

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0162a.10
(265 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
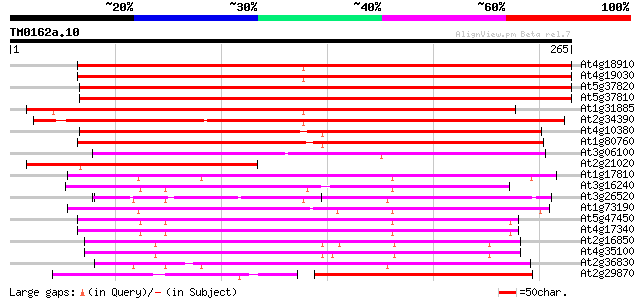
Score E
Sequences producing significant alignments: (bits) Value
At4g18910 major intrinsic protein (MIP)- like 319 1e-87
At4g19030 nodulin-26 like protein 314 3e-86
At5g37820 Membrane integral protein (MIP) -like 273 9e-74
At5g37810 Membrane integral protein (MIP) -like 271 3e-73
At1g31885 major intrinsic protein - like 258 2e-69
At2g34390 putative aquaporin (plasma membrane intrinsic protein) 231 3e-61
At4g10380 major intrinsic protein (MIP) - like 219 2e-57
At1g80760 nodulin-like protein 218 3e-57
At3g06100 putative major intrinsic protein 152 2e-37
At2g21020 putative major intrinsic (channel) protein 125 3e-29
At1g17810 putative protein 114 7e-26
At3g16240 delta tonoplast integral protein (delta-TIP) 110 6e-25
At3g26520 salt-stress induced tonoplast intrinsic protein 105 3e-23
At1g73190 tonoplast intrinsic protein, alpha (alpha-TIP) 103 1e-22
At5g47450 membrane channel protein-like; aquaporin (tonoplast in... 103 1e-22
At4g17340 membrane channel like protein 102 2e-22
At2g16850 putative aquaporin (plasma membrane intrinsic protein) 100 8e-22
At4g35100 plasma membrane intrinsic protein PIP3 100 1e-21
At2g36830 putative aquaporin (tonoplast intrinsic protein gamma) 100 1e-21
At2g29870 putative aquaporin (plasma membrane intrinsic protein) 100 1e-21
>At4g18910 major intrinsic protein (MIP)- like
Length = 294
Score = 319 bits (817), Expect = 1e-87
Identities = 155/240 (64%), Positives = 195/240 (80%), Gaps = 7/240 (2%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
SV F+QKL+AE +GT+FLIF GCA++ VN +D VTLPGIA+VWGL +MVL+YS+GHIS
Sbjct: 47 SVPFLQKLMAEVLGTYFLIFAGCAAVAVNTQHDKAVTLPGIAIVWGLTVMVLVYSLGHIS 106
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLF-------SGTHDQFS 145
GAHFNPAVT AFA+ RFP QV Y+ SQ++G+ LA+ L++LF SG HD F
Sbjct: 107 GAHFNPAVTIAFASCGRFPLKQVPAYVISQVIGSTLAAATLRLLFGLDQDVCSGKHDVFV 166
Query: 146 GTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITG 205
GT+PSG+NLQ+FVIEFI TF LMFVIS VATDNRAIGE+AG+A+GST+LLN++I+GP++G
Sbjct: 167 GTLPSGSNLQSFVIEFIITFYLMFVISGVATDNRAIGELAGLAVGSTVLLNVIIAGPVSG 226
Query: 206 ASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
ASMNP R+LGPA+ +S YR + +Y VS I GAV+GAWV+N++RYTDKPL EITK S LK
Sbjct: 227 ASMNPGRSLGPAMVYSCYRGLWIYIVSPIVGAVSGAWVYNMVRYTDKPLREITKSGSFLK 286
>At4g19030 nodulin-26 like protein
Length = 296
Score = 314 bits (805), Expect = 3e-86
Identities = 153/240 (63%), Positives = 191/240 (78%), Gaps = 7/240 (2%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
SV F+QKL+AEF+GT+FL+FTGCAS+VVN NDNVVTLPGIA+VWGL +MVLIYS+GHIS
Sbjct: 50 SVPFLQKLIAEFLGTYFLVFTGCASVVVNMQNDNVVTLPGIAIVWGLTIMVLIYSLGHIS 109
