

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0176a.2
(342 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
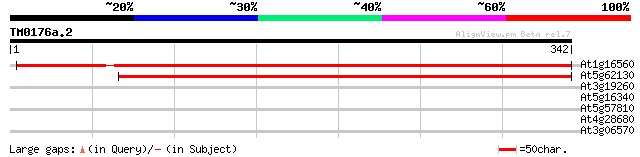
At1g16560 Unknown protein 532 e-152
At5g62130 unknown protein 366 e-101
At3g19260 longevity factor like protein 30 1.5
At5g16340 AMP-binding protein 28 5.5
At5g57810 unknown protein 28 7.2
At4g28680 aromatic amino-acid decarboxylase - like protein 28 7.2
At3g06570 unknown protein 28 9.5
>At1g16560 Unknown protein
Length = 342
Score = 532 bits (1371), Expect = e-152
Identities = 242/338 (71%), Positives = 276/338 (81%), Gaps = 4/338 (1%)
Query: 5 YVFAFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRP 64
Y A L+L + +ASAGD DP YR C+ +C+ +GCV Q CFP C SDG P
Sbjct: 5 YWTALFLLLPCLFCISNASAGDADPDYRTCVSECEISGCVGQLCFPQCNSSSDGG----P 60
Query: 65 WYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASV 124
WY+QEPLYLQWKKW CQ DCRY CM++RE ER+ L PVKYHGKWPF R+ G+QE ASV
Sbjct: 61 WYIQEPLYLQWKKWGCQGDCRYQCMVNRETERETLGQAPVKYHGKWPFKRVLGIQEPASV 120
Query: 125 AFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHS 184
AFS LNLAMHFHGW+SFFI++YYKLPLK + AYYEY GLWHIYGLLS+NSWFWSAVFHS
Sbjct: 121 AFSVLNLAMHFHGWLSFFIMIYYKLPLKQDRTAYYEYVGLWHIYGLLSMNSWFWSAVFHS 180
Query: 185 RDVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYK 244
RDVDLTE LDYSSAV +LG+SLILAILR+F++R EA RVMVSAP++AFV TH++YINFYK
Sbjct: 181 RDVDLTERLDYSSAVAILGFSLILAILRTFDIRVEAARVMVSAPILAFVTTHILYINFYK 240
Query: 245 LDYGWNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGL 304
LDYGWNMIVCV M V QL +WA WA +S HPS WKLW+VVIAGGLAMLLEIYDFPPYEG
Sbjct: 241 LDYGWNMIVCVAMGVSQLFLWARWAAVSSHPSNWKLWVVVIAGGLAMLLEIYDFPPYEGY 300
Query: 305 LDAHALWHATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 342
DAH++WHA TIPLT +WWSFIRDDAEFRTS +KK K
Sbjct: 301 FDAHSIWHAATIPLTILWWSFIRDDAEFRTSSLLKKTK 338
>At5g62130 unknown protein
Length = 276
Score = 366 bits (939), Expect = e-101
Identities = 158/276 (57%), Positives = 216/276 (78%)
Query: 67 MQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAF 126
MQEPLYL+WK+WDCQ+DC+Y CM+ RE+ERK P KY GKWP +YG+QE SVAF
Sbjct: 1 MQEPLYLRWKQWDCQSDCQYECMMTREEERKRNGERPTKYFGKWPLKHVYGIQEPVSVAF 60
Query: 127 SALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRD 186
SAL+LAM F GWVS+FIL+YYKLPL+ ++ YYEY G+ HIY ++ +NS FWS++ HSRD
Sbjct: 61 SALDLAMQFQGWVSYFILVYYKLPLQPNRKTYYEYNGIVHIYAIIVMNSLFWSSICHSRD 120
Query: 187 VDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLD 246
V+LTE LDYSSA +L G+SLILAILRSF+++D++ ++MV+AP++A V TH++Y+NFY LD
Sbjct: 121 VELTERLDYSSATVLAGFSLILAILRSFSIQDQSVKIMVTAPILAVVATHILYLNFYNLD 180
Query: 247 YGWNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLD 306
G + V + ++LV+W +WA L+ HPS+WKL +I+ L + L ++DFPPY+G +D
