

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.13
(433 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
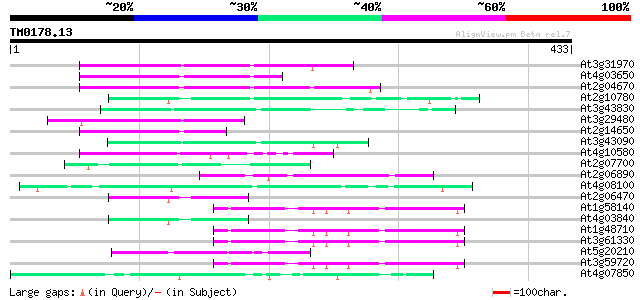
Score E
Sequences producing significant alignments: (bits) Value
At3g31970 hypothetical protein 92 7e-19
At4g03650 putative reverse transcriptase 88 8e-18
At2g04670 putative retroelement pol polyprotein 87 2e-17
At2g10780 pseudogene 82 6e-16
At3g43830 putative protein 81 1e-15
At3g29480 hypothetical protein 77 2e-14
At2g14650 putative retroelement pol polyprotein 73 3e-13
At3g43090 putative protein 69 4e-12
At4g10580 putative reverse-transcriptase -like protein 65 7e-11
At2g07700 putative retroelement pol polyprotein 65 9e-11
At2g06890 putative retroelement integrase 59 7e-09
At4g08100 putative polyprotein 57 2e-08
At2g06470 putative retroelement pol polyprotein 54 1e-07
At1g58140 hypothetical protein 52 8e-07
At4g03840 putative transposon protein 51 1e-06
At1g48710 hypothetical protein 51 1e-06
At3g61330 copia-type polyprotein 51 1e-06
At5g20210 putative protein 50 2e-06
At3g59720 copia-type reverse transcriptase-like protein 50 2e-06
At4g07850 putative polyprotein 50 3e-06
>At3g31970 hypothetical protein
Length = 1329
Score = 91.7 bits (226), Expect = 7e-19
Identities = 66/219 (30%), Positives = 92/219 (41%), Gaps = 12/219 (5%)
Query: 55 EQAAAQTRGLNDFKRQDPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLT 114
E+ + R + +R GG PEEAD W +E F R AE +VD A + L
Sbjct: 138 EEVPSYLRMMEQLQRIGTRYFFGGTSPEEADSWKSRVEHNFGSSRCPAEYRVDLAVHFLE 197
Query: 115 GEAEYWWRGARAMMEADHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEY 174
G+A WWR A H ++W F F KYFP A EA+FL L QG +V EY
Sbjct: 198 GDAHLWWRSVTARRRQAH--MSWADFVAEFNAKYFPQEALDRMEARFLELTQGERSVREY 255
Query: 175 ASKLESLAKHFRYFRGQIDEGYMCERFIDGLSYELQRAVQPLGLNRYQVLVEKTKGIE-- 232
+ L + RG D+ RF+ GL +L+ + L LVE +E
Sbjct: 256 EREFNRLLVYAG--RGMQDDQAQMRRFLRGLRPDLRVRCRVLQYATKAALVETAAEVEED 313
Query: 233 ------AIDNARGKFQSQNKPNQGNGGSAMTNQGRNDKG 265
+ A ++Q + GG Q R ++G
Sbjct: 314 LQRQVVGVSPAVKPKKTQQQVAPSKGGKPAQGQKRKERG 352
>At4g03650 putative reverse transcriptase
Length = 839
Score = 88.2 bits (217), Expect = 8e-18
Identities = 56/157 (35%), Positives = 75/157 (47%), Gaps = 6/157 (3%)
Query: 55 EQAAAQTRGLNDFKRQDPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLT 114
E+ + R + +R SGG PEEAD W ++E+ F R AE +VD + L
Sbjct: 136 EEVPSYLRMMKQLQRIGTEYFSGGTSPEEADSWRSQVERNFGSSRCPAEYRVDLTVHFLE 195
Query: 115 GEAEYWWRGARA-MMEADHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAE 173
G+A WWR A +AD ++W F F KYFP A EA+FL L QG +V E
Sbjct: 196 GDAHLWWRSVTARRRQAD---MSWADFMAEFNAKYFPREALDRMEARFLELTQGVRSVRE 252
Query: 174 YASKLESLAKHFRYFRGQIDEGYMCERFIDGLSYELQ 210
Y K L + RG D+ RF+ GL L+
Sbjct: 253 YDRKFNRLLVYAG--RGMEDDQAQMRRFLRGLRPNLR 287