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLF-------SGTHDQFS 145
GAH NPAVT AFA+ RFP QV Y+ SQ++G+ LA+ L++LF SG HD F
Sbjct: 110 GAHINPAVTIAFASCGRFPLKQVPAYVISQVIGSTLAAATLRLLFGLDHDVCSGKHDVFI 169
Query: 146 GTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITG 205
G+ P G++LQAF +EFI TF LMF+IS VATDNRAIGE+AG+AIGST+LLN+LI+ P++
Sbjct: 170 GSSPVGSDLQAFTMEFIVTFYLMFIISGVATDNRAIGELAGLAIGSTVLLNVLIAAPVSS 229
Query: 206 ASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
ASMNP R+LGPA+ + Y+ I +Y V+ GA+AGAWV+N +RYTDKPL EITK S LK
Sbjct: 230 ASMNPGRSLGPALVYGCYKGIWIYLVAPTLGAIAGAWVYNTVRYTDKPLREITKSGSFLK 289
>At5g37820 Membrane integral protein (MIP) -like
Length = 283
Score = 273 bits (697), Expect = 9e-74
Identities = 130/232 (56%), Positives = 172/232 (74%)
Query: 34 VFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISG 93
V QKL+AE +GT+F+IF+GC +VVN +T PGI + WGL++MV+IYS GHISG
Sbjct: 39 VCLTQKLIAEMIGTYFIIFSGCGVVVVNVLYGGTITFPGICVTWGLIVMVMIYSTGHISG 98
Query: 94 AHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTN 153
AHFNPAVT FA +RFPW QV YI +QL G++LAS L+++F+ T F GT P+ ++
Sbjct: 99 AHFNPAVTVTFAVFRRFPWYQVPLYIGAQLTGSLLASLTLRLMFNVTPKAFFGTTPTDSS 158
Query: 154 LQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPART 213
QA V E I +FLLMFVIS VATD+RA GE+AGIA+G T++LN+ ++GPI+GASMNPAR+
Sbjct: 159 GQALVAEIIISFLLMFVISGVATDSRATGELAGIAVGMTIILNVFVAGPISGASMNPARS 218
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
LGPAI +Y+ I VY V G AG +V+N +R+TDKPL E+TK +S L+
Sbjct: 219 LGPAIVMGRYKGIWVYIVGPFVGIFAGGFVYNFMRFTDKPLRELTKSASFLR 270
>At5g37810 Membrane integral protein (MIP) -like
Length = 283
Score = 271 bits (693), Expect = 3e-73
Identities = 125/232 (53%), Positives = 173/232 (73%)
Query: 34 VFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISG 93
V QKL+AE +GT+F++F+GC +VVN +T PGI + WGL++MV+IYS GHISG
Sbjct: 39 VCLTQKLIAEMIGTYFIVFSGCGVVVVNVLYGGTITFPGICVTWGLIVMVMIYSTGHISG 98
Query: 94 AHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTN 153
AHFNPAVT FA +RFPW QV YI +Q G++LAS L+++F T + F GT P+ +
Sbjct: 99 AHFNPAVTVTFAIFRRFPWHQVPLYIGAQFAGSLLASLTLRLMFKVTPEAFFGTTPADSP 158
Query: 154 LQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPART 213
+A V E I +FLLMFVIS VATDNRA+GE+AGIA+G T+++N+ ++GPI+GASMNPAR+
Sbjct: 159 ARALVAEIIISFLLMFVISGVATDNRAVGELAGIAVGMTIMVNVFVAGPISGASMNPARS 218
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
LGPA+ Y+ I VY V + G ++G +V+N++R+TDKPL E+TK +S L+
Sbjct: 219 LGPALVMGVYKHIWVYIVGPVLGVISGGFVYNLIRFTDKPLRELTKSASFLR 270
>At1g31885 major intrinsic protein - like
Length = 294
Score = 258 bits (659), Expect = 2e-69
Identities = 135/242 (55%), Positives = 178/242 (72%), Gaps = 11/242 (4%)
Query: 9 ETHDVVLSVNK----DASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNN 64
+T VVL + D S++ + S SV FVQKL+ EFVGTF +IF GC++IVVN+
Sbjct: 10 QTQTVVLDIENYQSIDDSRSSDLSAPLVSVSFVQKLIGEFVGTFTMIFAGCSAIVVNETY 69