Sbjct: 181 EGLHWKVIFGIGGIELVVWGLWAALTSHPSKWKLRAFLISSILTLCLRMFDFPPYKGYID 240
Query: 307 AHALWHATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 342
AHALW IPL+Y+WWSF+ DDA FRT++ +KK+K
Sbjct: 241 AHALWRGAGIPLSYLWWSFVCDDAVFRTTVNLKKSK 276
>At3g19260 longevity factor like protein
Length = 296
Score = 30.4 bits (67), Expect = 1.5
Identities = 22/86 (25%), Positives = 40/86 (45%), Gaps = 15/86 (17%)
Query: 128 ALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDV 187
A ++ ++FHGW + + L KL YY +++YG+ +L +W +R
Sbjct: 100 ARDIKLYFHGWPNQELKLSIKL--------YYMCQCGFYVYGVAALLAW------ETRRK 145
Query: 188 DLTETLDYS-SAVILLGYSLILAILR 212
D + + +ILL YS + + R
Sbjct: 146 DFAVMMSHHVITIILLSYSYLTSFFR 171
>At5g16340 AMP-binding protein
Length = 550
Score = 28.5 bits (62), Expect = 5.5
Identities = 20/71 (28%), Positives = 29/71 (40%), Gaps = 8/71 (11%)
Query: 256 VMAVVQLVIWAV-------WAGLSGHPSRW-KLWLVVIAGGLAMLLEIYDFPPYEGLLDA 307
VM+V L+ WAV W H + W W + GG + L +D P L+
Sbjct: 212 VMSVDSLIDWAVPKNPVYLWTLPIFHSNGWTNPWGIAAVGGTNVCLRKFDAPLIYRLIRD 271
Query: 308 HALWHATTIPL 318
H + H P+
Sbjct: 272 HGVTHMCGAPV 282
>At5g57810 unknown protein
Length = 317
Score = 28.1 bits (61), Expect = 7.2
Identities = 23/85 (27%), Positives = 44/85 (51%), Gaps = 6/85 (7%)
Query: 198 AVILLGYSLILAILRSFNVRD----EATRVMVSAPLIA-FVITHVMYINFYKLDYGWNMI 252
++ LLGY++ L +RS++ D + + S L+A FV+++ K ++
Sbjct: 70 SLTLLGYAVWLLYMRSYDCEDILGLPRVQTLASVGLLAVFVVSNAALFLRRKFPMP-ALV 128
Query: 253 VCVVMAVVQLVIWAVWAGLSGHPSR 277
V VV+ ++ L I +AG++ SR
Sbjct: 129 VMVVVLLLMLFIGLAYAGVNEMQSR 153
>At4g28680 aromatic amino-acid decarboxylase - like protein
Length = 545
Score = 28.1 bits (61), Expect = 7.2
Identities = 16/58 (27%), Positives = 26/58 (44%), Gaps = 11/58 (18%)
Query: 110 WPFARIYGMQEAASVAFSALNLAMHFHGWVS-----------FFILLYYKLPLKDGKE 156
W R+YG + + +NLA HF +V+ +F L+ ++L DG E
Sbjct: 421 WMVLRLYGSENLRNFIRDHVNLAKHFEDYVAQDPSFEVVTTRYFSLVCFRLAPVDGDE 478
>At3g06570 unknown protein
Length = 390
Score = 27.7 bits (60), Expect = 9.5
Identities = 9/23 (39%), Positives = 13/23 (56%)
Query: 246 DYGWNMIVCVVMAVVQLVIWAVW 268
DY W MI C +A+ + W +W
Sbjct: 342 DYKWKMIWCAEIALERRTSWEIW 364
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.142 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,252,206
Number of Sequences: 26719
Number of extensions: 350750
Number of successful extensions: 820
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 816
Number of HSP's gapped (non-prelim): 7
length of query: 342
length of database: 11,318,596
effective HSP length: 100
effective length of query: 242
effective length of database: 8,646,696
effective search space: 2092500432
effective search space used: 2092500432
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0176a.2


