

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0162a.1
(118 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
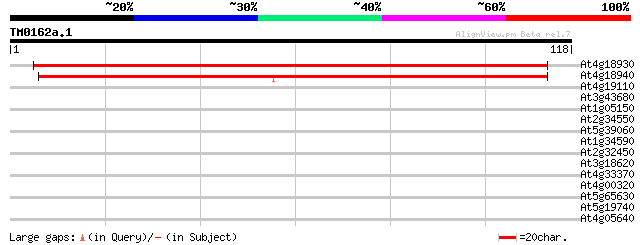
Sequences producing significant alignments: (bits) Value
At4g18930 unknown protein 131 5e-32
At4g18940 unknown protein (At4g18940) 105 3e-24
At4g19110 putative serine/threonine-protein kinase 28 0.73
At3g43680 putative protein 28 0.73
At1g05150 putative O-GlcNAc transferase 28 0.96
At2g34550 putative gibberellin 2-oxidase 27 1.2
At5g39060 transposase -like protein 27 1.6
At1g34590 hypothetical protein 27 2.1
At2g32450 putative O-GlcNAc transferase 26 2.8
At3g18620 unknown protein 25 4.7
At4g33370 putative protein 25 6.2
At4g00320 25 6.2
At5g65630 unknown protein 25 8.1
At5g19740 peptidase-like protein 25 8.1
At4g05640 putative protein 25 8.1
>At4g18930 unknown protein
Length = 181
Score = 131 bits (330), Expect = 5e-32
Identities = 61/108 (56%), Positives = 75/108 (68%)
Query: 6 IETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKL 65
+E KK+VYSVWA+P + PR KLM LRSEF GP F PH+TV + LTAD+A
Sbjct: 1 MEEVKKDVYSVWALPDEESEPRFKKLMEALRSEFTGPRFVPHVTVAVSAYLTADEAKKMF 60
Query: 66 RSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHF 113
SAC+ LKA+ TVDRV+ GTFF+QCV+LLL +P V+E HC NHF
Sbjct: 61 ESACDGLKAYTATVDRVSTGTFFFQCVFLLLQTTPEVMEAGEHCKNHF 108
>At4g18940 unknown protein (At4g18940)
Length = 196
Score = 105 bits (263), Expect = 3e-24
Identities = 48/109 (44%), Positives = 72/109 (66%), Gaps = 2/109 (1%)
Query: 7 ETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAI--TLTADDALNK 64
+ ++K +Y++WA+P D V R+ +LM LRSEFGGP F+PH+T+VG LTA +A
Sbjct: 18 DEEEKAMYAIWAVPEDDVEDRLQRLMEGLRSEFGGPAFDPHLTLVGPFPYKLTASEAKRM 77
Query: 65 LRSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHF 113
+SACE K + TVD+V+AGT ++QC+Y+ L + V+ + H HF
Sbjct: 78 FKSACEGFKVYPATVDQVSAGTSYFQCLYVSLRHTVEVMNAAGHFMAHF 126
>At4g19110 putative serine/threonine-protein kinase
Length = 461
Score = 28.1 bits (61), Expect = 0.73
Identities = 16/62 (25%), Positives = 26/62 (41%), Gaps = 2/62 (3%)
Query: 46 PHITVVGAITLTADDALNKLRSACEELKAFQVTVDRVAAGTFFYQCVYL--LLHPSPPVV 103
P + + + ++DA+N + C + + T V FF C Y+ L P P V
Sbjct: 241 PGVPLSSLMPSASEDAINLIERLCSWDPSSRPTAAEVLQHPFFQSCFYVPPSLRPKPSVA 300
Query: 104 ET 105
T
Sbjct: 301 RT 302
>At3g43680 putative protein
Length = 539
Score = 28.1 bits (61), Expect = 0.73
Identities = 17/52 (32%), Positives = 25/52 (47%), Gaps = 2/52 (3%)
Query: 64 KLRSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETS--AHCSNHF 113
K S+ E K + DRVA+ C Y LL P ++E+ A ++HF
Sbjct: 268 KKASSAEAGKELPIFEDRVASANLLGGCAYPLLSPPDTLLESRKYAETASHF 319
>At1g05150 putative O-GlcNAc transferase
Length = 808
Score = 27.7 bits (60), Expect = 0.96
Identities = 14/33 (42%), Positives = 20/33 (60%), Gaps = 2/33 (6%)
Query: 5 AIETQKKEVYSVWAIPPDH--VRPRIAKLMTDL 35
A+E QKK+ + WA+ P+H V KL+ DL
Sbjct: 120 AVEAQKKQRTAAWAVSPNHGIVFDETWKLVDDL 152
>At2g34550 putative gibberellin 2-oxidase
Length = 335
Score = 27.3 bits (59), Expect = 1.2
Identities = 13/36 (36%), Positives = 19/36 (52%)
Query: 42 PHFEPHITVVGAITLTADDALNKLRSACEELKAFQV 77
P +P ++ I LT DA ++ ACEE F+V
Sbjct: 18 PKCKPRPVLIPVIDLTDSDAKTQIVKACEEFGFFKV 53
>At5g39060 transposase -like protein
Length = 543