Query: 65 DNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLL 124
VTLPGIALVWGLV+ V+IYS+GH+SGAHFNPAV+ AFA++K+FP+ QV YIA+QLL
Sbjct: 70 GKPVTLPGIALVWGLVVTVMIYSIGHVSGAHFNPAVSIAFASSKKFPFNQVPGYIAAQLL 129
Query: 125 GAVLASGILKMLF-------SGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATD 177
G+ LA+ +L+++F S D + GT PS +N +FV+EFI TF LMFVISAVATD
Sbjct: 130 GSTLAAAVLRLVFHLDDDVCSLKGDVYVGTYPSNSNTTSFVMEFIATFNLMFVISAVATD 189
Query: 178 NRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGA 237
RA G AGIAIG+T++L+IL SGPI+GASMNPAR+LGPA+ Y+ + +Y VS + GA
Sbjct: 190 KRATGSFAGIAIGATIVLDILFSGPISGASMNPARSLGPALIWGCYKDLWLYIVSPVIGA 249
Query: 238 VA 239
++
Sbjct: 250 LS 251
>At2g34390 putative aquaporin (plasma membrane intrinsic protein)
Length = 288
Score = 231 bits (589), Expect = 3e-61
Identities = 125/258 (48%), Positives = 172/258 (66%), Gaps = 12/258 (4%)
Query: 12 DVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLP 71
D L NK S SS SV F+QKL+AE VGT++LIF GCA+I VN +++VVTL
Sbjct: 26 DTSLPSNKHES----SSPPLLSVHFLQKLLAELVGTYYLIFAGCAAIAVNAQHNHVVTLV 81
Query: 72 GIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASG 131
GIA+VWG+V+MVL+Y +GH+S AHFNPAVT A A+++RFP QV YI Q++G+ LAS
Sbjct: 82 GIAVVWGIVIMVLVYCLGHLS-AHFNPAVTLALASSQRFPLNQVPAYITVQVIGSTLASA 140
Query: 132 ILKMLF-------SGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEM 184
L++LF S HD F G+ PSG++LQAFV+EFI T LM V+ AV T R E+
Sbjct: 141 TLRLLFDLNNDVCSKKHDVFLGSSPSGSDLQAFVMEFIITGFLMLVVCAVTTTKRTTEEL 200
Query: 185 AGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVF 244
G+ IG+T+ LN++ +G ++GASMNPAR++GPA+ Y+ I +Y ++ GAV+GA +
Sbjct: 201 EGLIIGATVTLNVIFAGEVSGASMNPARSIGPALVWGCYKGIWIYLLAPTLGAVSGALIH 260
Query: 245 NILRYTDKPLHEITKGSS 262
+L E +K S
Sbjct: 261 KMLPSIQNAEPEFSKTGS 278
>At4g10380 major intrinsic protein (MIP) - like
Length = 304
Score = 219 bits (557), Expect = 2e-57
Identities = 115/220 (52%), Positives = 152/220 (68%), Gaps = 5/220 (2%)
Query: 34 VFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISG 93
V +KL AEFVGTF LIFT A +VN+ D TL G A GL +M++I S GHISG
Sbjct: 74 VSLTRKLGAEFVGTFILIFTATAGPIVNQKYDGAETLIGNAACAGLAVMIIILSTGHISG 133
Query: 94 AHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG--TIPSG 151
AH NP++T AFA + FPW V YIA+Q+ ++ AS LK +F H SG TIPS
Sbjct: 134 AHLNPSLTIAFAALRHFPWAHVPAYIAAQVSASICASFALKGVF---HPFMSGGVTIPSV 190
Query: 152 TNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPA 211
+ QAF +EFI TF+L+FV++AVATD RA+GE+AGIA+G+T++LNIL++GP TG SMNP
Sbjct: 191 SLGQAFALEFIITFILLFVVTAVATDTRAVGELAGIAVGATVMLNILVAGPSTGGSMNPV 250
Query: 212 RTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTD 251
RTLGPA+ YR++ VY V+ GA++GA V+ ++ D
Sbjct: 251 RTLGPAVASGNYRSLWVYLVAPTLGAISGAAVYTGVKLND 290
Score = 33.9 bits (76), Expect = 0.093