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 86.7 bits (213), Expect = 2e-17
Identities = 68/235 (28%), Positives = 100/235 (41%), Gaps = 10/235 (4%)
Query: 55 EQAAAQTRGLNDFKRQDPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLT 114
E+ + R + +R S G PEEAD W +++ F R AE +VD A + L
Sbjct: 118 EEVPSYLRMMEQLQRIGTWYFSSGTSPEEADSWRSRVQRNFGSSRCPAEYRVDLAVHFLE 177
Query: 115 GEAEYWWRGARA-MMEADHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAE 173
G+A WWR A +AD ++W F F KYFP A EA+FL L QG +V E
Sbjct: 178 GDAHLWWRSVTARRRQAD---MSWADFVAEFNAKYFPQEALDRMEARFLELTQGEWSVRE 234
Query: 174 YASKLESLAKHFRYFRGQIDEGYMCERFIDGLSYELQRAVQPLGLNRYQVLVEKTKGIEA 233
Y + L + RG D+ RF+ GL +L+ + LVE +E
Sbjct: 235 YDREFNRLLAYAG--RGMEDDQAQMRRFLRGLRPDLRVQCRVSQYATKAALVETAAEVE- 291
Query: 234 IDNARGKFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQ--GQGTASGSF 286
++ + + + Q + K QK+ + P GQG +G F
Sbjct: 292 -EDLQRQVVGVSPAVQTKNTQQQVTPSKGGKPAQGQKRKWDHPSRAGQGGRAGYF 345
>At2g10780 pseudogene
Length = 1611
Score = 82.0 bits (201), Expect = 6e-16
Identities = 82/303 (27%), Positives = 109/303 (35%), Gaps = 43/303 (14%)
Query: 77 GGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWW------RGARAMMEA 130
G DP AD W L++ F R + + D A + L G+A WW RG A
Sbjct: 184 GSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLEGDAHNWWLTVEKRRGDEVRSFA 243
Query: 131 DHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEYASKLESLAKHFRYFRG 190
D F F KYFP A E +L L QG TV EY + L ++ R
Sbjct: 244 D--------FEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRYVG--RE 293
Query: 191 QIDEGYMCERFIDGLSYELQRAVQPLGLNRYQVLVEKTKGIEAIDNARGKFQSQNKPNQG 250
+E RFI GL E++ N LVE+ IE + P +
Sbjct: 294 LEEEQAQLRRFIRGLRIEIRNHCLVRTFNSVSELVERAAMIEEGIEEERYLNREKAPIRN 353
Query: 251 NGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRPLAVSLDGVTCFK 310
N + DK R + K + + +G G N A+ C +
Sbjct: 354 NQSTKPA-----DKKRKFDKVDNTKSDAK---TGECVTCGKNHSGTCWKAIG----ACGR 401
Query: 311 CNQKGHYASHCP-----------EGPLMCWNCNKPGHLARDCRIPKVEDAANVAGARRPT 359
C K H CP E C+ C K GHL R+C PK+ A +R
Sbjct: 402 CGSKDHAIQSCPKMEPGQSKVLGEETRTCFYCGKTGHLKREC--PKL--TAEKQAGQRDN 457
Query: 360 AGG 362
GG
Sbjct: 458 RGG 460
>At3g43830 putative protein
Length = 329
Score = 80.9 bits (198), Expect = 1e-15
Identities = 73/274 (26%), Positives = 101/274 (36%), Gaps = 63/274 (22%)
Query: 71 DPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRGARAMMEA 130
D +GG DP +A W LE F +R E D A+ L GEA+ WW + E
Sbjct: 112 DIETFNGGTDPVKAYNWRNMLECKFKSMRCPVEFWRDLASCYLRGEAQEWWERVK-QREQ 170
Query: 131 DHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEYASKLESLAKHFRYFRG 190
W F+ F +Y P E +FLRL+QG TV +Y + SL + R RG
Sbjct: 171 VGCVDQWSFFKEEFTRRYLPEETIDDLEMKFLRLQQGTKTVRKYEKEFHSLERFERRKRG 230
