPIMLLRNLNQKEGLCNGTRLIVTHLGDKVLKAEILSDTTKERKKVL 1535
Query: 675 IPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVA 734
IPR+ L+P +S PF +RRQFPV +C+AMTINKSQGQ+L+ V LYLP+PVF+HGQLYVA
Sbjct: 1536 IPRIILSPQDSKHPFTLRRRQFPVRMCYAMTINKSQGQTLNRVALYLPKPVFSHGQLYVA 1595
Query: 735 LSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVFD 769
LSRV S KGL +L + T N++Y+EVF+
Sbjct: 1596 LSRVTSPKGLTVLDTSKKKEGKYVT-NIVYREVFN 1629
>At3g30420 hypothetical protein
Length = 837
Score = 307 bits (787), Expect = 1e-83
Identities = 179/395 (45%), Positives = 241/395 (60%), Gaps = 23/395 (5%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N K LIIWDE PM ++H FEA+DR+L DI+ GG V+LGGDFRQIL
Sbjct: 445 NVLSKVDLIIWDEAPMAHRHTFEAVDRTLRDILSVGDEKALTKTLGGKTVLLGGDFRQIL 504
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQVGD 505
PVI + +R E V +AIN SYLW+ C L+ NM +Q P EIK FA+W+LQ+GD
Sbjct: 505 PVIPQRTRQETVSAAINRSYLWESCHKYLLSQNMRVQ-------PEEIK-FAEWILQIGD 556
Query: 506 GTV--KTI----DEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAI 559
G KT D+EE I I NLL+ + + PL L P + Q + A+
Sbjct: 557 GEAPRKTHGIDDDQEEDNIIIDKNLLLPETENPLEVLCQSVSPDFTNTFQDLENLKGTAV 616
Query: 560 LAPTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTS----EFLNDFKC 615
L P E+V+EIN+++L+ +PG EY S D++ D+D + E F E+LN +
Sbjct: 617 LTPRNETVDEINDYLLSKVPGLAKEYFSADSI---DQDEALTEEGFEMSYPMEYLNSLEF 673
Query: 616 SEIPNHAIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGN-KMGETTF 674
+P H + LKVGVPIML+RN++Q GLCN TR+ V L ++ A IL+ K +
Sbjct: 674 PGLPAHRLCLKVGVPIMLLRNLNQKEGLCNGTRLTVTHLGDKVLKAEILSDTTKKRKKVL 733
Query: 675 IPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVA 734
IPR+ L+P +S PF +RRQFPV +C+AMT+NKSQGQ+L+ V LYLP+PVF+HGQLYVA
Sbjct: 734 IPRIILSPQDSKHPFTLRRRQFPVRMCYAMTVNKSQGQTLNRVALYLPKPVFSHGQLYVA 793
Query: 735 LSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVFD 769
LSRV S KGL +L + T N++Y+EVF+
Sbjct: 794 LSRVTSPKGLTVLDTSKKKEGKYVT-NIVYREVFN 827
>At5g34960 putative protein
Length = 1033
Score = 282 bits (722), Expect = 4e-76
Identities = 157/387 (40%), Positives = 222/387 (56%), Gaps = 74/387 (19%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N K+ASL+IWDE PM++ RQ+L
Sbjct: 717 NLVKEASLVIWDEAPMMS--------------------------------------RQVL 738
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQVGD 505
VI+ R+EIV S++N+SYLW HCKV+K+ + D
Sbjct: 739 LVINGAGRAEIVMSSLNASYLWDHCKVLKIN-------------------------EPND 773
Query: 506 GTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAPT 563
G +I+IP +LLI +P+ + N Y PK+ H + FFQ RAILAPT
Sbjct: 774 GV--------AIIDIPEDLLITNTDKPIESITNEIYGDPKILHEITDPKFFQGRAILAPT 825
Query: 564 LESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAI 623
E V IN ++L + +E YLS D++ +D +S +N T +FLN K + +PNH++
Sbjct: 826 NEDVNTINEYLLEQLHAEERIYLSADSIDPTDSNS-LNNPVITPDFLNSIKLAGLPNHSL 884
Query: 624 KLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPS 683
+LKV P+ML+RN+D GLCN TR+ + L I+ A ++ G+ +G IP ++LTP+
