

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0173b.3
(286 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
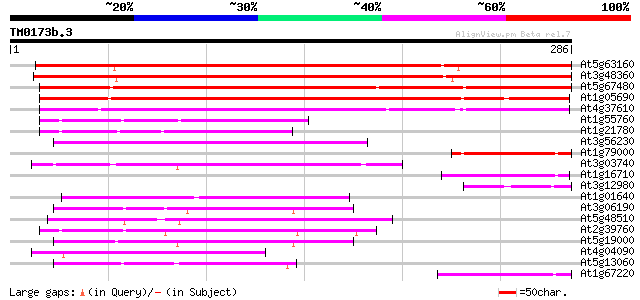
Score E
Sequences producing significant alignments: (bits) Value
At5g63160 unknown protein 350 7e-97
At3g48360 unknown protein 347 6e-96
At5g67480 unknown protein 232 2e-61
At1g05690 unknown protein 229 2e-60
At4g37610 unknown protein 221 3e-58
At1g55760 unknown protein 70 1e-12
At1g21780 unknown protein 65 4e-11
At3g56230 putative protein 64 9e-11
At1g79000 hypothetical protein 58 5e-09
At3g03740 unknown protein 56 3e-08
At1g16710 hypothetical protein 56 3e-08
At3g12980 putative CREB-binding protein 55 6e-08
At1g01640 hypothetical protein 55 6e-08
At3g06190 unknown protein 52 4e-07
At5g48510 putative protein 52 5e-07
At2g39760 unknown protein 50 1e-06
At5g19000 unknown protein 49 3e-06
At4g04090 hypothetical protein 47 1e-05
At5g13060 unknown protein 45 3e-05
At1g67220 hypothetical protein 43 2e-04
>At5g63160 unknown protein
Length = 365
Score = 350 bits (897), Expect = 7e-97
Identities = 173/282 (61%), Positives = 220/282 (77%), Gaps = 10/282 (3%)
Query: 14 EPDIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHR--SSERIIQIHGVPGDAVTAF 71
E D+ I TS IPAHS ILAS+SPVL ++I++PRK SS+++I+I GVP DAV+ F
Sbjct: 24 ETDVEIITSGRRSIPAHSGILASVSPVLTNIIEKPRKIHGGSSKKVIKILGVPCDAVSVF 83
Query: 72 LTFLYSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLC 131
+ FLYS TE+EM++YG+HLLALSHVYMV LKQRCTKG+ +RV ENVVD+LQLARLC
Sbjct: 84 VRFLYSPSVTENEMEKYGIHLLALSHVYMVTQLKQRCTKGVGERVTAENVVDILQLARLC 143
Query: 132 DAPDLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVLRFMDESETRREKSRRSREV 191
DAPDL L+CM + FKTVE+TEGW+FL +HDP+LEL++L+F+D++E+R+++ RR R
Sbjct: 144 DAPDLCLKCMRFIHYKFKTVEQTEGWKFLQEHDPFLELDILQFIDDAESRKKRRRRHRRE 203
Query: 192 QGLYAQLSEAMECLDHICTEGCTLVGPHHVEVDRKS------GPCNKFATCQGVQQLIRH 245
Q LY QLSEAMEC++HICTEGCTLVGP +D KS GPC+ F+TC G+Q LIRH
Sbjct: 204 QNLYLQLSEAMECIEHICTEGCTLVGPSS-NLDNKSTCQAKPGPCSAFSTCYGLQLLIRH 262
Query: 246 YATCTKRMGG-GCLRCKRMWQLFRLHSYGCEHADSCNVPLCR 286
+A C KR+ G GC+RCKRM QL RLHS C+ ++SC VPLCR
Sbjct: 263 FAVCKKRVDGKGCVRCKRMIQLLRLHSSICDQSESCRVPLCR 304
>At3g48360 unknown protein
Length = 364
Score = 347 bits (889), Expect = 6e-96
Identities = 174/286 (60%), Positives = 220/286 (76%), Gaps = 13/286 (4%)
Query: 13 PEPDIFIHTSDGTRIPAHSNILASMSPVLESMIDRP-RKHRS--SERIIQIHGVPGDAVT 69
P D+ I TSD RIPAHS +LAS SPVL +++ +P R++R S+R+I+I GVP DAV+
Sbjct: 32 PTSDVEIVTSDNRRIPAHSGVLASASPVLMNIMKKPMRRYRGCGSKRVIKILGVPCDAVS 91
Query: 70 AFLTFLYSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLAR 129
F+ FLYS TEDEM+RYG+HLLALSHVYMV LKQRC+KG+ QR+ TENVVD+LQLAR
Sbjct: 92 VFIKFLYSSSLTEDEMERYGIHLLALSHVYMVTQLKQRCSKGVVQRLTTENVVDVLQLAR 151