Query: 191 QIDEGYMCERFIDGLSYELQRAVQPLGLNRYQVLVEKTKGIEAIDNARGKFQSQNKPNQG 250
E + +FI GL +++ + LVEK +E
Sbjct: 231 ---EHELIHKFISGLRVDIRLCCHVRDFDNMIDLVEKAASLEI----------------- 270
Query: 251 NGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRPLAVSLDGVTCFK 310
G ++ R Y R + A GS+ TG
Sbjct: 271 ---------GLEEEAR------YKRIAQEKEAMGSYGQTG-------------------- 295
Query: 311 CNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIP 344
H C E + C+ C GH+ARDCR+P
Sbjct: 296 -----HSKRRCQE--VTCYRCGVAGHIARDCRMP 322
>At3g29480 hypothetical protein
Length = 718
Score = 76.6 bits (187), Expect = 2e-14
Identities = 50/154 (32%), Positives = 66/154 (42%), Gaps = 4/154 (2%)
Query: 30 AATAALVAAQNHAEENARRVQREER--EQAAAQTRGLNDFKRQDPPKLSGGFDPEEADLW 87
AA A V Q VQ+ E+ + R + +R SGG PEEAD W
Sbjct: 4 AAGGAQVPVQAPVAPRVAEVQQRAAVAEEVPSYLRMMEQLQRIGIGYFSGGTSPEEADSW 63
Query: 88 LQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRGARAMMEADHQAITWECFRGAFLDK 147
+E+ F R E +VD A + L G+A WW+ + ++W F F K
Sbjct: 64 RSRVERNFGSSRCPVEYRVDLAVHFLEGDAHLWWKSVTTRRRQAN--MSWADFVAEFNAK 121
Query: 148 YFPNSARAAKEAQFLRLRQGGMTVAEYASKLESL 181
YFP A EA FL L QG +V EY + L
Sbjct: 122 YFPQEALDRMEAHFLELTQGERSVREYDREFNRL 155
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 73.2 bits (178), Expect = 3e-13
Identities = 43/114 (37%), Positives = 55/114 (47%), Gaps = 4/114 (3%)
Query: 55 EQAAAQTRGLNDFKRQDPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLT 114
E+ + R + +R SGG PE AD W +E+ F R AE ++D A + L
Sbjct: 136 EEVPSYLRMMEQLQRIGTGYFSGGTSPEVADSWRSRVERNFGSSRCPAEYRIDLAVHFLE 195
Query: 115 GEAEYWWRGARA-MMEADHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQG 167
G+A WWR A +AD I+W F F KYFP A EA FL L QG
Sbjct: 196 GDAHLWWRSVTARRRQAD---ISWADFVAEFNAKYFPQEALDRMEAHFLELTQG 246
>At3g43090 putative protein
Length = 357
Score = 69.3 bits (168), Expect = 4e-12
Identities = 62/214 (28%), Positives = 87/214 (39%), Gaps = 16/214 (7%)
Query: 76 SGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRGARAMMEADHQAI 135
+GG DP +AD W + LE F R E + D + L EA +WW G H I
Sbjct: 132 TGGIDPVKADDWRKLLENNFENARCPVEFQKDIIVHFLREEASHWWDGVLGNTPVQH-VI 190
Query: 136 TWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEYASKLESLAK-HFRYFRGQIDE 194
WE FR F K+FP A + E F LRQ V E +L L++ R RG E
Sbjct: 191 FWEDFREEFNRKFFPQEAMDSLEDDFEELRQDTKKVRENERELSHLSRFSVRAGRG---E 247
Query: 195 GYMCERFIDGLSYELQRAVQPLGLNRYQVLVEKTKGIEA--IDNARGKFQSQNKPNQGN- 251
M R + GL ++ LVEK IE+ + A+ +Q + Q N
Sbjct: 248 QSMIRRLMRGLRPDILTRCASREFFSMIDLVEKAARIESGLVLEAKYLKNTQARTTQANV 307
Query: 252 --------GGSAMTNQGRNDKGRHYQKKPYVRPQ 277
G + + ND + K+ + P+
Sbjct: 308 FSVVKWVISGRIVRKRNINDGNQTLPKRQAIGPR 341
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 65.1 bits (157), Expect = 7e-11
Identities = 57/211 (27%), Positives = 88/211 (41%), Gaps = 31/211 (14%)