Sbjct: 885 RLKVSAPVMLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDIIGHIVLIPTVNLTPT 944
Query: 684 NSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKG 743
++ +PFK +RRQFP+++ FAMTIN SQGQSL HVGLYLP+ VF+HGQLYVALSRV S+KG
Sbjct: 945 DTKLPFKMRRRQFPLSVAFAMTINTSQGQSLEHVGLYLPKAVFSHGQLYVALSRVTSKKG 1004
Query: 744 LKMLIIDDEGVVSNTTRNVMYQEVFDN 770
LK LI+D +G + T NV+++EVF N
Sbjct: 1005 LKFLILDKDGKLQKQTTNVVFKEVFQN 1031
>At3g43350 putative protein
Length = 830
Score = 279 bits (714), Expect = 4e-75
Identities = 156/358 (43%), Positives = 212/358 (58%), Gaps = 45/358 (12%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
K+A LIIWDE PM++KHCFE+LDR+L DI+ D P GG V+V GGDFRQ+LPVI
Sbjct: 275 KEAKLIIWDEAPMMSKHCFESLDRTLKDIVNNPG----DKPLGGKVIVFGGDFRQVLPVI 330
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQVGDGTV 508
+ R EIV +A+NSSY+W+H KV++LT NM L S +I++F+ W+L VGDG +
Sbjct: 331 NGAGREEIVFAALNSSYIWEHSKVLELTKNMRLLADISEHEKRDIEDFSKWILDVGDGKI 390
Query: 509 KTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKL---AHNLQKNSF--FQERAILAPT 563
++ LI+IP LI +P+ ++ Y + +K + +Q RAIL PT
Sbjct: 391 SQPNDGIALIDIPEEFLINGDNDPVESIIEAVYGNTFMEEKDPKKTDYPQYQGRAILCPT 450
Query: 564 LESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAI 623
E V IN M+ M+ G+E YLS D++ +D S NA + ++FLN+ + +PNH +
Sbjct: 451 NEDVNSINEHMMRMLDGEERIYLSSDSIDPADISSANNAA-YLADFLNNVRVYGLPNHCL 509
Query: 624 KLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPS 683
+LKVG P+ML+RN+D GLCN TR+ V +T II A +
Sbjct: 510 RLKVGCPVMLLRNMDPNKGLCNGTRLQVTQMTDTIIQARFIT------------------ 551
Query: 684 NSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSR 741
FAMTINKSQGQ+L VGLYLPRPVF+HGQLYVA+SRV S+
Sbjct: 552 -----------------AFAMTINKSQGQTLESVGLYLPRPVFSHGQLYVAISRVTSK 592
Score = 144 bits (363), Expect = 2e-34
Identities = 67/119 (56%), Positives = 90/119 (75%)
Query: 627 VGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSD 686
VG P+ML+RN+D GLCN TR+ V + +I A + GN++G+ IPRM +TP ++
Sbjct: 710 VGCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPLDTR 769
Query: 687 IPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLK 745
+PFK +R+QF +++ FAMTINKSQGQ+L VGLYLPRPVF+HGQLYVA+SRV S+ G K
Sbjct: 770 LPFKMRRKQFALSVAFAMTINKSQGQTLESVGLYLPRPVFSHGQLYVAISRVTSKTGTK 828
Score = 142 bits (359), Expect = 5e-34
Identities = 65/115 (56%), Positives = 89/115 (76%)
Query: 627 VGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSD 686
+G P+ML+RN+D GLCN TR+ V + +I A + GN++G+ IPRM +TPS++
Sbjct: 594 IGCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTR 653
Query: 687 IPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSR 741
+PFK +R+QF +++ FAMTINKSQGQ+L VGLYLPRPVF+HGQLYVA+SRV S+
Sbjct: 654 LPFKMRRKQFALSVAFAMTINKSQGQTLESVGLYLPRPVFSHGQLYVAISRVTSK 708
>At5g37110 putative helicase
Length = 1307
Score = 272 bits (696), Expect = 5e-73
Identities = 153/365 (41%), Positives = 212/365 (57%), Gaps = 52/365 (14%)
Query: 384 KPNY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQ 443
+ N K+ASLIIWDE PM++++CFE+LDRSL+DI +G + PFGG VVV GG