Query: 130 LCDAPDLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVLRFMDESETRREKSRRSR 189
LCDAPD+ LR M + FKTVE+TEGW+F+ +HDP+LEL++L+F+D++E+R+++ RR R
Sbjct: 152 LCDAPDVCLRSMRLIHSQFKTVEQTEGWKFIQEHDPFLELDILQFIDDAESRKKRRRRHR 211
Query: 190 EVQGLYAQLSEAMECLDHICTEGCTLVGPHHVEVD--------RKSGPCNKFATCQGVQQ 241
+ Q LY QLSEAMEC++HICT+GCTLVGP +V VD KS PC F+TC G+Q
Sbjct: 212 KEQDLYMQLSEAMECIEHICTQGCTLVGPSNV-VDNNKKSMTAEKSEPCKAFSTCYGLQL 270
Query: 242 LIRHYATCTKRMGG-GCLRCKRMWQLFRLHSYGCEHADSCNVPLCR 286
LIRH+A C +R GCLRCKRM QLFRLHS C+ DSC VPLCR
Sbjct: 271 LIRHFAVCKRRNNDKGCLRCKRMLQLFRLHSLICDQPDSCRVPLCR 316
>At5g67480 unknown protein
Length = 372
Score = 232 bits (591), Expect = 2e-61
Identities = 123/273 (45%), Positives = 168/273 (61%), Gaps = 5/273 (1%)
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
D+ I+T +G+ I AH+NIL + S V++ M+ + ++H I I GVP DAV F+ FL
Sbjct: 61 DVVIYTDNGSIIYAHANILGTASTVIKGMLKQAKRH-GKWHTISIRGVPHDAVRVFIRFL 119
Query: 76 YSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRV-NTENVVDMLQLARLCDAP 134
YS ++EM+ + MHLL LSH Y+VP LK+ C L + TENVVD+ QLA LCD P
Sbjct: 120 YSSCYEKEEMNEFIMHLLLLSHAYVVPQLKRVCEWHLEHGLLTTENVVDVFQLALLCDFP 179
Query: 135 DLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVL-RFMDESETRREKSRRSREVQG 193
L L + ++F + TE W + K P+LE EV + E+ TR+E+ R+ R Q
Sbjct: 180 RLSLISHRMIMKHFNELSATEAWTAMKKSHPFLEKEVRDSVIIEANTRKERMRK-RNDQR 238
Query: 194 LYAQLSEAMECLDHICTEGCTLVGPHHVEVDRKSGPCNKFATCQGVQQLIRHYATCTKRM 253
+Y+QL EAME L HIC +GC +GPH + CN + C+G++ LIRH+A C R+
Sbjct: 239 IYSQLYEAMEALVHICRDGCKTIGPHDKDFKPNHATCN-YEACKGLESLIRHFAGCKLRV 297
Query: 254 GGGCLRCKRMWQLFRLHSYGCEHADSCNVPLCR 286
GGC+ CKRMWQL LHS C +D C VPLCR
Sbjct: 298 PGGCVHCKRMWQLLELHSRVCAGSDQCRVPLCR 330
>At1g05690 unknown protein
Length = 364
Score = 229 bits (583), Expect = 2e-60
Identities = 113/271 (41%), Positives = 173/271 (63%), Gaps = 5/271 (1%)
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
D ++ T + + PAHS++LA+ SPV+ +++++ R ++ ++IHGVP +AV F+ FL
Sbjct: 55 DTYVETDNKSHFPAHSSVLAAASPVIATLLNQSRD-KNGNTYLKIHGVPCEAVYMFIRFL 113
Query: 76 YSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQR-VNTENVVDMLQLARLCDAP 134
YS E+EM ++ +HLL LSH Y VP LK+ C + L Q +N ENV+D+LQLAR CD
Sbjct: 114 YSSCYEEEEMKKFVLHLLVLSHCYSVPSLKRLCVEILDQGWINKENVIDVLQLARNCDVT 173
Query: 135 DLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVLRFMDESETRREKSRRSREVQGL 194
+ C++ + ++FK+V TEGW+ + + +P LE E++ + ES++R+++ RR E + +
Sbjct: 174 RICFVCLSMVIKDFKSVSSTEGWKVMKRSNPLLEQELIEAVIESDSRKQERRRKLEEREV 233
Query: 195 YAQLSEAMECLDHICTEGCTLVGPHHVEVDRKSGPCNKFATCQGVQQLIRHYATCTKRMG 254
Y QL EAME L HIC EGC +GP + C KF C+G++ +RH+ C R
Sbjct: 234 YLQLYEAMEALVHICREGCGTIGPRDKALKGSHTVC-KFPACKGLEGALRHFLGCKSR-- 290
Query: 255 GGCLRCKRMWQLFRLHSYGCEHADSCNVPLC 285