Identities = 32/106 (30%), Positives = 45/106 (42%), Gaps = 12/106 (11%)
Query: 30 TYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVG 89
T SV Q EF+ TF L+F A V + V L GIA+ G +M+ I G
Sbjct: 186 TIPSVSLGQAFALEFIITFILLFVVTA---VATDTRAVGELAGIAV--GATVMLNILVAG 240
Query: 90 HISGAHFNPAVTFAFATTK---RFPWIQVAPYIASQLLGAVLASGI 132
+G NP T A R W+ Y+ + LGA+ + +
Sbjct: 241 PSTGGSMNPVRTLGPAVASGNYRSLWV----YLVAPTLGAISGAAV 282
>At1g80760 nodulin-like protein
Length = 305
Score = 218 bits (554), Expect = 3e-57
Identities = 117/222 (52%), Positives = 152/222 (67%), Gaps = 5/222 (2%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
+V +KL AEFVGT LIF G A+ +VN+ D TL G A GL +M++I S GHIS
Sbjct: 75 NVSLYRKLGAEFVGTLILIFAGTATAIVNQKTDGAETLIGCAASAGLAVMIVILSTGHIS 134
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG--TIPS 150
GAH NPAVT AFA K FPW V YI +Q++ +V A+ LK +F T SG T+P+
Sbjct: 135 GAHLNPAVTIAFAALKHFPWKHVPVYIGAQVMASVSAAFALKAVFEPT---MSGGVTVPT 191
Query: 151 GTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNP 210
QAF +EFI +F LMFV++AVATD RA+GE+AGIA+G+T++LNILI+GP T ASMNP
Sbjct: 192 VGLSQAFALEFIISFNLMFVVTAVATDTRAVGELAGIAVGATVMLNILIAGPATSASMNP 251
Query: 211 ARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDK 252
RTLGPAI + YRAI VY + I GA+ GA + I++ ++
Sbjct: 252 VRTLGPAIAANNYRAIWVYLTAPILGALIGAGTYTIVKLPEE 293
>At3g06100 putative major intrinsic protein
Length = 275
Score = 152 bits (383), Expect = 2e-37
Identities = 80/215 (37%), Positives = 129/215 (59%), Gaps = 2/215 (0%)
Query: 40 LVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPA 99
++AE VGTF L+F+ C I + + V L A+ GL ++V++YS+GHISGAH NP+
Sbjct: 48 VMAELVGTFILMFSVCGVISSTQLSGGHVGLLEYAVTAGLSVVVVVYSIGHISGAHLNPS 107
Query: 100 VTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVI 159
+T AFA FPW QV YI +Q LGA A+ ++ + G + T P+ + + AF +
Sbjct: 108 ITIAFAVFGGFPWSQVPLYITAQTLGATAAT-LVGVSVYGVNADIMATKPALSCVSAFFV 166
Query: 160 EFITTFLLMFVISAV-ATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAI 218
E I T +++F+ SA+ ++ +G + G IG+ + L +LI+GPI+G SMNPAR+LGPA+
Sbjct: 167 ELIATSIVVFLASALHCGPHQNLGNLTGFVIGTVISLGVLITGPISGGSMNPARSLGPAV 226
Query: 219 FHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKP 253
+ + +Y + + GA+ G + + +P
Sbjct: 227 VAWDFEDLWIYMTAPVIGAIIGVLTYRSISLKTRP 261
Score = 40.0 bits (92), Expect = 0.001
Identities = 25/96 (26%), Positives = 47/96 (48%), Gaps = 6/96 (6%)
Query: 154 LQAFVIEFITTFLLMFVISAVATDNRAIGEMAG-----IAIGSTLLLNILISGPITGASM 208
L+ + E + TF+LMF + V + + G G + G ++++ + G I+GA +
Sbjct: 45 LRIVMAELVGTFILMFSVCGVISSTQLSGGHVGLLEYAVTAGLSVVVVVYSIGHISGAHL 104
Query: 209 NPARTLGPAIFHS-KYRAIVVYFVSTIFGAVAGAWV 243
NP+ T+ A+F + + +Y + GA A V
Sbjct: 105 NPSITIAFAVFGGFPWSQVPLYITAQTLGATAATLV 140
>At2g21020 putative major intrinsic (channel) protein
Length = 262
Score = 125 bits (313), Expect = 3e-29