Sbjct: 978 RANLVKEASLIIWDEAPMMSRYCFESLDRSLSDICG----NGDNKPFGGKVVVFGG---- 1029
Query: 444 ILPVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQV 503
S S +IKEF++W+L V
Sbjct: 1030 -----------------------------------------LSVSEAKDIKEFSEWILAV 1048
Query: 504 GDGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNS--FFQERAILA 561
GDG + ++ E LI IP LI + K+P+ + Y + ++N FFQE+AIL
Sbjct: 1049 GDGRIVEPNDGEALIVIPSEFLITKAKDPIEAICTEIYGDITKIHEQNDPIFFQEKAILC 1108
Query: 562 PTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNH 621
PT E V +IN ML + G+E +LS D+L +D G N T +FLN K S +PNH
Sbjct: 1109 PTNEDVNQINETMLDNLQGEEFTFLSSDSLDPADI-GGKNNPALTPDFLNSVKVSRLPNH 1167
Query: 622 AIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLT 681
++LK+G P+ML+RNID GL N TR+ + + +I+ A IL G++ G IPR+ L
Sbjct: 1168 KLRLKIGCPVMLLRNIDPIGGLMNGTRLRITQMGPFILQAMILTGDRAGHLVLIPRLKLA 1227
Query: 682 PSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSR 741
PS++ +PF+ +R Q P+ +CFAMTINKSQGQSL VG++L RP F+HGQLYVA+SRV S+
Sbjct: 1228 PSDTKLPFRMRRTQLPLAVCFAMTINKSQGQSLKRVGIFLLRPCFSHGQLYVAISRVTSK 1287
Query: 742 KGLKM 746
LK+
Sbjct: 1288 TRLKI 1292
>At2g14300 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 1230
Score = 263 bits (671), Expect = 4e-70
Identities = 136/303 (44%), Positives = 192/303 (62%), Gaps = 1/303 (0%)
Query: 427 DIPFGGIVVVLGGDFRQILPVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATS 486
D+PFG V++ GGDFRQIL VI R IV S++NSSYLW+HCKV+KLT NM L
Sbjct: 929 DMPFGRKVILFGGDFRQILHVIPAAGRELIVKSSLNSSYLWQHCKVLKLTKNMRLLQDID 988
Query: 487 TSSPAEIKEFADWLLQVGDGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAH 546
+ EI++F W+L VG+G + + T I+IP ++LI + P+ ++ Y
Sbjct: 989 INEAREIEDFLKWILTVGEGKLNEPSDGVTHIQIPDDILIPEGDNPIESIIKAVYGTTFA 1048
Query: 547 NLQKNSFFQERAILAPTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFT 606
+ FFQ +AIL PT + V IN+ ML+ + G+E Y S +++ SD + N +T
Sbjct: 1049 QKRDPKFFQHKAILCPTNDDVNSINDHMLSKLTGEERIYRSSNSIDPSDTRADKNPI-YT 1107
Query: 607 SEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNG 666
+FLN K S + NH ++LKVG P+ML+RN GL N TR+ + L ++ IL G
Sbjct: 1108 PDFLNKIKISGLANHLLRLKVGCPVMLLRNFYPHGGLMNGTRLQIVRLGDKLVQGRILTG 1167
Query: 667 NKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVF 726
++G+ IPRMSLTPS+ +PFK +RR FP+++ FAMTINKSQGQSL +VG+YLP+ VF
Sbjct: 1168 TRVGKLVIIPRMSLTPSDRRLPFKMKRRHFPLSVAFAMTINKSQGQSLGNVGMYLPKAVF 1227
Query: 727 THG 729
+HG
Sbjct: 1228 SHG 1230
>At3g51700 unknown protein
Length = 344
Score = 206 bits (525), Expect = 3e-53
Identities = 112/295 (37%), Positives = 176/295 (58%), Gaps = 3/295 (1%)
Query: 476 TVNMILQNATSTSSPAEIKE-FADWLLQVGDGTVKTIDEEETLIEIPPNLLIEQCKEPLL 534
+++M+L +++ T I E F W+ +G + ++ ET I+I +LLI +CK+P+
Sbjct: 39 SIDMVLLDSSGTKIHTTIDEAFTKWITNIGGENINKPNDGETKIDIHEDLLITECKDPIK 98
Query: 535 ELVNFAYPKLAHNLQKNSFFQERAILAPTLESVEEINNFMLAMIPGDETEYLSYDTLCKS 594
+V+ Y + F+QERAIL T + +EIN++ML+ + G+ET+ DT+ +
Sbjct: 99 TIVDEVYGESFTESYNPDFYQERAILCHTNDVADEINDYMLSQLQGEETKCYGADTIYPT 158