C CKRMWQL +LHS C+ ++SC V LC
Sbjct: 291 ASCSHCKRMWQLLQLHSCICDDSNSCKVSLC 321
>At4g37610 unknown protein
Length = 368
Score = 221 bits (563), Expect = 3e-58
Identities = 119/273 (43%), Positives = 165/273 (59%), Gaps = 8/273 (2%)
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
D+ IHT D I AHSN++ S V+ M+ + K +S + I I GVP A+ F+ FL
Sbjct: 56 DVLIHTDDNGLIYAHSNVIGMASDVIRGMM-KQHKRKSHRKSISILGVPHHALRVFIRFL 114
Query: 76 YSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGL-SQRVNTENVVDMLQLARLCDAP 134
YS + +M+ + +HLL LSHVY+VPHLK+ C S +N ENV+D+ QLA LCDAP
Sbjct: 115 YSSCYEKQDMEDFAIHLLVLSHVYVVPHLKRVCESEFESSLLNKENVIDVFQLALLCDAP 174
Query: 135 DLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVLRFMD-ESETRREKSRRSREVQG 193
L L C + NF+ V +EGW+ + + P L+ E+LR + E + ++++R+ +E+Q
Sbjct: 175 RLGLLCHRMILNNFEEVSTSEGWQAMKESHPRLQKELLRSVAYELNSLKQRNRKQKEIQ- 233
Query: 194 LYAQLSEAMECLDHICTEGCTLVGPHHVEVDRKSGPCNKFATCQGVQQLIRHYATCTKR- 252
Y QL EAME HIC +GC +GP E S C F C G++QL++H A C R
Sbjct: 234 TYTQLYEAMEAFVHICRDGCREIGPTKTETPHMS--CG-FQACNGLEQLLKHLAGCKLRS 290
Query: 253 MGGGCLRCKRMWQLFRLHSYGCEHADSCNVPLC 285
+ GGC RCKRMWQL LHS C ++ C VPLC
Sbjct: 291 IPGGCSRCKRMWQLLELHSRICVDSEQCKVPLC 323
>At1g55760 unknown protein
Length = 329
Score = 70.5 bits (171), Expect = 1e-12
Identities = 48/137 (35%), Positives = 72/137 (52%), Gaps = 3/137 (2%)
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
DI I+ SDG+ I AH +LA+ SPV SM K + I + +P DA AFL+++
Sbjct: 165 DITINASDGS-IGAHRAVLAARSPVFRSMFLHDLKEKELSEI-NVLDMPLDACQAFLSYI 222
Query: 76 YSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPD 135
Y ED + + + LL + Y + LK+ C L ++T+NV++ LQ A L P+
Sbjct: 223 YGNIQNEDFLI-HRLALLQAAEKYDIADLKEACHLSLLDDIDTKNVLERLQNAYLYQLPE 281
Query: 136 LHLRCMNHLTRNFKTVE 152
L CM +L + K E
Sbjct: 282 LKASCMRYLVKFGKIFE 298
>At1g21780 unknown protein
Length = 326
Score = 65.1 bits (157), Expect = 4e-11
Identities = 41/129 (31%), Positives = 69/129 (52%), Gaps = 3/129 (2%)
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
D+ IHT+DGT + AH IL++ S V +SM + S I I + ++ A L++L
Sbjct: 162 DVIIHTADGT-LSAHKAILSASSTVFKSMFHHDLMEKESS-TIHIDDMSRESCMALLSYL 219
Query: 76 YSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPD 135
Y T++E ++ + LL ++ Y + LK C + L + +N+ NV++ LQ A L
Sbjct: 220 YG-NITQEEFWKHRLALLGAANKYDITDLKAACEESLMEDINSSNVLERLQEAWLYQLEK 278
Query: 136 LHLRCMNHL 144
L C+ +L
Sbjct: 279 LKKGCLMYL 287
>At3g56230 putative protein
Length = 282
Score = 63.9 bits (154), Expect = 9e-11
Identities = 38/161 (23%), Positives = 84/161 (51%), Gaps = 1/161 (0%)
Query: 23 DGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTE 82
DG IPAH +LAS S + ++++D + E I + + + + A L FLY+
Sbjct: 120 DGPPIPAHRALLASKSEIFKNILDSDGCKTAPEYAITLQELNSEQLQALLEFLYTGTLAS 179