Identities = 63/113 (55%), Positives = 80/113 (70%), Gaps = 4/113 (3%)
Query: 9 ETHDVVLSVNKDASKTIESSDTYT----SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNN 64
+T VV + D S S + SV FVQKL+ EFVGTF +IF GC++IVVN+
Sbjct: 21 QTQTVVSDIENDRSIDDSQSSVLSGPLVSVSFVQKLIGEFVGTFSMIFAGCSAIVVNETY 80
Query: 65 DNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAP 117
VTLPGIALVWGLV+ V+IYS+GH+SGAHFNPAV+ AFA++K+FP+ Q P
Sbjct: 81 GKPVTLPGIALVWGLVVTVMIYSIGHVSGAHFNPAVSIAFASSKKFPFNQFHP 133
>At1g17810 putative protein
Length = 267
Score = 114 bits (284), Expect = 7e-26
Identities = 78/245 (31%), Positives = 125/245 (50%), Gaps = 14/245 (5%)
Query: 28 SDTYTSVFFVQKLVAEFVGTFFLIFTGCASIV-VNKNNDNVVTLPGIALVWGLVLMVLIY 86
+D T ++ +AEF+ TF +F G SI+ ++K + G GLVL+ L +
Sbjct: 14 ADEATHPDSIRATLAEFLSTFVFVFAGEGSILALDKLYWDTAAHTGTNTPGGLVLVALAH 73
Query: 87 SVG---------HISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLF 137
++ ++SG H NPAVTFA R I+ Y +QL+GA+LA +L++
Sbjct: 74 ALALFAAVSAAINVSGGHVNPAVTFAALIGGRISVIRAIYYWVAQLIGAILACLLLRLAT 133
Query: 138 SGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLL 195
+G + L ++E I TF L++V+ + A D + +IG +A +AIG +
Sbjct: 134 NGLRPVGFHVASGVSELHGLLMEIILTFALVYVVYSTAIDPKRGSIGIIAPLAIGLIVGA 193
Query: 196 NILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFN--ILRYTDKP 253
NIL+ GP GASMNPAR GPA+ ++ +Y+V G A ++ I+ ++P
Sbjct: 194 NILVGGPFDGASMNPARAFGPALVGWRWSNHWIYWVGPFIGGALAALIYEYMIIPSVNEP 253
Query: 254 LHEIT 258
H T
Sbjct: 254 PHHST 258
>At3g16240 delta tonoplast integral protein (delta-TIP)
Length = 250
Score = 110 bits (276), Expect = 6e-25
Identities = 75/221 (33%), Positives = 113/221 (50%), Gaps = 15/221 (6%)
Query: 27 SSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVV--NKNNDNVVTLPG---IALVWGLVL 81
S D S+ ++ +AEF+ T +F G S + +D + PG IA+ G L
Sbjct: 8 SFDDSFSLASLRAYLAEFISTLLFVFAGVGSAIAYAKLTSDAALDTPGLVAIAVCHGFAL 67
Query: 82 MVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSG-- 139
V + +ISG H NPAVTF A + I Y +QLLG+ A +LK + G
Sbjct: 68 FVAVAIGANISGGHVNPAVTFGLAVGGQITVITGVFYWIAQLLGSTAACFLLKYVTGGLA 127
Query: 140 --THDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLL 195
TH +G +++ V+E I TF L++ + A A D + ++G +A +AIG +
Sbjct: 128 VPTHSVAAGL----GSIEGVVMEIIITFALVYTVYATAADPKKGSLGTIAPLAIGLIVGA 183
Query: 196 NILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFG 236
NIL +GP +G SMNPAR+ GPA+ + VY+V + G
Sbjct: 184 NILAAGPFSGGSMNPARSFGPAVAAGDFSGHWVYWVGPLIG 224
>At3g26520 salt-stress induced tonoplast intrinsic protein
Length = 253
Score = 105 bits (262), Expect = 3e-23
Identities = 75/223 (33%), Positives = 105/223 (46%), Gaps = 8/223 (3%)
Query: 41 VAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPG----IALVWGLVLMVLIYSVGHISGAH 95
+AEF+ T +F G S I NK DN T P AL L V + +ISG H
Sbjct: 25 LAEFISTLIFVFAGSGSGIAFNKITDNGATTPSGLVAAALAHAFGLFVAVSVGANISGGH 84