Query: 55 EQAAAQTRGLNDFKRQDPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLT 114
E+ + R + +R D SGG PEEAD W + + F R AE +VD A + L
Sbjct: 132 EEVLSYLRMMEQLQRIDTGYFSGGTSPEEADSWRSRVGRNFGSSRCPAEYRVDLAVHFLE 191
Query: 115 GEAEYWWRGARAMMEADHQAITWECFRGAFLDKYFPNSA--------RAAKEAQFLRLRQ 166
G+A WWR A ++W F F KYFP A +AQ R +
Sbjct: 192 GDAHLWWRSVTARRR--QTDMSWADFVAEFKAKYFPQEALDPYAGQGMEDDQAQMRRFLR 249
Query: 167 G-------GMTVAEYASKLESLAKHFRYFRGQIDEGYMCERFIDGLSYELQRAVQPLGLN 219
G V++YA+K + +++E + +R + G+S VQP
Sbjct: 250 GLRPDLRVRCRVSQYATKAALVET-----AAEVEEDF--QRQVVGVS----PVVQP---K 295
Query: 220 RYQVLVEKTKGIEAIDNARGKFQSQNKPNQG 250
+ Q V +KG + + K+ ++ QG
Sbjct: 296 KTQQQVTPSKGGKPAQGQKRKWDHPSRAGQG 326
>At2g07700 putative retroelement pol polyprotein
Length = 543
Score = 64.7 bits (156), Expect = 9e-11
Identities = 53/201 (26%), Positives = 81/201 (39%), Gaps = 43/201 (21%)
Query: 43 EENARRVQREEREQAAA----------QTRGLNDFKRQDPPKLSGGFDPEEADLWLQELE 92
+++ +++Q +RE++ + + G+ FK G + AD WL++LE
Sbjct: 19 QQHQQQIQESQREESTSAKFLKLVMMMRDLGIRVFK--------GEHNTVVADKWLRDLE 70
Query: 93 KIFTFLRTTAEMKVDYATYLLTGEAEYWWRGARAMMEADHQAITWECFRGAFLDKYFPNS 152
F R K A L +A W A + Q I+WE FR F KYFP
Sbjct: 71 MNFEISRCPDNFKRQIAVNFLDKDARIWRESVTARNQG--QMISWEAFRREFEKKYFPPE 128
Query: 153 ARAAKEAQFLRLRQGGMTVAEYASKLESLAKHFRY-FRGQIDEGYMCERFIDGLSYELQR 211
AR E QF++ RY + GQ DE M RF+ GL E+
Sbjct: 129 ARDRLEQQFMK----------------------RYLYNGQDDEALMVRRFLRGLRLEIWG 166
Query: 212 AVQPLGLNRYQVLVEKTKGIE 232
+Q + + L E+ +E
Sbjct: 167 RLQAVKYDSVNELTERADNVE 187
>At2g06890 putative retroelement integrase
Length = 1215
Score = 58.5 bits (140), Expect = 7e-09
Identities = 45/184 (24%), Positives = 80/184 (43%), Gaps = 16/184 (8%)
Query: 147 KYFPNSARAAKEAQFLRLRQGGMTVAEYASKLESLAKHFRYFRGQIDEGYMC--ERFIDG 204
++ P+ + L QG +V EY ++E+L R I E RF+
Sbjct: 8 RFVPSHYHRELHLKLRNLTQGNRSVEEYYKEMETLM-----LRADISEDREATLSRFLGD 62
Query: 205 LSYELQRAVQPLGLNRYQVLVEKTKGIEAIDNARGKFQSQNKPNQGNGGSAMTNQGRNDK 264
L+ ++Q ++ +Y V +E+ + + K +S ++ + G+G A R ++
Sbjct: 63 LNRDIQDRLE----TQYYVQIEEMLHKAILFEQQVKRKSSSRSSYGSGTIAKPTYQREER 118
Query: 265 GRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRPLAVS-LDGVTCFKCNQKGHYASHCPE 323
Y KP V P+ + + G A ++ S + V C+KC KGHYA+ CP
Sbjct: 119 TSSYHNKPIVSPRAESKPYAAVQDHKGKA----EISTSRVRDVRCYKCQGKGHYANECPN 174
Query: 324 GPLM 327
+M
Sbjct: 175 KRVM 178
>At4g08100 putative polyprotein
Length = 1054
Score = 57.4 bits (137), Expect = 2e-08
Identities = 83/361 (22%), Positives = 122/361 (32%), Gaps = 32/361 (8%)
Query: 8 LTALMAQQVTNSA----AREEAEAQRAATAALVAAQNHAEENARRVQREEREQAAAQTRG 63
L+AL QV A +EE A A +H+ + R R ER G
Sbjct: 35 LSALNLHQVNPDANPCLRQEEPREDEEARNYYSHASSHSSQ---RRHRRERPPPRDPLGG 91
Query: 64 LNDFKRQDPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRG 123