Score = 26.9 bits (58), Expect = 1.6
Identities = 21/80 (26%), Positives = 36/80 (44%), Gaps = 10/80 (12%)
Query: 10 KKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSAC 69
K ++ S A PP H IA +++L ++G E + TLT D+A
Sbjct: 90 KTKILSFCAFPPPHSGVAIAMKLSELLKDWG---IEKKV-----FTLTVDNA--SANDTM 139
Query: 70 EELKAFQVTVDRVAAGTFFY 89
+ + ++ D V +G FF+
Sbjct: 140 QSILKRKLQKDLVCSGEFFH 159
>At1g34590 hypothetical protein
Length = 820
Score = 26.6 bits (57), Expect = 2.1
Identities = 16/52 (30%), Positives = 24/52 (45%), Gaps = 2/52 (3%)
Query: 64 KLRSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETS--AHCSNHF 113
K S E K + DR+A+ CV LL P ++E+ A ++HF
Sbjct: 545 KKTSGSEAEKVLPIFEDRIASANLLVGCVGTLLPPPDTLLESRKYAETASHF 596
>At2g32450 putative O-GlcNAc transferase
Length = 802
Score = 26.2 bits (56), Expect = 2.8
Identities = 13/33 (39%), Positives = 20/33 (60%), Gaps = 2/33 (6%)
Query: 5 AIETQKKEVYSVWAIPPDH--VRPRIAKLMTDL 35
A+E QK++ + WA+ P+H V KL+ DL
Sbjct: 117 AVEAQKQQRTAAWAVSPNHGIVFDETWKLVDDL 149
>At3g18620 unknown protein
Length = 345
Score = 25.4 bits (54), Expect = 4.7
Identities = 9/21 (42%), Positives = 15/21 (70%)
Query: 91 CVYLLLHPSPPVVETSAHCSN 111
CVY L+H PP+ + +A+ S+
Sbjct: 211 CVYTLIHILPPIEKGAAYASD 231
>At4g33370 putative protein
Length = 542
Score = 25.0 bits (53), Expect = 6.2
Identities = 16/41 (39%), Positives = 19/41 (46%), Gaps = 7/41 (17%)
Query: 12 EVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVG 52
E S W PP HVR K M +R ++ HITV G
Sbjct: 56 EPLSTWWKPPLHVRKMSTKQMDLIRKQW-------HITVNG 89
>At4g00320
Length = 773
Score = 25.0 bits (53), Expect = 6.2
Identities = 11/32 (34%), Positives = 20/32 (62%)
Query: 51 VGAITLTADDALNKLRSACEELKAFQVTVDRV 82
+G +T + +D+L++L S C L+ V DR+
Sbjct: 162 LGRVTYSDEDSLHRLLSNCPVLEDLVVERDRI 193
>At5g65630 unknown protein
Length = 590
Score = 24.6 bits (52), Expect = 8.1
Identities = 17/47 (36%), Positives = 25/47 (53%), Gaps = 3/47 (6%)
Query: 71 ELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHFGYKS 117
ELK ++ +R+ +GTF Q Y + P P V SA +N G K+
Sbjct: 90 ELKQIRILRERIESGTFETQQGYTI--PEVPAVR-SAPLNNFTGEKN 133
>At5g19740 peptidase-like protein
Length = 681
Score = 24.6 bits (52), Expect = 8.1
Identities = 14/41 (34%), Positives = 20/41 (48%), Gaps = 6/41 (14%)
Query: 38 EFGGPHFEPHITVVGAITLTADDALNKLRSACEELKAFQVT 78
+FG P F+ H+ + + L A LR A EE+ F T
Sbjct: 504 KFGDPMFQRHVAMASVLGLVA------LRLADEEIIPFNYT 538
>At4g05640 putative protein
Length = 207
Score = 24.6 bits (52), Expect = 8.1
Identities = 11/31 (35%), Positives = 14/31 (44%)
Query: 20 PPDHVRPRIAKLMTDLRSEFGGPHFEPHITV 50
PPD+ RP + RS PH H T+
Sbjct: 100 PPDNTRPFTLPHHSTPRSSITSPHHHHHSTI 130
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.134 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,656,399
Number of Sequences: 26719
Number of extensions: 90238
Number of successful extensions: 227
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 216
Number of HSP's gapped (non-prelim): 15
length of query: 118
length of database: 11,318,596
effective HSP length: 94
effective length of query: 24
effective length of database: 8,807,010
effective search space: 211368240
effective search space used: 211368240
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0162a.1