Query: 96 FNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQ 155
NPAVTF ++ Y +QLLG+V A +L G G +L
Sbjct: 85 VNPAVTFGVLLGGNITLLRGILYWIAQLLGSVAACFLLSFATGGEPIPAFGLSAGVGSLN 144
Query: 156 AFVIEFITTFLLMFVISAVATD--NRAIGEMAGIAIGSTLLLNILISGPITGASMNPART 213
A V E + TF L++ + A A D N ++G +A IAIG + NIL G +GASMNPA
Sbjct: 145 ALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFIVGANILAGGAFSGASMNPAVA 204
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHE 256
GPA+ + VY+ + G +++ + + D+ HE
Sbjct: 205 FGPAVVSWTWTNHWVYWAGPLIGGGLAGIIYDFV-FIDENAHE 246
Score = 44.7 bits (104), Expect = 5e-05
Identities = 38/110 (34%), Positives = 53/110 (47%), Gaps = 6/110 (5%)
Query: 40 LVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPA 99
LV E V TF L++T A+ V+ N ++ T+ IA+ G ++ I + G SGA NPA
Sbjct: 146 LVFEIVMTFGLVYTVYAT-AVDPKNGSLGTIAPIAI--GFIVGANILAGGAFSGASMNPA 202
Query: 100 VTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLF--SGTHDQFSGT 147
V F A + W Y A L+G LA I +F H+Q T
Sbjct: 203 VAFGPAVVS-WTWTNHWVYWAGPLIGGGLAGIIYDFVFIDENAHEQLPTT 251
>At1g73190 tonoplast intrinsic protein, alpha (alpha-TIP)
Length = 268
Score = 103 bits (257), Expect = 1e-22
Identities = 73/242 (30%), Positives = 119/242 (49%), Gaps = 15/242 (6%)
Query: 28 SDTYTSVFFVQKLVAEFVGTFFLIFTGCASIV----------VNKNNDNVVTLPGIALVW 77
+D T ++ +AEF+ TF +F SI+ + + L +AL
Sbjct: 14 ADEATHPDSIRATLAEFLSTFVFVFAAEGSILSLDKLYWEHAAHAGTNTPGGLILVALAH 73
Query: 78 GLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLF 137
L + + ++SG H NPAVTF R I+ Y +QLLGA+LA +L++
Sbjct: 74 AFALFAAVSAAINVSGGHVNPAVTFGALVGGRVTAIRAIYYWIAQLLGAILACLLLRLTT 133
Query: 138 SGTHDQFSGTIPSGTN-LQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLL 194
+G + SG + V+E I TF L++V+ + D + ++G +A +AIG +
Sbjct: 134 NGMRP-VGFRLASGVGAVNGLVLEIILTFGLVYVVYSTLIDPKRGSLGIIAPLAIGLIVG 192
Query: 195 LNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRY-TDKP 253
NIL+ GP +GASMNPAR GPA+ ++ +Y+V G+ A ++ + T+ P
Sbjct: 193 ANILVGGPFSGASMNPARAFGPALVGWRWHDHWIYWVGPFIGSALAALIYEYMVIPTEPP 252
Query: 254 LH 255
H
Sbjct: 253 TH 254
>At5g47450 membrane channel protein-like; aquaporin (tonoplast
intrinsic protein)-like
Length = 250
Score = 103 bits (256), Expect = 1e-22
Identities = 70/216 (32%), Positives = 108/216 (49%), Gaps = 8/216 (3%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVV--NKNNDNVVTLPG---IALVWGLVLMVLIYS 87
SV ++ ++EF+ T +F G S V +D + G IA+ L V +
Sbjct: 14 SVSSLKAYLSEFIATLLFVFAGVGSAVAFAKLTSDGALDPAGLVAIAIAHAFALFVGVSI 73
Query: 88 VGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGT 147
+ISG H NPAVT A I Y +Q LG+++A +L + +G G
Sbjct: 74 AANISGGHLNPAVTLGLAIGGNITLITGFFYWIAQCLGSIVACLLLVFVTNGKSVPTHGV 133
Query: 148 IPSGTNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISGPITG 205
++ V+E + TF L++ + A A D + ++G +A IAIG + NIL +GP +G