L + P G +P+ W Q++E +F KV A A WW
Sbjct: 92 L----KLKIPAFHGTDNPDTYLEWEQKIELVFLCQECLQSNKVKIAATKFYNYALSWWDQ 147
Query: 124 --ARAMMEADHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEYASKLESL 181
D+ TW + ++ P+ + L QG TV EY ++E+L
Sbjct: 148 LVTSRRRTRDYPIKTWNQLKFVMRKRFVPSYYHRELHQRLRNLVQGSKTVEEYFLEMETL 207
Query: 182 AKHFRYFRGQIDEGYMCERFIDGLSYELQRAVQPLGLNRYQVLVEKTKGIEAIDNARGKF 241
Q D + RF+ GL+ E+Q ++ + ++ K E +
Sbjct: 208 MLRADL---QEDGEAVMSRFMGGLNREIQDRLETQHYVELEEMLHKAVMFEQQIKRKNAR 264
Query: 242 QSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRPLAV 301
S K N +G + Q G KP+V+P+ P G + +
Sbjct: 265 SSHTKTNYSSGKPSY--QKEEKFGYQNDSKPFVKPK-----PVDLDPKGKG----KEVIT 313
Query: 302 SLDGVTCFKCNQKGHYASHCPEGPLMCWNCN-----KPGHLARDCRIPKVEDAANVAGAR 356
+ CFK HYAS C +M N K D VE A AR
Sbjct: 314 RARDIRCFKSQGLRHYASKCSNKRIMVHRDNGEVESKDESAEEDSMEEDVESPARGEFAR 373
Query: 357 R 357
R
Sbjct: 374 R 374
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 54.3 bits (129), Expect = 1e-07
Identities = 37/114 (32%), Positives = 47/114 (40%), Gaps = 14/114 (12%)
Query: 77 GGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWW------RGARAMMEA 130
G DP AD W L++ F R + + D A + L G+A WW RG A
Sbjct: 149 GSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLEGDAHNWWLTVEKRRGDEVRSFA 208
Query: 131 DHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEYASKLESLAKH 184
D F F KYFP A E +L L QG TV EY + L ++
Sbjct: 209 D--------FEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRY 254
>At1g58140 hypothetical protein
Length = 1320
Score = 51.6 bits (122), Expect = 8e-07
Identities = 45/203 (22%), Positives = 93/203 (45%), Gaps = 23/203 (11%)
Query: 158 EAQFLRLRQGGMTVAEYASKLESLAKHFRYFRGQIDEGYMCERFIDGLSYELQRAVQPLG 217
E + L++++G + V++Y S++ ++ + + ++D+ + E+ + L + + V
Sbjct: 119 EFEALQMKEGEL-VSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIV---- 173
Query: 218 LNRYQVLVEKTKGIEA--IDNARGKFQS--QNKPNQGNGGSAMTNQG--RNDKGRHYQKK 271
++E+TK +EA I+ G Q+ + K + + + N + + G+ YQ++
Sbjct: 174 -----TVIEETKDLEAMTIEQLLGSLQAYEEKKKKKEDIVEQVLNMQITKEENGQSYQRR 228
Query: 272 PYVRPQGQGTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNC 331
+ +G+G G P + G + KGH S + + C+NC
Sbjct: 229 GGGQVRGRGRGGYG----NGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKSSVKCYNC 284
Query: 332 NKPGHLARDCRIP---KVEDAAN 351
K GH A +C+ P K E+ AN
Sbjct: 285 GKFGHYASECKAPSNKKFEEKAN 307
>At4g03840 putative transposon protein
Length = 973
Score = 51.2 bits (121), Expect = 1e-06
Identities = 36/114 (31%), Positives = 46/114 (39%), Gaps = 14/114 (12%)
Query: 77 GGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWW------RGARAMMEA 130
G DP AD W L++ F R + + D A + L +A WW RG A
Sbjct: 149 GSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLERDAHNWWLTVEKRRGDEVRSFA 208