Query: 83 DEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMN 142
D++++ L + YM+ +L++ C + + ++ +V+++L ++ L + L C+
Sbjct: 180 DKLEKNVYALFIAADKYMIHYLQELCEQYMLSSLDISSVLNVLDVSDLGSSKTLKEACVG 239
Query: 143 HLTRNFKTVEETEGWRFLTKHDPWLELEVLR-FMDESETRR 182
+ RN V ++ + ++ + L +E+ R F+ E+ ++R
Sbjct: 240 FVVRNMDDVVFSDKYEPFSQKNQHLCVEITRAFLMETRSKR 280
>At1g79000 hypothetical protein
Length = 1691
Score = 58.2 bits (139), Expect = 5e-09
Identities = 25/61 (40%), Positives = 38/61 (61%), Gaps = 2/61 (3%)
Query: 226 KSGPCNKFATCQGVQQLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSCNVPLC 285
+S C ++ C+ V+ L RH C R GGC+ CK+MW L +LH+ C+ ++ C+VP C
Sbjct: 1596 RSAHC-QYPNCRKVKGLFRHGINCKVRASGGCVLCKKMWYLLQLHARACKESE-CHVPRC 1653
Query: 286 R 286
R
Sbjct: 1654 R 1654
>At3g03740 unknown protein
Length = 465
Score = 55.8 bits (133), Expect = 3e-08
Identities = 51/204 (25%), Positives = 88/204 (43%), Gaps = 21/204 (10%)
Query: 12 EPEPDIFIHTSDGTRIPAHSNILASMSPVLES-MIDRPRKHRSSERIIQIHGVPGDAVTA 70
E DI + S G + AH +LA+ SPV ES +D + +R I++ + A
Sbjct: 213 EDGSDITFNVS-GEKFRAHRLVLAARSPVFESEFLDVTGEE---DRDIEVTDMEPKVFKA 268
Query: 71 FLTFLYSRRCTEDE--------------MDRYGMHLLALSHVYMVPHLKQRCTKGLSQRV 116
L ++Y ED D LL + Y +P L C L + +
Sbjct: 269 LLHYIYKDALIEDAESSSSSGSSVGPSASDTLAAKLLGAADKYKLPRLSLMCESVLCKDI 328
Query: 117 NTENVVDMLQLARLCDAPDLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVLRFMD 176
+ ++V ++L LA +A L C+ N V ++G+ +L +H P L+ E+L+ +
Sbjct: 329 SVDSVANILALADRYNASALKSVCLKFAAENLIAVMRSDGFDYLREHCPSLQSELLKTVA 388
Query: 177 ESETRREKSRRSREVQGLYAQLSE 200
E E S + + ++ Q S+
Sbjct: 389 GCE--EELSGGGGKTRSVWGQFSD 410
>At1g16710 hypothetical protein
Length = 1706
Score = 55.8 bits (133), Expect = 3e-08
Identities = 24/65 (36%), Positives = 36/65 (54%), Gaps = 1/65 (1%)
Query: 221 VEVDRKSGPCNKFATCQGVQQLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSC 280
V + P + C+ V+ L RH C R GGC+ CK+MW L +LH+ C+ ++ C
Sbjct: 1605 VHASQCRSPVCLYPNCRKVKGLFRHGLRCKVRASGGCVLCKKMWYLLQLHARACKESE-C 1663
Query: 281 NVPLC 285
+VP C
Sbjct: 1664 DVPRC 1668
>At3g12980 putative CREB-binding protein
Length = 1670
Score = 54.7 bits (130), Expect = 6e-08
Identities = 25/55 (45%), Positives = 32/55 (57%), Gaps = 4/55 (7%)
Query: 232 KFATCQGVQQLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSCNVPLCR 286
++ C+ ++ LIRH C R GC+ CK+MW LFRLHS C C VP CR
Sbjct: 1582 QYPRCRVIKGLIRHGLVCKTR---GCIACKKMWSLFRLHSRNCRD-PQCKVPKCR 1632
>At1g01640 hypothetical protein
Length = 207
Score = 54.7 bits (130), Expect = 6e-08
Identities = 37/147 (25%), Positives = 67/147 (45%), Gaps = 2/147 (1%)
Query: 27 IPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEMD 86
IP H +LA+ S V +M+D S E I + + D + + L FLYS
Sbjct: 37 IPTHKAVLAARSKVFRNMLDSDECKTSPEESITLPDLSHDELKSLLEFLYSGNLKAPYNQ 96