Sbjct: 134 SAGLGAVEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFIVGANILAAGPFSG 193
Query: 206 ASMNPARTLGPAIFHSKYRAIVVYFVSTIF-GAVAG 240
SMNPAR+ GPA+ I +Y+V + GA+AG
Sbjct: 194 GSMNPARSFGPAVVSGDLSQIWIYWVGPLVGGALAG 229
>At4g17340 membrane channel like protein
Length = 250
Score = 102 bits (255), Expect = 2e-22
Identities = 68/216 (31%), Positives = 109/216 (49%), Gaps = 8/216 (3%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVV--NKNNDNVVTLPG---IALVWGLVLMVLIYS 87
SV ++ ++EF+ T +F G S + +D + G +A+ L V +
Sbjct: 14 SVASLKAYLSEFIATLLFVFAGVGSALAFAKLTSDAALDPAGLVAVAVAHAFALFVGVSI 73
Query: 88 VGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGT 147
+ISG H NPAVT A I Y +Q LG+++A +L + +G G
Sbjct: 74 AANISGGHLNPAVTLGLAVGGNITVITGFFYWIAQCLGSIVACLLLVFVTNGESVPTHGV 133
Query: 148 IPSGTNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISGPITG 205
++ V+E + TF L++ + A A D + ++G +A IAIG + NIL +GP +G
Sbjct: 134 AAGLGAIEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFIVGANILAAGPFSG 193
Query: 206 ASMNPARTLGPAIFHSKYRAIVVYFVSTIF-GAVAG 240
SMNPAR+ GPA+ + I +Y+V + GA+AG
Sbjct: 194 GSMNPARSFGPAVVSGDFSQIWIYWVGPLVGGALAG 229
>At2g16850 putative aquaporin (plasma membrane intrinsic protein)
Length = 278
Score = 100 bits (249), Expect = 8e-22
Identities = 64/221 (28%), Positives = 112/221 (49%), Gaps = 15/221 (6%)
Query: 36 FVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV---VTLPGIALVWGLVLMVLIYSVGHIS 92
F + ++AEF+ T ++ A+++ +KN V L GIA +G ++ VL+Y IS
Sbjct: 34 FYRAIIAEFIATLLFLYVTVATVIGHKNQTGPCGGVGLLGIAWAFGGMIFVLVYCTAGIS 93
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG---TIP 149
G H NPAVTF ++ + Y+ +Q LGA+ G++K + + G T+
Sbjct: 94 GGHINPAVTFGLFLARKVSLPRAVAYMVAQCLGAICGVGLVKAFMMTPYKRLGGGANTVA 153
Query: 150 SGTNL-QAFVIEFITTFLLMFVISAVATDNRA-----IGEMAGIAIGSTLLLNILISGPI 203
G + A E I TF+L++ + + R+ + +A + IG + + L + PI
Sbjct: 154 DGYSTGTALGAEIIGTFVLVYTVFSATDPKRSARDSHVPVLAPLPIGFAVFMVHLATIPI 213
Query: 204 TGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
TG +NPAR+ G A+ ++ +A +++V GA+A A
Sbjct: 214 TGTGINPARSFGAAVIYNNEKAWDDHWIFWVGPFVGALAAA 254
>At4g35100 plasma membrane intrinsic protein PIP3
Length = 280
Score = 100 bits (248), Expect = 1e-21
Identities = 65/221 (29%), Positives = 111/221 (49%), Gaps = 15/221 (6%)
Query: 36 FVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV---VTLPGIALVWGLVLMVLIYSVGHIS 92
F + L+AEF+ T ++ A+++ +K V L GIA +G ++ VL+Y IS
Sbjct: 36 FYRALIAEFIATLLFLYVTVATVIGHKKQTGPCDGVGLLGIAWAFGGMIFVLVYCTAGIS 95
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG---TIP 149
G H NPAVTF ++ ++ Y+ +Q LGA+ G +K ++ G T+
Sbjct: 96 GGHINPAVTFGLFLARKVSLVRALGYMIAQCLGAICGVGFVKAFMKTPYNTLGGGANTVA 155
Query: 150 SG-TNLQAFVIEFITTFLLMFVISAVATDNRA-----IGEMAGIAIGSTLLLNILISGPI 203
G + A E I TF+L++ + + R+ I +A + IG + + L + PI