Query: 595 DEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNAL 654
+ + EFLN K P+ ++LKVG P+ML+R++ L TR+ + +
Sbjct: 159 HASPN-DKMLYPLEFLNSIKIPGFPDFKLRLKVGAPVMLLRDLAPYGWLRKGTRLQITRV 217
Query: 655 TKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSL 714
+++ A I+ GN GE IPR+ + P K +RRQFPV L FAMTI++SQ Q+L
Sbjct: 218 ETFVLEAMIITGNNHGEKVLIPRIPSDLREAKFPIKMRRRQFPVKLAFAMTIDESQRQTL 277
Query: 715 SHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVV-SNTTRNVMYQEVF 768
S VG+YLPR + HGQ YVA+S+VKSR GLK+LI D +G T+NV+++E+F
Sbjct: 278 SKVGIYLPRQLLFHGQRYVAISKVKSRAGLKVLITDKDGKPDQEETKNVVFKELF 332
>At3g42420 putative protein
Length = 1018
Score = 196 bits (498), Expect = 4e-50
Identities = 120/326 (36%), Positives = 174/326 (52%), Gaps = 69/326 (21%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N +KASLIIWDE P+++K CFE+LD+S +DI
Sbjct: 761 NLIRKASLIIWDEAPVMSKWCFESLDKSFSDI---------------------------- 792
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVG 504
+G+ N HCKV+KLT NM +L S +IKEF+DWLL VG
Sbjct: 793 -----------IGNKDNKD----HCKVLKLTKNMRLLSKDLSEEEAKDIKEFSDWLLAVG 837
Query: 505 DGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTL 564
+ ++ I E +P + VN + FF +R IL+PT
Sbjct: 838 N-PIEAISHEIY-------------GDPAMLKVN----------EDQKFFLKRVILSPTN 873
Query: 565 ESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIK 624
+ V IN +ML + G+E +LS D++ SD DS N T + LN K S +P+H ++
Sbjct: 874 DDVHTINQYMLNNMEGEERIFLSADSIDPSDSDSLKNPV-ITPDLLNSIKLSGLPHHELR 932
Query: 625 LKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSN 684
LK+G PI+L+RN+D GLCN TR+ + +T ++ A ++ G++ G+ IP +++TPSN
Sbjct: 933 LKIGAPIILLRNLDPKGGLCNGTRLQITQMTIQVLQAKVIIGDRSGDIVLIPLINITPSN 992
Query: 685 SDIPFKFQRRQFPVTLCFAMTINKSQ 710
+PF+ +RRQFPV+L FAMTINKSQ
Sbjct: 993 MKLPFRMRRRQFPVSLAFAMTINKSQ 1018
>At5g28780 putative protein
Length = 337
Score = 191 bits (484), Expect = 2e-48
Identities = 106/269 (39%), Positives = 161/269 (59%), Gaps = 3/269 (1%)
Query: 503 VGDGTVKTIDEEE-TLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILA 561
VG+G T + ++ ++E+ + ++ L ++ AY + + + + ER IL
Sbjct: 63 VGNGNAPTCETDDGEVVEMDTSFFLKHNGNRLQQVTKGAYVQFSVSQPNFQYLTERGILT 122
Query: 562 PTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNH 621
P E V+EIN +ML+ + GD EYLS ++ K+D + ++LN + +P H
Sbjct: 123 PHNEYVDEINAYMLSQVGGDSKEYLSSYSIGKADTIGADYEALYHVKYLNSLEFPSLPKH 182
Query: 622 AIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLT 681
I LK GVPIM +RN +Q GLCN TR+IV L + +I A I+ G G+ IPR L+
Sbjct: 183 KISLKKGVPIMQMRNFNQKEGLCNGTRLIVTNLGEQVIEAQIVTGTHAGKMVSIPRFILS 242
Query: 682 PSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSR 741
P S+ PF +R+QFP+ +C+AMTI K+QGQSL LYLP PVF+H QLYVALSRV S
Sbjct: 243 PPQSEHPFTLRRQQFPMRVCYAMTIIKNQGQSLKSDVLYLPNPVFSHVQLYVALSRVTSP 302
Query: 742 KGLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
GL +L DD+ ++ +N++Y+E +++
Sbjct: 303 IGLTILHGDDQ--KNDEVKNIVYKEFYND 329
>At2g07620 putative helicase
Length = 1241