Query: 87 RYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTR 146
++L A Y + +L+ C +++ NV+D+L+LA + L +NH+ +
Sbjct: 97 YRSLYLAA--DKYDISYLQDVCRNHFIASLSSRNVLDILELASIPCDTILKDAAINHIVK 154
Query: 147 NFKTVEETEGWRFLTKHDPWLELEVLR 173
+ + V + + +P L +E+ R
Sbjct: 155 HMEEVVVPMKYETFVQRNPDLSVEITR 181
>At3g06190 unknown protein
Length = 406
Score = 52.0 bits (123), Expect = 4e-07
Identities = 46/166 (27%), Positives = 68/166 (40%), Gaps = 15/166 (9%)
Query: 23 DGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTE 82
DG PAH +LA+ S V + + P + ++ II I V L F+Y
Sbjct: 209 DGETFPAHKLVLAARSAVFRAQLFGPLRSENTNCII-IEDVQAPIFKMLLHFIYWDE-MP 266
Query: 83 DEMDRYG-----------MHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLC 131
D D G HLLA + Y + L+ C L + ++ V L LA
Sbjct: 267 DMQDLIGTDLKWASTLVAQHLLAAADRYALERLRTICESKLCEGISINTVATTLALAEQH 326
Query: 132 DAPDLHLRCMNH--LTRNFKTVEETEGWRFLTKHDPWLELEVLRFM 175
L C+ L N K V ET+G+ +L + P L E+L ++
Sbjct: 327 HCFQLKAACLKFIALPENLKAVMETDGFDYLKESCPSLLSELLEYV 372
>At5g48510 putative protein
Length = 224
Score = 51.6 bits (122), Expect = 5e-07
Identities = 42/184 (22%), Positives = 88/184 (47%), Gaps = 12/184 (6%)
Query: 20 HTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERI--IQIHGVPGDAVTAFLTFLYS 77
++ +G+ I AH ILAS S V ++M + S++ + I + + + + AF+ F+
Sbjct: 32 NSDEGSAISAHKLILASRSEVFKNMFELDEFKTSTKHVETITLSEMKHEELEAFVEFI-- 89
Query: 78 RRCTEDEM-----DRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCD 132
C++ M ++ L + Y + HL+ C L ++ N +D+L LA++
Sbjct: 90 --CSDGSMLSANVKQHARSLYLAADKYEILHLRDLCRTELISSLSLLNSLDLLVLAQIPF 147
Query: 133 APDLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVLRFMDESETRREKSRRS-REV 191
L+ +++ N T+ ++ ++ +P L +E+++ + T R S S +
Sbjct: 148 DKVLNDESFSYIKNNISTIASSDEFKLFVAGNPNLAVEIMKVSLITLTLRCSSCGSNHSL 207
Query: 192 QGLY 195
QG+Y
Sbjct: 208 QGMY 211
>At2g39760 unknown protein
Length = 408
Score = 50.1 bits (118), Expect = 1e-06
Identities = 48/187 (25%), Positives = 81/187 (42%), Gaps = 17/187 (9%)
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
DI D T AH ILA+ SPV + P + + +RI+ I + A L+F+
Sbjct: 195 DIAFQVGDET-YKAHKLILAARSPVFRAQFFGPIGNNNVDRIV-IDDIEPSIFKAMLSFI 252
Query: 76 YSR----------RCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDML 125
Y+ + HLLA + +Y + LK C L ++++ +NV L
Sbjct: 253 YTDVLPNVHEITGSTSASSFTNMIQHLLAAADLYDLARLKILCEVLLCEKLDVDNVATTL 312
Query: 126 QLARLCDAPDLHLRCMNHLT--RNFKTVEETEGWRFLTKHDPWLELEVLRFM---DESET 180
LA L C+ + N V ++EG++ L + P L E+L + D+S T
Sbjct: 313 ALAEQHQFLQLKAFCLEFVASPANLGAVMKSEGFKHLKQSCPTLLSELLNTVAAADKSST 372
Query: 181 RREKSRR 187
+ +++
Sbjct: 373 SGQSNKK 379
>At5g19000 unknown protein
Length = 407
Score = 48.9 bits (115), Expect = 3e-06
Identities = 43/165 (26%), Positives = 67/165 (40%), Gaps = 13/165 (7%)