Sbjct: 156 DGYSKGTALGAEIIGTFVLVYTVFSATDPKRSARDSHIPVLAPLPIGFAVFMVHLATIPI 215
Query: 204 TGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
TG +NPAR+ G A+ ++ +A +++V GA+A A
Sbjct: 216 TGTGINPARSFGAAVIYNNEKAWDDQWIFWVGPFLGALAAA 256
>At2g36830 putative aquaporin (tonoplast intrinsic protein gamma)
Length = 251
Score = 100 bits (248), Expect = 1e-21
Identities = 75/216 (34%), Positives = 105/216 (47%), Gaps = 13/216 (6%)
Query: 41 VAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPG------IALVWGLVLMVLIYSVG-HIS 92
+AEF+ T + G S + NK +N T P +A +GL + V SVG +IS
Sbjct: 24 LAEFISTLIFVVAGSGSGMAFNKLTENGATTPSGLVAAAVAHAFGLFVAV---SVGANIS 80
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGT 152
G H NPAVTF ++ Y +QLLG+V+A ILK G G
Sbjct: 81 GGHVNPAVTFGAFIGGNITLLRGILYWIAQLLGSVVACLILKFATGGLAVPAFGLSAGVG 140
Query: 153 NLQAFVIEFITTFLLMFVISAVATD--NRAIGEMAGIAIGSTLLLNILISGPITGASMNP 210
L AFV E + TF L++ + A A D N ++G +A IAIG + NIL G +GASMNP
Sbjct: 141 VLNAFVFEIVMTFGLVYTVYATAIDPKNGSLGTIAPIAIGFIVGANILAGGAFSGASMNP 200
Query: 211 ARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNI 246
A GPA+ + VY+ + G ++ +
Sbjct: 201 AVAFGPAVVSWTWTNHWVYWAGPLVGGGIAGLIYEV 236
>At2g29870 putative aquaporin (plasma membrane intrinsic protein)
Length = 139
Score = 99.8 bits (247), Expect = 1e-21
Identities = 48/103 (46%), Positives = 71/103 (68%)
Query: 145 SGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPIT 204
SG+ PSG++LQAFV+EFI T LM V+ AV T R E+ G+ IG+T+ LN++ G ++
Sbjct: 12 SGSSPSGSDLQAFVMEFIITGFLMLVVCAVTTTKRTTEELEGLIIGATVTLNVIFVGEVS 71
Query: 205 GASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNIL 247
GASMNPAR++GPA+ Y+ I +Y ++ GAV+ A + +L
Sbjct: 72 GASMNPARSIGPALVWGCYKGIWIYLLAPTLGAVSRALIHKML 114
Score = 41.6 bits (96), Expect = 4e-04
Identities = 36/119 (30%), Positives = 54/119 (45%), Gaps = 12/119 (10%)
Query: 21 ASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLV 80
A T+ SS + S +Q V EF+ T FL+ CA + + + L+ G
Sbjct: 5 ARNTMSSSGSSPSGSDLQAFVMEFIITGFLMLVVCAVTTTKRTTEELE-----GLIIGAT 59
Query: 81 LMVLIYSVGHISGAHFNPAVTFAFATT---KRFPWIQVAPYIASQLLGAVLASGILKML 136
+ + + VG +SGA NPA + A + WI Y+ + LGAV + I KML
Sbjct: 60 VTLNVIFVGEVSGASMNPARSIGPALVWGCYKGIWI----YLLAPTLGAVSRALIHKML 114
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.138 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,037,887
Number of Sequences: 26719
Number of extensions: 185795
Number of successful extensions: 730
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 570
Number of HSP's gapped (non-prelim): 56
length of query: 265
length of database: 11,318,596
effective HSP length: 97
effective length of query: 168
effective length of database: 8,726,853
effective search space: 1466111304
effective search space used: 1466111304
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0162a.10