Query: 131 DHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEYASKLESLAKH 184
D F F KYFP A E +L L QG TV EY + L ++
Sbjct: 209 D--------FEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRY 254
>At1g48710 hypothetical protein
Length = 1352
Score = 51.2 bits (121), Expect = 1e-06
Identities = 45/203 (22%), Positives = 93/203 (45%), Gaps = 23/203 (11%)
Query: 158 EAQFLRLRQGGMTVAEYASKLESLAKHFRYFRGQIDEGYMCERFIDGLSYELQRAVQPLG 217
E + L++++G + V++Y S++ ++ + + ++D+ + E+ + L + + V
Sbjct: 119 EFEALQMKEGEL-VSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIV---- 173
Query: 218 LNRYQVLVEKTKGIEA--IDNARGKFQS--QNKPNQGNGGSAMTNQG--RNDKGRHYQKK 271
++E+TK +EA I+ G Q+ + K + + + N + + G+ YQ++
Sbjct: 174 -----TVIEETKDLEAMTIEQLLGSLQAYEEKKKKKEDIIEQVLNMQITKEENGQSYQRR 228
Query: 272 PYVRPQGQGTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNC 331
+ +G+G G P + G + KGH S + + C+NC
Sbjct: 229 GGGQVRGRGRGGYG----NGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKSSVKCYNC 284
Query: 332 NKPGHLARDCRIP---KVEDAAN 351
K GH A +C+ P K E+ AN
Sbjct: 285 GKFGHYASECKAPSNKKFEEKAN 307
>At3g61330 copia-type polyprotein
Length = 1352
Score = 50.8 bits (120), Expect = 1e-06
Identities = 44/203 (21%), Positives = 93/203 (45%), Gaps = 23/203 (11%)
Query: 158 EAQFLRLRQGGMTVAEYASKLESLAKHFRYFRGQIDEGYMCERFIDGLSYELQRAVQPLG 217
E + L++++G + V++Y S++ ++ + + ++D+ + E+ + L + + V
Sbjct: 119 EFEALQMKEGEL-VSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIV---- 173
Query: 218 LNRYQVLVEKTKGIEA--IDNARGKFQS--QNKPNQGNGGSAMTNQG--RNDKGRHYQKK 271
++E+TK +EA I+ G Q+ + K + + + N + + G+ YQ++
Sbjct: 174 -----TVIEETKDLEAMTIEQLLGSLQAYEEKKKKKEDIAEQVLNMQITKEENGQSYQRR 228
Query: 272 PYVRPQGQGTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNC 331
+ +G+G G P + G + KGH S + + C+NC
Sbjct: 229 GGGQVRGRGRGGYG----NGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKSSVKCYNC 284
Query: 332 NKPGHLARDCRIP---KVEDAAN 351
K GH A +C+ P K E+ A+
Sbjct: 285 GKFGHYASECKAPSNKKFEEKAH 307
>At5g20210 putative protein
Length = 256
Score = 50.1 bits (118), Expect = 2e-06
Identities = 38/154 (24%), Positives = 70/154 (44%), Gaps = 10/154 (6%)
Query: 79 FDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRGARAMMEADHQAITWE 138
FD + +WL+ +E FT KVD A + G+A W + + ++ +TW
Sbjct: 70 FDGDHLFIWLEIIEVYFTLQLAPEVCKVDTAFRSMEGDALLWLQ----CLRQENPTLTWS 125
Query: 139 CFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEYASKLESLAKHFRYFRGQIDEGYMC 198
+ ++++ A+ E Q LRQ G +V EYA++ + A + Q+ +G
Sbjct: 126 QLKTELIEEFGGGDITASPEEQLASLRQTG-SVDEYANEFRARAAQIGN-KDQLVKG--- 180
Query: 199 ERFIDGLSYELQRAVQPLGLNRYQVLVEKTKGIE 232
F++GL E++ +P ++ + K IE
Sbjct: 181 -MFLNGLKEEIRVMFRPNDAEGPEMAIRTAKAIE 213
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 50.1 bits (118), Expect = 2e-06