Query: 23 DGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTE 82
DG AH +LA+ SPV + + P R++ + I I + L F+Y +
Sbjct: 210 DGETFNAHKLVLATRSPVFNAQLFGPLGDRNT-KCITIEDMEAPIFKVLLHFIYWDELPD 268
Query: 83 DE----------MDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCD 132
+ HLLA + Y + LK C L + V V L LA
Sbjct: 269 MQELIGTDSTLASTLVAQHLLAAADRYALERLKAICESKLCEGVAINTVATTLALAEQHH 328
Query: 133 APDLHLRCMNH--LTRNFKTVEETEGWRFLTKHDPWLELEVLRFM 175
L C+ L N K V +T+G+ +L + P L E+L+++
Sbjct: 329 CLQLKAVCLKFVALPENLKAVMQTDGFDYLKESCPSLLTELLQYV 373
>At4g04090 hypothetical protein
Length = 192
Score = 47.0 bits (110), Expect = 1e-05
Identities = 34/124 (27%), Positives = 60/124 (47%), Gaps = 5/124 (4%)
Query: 12 EPEPDIFIHTSDGTR---IPAHSNILASMSPVLESMIDRPRKHRSSE-RIIQIHGVPGDA 67
E + D+ + D + I AH IL++ S V E M + + SS+ I + + +
Sbjct: 20 EQQVDVRLMARDSEKDAAISAHKRILSARSEVFEEMFESDKYKASSKLETITLSEMKHEV 79
Query: 68 VTAFLTFLYSR-RCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQ 126
+ AF+ F YS ++ ++ M L + + Y +P L C K L +N N + +LQ
Sbjct: 80 LEAFVDFTYSDGSMLSEKAKQHAMSLYSAAKDYEIPRLWCLCRKELIASLNMSNALRVLQ 139
Query: 127 LARL 130
LA++
Sbjct: 140 LAQI 143
>At5g13060 unknown protein
Length = 709
Score = 45.4 bits (106), Expect = 3e-05
Identities = 31/128 (24%), Positives = 60/128 (46%), Gaps = 7/128 (5%)
Query: 23 DGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTE 82
DG + AH L + S + +M D K R+++ + +I + + + F+YS R
Sbjct: 547 DGKQFYAHKIGLVASSDIFRAMFDGLYKERNAQNV-EIPNIRWEVFELMMKFIYSGRINI 605
Query: 83 DEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRC-- 140
+ LL + Y++ LK++C ++Q + +N+ +M +LA +A L C
Sbjct: 606 AK--HLAKDLLVAADQYLLEGLKRQCEYTIAQEICLDNIPEMYELADTFNASALRRACTL 663
Query: 141 --MNHLTR 146
+ H T+
Sbjct: 664 FVLEHFTK 671
>At1g67220 hypothetical protein
Length = 1357
Score = 43.1 bits (100), Expect = 2e-04
Identities = 21/69 (30%), Positives = 32/69 (45%), Gaps = 2/69 (2%)
Query: 219 HHVEVDRKSGPCNKFATCQGVQQLIRHYATCT-KRMGGGCLRCKRMWQLFRLHSYGCEHA 277
H + K+ + C V+ L H C ++ G C C ++WQ R+H Y C+
Sbjct: 1290 HALLCQHKTTKSCSYPKCHEVKALFTHNVQCKIRKKGTRCNTCYKLWQTIRIHVYHCQDL 1349
Query: 278 DSCNVPLCR 286
+C VP CR
Sbjct: 1350 -NCPVPQCR 1357
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.136 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,519,159
Number of Sequences: 26719
Number of extensions: 260693
Number of successful extensions: 891
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 833
Number of HSP's gapped (non-prelim): 45
length of query: 286
length of database: 11,318,596
effective HSP length: 98
effective length of query: 188
effective length of database: 8,700,134
effective search space: 1635625192
effective search space used: 1635625192
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0173b.3