Score = 189 bits (479), Expect = 7e-48
Identities = 91/186 (48%), Positives = 130/186 (68%), Gaps = 1/186 (0%)
Query: 585 YLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLC 644
YLS D++ D S +N FT FLN K S + NH + LK+G P+ML++NID GLC
Sbjct: 1054 YLSADSIDPQDPAS-LNNPVFTPYFLNSIKLSGLSNHNLTLKIGTPVMLLKNIDPKGGLC 1112
Query: 645 NDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAM 704
N TR+ V + +I+ A ++ G+++ + I + ++PS++ +PF+ +RRQFP+ + FAM
Sbjct: 1113 NGTRLQVTQMGNHILEARVITGDRVRDKVIIIKAQISPSDTKLPFRMRRRQFPIAVAFAM 1172
Query: 705 TINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMY 764
I KSQGQSL V +YLPRPVF+HGQLYVALSRV S+KGLK+LI+D EG + T NV++
Sbjct: 1173 RIKKSQGQSLKEVEIYLPRPVFSHGQLYVALSRVTSKKGLKVLIVDKEGNTQSQTMNVVF 1232
Query: 765 QEVFDN 770
+E+F N
Sbjct: 1233 KEIFQN 1238
Score = 115 bits (288), Expect = 9e-26
Identities = 58/96 (60%), Positives = 71/96 (73%), Gaps = 4/96 (4%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N K+ASLIIWDE PM++++CFE+LDRSLND+I PFGG VVV GGDFRQ+L
Sbjct: 943 NMLKEASLIIWDEAPMMSRYCFESLDRSLNDVIGNVDGK----PFGGKVVVFGGDFRQVL 998
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNMIL 481
VI R+EIV +A+NSSYLW+HC V+ LT NM L
Sbjct: 999 HVIHGAGRAEIVLAALNSSYLWEHCNVLTLTKNMSL 1034
>At3g51690 putative protein
Length = 374
Score = 171 bits (432), Expect = 2e-42
Identities = 98/295 (33%), Positives = 165/295 (55%), Gaps = 30/295 (10%)
Query: 476 TVNMILQNATSTSSPAEIKE-FADWLLQVGDGTVKTIDEEETLIEIPPNLLIEQCKEPLL 534
+++++L +++ T I E + W++ + + ++ ET I+I +LLI + K+P+
Sbjct: 38 SIDLVLLDSSETRIHTTIDEALSRWIMNISGDNINNPNDGETEIDISKDLLITESKDPIK 97
Query: 535 ELVNFAYPKLAHNLQKNSFFQERAILAPTLESVEEINNFMLAMIPGDETEYLSYDTLCKS 594
L+ Y + F + AIL + V++IN++ML+++PG+E E LS D++ S
Sbjct: 98 TLLKEVYGEYFAKSYNPDFCHDSAILCHRDDDVDQINDYMLSLLPGEEKECLSTDSISPS 157
Query: 595 DEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNAL 654
D + E LN K +P+ ++LKVG P+ML+R++D +
Sbjct: 158 PNDD----MFVPLEVLNSIKVPGLPDFKLRLKVGAPVMLLRDLDPS-------------- 199
Query: 655 TKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSL 714
GNK G+ +IPR++ P+ ++ P + +R Q+P+ L FAMTI++SQ +L
Sbjct: 200 ----------RGNKHGKKIWIPRIASYPTETNFPLQMRRTQYPLKLAFAMTIDESQVHTL 249
Query: 715 SHVGLYLPRPVFTHG-QLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVF 768
S VGLYLPR VF+HG Q++VA+S+VKSR GLK+LI D +G +N + F
Sbjct: 250 SKVGLYLPRQVFSHGRQMFVAISKVKSRAGLKVLITDKDGNPQEEAKNYPFTLAF 304
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.347 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,199,276
Number of Sequences: 26719
Number of extensions: 610675
Number of successful extensions: 2679
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 2559
Number of HSP's gapped (non-prelim): 54
length of query: 770
length of database: 11,318,596
effective HSP length: 107
effective length of query: 663
effective length of database: 8,459,663
effective search space: 5608756569
effective search space used: 5608756569
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0178.5