Identities = 44/203 (21%), Positives = 93/203 (45%), Gaps = 23/203 (11%)
Query: 158 EAQFLRLRQGGMTVAEYASKLESLAKHFRYFRGQIDEGYMCERFIDGLSYELQRAVQPLG 217
E + L++++G + V++Y S++ ++ + + ++D+ + E+ + L + + V
Sbjct: 119 EFEALQMKEGEL-VSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIV---- 173
Query: 218 LNRYQVLVEKTKGIEA--IDNARGKFQS--QNKPNQGNGGSAMTNQG--RNDKGRHYQKK 271
++E+TK +EA I+ G Q+ + K + + + N + + G+ YQ++
Sbjct: 174 -----TVIEETKDLEAMTIEQLLGSLQAYEEKKKKKEDIVEQVLNMQITKEENGQSYQRR 228
Query: 272 PYVRPQGQGTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNC 331
+ +G+G G P + G + KGH S + + C+NC
Sbjct: 229 GGGQVRGRGRGGYG----NGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKSSVKCYNC 284
Query: 332 NKPGHLARDCRIP---KVEDAAN 351
K GH A +C+ P K ++ AN
Sbjct: 285 GKFGHYASECKAPSNKKFKEKAN 307
>At4g07850 putative polyprotein
Length = 1138
Score = 49.7 bits (117), Expect = 3e-06
Identities = 72/336 (21%), Positives = 121/336 (35%), Gaps = 46/336 (13%)
Query: 1 MEQAIANLTALMAQQVTNSAAREEAEAQRAATAALVAAQNHAEENARRVQREEREQAAAQ 60
ME A LTA + + A+ +N+++ A + +
Sbjct: 1 MEAITATLTATLTTNMAKMMDERFEAARSNQERKNRTMRNNSDREATLSYYSQSSTRTSH 60
Query: 61 TRGLNDFKRQDPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYW 120
R + + PP+ A+L L +I F T KV A A W
Sbjct: 61 RRQRRGQEERHPPR------DNLANLKL----RIPPFHEYTEANKVKVAATEFYDYALSW 110
Query: 121 WRGARAMME--ADHQAITWECFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEYASKL 178
W D TW + ++ P+ + L QG TV EY ++
Sbjct: 111 WDQVVTTKRRLGDDSIETWNQLKNIMKRRFVPSHYHRELHQRLRNLVQGNRTVEEYFKEM 170
Query: 179 ESLAKHFRYFRGQIDEGYMCE----RFIDGLSYELQRAVQPLGLNRYQVLVEKTKGIEAI 234
E+L R + E CE RF+ GL+ ++ L+R++V+ + +E +
Sbjct: 171 ETLM-----LRADVQEE--CEATMSRFMGGLNRDI--------LDRFEVI--HYENLEEL 213
Query: 235 DNARGKFQSQNKPNQGN---GGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPTGG 291
+ F+ Q K S + Q G + KP+V+P+ + +S
Sbjct: 214 FHKAVMFEKQIKRRSAKPSYNSSKPSYQREEKSGFQKEYKPFVKPKVEEISSKG------ 267
Query: 292 NAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLM 327
+ + + D + CFKC+ GHYAS C +M
Sbjct: 268 ---KEKEVTRTRD-LKCFKCHGLGHYASECSNKRIM 299
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,665,840
Number of Sequences: 26719
Number of extensions: 431802
Number of successful extensions: 1550
Number of sequences better than 10.0: 103
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 29
Number of HSP's that attempted gapping in prelim test: 1301
Number of HSP's gapped (non-prelim): 205
length of query: 433
length of database: 11,318,596
effective HSP length: 102
effective length of query: 331
effective length of database: 8,593,258
effective search space: 2844368398
effective search space used: 2844368398
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0178.13


